Abstract
Background:
The genetic variant rs9923231 (
Objective:
Our study aimed to investigate whether the relationship between rs9923231 and dementia persists only for certain dementia sub-types, and if those taking warfarin are at greater risk.
Methods:
We used logistic regression and data from 238,195 participants from UK Biobank to examine the relationship between
Results:
Parental history of dementia,
Conclusion:
Our study reports for the first time an association between rs9923231 and vascular dementia, but further research is warranted to explore potential mechanisms and specify the relationship between rs9923231 and features of vascular dementia.
INTRODUCTION
Warfarin is the most prescribed anticoagulant worldwide [1] and is commonly used as a treatment for atrial fibrillation (AF) [2]. The drug functions by inhibiting the enzyme vitamin K epoxide reductase (VKOR), effectively interfering with the vitamin K cycle required for coagulation of blood [3]. As a result of variations in age, height, weight, genotype, and other factors [4–6], patients vary up to 20-fold in their sensitivity to warfarin [7]. Clinically, the optimum dose is estimated using tests of blood coagulation, commonly the International Normalized Ratio (INR). The strongest genetic predictor of warfarin sensitivity is the gene
In a recent genome-wide association study (GWAS) meta-analysis of parental dementia and case-control Alzheimer’s dementia (ADem) [9],
Most strokes in western countries are due to occlusions in blood vessels (ischemic), and some are due to ruptures in blood vessels (hemorrhagic) [12]. If carriers of the T-allele in rs9923231 experience a reduction of blood coagulation and subsequent seq-uential minor hemorrhagic strokes, the resulting pa-thology could manifest in dementia and explain the observed link. Furthermore, compared to non-carriers of the T-allele, patients with AF that carry the T-allele could be at an increased risk of intracerebral hemorrhage and consequentially VaD when prescribed warfarin. Here, we study the same UK Biobank cohort as previously [9], but consider both individual and parental dementia status. We test whether T-allele status is associated with an increased risk of VaD and explore whether carriers of the T-allele are at a greater risk of VaD than non-carriers when prescribed warfarin.
METHODS
Sample
We used data from UK Biobank, a large and detailed prospective study of over 500,000 participants aged 37–73 that were recruited between the years 2006 and 2010. UK Biobank has been described in detail before [13]. The Research Ethics Committee (REC) granted ethical approval for the study (reference 11/NW/0382) and the current analysis was conducted under data application 10279.
Genotyping
Details on genotyping in the UK Biobank have been reported before [14, 15]. Briefly, for 49,950 participants, genotyping was performed using the UK BiLEVE Axiom Array, and for 438,427 participants, genotyping was performed using the UK Biobank Axiom Array. The released data contained 805,426 markers for 488,377 participants. Further quality control steps were performed as previously reported [9]. They included the removal of outliers, of incongruent data points, and of related participants using a relationship cut-off of 0.025 (GCTA GREML) [16]. This left an unrelated cohort of 314,278 individuals of white British ancestries (Fig. 1).
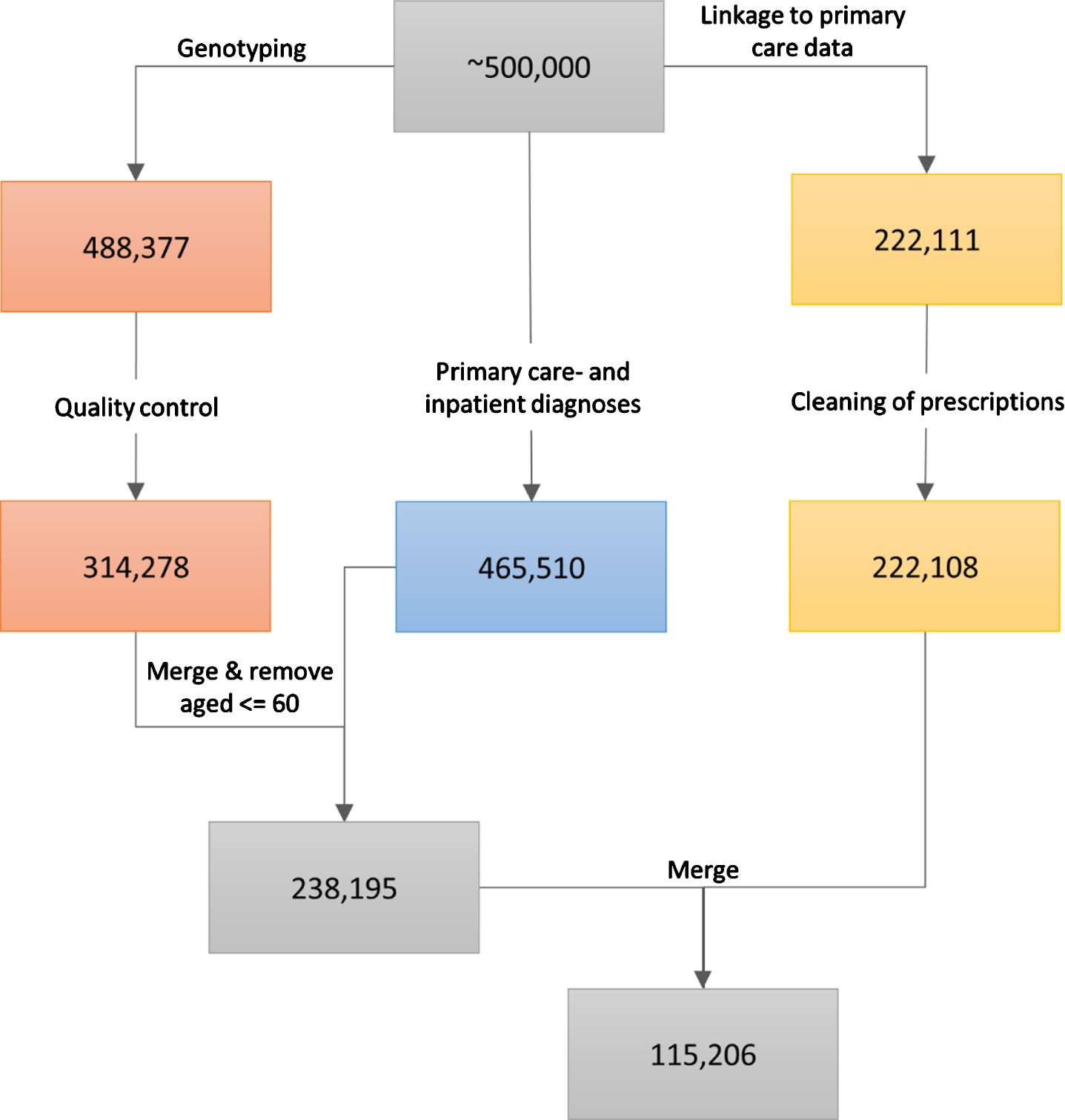
The data cleaning procedure. The left path (orange boxes) represents the genotyping and associated quality control, the middle path (blue box) represents the ascertainment of primary care- and inpatient diagnoses, and the right path (yellow boxes) represents the linkage to primary care prescriptions and the cleaning of the latter. The last two steps (grey boxes) involve the inclusion of only those participants that were older than 60 at the end of sampling and who passed through the left and middle paths (first grey box, 238,195 participants), or through all three data-cleaning paths (second grey box, 115,206 participants). All analyses that did not include prescribing data in the models were performed using the 238,195 participants, while the analyses that utilized warfarin prescription history used the 115,206 participants.
Warfarin prescription data
The UK Biobank obtained data on prescriptions for 222,111 participants via primary care computer system suppliers (EMIS Health and Vision for Scotland, and Wales, Vision and The Phoenix Partnership for England) and has engaged other intermediaries (Albasoft, a third-party data processor, for Scotland and the SAIL databank for Wales). All participants provided written consent for linkage to their health records upon recruitment to UK Biobank. The data were extracted in May 2017 for Scotland, in September 2017 for Wales, and in June, in July, and in August 2017 for England. The data include the exact dates of prescriptions, drug codes (BNF, Read v2, CTV3, and dm + d), names of drugs as written on the prescription, and, where available, the dosages of prescribed drugs. Empty prescriptions, prescriptions without a date, and duplicate prescriptions (defined as identical prescriptions issued to the same person on the same day) were removed from the sample. This resulted in the removal of 1,467,547 prescriptions. Three participants were completely removed from the dataset (Fig. 1). Warfarin prescriptions were extracted by searching for the word “warfarin” under the name/content of each prescription. For each participant, we calculated warfarin use by summing the number of days on which warfarin was prescribed, and warfarin dose by averaging the prescribed dose over all prescriptions of warfarin.
Disease status
Data on diagnoses for 465,510 participants were obtained by the UK Biobank from two sources: 1) from primary care similarly to the prescriptions described above, and 2) from hospital inpatient admissions data. Inpatients are defined as people who are admitted to hospital and occupy a hospital bed. These data included Hospital Episode Statistics for England, Scottish Morbidity Records for Scotland, and the Patient Episode Database for Wales. People with record of any dementia were included in a broad dementia category of “general dementia” that included ADem and VaD, as well as other types of dementia. Furthermore, narrower, more specific categories (ADem, VaD) were also identified. Information on the codes used in the extraction of each diagnosis is provided in Supplementary Table 1. We excluded from our analyses all participants that were 60 years old or younger on the last date of sampling (June 30, 2020) since dementia risk increases steeply with age. Parental diagnoses were ascertained during the initial assessment by asking participants about the presence of “Alzheimer’s disease/dementia” for both mother and father. In our analyses, the parental diagnosis of dementia was considered positive if at least one parent was reported to have suffered from the disorder.
Models
All analyses where the outcome variable was continuous were performed using linear regression; all models where the outcome variable was binary were performed using logistic regression. All models were controlled for the assessment center in which the par-ticipant was tested, the genotyping- batch and array, 40 genetic principal components, the age, sex, edu-cation, socioeconomic deprivation, alcohol consumption, smoking, physical activity, and body mass index (BMI) of the participants. The models predicting VaD were subsequently additionally controlled for
RESULTS
Sample characteristics
Among the 238,195 participants, 129,034 (54.2%) were female and 109,161 (45.8%) were male (Table 1). The age range at recruitment was 46–74 years and the median age was 60.9 years (IQR = 9.1). The demographic characteristics of the sub-sample used for analyses utilizing prescription history (Fig. 1) were very similar to the entire sample (Supplementary Table 2). A total of 13,361 (5.6%) participants had been diagnosed with AF and among the 115,206 participants with data on prescriptions, 5,513 (4.8%) had a history of being prescribed warfarin (Supplementary Table 3).
Demographic characteristics of the sample
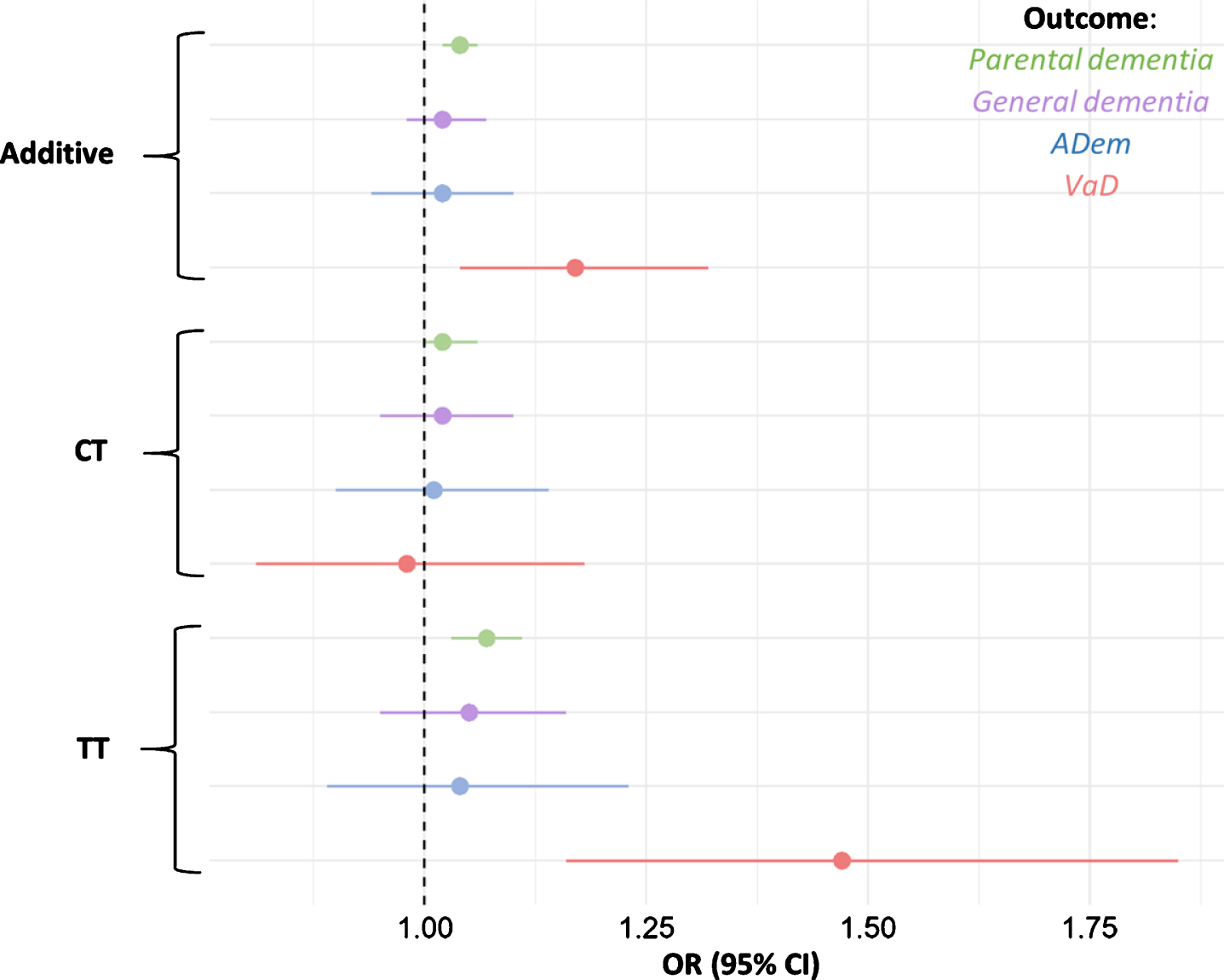
Odds ratios for parental dementia, ADem, and VaD per rs9923231 genotype status. Depicted are the additive effect and the effects of each allele group. The tails represent 95% confidence intervals for the ORs.
There were 145,186 (61.0%) carriers of the T-allele in the sample: 111,756 (46.9%) were heterozygous for the T-allele, and 33,430 (14.0%) were homozygous for the T-allele; the allele frequencies were in Hardy-Weinberg equilibrium (χ
rs9923231 polymorphism and warfarin dose
Carrying the T-allele was negatively associated with the average dose of warfarin (βperT - allele = –0.29, SE = 0.015,
rs9923231 polymorphism and dementia risk
Parents of carriers of the T-allele were more li-kely to have developed dementia (additive effect per T-allele: OR = 1.04, 95% CI = 1.02–1.06,
Results of the additive models with T as the effect allele, using rs9923231 to predict parental dementia, general dementia, ADem, and VaD
When limited to the specific outcome of VaD, the additive effect of the T-allele was much larger (OR = 1.17, 95% CI = 1.04–1.32,
Warfarin use and VaD in carriers of the T-allele
In our sample, participants diagnosed with AF were at greater risk for ADem (OR = 1.55, 95% CI = 1.31–1.81,
DISCUSSION
In this study, we explored the relationship between suspected dementia, atrial fibrillation, warfarin use, and rs9923231, whose T-allele is associated with a reduction in the dose of warfarin [3, 8]. We found a significant association between rs9923231 and suspected VaD, but not between rs9923231 and either suspected general dementia or suspected ADem. While AF was linked to VaD, the use of warfarin in patients that have AF and carry the T-allele did not increase the risk for VaD.
While there have been reports of variants for monogenic forms of VaD [20], data on the genetics of sporadic VaD are sparse. To our knowledge, only two GWAS have been conducted to investigate this: One (
Previous research has associated variation in rs9923231 with warfarin dose [3, 24], and with various adiposity-related traits, such as hip circumference, arm- and leg fat mass, and BMI (Gene Atlas [25]). To our knowledge the present study for the first time describes an association between rs9923231 and VaD, although it is important to note that this is not at a genome-wide significant threshold. The lack of a relationship between rs9923231 and either ADem or general dementia in the present study suggests that the association between the T-allele and ADem, as reported previously [9], might have been partly due to the classification of parental dementia. The UK Biobank questionnaire administered to participants did not distinguish between different types of dementia and it is not known how many of the 42,034 parents that were reportedly diagnosed with “Alzheimer’s/dementia” [9] may have suffered from VaD. This hypothesis is further supported by the estimated effect sizes for the association between rs9923231 and ADem, which were not num-erically larger than those for the association between rs9923231 genotype and parental dementia. Since parental dementia was used as a proxy for ADem in participants, the effect for ADem in participants sho-uld have been substantially greater than for parental dementia (even in the absence of statistical significance) if there truly was an association between rs9923231 and ADem (as opposed to an association between rs9923231 and VaD). Furthermore, in a rec-ent GWAS of clinically diagnosed ADem (
Based on our results and considering the impo-rtance of cardiovascular abnormalities in the pathology of dementia [10, 11], any future studies exploring the association between rs9923231 and dementia must strongly consider the role of cardiovascular factors; The relationship between genotype and dem-entia might hold only for cases of pure VaD or for those in which vascular pathology represents the main cause of the disorder.
There is an established association between dem-entia and both stroke [27] and WMH [28]; thus, stroke or WMH could act as mediators between rs9923231 genotype and VaD. However, we found no evidence for a positive association between either rs9923231 and stroke or rs9923231and WMH. Moreover, the latter association was statistically significant and negative in direction, suggesting participants carrying the T-allele were less likely to exhibit WMH. While we did not directly test for the effects of other relevant processes, including microbleeds and covert stroke, in the relationship between rs9923231 and VaD, given the lack of evidence for an association between rs9923231 and stroke, they are unlikely to act as prominent mediators. These results further complicate the potential relationship between rs9923231 and VaD and reinforce the need for additional studies to confirm this association and to test alternative mechanism distinct from stroke or WMH.
Interplay between AF, VaD and warfarin use
AF has been previously associated with cognitive decline and dementia. In our study, AF was associated with VaD and with ADem, even after controlling for hypertension and hypercholesterolemia. The association between AF and VaD is unsurprising, considering the inclusion of either vascular disease or history of stroke in almost all definitions of VaD [29]. Despite a substantial overlap of risk factors for AF and ADem, there is some evidence for an independent relationship between the two disorders [30, 31]
Due to the positive association between AF and VaD, the relationship between rs9923231 and VaD, and between rs9923231 and required warfarin dose, T-allele carriers that take warfarin to treat their AF might be at an increased risk of VaD than non-carriers due to warfarin-related brain hemorrhages. To test this, we studied an interaction between warfarin use, AF, and
Limitations and future directions
The present study has the advantages of having used a well-characterized sample with access to both inpatient- and primary-care diagnoses. However, we acknowledge several limitations. First, despite the large number of people recruited to UK Biobank, the age range at the end of the sampling period for the cohort is 60–83 years, resulting in a low incidence and prevalence of dementia. This heavily reduced the size of our sample, especially when testing for interactions, and led to wide confidence intervals for the estimated odds ratios. Second, despite it not being the only vitamin K antagonist anticoagulant on the UK market, only warfarin was included in the analysis. Third, the dose of warfarin ingested by participants was assumed to correspond to the average of their prescribed dose, despite some individuals possibly taking more or less of the medicine depending on their individual drug regimes. Fourth, clinical diagnoses of dementia subtypes are difficult and are prone to errors due to the presence of comorbidities and cardiovascular factors [32]. In the present paper, imaging data to confirm the diagnoses was unavailable and all diagnoses were based solely on records from primary care and hospitals. Finally, while most definitions of VaD include both dementia and a history of stroke or cardiovascular disease, VaD is very heterogeneous [33]; in the present study, we did not explore potential mechanisms and mediators of the association between rs9923231 and VaD, nor did we test the relationship for different subtypes of VaD.
The knowledge of genetic risk factors for diseases enables the generation of more accurate hypotheses about underlying biological mechanisms and illuminates potential targets for pharmacological intervention. Moreover, it allows for more informed stratification of participants in clinical trials. Studies that build on our research should aim to replicate the findings in a bigger sample and with greater precision determine the effect size for the association between rs9923231 and VaD. Additionally, further work is required to identify possible associations between rs9923231 and features of VaD, such as lacunar infarction, intracerebral hemorrhage, and white matter hyperintensities.
Footnotes
ACKNOWLEDGMENTS
DLM and REM are supported by Alzheimer’s Research UK major project grant ARUK-PG2017B-10. JM is supported by funding from the Wellcome Trust 4-year PhD in Translational Neuroscience—training the next generation of basic neuroscientists to embrace clinical research [108890/Z/15/Z]. PMV acknowledges funding from the Australian National Health and Medical Research Council (1113400) and the Australian Research Council (FL180100072). JM and TCR are members of the Alzheimer Scotland Dementia Research Centre funded by Alzheimer Scotland. TCR is employed by NHS Lothian and the Scottish Government. SRC is supported by Age UK (Disconnected Mind project), the UK Medical Research Council [MR/R024065/1] and a National Institutes of Health (NIH) research grant R01AG054628. The authors thank all participants of the UK Biobank for providing data for the study and Dr Michelle Luciano (Department of Psychology, University of Edinburgh) for managing UK Biobank data application 10279.
