Abstract
Background:
There is limited data on cerebrospinal fluid (CSF) biomarkers in sporadic amyloid-β (Aβ) cerebral amyloid angiopathy (CAA).
Objective:
To determine the profile of biomarkers relevant to neurodegenerative disease in the CSF of patients with CAA.
Methods:
We performed a detailed comparison of CSF markers, comparing patients with CAA, Alzheimer’s disease (AD), and control (CS) participants, recruited from the Biomarkers and Outcomes in CAA (BOCAA) study, and a Specialist Cognitive Disorders Service.
Results:
We included 10 CAA, 20 AD, and 10 CS participants (mean age 68.6, 62.5, and 62.2 years, respectively). In unadjusted analyses, CAA patients had a distinctive CSF biomarker profile, with significantly lower (
Conclusion:
CAA has a characteristic biomarker profile, suggestive of a global, rather than selective, accumulation of amyloid species; we also provide evidence of different phenotypes according to amyloid-PET positivity. Further replication and validation of these preliminary findings in larger cohorts is needed.
INTRODUCTION
Sporadic amyloid-β (Aβ) cerebral amyloid angiopathy (CAA) can be reliably diagnosed during life using the clinico-radiological Boston criteria [1, 2], but nearly all of its associated imaging features are likely to be irreversible markers of late stage disease [3]. Body fluid biomarkers, using either cerebrospinal fluid (CSF) or blood, are of interest as important candidate biomarkers which can be sampled repeatedly, allow for measurement of a variety of different disease-related processes, and provide insights into disease dynamics.
To date, most data on CSF biomarkers in CAA have focused on Aβ (Aβ40, Aβ42) and tau (total tau, t-tau, and phospho-tau, p-tau) measures. However, there is limited information on other amyloid species in CAA, including smaller proteins such as Aβ38, soluble amyloid-β protein precursor (sAβPP)
Our aim was to perform a detailed comparison of amyloid markers (Aβ38, Aβ40, Aβ42, sAβPP
MATERIALS AND METHODS
Patient selection
Participants were included from two sources. Firstly, we included participants from the cross-sectional prospective observational BOCAA (Biomarkers and Outcomes in Cerebral Amyloid Angiopathy) study (10 patients with CAA, 5 CS participants). Ethical approval for the BOCAA study was granted in October 2015 by the NHS Health Research Authority London (REC reference 15/LO/1443). Secondly, we included samples collected by the Specialist Cognitive Disorders Service at the NHNN, University College London Hospitals (UCLH) NHS Trust, London, UK (20 samples from patients with AD, 5 samples from age-matched CS participants). This study was approved by the Regional Ethics Committee at UCL.
In all cases, informed written consent was obtained for each participant, and inclusion criteria were standardized to be consistent with the BOCAA study (further details below).
Patients with CAA
All patients with CAA were recruited from the BOCAA study [4]. Consecutive patients with CAA were identified from a prospectively collected research database. Patients with CAA all met at least probable modified Boston Criteria [2], and were not included if they had evidence of co-existing AD or deep perforator (hypertensive) arteriopathy [4]. Further inclusion criteria were: age ≥55 years, Mini-Mental State Examination (MMSE) score ≥23, modified Rankin scale (mRS) ≤3 and capacity to give informed consent. Those with contraindications to PET or MRI scanning or lumbar puncture were excluded.
Control (CS) participants
CS participants were included from two sources. In all cases, CS participants were required to have no prior history of significant neurological disease. Further inclusion criteria were: age ≥55 years, MMSE score ≥23, and modified Rankin scale (mRS) ≤3.
Firstly, we included CS participants recruited as part of the BOCAA study, where patient partners were invited to participate as healthy volunteers. CS participants were also identified from a prospectively collected database of patients attending the ambulatory transient ischemic attack (TIA) service, provided by the NHNN; patients whose final diagnosis was not stroke, TIA, or any other significant neurological condition (and met the inclusion and exclusion criteria) were invited to participate. Those with contraindications to PET or MRI scanning or lumbar puncture were excluded.
Secondly, we included samples collected by the Specialist Cognitive Disorders Service at the National Hospital of Neurology and Neurosurgery (NHNN). Samples for age-matched CS participants were included if their final diagnosis, made on the basis of clinical assessment, imaging, and CSF, was not one of dementia or any other neurodegenerative condition [5]. MR imaging (acquired as part of routine clinical care) was reviewed for evidence of previous infarction (including lacunes), cerebral microbleeds, and cortical superficial siderosis; participant samples were only included in the absence of all of these features. Atrophy (medial temporal [6] and global cortical [7]) and white matter hyperintensities [8] were assessed on brain imaging, and those with evidence of moderate or severe grades of these imaging features were excluded.
Patients with Alzheimer’s disease
Patients with AD presented with “typical” [9] amnestic symptoms, were aged ≥55 years, and had a final diagnosis (on the basis of clinical assessment, imaging, and CSF) that was in keeping with AD; additionally, all imaging was reviewed for the presence of cerebral microbleeds and cortical superficial siderosis, and samples from patients with these features were not included (in order to avoid patients with mixed CAA and AD pathology). The CSF criteria for AD diagnosis were the presence of a t-tau/Aβ42 ratio >0.88 together with Aβ42 < 630 pg/ml [5].
CSF analysis
All CSF analyses were performed by the Biomarker Laboratory of the UK Dementia Research Institute at UCL (Group Lead: Professor Henrik Zetterberg). CSF was collected, processed, and stored according to standardized procedures, and was identical for all diagnostic groups [10]. Samples were collected in polypropylene tubes, immediately transported to the laboratory by hand, and centrifuged (at 1,750 for 5 min at 4°C) within 30 min from collection; samples were then aliquoted and stored at –80°C until testing. All biochemical assays were performed by operators blinded to the clinical diagnosis.
Amyloid markers
Aβ38, Aβ40, and Aβ42 were measured by electrochemiluminescence (ECL) using a Meso Scale Discovery V-PLEX Aβ peptide panel 1(6E10) kit, according to the manufacturer’s instructions (MSD, Rockville, MD). Briefly, samples were diluted 1 : 2 with diluent 35 and added in duplicate to microplate wells coated with mouse monoclonal peptide specific capture antibodies for human Aβx-38/x-40/x-42. Samples were incubated with anti-Aβ (amino acids 1–16 epitope) antibody (6E10 clone) as the detection antibody conjugated with an electrically excitable SULFO-TAG. Concentrations were calculated from ECL signal using a four-parameter logistic curve fitting method with the MSD Workbench software package. Intra-assay CVs (coefficients of variation) were less than 10%. All samples were measured on the same day by a single operator using the same reagents.
sAβPP
Tau markers (t-tau and p-tau)
The levels of CSF t-tau and p-tau (181P) were determined using a sandwich ELISA (INNOtest® hTAU-Ag p-Tau (181P) ; Fujirebio Europe N.V., Gent, Belgium) constructed to measure both normal tau and phosphorylated tau. Briefly, for the hTAU Ag assay, tau protein is captured from CSF samples by a monoclonal anti-tau antibody (AT120) bound to a microtiter plate. Captured tau is detected with two biotinylated tau-specific monoclonal antibodies (HT7 and BT2). Similarly, for the t-tau assay, p-tau (181P) is captured from CSF samples by anti-tau antibody HT7 bound onto a microtiter plate. Captured p-tau (181P) is detected with a biotinylated monoclonal anti-p-tau antibody (AT270). In both assays, peroxidase-labelled streptavidin and tetramethylbenzidine (TMB) substrate are also added. Peroxidase-catalyzed hydrolysis produces a colorimetric signal. Sample concentrations are extrapolated from a standard curve, fitted using a 4-parameter logistic algorithm. Intra-assay CVs were less than 20%.
Neurofilament light
NFL was measured using the commercially available NF-Light ELISA, according to the manufacturer’s instructions (UmanDiagnostics, Umeå, Sweden). Briefly, samples were diluted 1 : 2 with sample diluent and added in duplicate to microplate wells coated with a monoclonal capture antibody specific for NFL. Samples were incubated with a biotinylated NFL-specific monoclonal detection antibody. The detection complex was completed with the addition of horseradish peroxidase-labelled streptavidin and TMB substrate. Peroxidase-catalyzed hydrolysis produces a colorimetric signal. Sample concentrations were extrapolated from a standard curve, fitted using a 4-parameter logistic algorithm. Intra-assay CVs were less than 10%. Samples were run on two different days by different operators; the inter-assay CV was below 16%.
sTREM2
Samples were analyzed using an immunoassay protocol adapted from a previously published protocol [11]. Streptavidin-coated 96-well plates (Meso-Scale Discovery (MSD), Rockville, MD, USA) were blocked overnight at 4°C in block buffer (0.5% bovine serum albumin (BSA) and 0.05% Tween 20 in PBS; pH 7.4). The plates were then incubated with the biotinylated polyclonal goat anti-human TREM2 capture antibody (0.25
Neurogranin
Neurogranin was measured with the EUROIMMUN Elisa (EQ6551-9601-L) according to manufacturer’s instructions (EUROIMMUN, Lübeck, Germany). Briefly, samples were incubated with biotinylated monoclonal anti-neurogranin antibody, followed by addition to microplate wells coated with monoclonal antibodies specific for human neurogranin truncated at P75. Finally, streptavidin peroxidase conjugate was added to initiate the color-changing reaction. Intra-assay CVs were less than 10%, and all samples were measured on the same day by a single operator using the same reagents. Sample concentrations are extrapolated from a standard curve, fitted using a 5-parameter logistic algorithm.
Amyloid-PET acquisition and interpretation
All participants in the BOCAA study underwent PET (using the amyloid ligand 18F-Florbetapir, Amyvid) and MR scanning, acquired using the same hybrid Siemens Biograph PET/MR scanner; the protocol for acquisition and initial processing has been previously described [12]. Visual reads were performed by a trained individual who was blind to all participant clinical details (including diagnosis).
Statistics
Statistical analysis was performed using Stata (Version 15.1). Group characteristics were compared using one-way ANOVA (age), chi squared (sex), or Kruskal-Wallis (MMSE) tests. Median and interquartile range values were calculated for each biomarker, and, given the non-normal distribution of the data, comparisons between groups were made using the Kruskal-Wallis test. If a significant difference was identified (defined as p < 0.05), Dunn’s test was used for
In order to perform age-adjusted analyses, we used quantile regression (comparing group medians) and calculated predicted medians. We then performed
Finally, we performed exploratory
RESULTS
We included 20 patients with AD, 10 patients with CAA, and 10 CS participants in this analysis; baseline characteristics are shown in Table 1. Patients with CAA were older than the two other groups (mean age±SD: CAA 68.6±3.0 years, AD 62.5± 4.1 years, and CS 62.2±5.4 years). As expected, those in the AD group had a lower MMSE (median score 24, compared with 29 for CAA and CS groups).
Comparison of characteristics and biomarkers by group. Group comparison p values were obtained using one-way ANOVA (age), chi squared tests (sex), or Kruskal-Wallis tests (remainder).
Of the patients with CAA, 5 (50%) presented with intracerebral hemorrhage, and the remainder presented with transient focal neurological episodes associated with convexity subarachnoid hemorrhage. All CAA patients had been clinically asymptomatic in the 6 months prior to their study visits. Amongst the CAA group, 7 patients (70%) showed evidence of cortical superficial siderosis (focal in three cases, disseminated in four cases) and 9 (90%) had lobar microbleeds (median number, 3.5). All CAA patients showed evidence of both deep and periventricular white matter hyperintensities on MRI (median Fazekas scores, 1 in deep and 2 in periventricular regions [8]).
Individual markers
Univariable comparisons
Univariable comparisons are shown in Table 1 and Fig. 1. There were significant differences between the three groups for the following markers: Aβ38, Aβ40, Aβ42, sAβPP
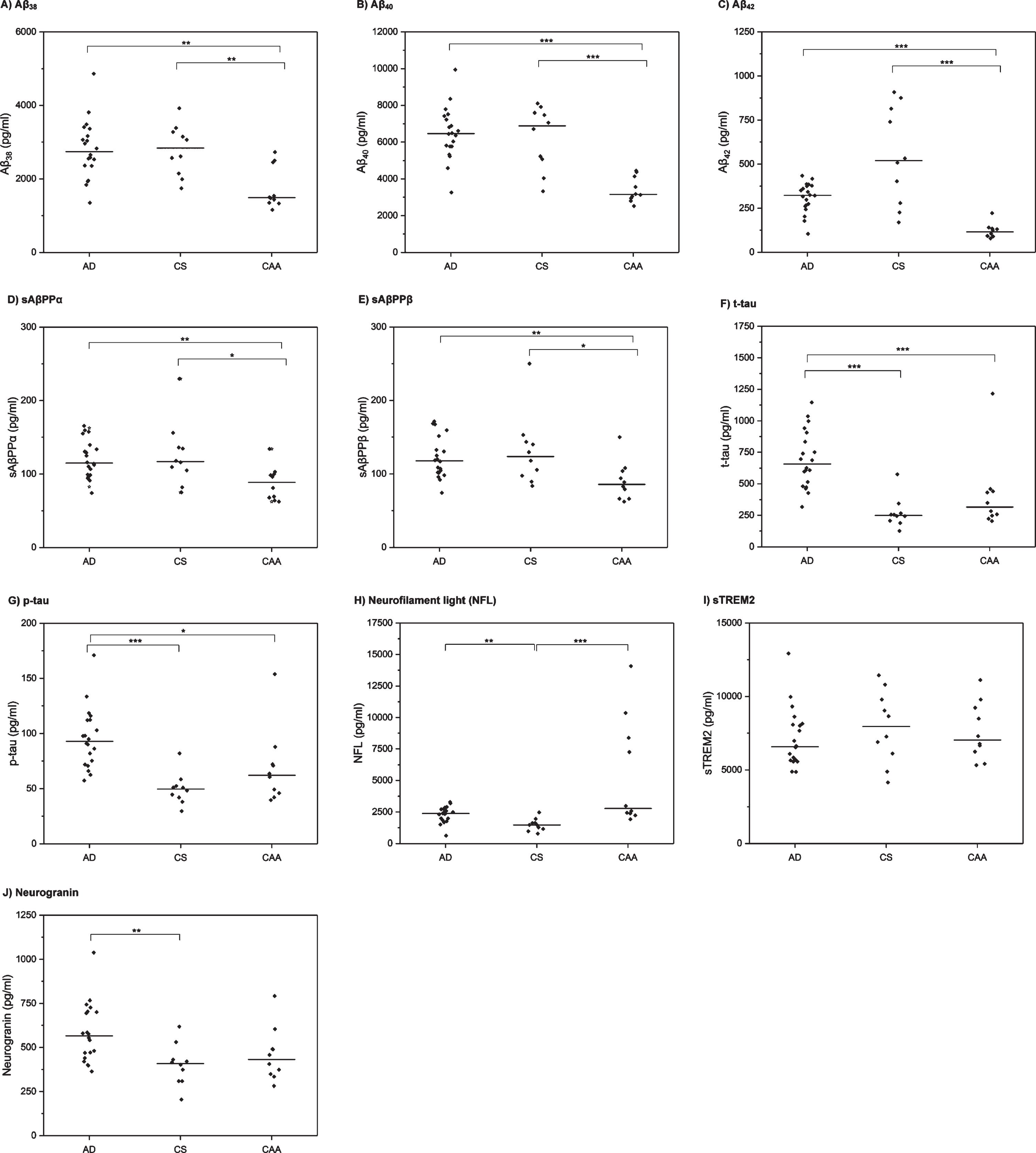
CSF biomarker profiles in AD, CAA, and control (CS) participants. Horizontal line indicates median value per group. Each diamond indicates an individual data point.
In
Age-adjusted quantile regression
Results from the age-adjusted quantile regression are shown in Table 2; scatter plots demonstrating the distribution of each biomarker by age are shown in Supplementary Figure 1. For the amyloid markers, there were significant differences between the three groups for Aβ38, Aβ40, Aβ42, and sAβPPβ, but not sAβPP
Age-adjusted quantile regression (comparing medians)
There were significant differences between the three groups for both t-tau and p-tau. Pairwise comparisons of the age-adjusted medians found significant differences for both t-tau (higher in AD group; median difference 357 pg/ml) and p-tau (higher in AD group; median difference 40.9 pg/ml) between the CAA and AD groups. There was no significant difference between the CAA and CS groups for either t-tau (median difference 53.1 pg/ml) or p-tau (median difference 2.2 pg/ml). There were significant differences between the AD and CS groups for t-tau (higher in AD group; median difference 410 pg/ml) and p-tau (higher in AD group; median difference 43.1 pg/ml).
There was a significant difference in neurogranin between the three groups, but no significant differences were identified in pairwise
Aβ ratios
Results for comparison of Aβ ratios are provided in the Supplementary Material.
Associations with amyloid-PET positivity
We then performed
Comparison of PET positive and negative patients with CAA.
Given these results, we performed a further comparison, in which amyloid-PET negative CAA patients were compared with AD and CS groups; these results are provided in the Supplementary Material. Patients with amyloid-PET negative CAA had lower Aβ38, Aβ40, and Aβ42 than both AD and CS groups, with no statistically significant differences between the AD and CS groups for these markers. There were no significant differences in sAβPP
DISCUSSION
In this exploratory, hypothesis-generating study, we found that patients with CAA had a distinctive CSF profile compared with CS participants and patients with AD. In unadjusted analyses, patients with CAA showed lower levels of all amyloid components measured (Aβ38, Aβ40, Aβ42, sAβPP
Our results for Aβ40, Aβ42, t-tau, and p-tau in CAA are in keeping with previously reported data [13–19]; however, we extend this earlier work further by demonstrating that Aβ40 and Aβ42 are not the only amyloid species to be reduced in CAA. The processing pathway from AβPP to pathological Aβ is well described [20–22]. However, CAA differs from AD in that parenchymal Aβ plaques are predominantly composed of Aβ42, whereas the vascular Aβ deposits in CAA are a mixture of Aβ40 and Aβ42, with the former being more common [16, 23]. The reduced levels of CSF Aβ40 and Aβ42 previously described in CAA are thought to be secondary to “selective trapping” of both these species in the vasculature, in contrast with AD, where only Aβ42 is found (“trapped”) in the parenchyma [16]. Aβ peptides of other lengths, including Aβ38, have also been shown to be deposited in the leptomeningeal vasculature [24]. Our finding of reductions in CSF Aβ38 and sAβPPβ (and sAβPP
Our results also provide new information on non-amyloid biomarkers in CAA. We found significant elevations in t-tau, p-tau, and neurogranin in patients with AD, but not in patients with CAA compared with healthy controls. This is in contrast with other studies which have found that t-tau and p-tau levels in CAA are higher than controls but lower than patients with AD [13, 27], and may reflect our small sample size. Pathological aggregation of tau protein is important in AD [28, 29]; tau aggregation is thought to result in synaptic dysfunction and subsequent neuronal loss, and in AD, it is tau (rather than Aβ) pathology that most closely correlates with cognition [29]. The intermediate tau levels described in CAA might thus be reflective of coexistent AD pathology. Cognitive impairment is a recognized feature of CAA [30] and while there is evidence that CAA is associated with atrophy (presumably secondary to neuronal loss [31]), cognitive impairment in these patients might be secondary to other mechanisms, such as network disruption [32] or impaired blood flow responses [33]. Our CSF findings suggest that synaptic dysfunction is a less prominent feature of CAA compared with AD, and that these markers might be useful for distinguishing these two Aβ pathologies in patients with cognitive impairment.
We also provide new data on other markers that have not been tested in CAA. NFL has shown great promise as a biomarker in a large number of neurological conditions [34], including sporadic cerebral small vessel disease [35–37], although age-adjusted analyses were only performed in one study [36]. We did not find a difference between the AD, CAA, and CS groups in age-adjusted analyses for CSF NFL, but in unadjusted analyses, the CAA group had higher NFL than the control participant group, and in the
Our exploratory
Our work has a number of strengths. We were able to evaluate a large number of markers and to reduce false-positive error we have used two separate statistical methods, one rank-based (Kruskal-Wallis) and the other an age-adjusted analysis based upon comparison of medians. However, there are some limitations. As mentioned earlier, this was a small exploratory study which may not have been powered to detect differences for all the biomarkers considered. Given our small sample size, we acknowledge the possibility of false-positive results due to multiple comparisons as a limitation of our work. While larger studies have considered some of the CSF markers we describe (in particular Aβ40, Aβ42, t-tau, and p-tau), we provide new data in CAA for a number of other markers, which we hope will inform future research with regard to effect sizes and sample size calculations. We acknowledge that there can be discrepancies between clinical and pathological diagnoses of CAA and AD, as well as significant overlap in these pathologies. While we attempted to avoid this on the basis of clinical and radiological findings, we did not have autopsy data, which would be the gold standard. However, we would argue that most clinicians who review patients with AD or CAA make their diagnosis on the basis of clinical and radiological (rather than autopsy) findings, and therefore our results remain of relevance. We specifically selected non-demented CAA patients with “early” or mild disease; these patients may not have such marked biomarker perturbations as those with more severe disease. The AD patients selected from the Specialist Cognitive Disorders Service may not be fully representative of AD patients more generally; this is a highly specialist tertiary service, which often sees younger patients or those with atypical presentations. We used CSF criteria (t-tau and Aβ42) in order to identify patients with AD and half of the control group, and so these markers are not fully independent in our analyses. Many of our control participants presented to neurology services (either the TIA clinic, or the Specialist Cognitive Disorders Service), and although we only included individuals without clinical, radiological, or CSF evidence of significant neurological disease, we acknowledge that they cannot be described as true “healthy” controls; this is a further limitation of our work. We selected control participants with minimal radiological evidence of brain pathology (WMH, atrophy), and this may not be truly representative of unselected age-matched individuals without AD or CAA. We determined PET positivity on qualitative (i.e., based on visual reads) rather than quantitative grounds, and the group sizes are particularly small in this
Conclusions
Our findings suggest that patients with CAA appear to have a CSF profile distinct from both patients with AD and age-matched control participants, characterized by a global reduction in secreted AβPP and Aβ species, but normal neuronal protein levels (tau, neurogranin) compared with AD patients. Replication of these findings in larger cohorts with longitudinal cohort measurements is required to confirm these markers as effective biomarkers for CAA, and their value for monitoring disease progression in clinical trials.
Footnotes
ACKNOWLEDGMENTS
We would like to acknowledge our consultant colleagues from the Dementia Research Centre, who contributed patient samples, and the clinicians who performed the lumbar punctures.
The BOCAA study was funded by the Rosetrees Trust. GB receives funding from the NIHR (as an Academic Clinical Fellow) and the Rosetrees Trust. AK acknowledges the support of the Wolfson Foundation and Weston Brain Institute. HZ is a Wallenberg Academy Fellow supported by grants from the Swedish Research Council, the European Research Council and the UK Dementia Research Institute at UCL. JMS acknowledges the support of the National Institute for Health Research University College London Hospitals Biomedical Research Centre, Wolfson Foundation, EPSRC (EP/J020990/1), MRC Dementias Platform UK (MR/L023784/1), ARUK (ARUK-Network 2012-6-ICE; ARUK-PG2017-1946; ARUK-PG2017-1946), Brain Research UK (UCC14191), Weston Brain Institute (UB170045) and European Union’s Horizon 2020 research and innovation programme (Grant 666992); in addition NCF acknowledges support from the UK Dementia Research Institute. DJW receives research support from the Stroke Association, the British Heart Foundation and the Rosetrees Trust. This work was undertaken at UCLH/UCL which receives a proportion of funding from the Department of Health’s National Institute for Health Research (NIHR) Biomedical Research Centres funding scheme.
