Abstract
An increasing number of researchers support reproducibility by including pointers to and descriptions of datasets, software and methods in their publications. However, scientific articles may be ambiguous, incomplete and difficult to process by automated systems. In this paper we introduce RO-Crate, an open, community-driven, and lightweight approach to packaging research artefacts along with their metadata in a machine readable manner. RO-Crate is based on Schema.org annotations in JSON-LD, aiming to establish best practices to formally describe metadata in an accessible and practical way for their use in a wide variety of situations.
An RO-Crate is a structured archive of all the items that contributed to a research outcome, including their identifiers, provenance, relations and annotations. As a general purpose packaging approach for data and their metadata, RO-Crate is used across multiple areas, including bioinformatics, digital humanities and regulatory sciences. By applying “just enough” Linked Data standards, RO-Crate simplifies the process of making research outputs FAIR while also enhancing research reproducibility.
An RO-Crate for this article
1
Introduction
The move towards Open Science has increased the need and demand for the publication of artefacts of the research process [104]. This is particularly apparent in domains that rely on computational experiments; for example, the publication of software, datasets and records of the dependencies that such experiments rely on [113].
It is often argued that the publication of these assets, and specifically software [80], workflows [55] and data, should follow the FAIR principles [123]; namely, that they are Findable, Accessible, Interoperable and Reusable. These principles are agnostic to the
Important examples include data publication with rich metadata (e.g. Zenodo [40]), domain-specific data deposition (e.g. PDB [16]) and following practices for reproducible research software [101] (e.g. use of containers). While these platforms are useful, experience has shown that it is important to put greater emphasis on the interconnection of the multiple artefacts that make up the research process [71].
The notion of
A Research Object combines the ability to bundle multiple types of artefacts together, such as spreadsheets, code, examples, and figures. The RO is augmented with annotations and relationships that describe the artefacts’
This notion of ROs provides a compelling vision as an approach for implementing FAIR data. However, existing Research Object implementations require a large technology stack [14], are typically tailored to a particular platform and are also not easily usable by end-users.
To address this gap, a new community came together [23] to develop
An introduction to RO-Crate, its purpose and context;
A guide to the RO-Crate community and tooling;
Examples of RO-Crate usage, demonstrating its value as connective tissue for different artefacts from different communities.
The rest of this paper is organised as follows. We first describe RO-Crate through its development methodology that formed the RO-Crate concept, showing its foundations in Linked Data and emerging principles. We then define RO-Crate technically, before we introduce the community and tooling. We move to analyse RO-Crate with respect to usage in a diverse set of domains. Finally, we present related work and conclude with some remarks including RO-Crate highlights and future work. The appendix adds a formal definition of RO-Crate using First-Order logic.
RO-Crate
RO-Crate aims to provide an approach to packaging research artefacts with their metadata that can be easily adopted. To illustrate this, let us imagine a research paper reporting on the sequence analysis of proteins obtained from an experiment on mice. The sequence output files, sequence analysis code, resulting data and reports summarising statistical measures are all important and inter-related research artefacts, and consequently would ideally all be co-located in a directory and accompanied with their corresponding metadata. In reality, some of the artefacts (e.g. data or software) will be recorded as external reference to repositories that are not necessarily following the FAIR principles. This conceptual directory, along with the relationships between its constituent digital artefacts, is what the RO-Crate model aims to represent, linking together all the elements of an experiment that are required for the experiment’s reproducibility and reusability.
The question then arises as to how the directory with all this material should be packaged in a manner that is accessible and usable by others. This means programmatically and automatically accessible by machines and human readable. A de facto approach to sharing collections of resources is through compressed archives (e.g. a zip file). This solves the problem of “packaging”, but it does not guarantee downstream access to all artefacts in a programmatic fashion, nor describe the role of each file in that particular research. Both features, the ability to automatically access and reason about an object, are crucial and lead to the need for explicit metadata about the contents of the folder, describing each and linking them together.
Examples of metadata descriptions across a wide range of domains
2
RO-Crate seeks to address this complexity by:
being conceptually simple and easy to understand for developers;
providing strong, easy tooling for integration into community projects;
providing a strong and opinionated guide regarding current best practices;
adopting de-facto standards that are widely used on the Web.
In the following sections we demonstrate how the RO-Crate specification and ecosystem achieve these goals.
It is a good question as to what base level we assume for ‘conceptually simple’. We take simplicity to apply at two levels: for the
For our development methodology we followed the mantra of working closely with a small group to get a deep understanding of requirements and ensure rapid feedback loops. We created a pool of early adopter projects from a range of disciplines and groups, primarily addressing developers of platforms. Thus the base level for simplicity was
We assumed a developer familiar with making Web applications with JSON data (who would then learn how to make
Addressing the simplicity of understanding and engaging with RO-Crate by data practitioners is through the platforms, for example with interactive tools (Section 3) like Describo
4
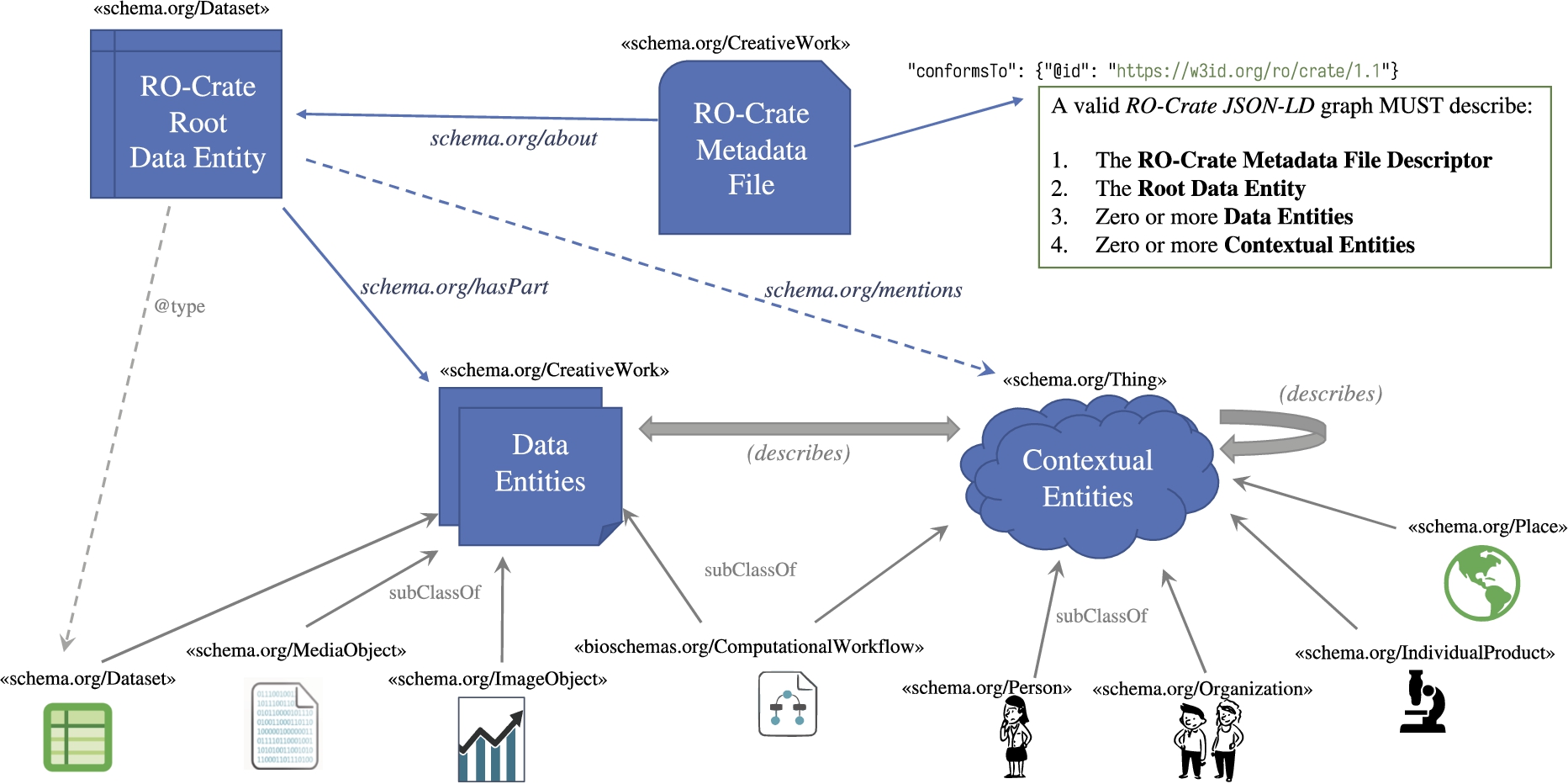
A key premise of RO-Crate is the existence of a wide variety of resources on the Web that can help describe research. As such, RO-Crate relies on the Linked Data principles [63]. Figure 1 shows the main conceptual elements involved in an RO-Crate: The RO-Crate Metadata File (top) describes the Research Object using structured metadata including external references, coupled with the contained artefacts (bottom) bundled and described by the RO-Crate.
The conceptual notion of a

Conceptual overview of RO-Crate. A
The
RO-Crates are
The foundation of Linked Data and shared vocabularies also means that multiple RO-Crates and other Linked Data resources can be indexed, combined, queried, validated or transformed using existing Semantic Web technologies such as SPARQL,
6
The possibilities of consuming
10
Some consideration is needed in processing of RO-Crates as knowledge graphs, e.g. establishing absolute IRIs for files inside a ZIP archive, detailed in the RO-Crate specification: Note that an RO-Crate is not required to be published on the Web, see Section 2.2.2.
An RO-Crate is defined
12
The Root Data Entity is a directory, the
The minimal requirements for the root data entity metadata
14
RO-Crates can be stored, transferred or published in multiple ways, e.g. BagIt [74], Oxford Common File Layout [96] (OCFL), downloadable ZIP archives in Zenodo or through dedicated online repositories, as well as published directly on the Web, e.g. using GitHub Pages.
15
RO-Crate distinguishes between data and contextual entities
16
As both types of entities are identified by IRIs, their distinction is allowed to be blurry; data entities can be located anywhere and be complex, while contextual entities can have a Web presence beyond their description inside the RO-Crate. For instance
A particular IRI may appear as a contextual entity in one RO-Crate and as a data entity in another; the distinction lies in the fact that data entities can be considered to be
In RO-Crate, a referenced contextual entity (e.g. a person identified by ORCID) should always be described within the RO-Crate Metadata File with at least a
Figure 2 shows a simplified UML class diagram of RO-Crate, highlighting the different types of data entities and contextual entities that can be aggregated and related. While an RO-Crate would usually contain one or more data entities (
Simplified UML class diagram of RO-Crate. The
RO-Crate as a specification aims to build a set of recommended practices on how to practically apply existing standards in a common way to describe research outputs and their provenance, without having to learn each of the underlying technologies in detail.
As such, the RO-Crate 1.1
18
However the primary purpose of the RO-Crate specification is to assist developers in leveraging Linked Data principles for the focused purpose of describing Research Objects in a structured language, while reducing the steep learning curve otherwise associated with Semantic Web adaptation, like development of ontologies, identifiers, namespaces, and RDF serialization choices.
One aim of RO-Crate is to be conceptually simple. This simplicity has been repeatedly checked and confirmed through an informal community review process. For instance, in the discussion on supporting ad-hoc vocabularies
20
To further verify this idea of simplicity, we have formalised the RO-Crate definition (see
The RO-Crate specification provides a core set of conventions to describe research outputs using types and properties applicable across scientific domains. However we have found that domain-specific use of RO-Crate will, implicitly or explicitly, form a specialised
Making such profiles explicit allow further reliable programmatic consumption and generation of RO-Crates beyond the core types defined in the RO-Crate specification. Following the RO-Crate mantra of
The next version of the RO-Crate specification 1.2 will define a formalization
26
In addition, there are sometimes existing domain-specific metadata formats, but they are either not RDF-based (and thus time-consuming to construct terms for in JSON-LD) or are at a different granularity level that might become overwhelming if represented directly in the RO-Crate Metadata file (e.g. W3C PROV bundle detailing every step execution of a workflow run [68]). RO-Crate allows such
Section 4 examines the observed specializations of RO-Crate use in several domains and their emerging profiles.
The RO-Crate conceptual model has been realised using JSON-LD and Schema.org in a prescriptive form as discussed in Section 2.2. These technical choices were made to cater for simplicity from a developer perspective (as introduced in Section 2.1).
JSON-LD
28
However, JSON-LD alone has too many degrees of freedom and hidden complexities for software developers to reliably produce and consume without specialised expertise or large RDF software frameworks. A large part of the RO-Crate specification is therefore dedicated to describing the acceptable subset of JSON structures.
RO-Crate mandates
29
The avid reader may spot that the RO-Crate Metadata file use the extension
Simplified
31
RO-Crate metadata file showing the flattened compacted JSON-LD
In Recommended properties for types shown in Listing 1 also include
When JSON-LD 1.0 [112] was proposed, one of the motivations was to seamlessly apply an RDF nature on top of regular JSON as frequently used by Web APIs. JSON objects in APIs are frequently nested with objects at multiple levels, and the perhaps most common form of JSON-LD is the compacted form
32
While this feature of JSON-LD can be seen as a way to “hide” its RDF nature, we found that the use of nested trees (e.g. a
By comparison, a single flat
In JSON-LD, the
RO-Crate reuses vocabulary terms and IRIs from Schema.org, but provides its own versioned JSON-LD context,
35
The rationale behind this decision is to support JSON-based RO-Crate applications that are largely unaware of JSON-LD, that still may want to process the
Similarly, while the Schema.org context currently
37
The RO-Crate conceptual model, implementation and best practices are developed by a growing community of researchers, developers and publishers. RO-Crate’s community is a key aspect of its effectiveness in making research artefacts FAIR. Fundamentally, the community provides the overall context of the implementation and model and ensures its interoperability.
The RO-Crate community consists of:
a diverse set of people representing a variety of stakeholders; a set of collective norms; an open platform that facilitates communication (GitHub, Google Docs, monthly teleconferences).
People
The initial concept of RO-Crate was formed at the first Workshop on Research Objects (RO2018
38
An important outcome of discussions that took place at RO2018 was the conclusion that the original Wf4Ever Research Object ontologies [14], in principle sufficient for packaging research artefacts with rich descriptions, were, in practice, considered inaccessible for regular programmers (e.g., Web developers) and in danger of being incomprehensible for domain scientists due to their reliance on Semantic Web technologies and other ontologies.
DataCrate [103] was presented at RO2018 as a promising lightweight alternative approach, and an agreement was made by a group of volunteers to attempt building what was initially called
This group, originally made up of library and Semantic Web experts, has subsequently grown to include domain scientists, developers, publishers and more. This perspective of multiple views led to the specification being used in a variety of domains, from bioinformatics and regulatory submissions to humanities and cultural heritage preservation.
The RO-Crate community is strongly engaged with the European-wide biology/bioinformatics collaborative e-Infrastructure ELIXIR [34], along with European Open Science Cloud
40
A key set of stakeholders are developers: the RO-Crate community has made a point of attracting developers who can implement the specifications but, importantly, keeps “developer user experience” in mind. This means that the specifications are straightforward to implement and thus do not require expertise in technologies that are not widely deployed.
This notion of catering to “developer user experience” is an example of the set of norms that have developed and now define the community.
The RO-Crate community is driven by informal conventions and notions that are prevalent but not neccessarily written down. Here, we distil what we as authors believe are the critical set of norms that have facilitated the development of RO-Crate and contributed to the ability for RO-Crate research packages to be FAIR. This is not to say that there are no other norms within the community nor that everyone in the community holds these uniformly. Instead, what we emphasise is that these norms are helpful and also shaped by community practices.
Simplicity
Developer friendliness
Focus on examples and best practices rather than rigorous specification
Reuse “just enough” Web standards
A core norm of RO-Crate is that of
While the above norms alone could easily lead to the creation of “yet another” JSON format, we keep the goal of
Open platforms
The critical infrastructure that enables the community around RO-Crate is the use of open development platforms. This underpins the importance of open community access to supporting FAIR. Specifically, it is difficult to build and consume FAIR research artefacts without being able to access the specifications, understand how they are developed, know about any potential implementation issues, and discuss usage to evolve best practices.
The development of RO-Crate was driven by capturing documentation of real-life examples and best practices rather than creating a rigorous specification. At the same time, we agreed to be opinionated on the syntactic form to reduce the jungle of implementation choices; we wanted to keep the important aspects of Linked Data to adhere to the FAIR principles while retaining the option of combining and extending the structured metadata using the existing Semantic Web stack, not just build a standalone JSON format.
Further work during 2019 started adapting the DataCrate documentation through a more collaborative and exploratory
In addition to the typical Open Source-style development with GitHub issues and pull requests, the RO-Crate Community have, at time of writing, two regular monthly calls, a Slack channel and a mailing list for coordinating the project; also many of its participants collaborate on RO-Crate at multiple conferences and coding events such as the ELIXIR BioHackathon.
46
Applications and libraries implementing RO-Crate, targeting different types of users across multiple programming languages. Status is indicative as assessed by this work (Alpha < Beta < Release Candidate (RC) < Release)
The work of the community has led to the development of a number of tools for creating and using RO-Crates. Table 1 shows the current set of implementations. Reviewing this list, one can see support for commonly used programming languages, including Python, JavaScript, and Ruby. Additionally, the tools can be integrated into commonly used research environments, in particular, the command line tool
While the development of these tools is promising, our analysis of their maturity status shows that the majority of them are in the Beta stage. This is partly due to the fact that the RO-Crate specification itself only recently reached 1.0 status, in November 2019 [105]. Now that there is a fixed point of reference: With version 1.1 (October 2020) [107] RO-Crate has stabilised based on feedback from application development, and now we are seeing a further increase in the maturity of these tools, along with the creation of new ones.
Given the stage of the specification, these tools have been primarily targeting developers, essentially providing them with the core libraries for working with RO-Crate. Another target has been that of research data managers who need to manage and curate large amounts of data.
RO-Crate fundamentally forms part of an infrastructure to help build FAIR research artefacts. In other words, the key question is whether RO-Crate can be used to share and (re)use research artefacts. Here we look at three research domains where RO-Crate is being applied: Bioinformatics, Regulatory Science and Cultural Heritage. In addition, we note how RO-Crate may have an important role as part of machine-actionable data management plans and institutional repositories.
From these varied uses of RO-Crate we observe natural differences in their detail level and the type of entities described by the RO-Crate. For instance, on submission of an RO-Crate to a workflow repository, it is reasonable to expect the RO-Crate to contain at least one workflow, ideally with a declared licence and workflow language. Specific additional recommendations such as on identifiers is also needed to meet the emerging requirements of FAIR Digital Objects.
49
WorkflowHub.eu
51
We here describe three different RO-Crate profiles developed for use with WorkflowHub.
Being cross-domain, WorkflowHub has to cater for many different workflow systems. Many of these, for instance Nextflow [39] and Snakemake [73], by virtue of their script-like nature, reference multiple neighbouring files typically maintained in a GitHub repository. This calls for a data exchange method that allows keeping related files together. WorkflowHub has tackled this problem by adopting RO-Crate as the packaging mechanism [17], typing and annotating the constituent files of a workflow and – crucially – marking up the workflow language, as many workflow engines use common file extensions like
RO-Crate acts therefore as an interoperability layer between registries, repositories and users in WorkflowHub. The iterative development between WorkflowHub developers and the RO-Crate community heavily informed the creation of the Bioschemas [58] profile for Computational Workflows,
54
RO-Crates in WorkflowHub have so far been focused on workflows that are ready to be run, and development of WorkflowHub is now creating a
This workflow run profile is a continuation of our previous work with capturing workflow provenance in a Research Object in CWLProv [68] and TavernaPROV [110]. In both cases, we used the PROV Ontology [81], including details of every task execution with all the intermediate data, which required significant workflow engine integration.
57
CWLProv and TavernaProv predate RO-Crate, but use RO-Bundle[111], a similar Research Object packaging method with JSON-LD metadata.
Simplifying from the CWLProv approach, the planned Workflow Run RO-Crate profile will use a high level Schema.org provenance
58
WorkflowHub has recently enabled minting of Digital Object Identifiers (DOIs), a PID commonly used for scholarly artefacts, for registered workflows, e.g.
The value of computational workflows, however, is potentially undermined by the “collapse” over time of the software and services they depend upon: for instance, software dependencies can change in a non-backwards-compatible manner, or active maintenance may cease; an external resource, such as a reference index or a database query service, could shift to a different URL or modify its access protocol; or the workflow itself may develop hard-to-find bugs as it is updated. This
For this reason, WorkflowHub is complemented by a monitoring and testing service called LifeMonitor [35], also supported by EOSC-Life. LifeMonitor’s main goal is to assist in the creation, periodic execution and monitoring of workflow tests, enabling the early detection of software collapse in order to minimise its detrimental effects. The communication of metadata related to workflow testing is achieved through the adoption of a
In addition to showcasing RO-Crate’s extensibility, the testing profile is an example of the format’s flexibility and adaptability to the different needs of the research community. Though ultimately related to a computational workflow, in fact, most of the testing-specific entities are more about describing a protocol for interacting with a monitoring service than a set of research outputs and its associated metadata. Indeed, one of LifeMonitor’s main functionalities is monitoring and reporting on test suites running on existing Continuous Integration (CI) services, which is described in terms of service URLs and job identifiers in the testing profile. In principle, in this context, data could disappear altogether, leading to an RO-Crate consisting entirely of contextual entities. Such an RO-Crate acts more as an exchange format for communication between services (WorkflowHub and LifeMonitor) than as an aggregator for research data and metadata, providing a good example of the format’s high versatility.
BioCompute Objects
62
BCOs provide a structured view over a particular workflow, informing regulators about its workings independently of the underlying workflow definition language. However, BCOs have only limited support for additional metadata.
64
IEEE 2791-2020 do permit user extensions in the
As a custom JSON format, BCOs cannot be extended with Linked Data concepts, except by adding an additional top-level JSON object formalised in another JSON Schema. A BCO and workflow submitted by upload to a regulator will also frequently consist of multiple cross-related files. Crucially, there is no way to tell whether a given
We can then consider how a BCO and its referenced artefacts can be packaged and transferred following FAIR principles.
Here the BCO is responsible for describing the
A similar separation of concerns can be found if considering the RO-Crate as a set of files, where the
Specifically, a BCO description alone is insufficient for reliable re-execution of a workflow, which would need a compatible workflow engine depending on the original workflow definition language, so IEEE 2791 recommends using Common Workflow Language (CWL) [36] for interoperable pipeline execution. CWL itself relies on tool packaging in software containers using Docker
67

Separation of Concerns in BCO RO-Crate. BioCompute Object (IEEE2791) is a JSON file that structurally explains the purpose and implementation of a computational workflow, for instance implemented in Common Workflow Language (CWL), that installs the workflow’s software dependencies as Docker containers or BioConda packages. An example execution of the workflow shows the different kinds of result outputs, which may be external, using GitHub LFS [85] to support larger data. RO-Crate gathers all these local and external resources, relating them and giving individual descriptions, for instance permanent DOI identifiers for reused datasets accessed from Zenodo, but also adding external identifiers to attribute authors using ORCID or to identify which licences apply to individual resources. The RO-Crate and its local files are captured in a BagIt whose checksum ensures completeness, combined with Big Data Bag [25] features to “complete” the bag with large external files such as the workflow outputs.
The Pacific And Regional Archive for Digital Sources in Endangered Cultures (PARADISEC
69
The Modern PARADISEC demonstrator
70
The PARADISEC use case takes advantage of several RO-Crate features and principles. Firstly, the transcribed metadata are now independent of the PARADISEC platform and can be archived, preserved and processed in its own right, using Schema.org as base vocabulary and extended with PARADISEC-specific terms.
In this approach, RO-Crate is the holder of itemised metadata, stored in regular files that are organised using Oxford Common File Layout
72
The Endings Project
Machine-actionable Data Management Plans (maDMPs) have been proposed as an improvement to automate FAIR data management tasks in research [88]; maDMPs use PIDs and controlled vocabularies to describe what happens to data over the research life cycle [22]. The Research Data Alliance’s
A mapping has been produced between Research Object Crates and Machine-actionable Data Management Plans [87], implemented by the RO-Crate RDA maDMP Mapper [7]. A similar mapping has been implemented by
Start a skeleton data management plan based on an existing RO-Crate dataset, e.g. an RO-Crate from WorkflowHub.
Instantiate an RO-Crate based on a data management plan.
An important nuance here is that data management plans are (ideally) written in
Institutional data repositories – Harvard Data Commons
The concept of a
the integration of Harvard Research Computing with Harvard Dataverse by leveraging Globus endpoints [27]; this will allow an automatic transfer of large datasets to the repository. In some cases, only the metadata will be transferred while the data stays stored in remote storage;
support for advanced research workflows and providing packaging options for assets such as code and workflows in the Harvard Dataverse repository to enable reproducibility and reuse, and
interation of repositories supported by Harvard, which include DASH,
79
Particularly relevant to this article is the second objective of the Harvard Data Commons, which aims to support the deposit of research artefacts to Harvard Dataverse with sufficient information in the metadata to allow their future reuse (Fig. 4). To support the incorporation of data, code, and other artefacts from various institutional infrastructures, Harvard Data Commons is currently working on RO-Crate adaptation. The RO-Crate metadata provides the necessary structure to make all research artefacts FAIR. The Dataverse software already has extensive support for metadata
80
Even though the Harvard Data Commons is specific to Harvard University, the overall vision and the three objectives can be abstracted and applied to other universities or research organisations. The Commons will be designed and implemented using standards and commonly-used approaches to make it interoperable and reusable by others.

One aspect of Harvard Data Commons. Automatic encapsulation and deposit of artefacts from data management tools used during active research at the Harvard Dataverse repository.
With the increasing digitisation of research processes, there has been a significant call for the wider adoption of interoperable sharing of data and its associated metadata. We refer to [72] for a comprehensive overview and recommendations, in particular for data; notably that review highlights the wide variety of metadata and documentation that the literature prescribes for enabling data reuse. Likewise, we suggest [82] that covers the importance of metadata standards in reproducible computational research.
Here we focus on approaches for bundling research artefacts along with their metadata. This notion of publishing compound objects for scholarly communication has a long history behind it [29,117], but recent approaches have followed three main strands: (1) publishing to centralised repositories; (2) packaging approaches similar to RO-Crate; and (3) bundling the computational workflow around a scientific experiment.
Bundling and packaging digital research artefacts
Early work making the case for publishing compound scholarly communication units [117] led to the development of the Object Re-Use and Exchange model
81
The challenge of describing computational workflows was one of the main motivations for the early proposal of
Considering the FAIR principles [123], we can say with hindsight that the initial RO approaches strongly targeted
The first implementation of Research Objects for sharing workflows in myExperiment [57] was based on RDF ontologies [93], building on Dublin Core, FOAF, SIOC, Creative Commons and OAI-ORE to form myExperiment ontologies for describing social networking, attribution and credit, annotations, aggregation packs, experiments, view statistics, contributions, and workflow components [92].
This initially workflow-centric approach was further formalised as the Wf4Ever Research Object Model [14], which is a general-purpose research artefact description framework. This model is based on existing ontologies (FOAF, Dublin Core Terms, OAI-ORE and AO/OAC precursors to the W3C Web Annotation Model [28]) and adds specializations for workflow models and executions using W3C PROV-O [81]. The Research Object statements are saved in a
We now claim that one barrier for wider adoption of the Wf4Eer Research Object model for general packaging digital research artefacts was exactly this re-use of multiple existing vocabularies (FAIR principle I2:
Several developments for Research Objects improved on this situation, such as ROHub used by Earth Sciences [48], which provides a user-interface for making Research Objects, along with Research Object Bundle [111] (RO Bundle), which is a ZIP-archive embedding data files and a JSON-LD serialization of the manifest with mappings for a limited set of terms. RO Bundle was also used for storing detailed workflow run provenance (TavernaPROV [110]).
RO-Bundle evolved to Research Object BagIt archives,
82
FAIR Digital Objects (FDO) [38] have been proposed as a conceptual framework for making digital resources available in a Digital Objects (DO) architecture which encourages active use of the objects and their metadata. In particular, an FDO has five parts: (i) The FDO
The Digital Object Interface Protocol [47] can be considered an “abstract protocol” of requirements, DOs could be implemented in multiple ways. One suggested implementation is the FAIR Digital Object Framework,
83
By providing a predictable and extensible serialisation of structured metadata.
By formalising how to aggregate digital objects as collections (and adding their context).
By providing a natural Metadata FDO in the form of the RO-Crate Metadata File.
By being based on Linked Data and the Schema.org vocabulary, meaning that PIDs already exist for common types and properties.
At the same time, it is clear that the goal of FDO is broader than that of RO-Crate; namely, FDOs are active objects with distributed operations, and add further constraints such as PIDs for every element. These features improve FAIR features of digital objects and are also useful for RO-Crate, but they also severely restrict the infrastructure that needs to be implemented and maintained in order for FDOs to remain accessible. RO-Crate, on the other hand, is more flexible: it can minimally be used within any file system structure, or ideally exposed through a range of Web-based scenarios. A
The use of computational workflows, typically combining a chain of tools in an analytical pipeline, has gained prominence in particular in the life sciences. Workflows might be used primarily to improve computational scalability, as well as to assist in making computed data results FAIR [55], for instance by improving reproducibility [30], but also because programmatic data usage help propagate their metadata and provenance [69]. At the same time, workflows raise additional FAIR challenges, since they can be considered important research artefacts themselves. This viewpoint poses the problem of capturing and explaining the computational methods of a pipeline in sufficient machine-readable detail [80].
Even when researchers follow current best practices for workflow reproducibility [30,60], the communication of computational outcomes through traditional academic publishing routes effectively adds barriers as authors are forced to rely on a textual manuscript representations. This hinder reproducibility and FAIR use of the knowledge previously captured in the workflow.
As a real-life example, let us look at a metagenomics article [4] that describes a computational pipeline. Here the authors have gone to extraordinary efforts to document the individual tools that have been reused, including their citations, versions, settings, parameters and combinations. The
This attention to reporting detail for computational workflows is unfortunately not yet the norm, and although bioinformatics journals have strong
However detailed this additional information might be, another researcher who wants to reuse a particular computational method may first want to assess if the described tool or workflow is Re-runnable (executable at all), Repeatable (same results for original inputs on same platform), Reproducible (same results for original inputs with different platform or newer tools) and ultimately Reusable (similar results for different input data), Repurposable (reusing parts of the method for making a new method) or Replicable (rewriting the workflow following the method description) [15,54].
Following the textual description alone, researchers would be forced to jump straight to evaluate “Replicable” by rewriting the pipeline from scratch. This can be expensive and error-prone. They would firstly need to install all the software dependencies and download reference datasets. This can be a daunting task, which may have to be repeated multiple times as workflows typically are developed at small scale on desktop computers, scaled up to local clusters, and potentially put into production using cloud instances, each of which will have different requirements for software installations.
In recent years the situation has been greatly improved by software packaging and container technologies like Docker and Conda, these technologies have been increasingly adopted in life sciences [90] thanks to collaborative efforts such as BioConda [61] and BioContainers [37], and support by Linux distributions (e.g. Debian Med [89]). As of November 2021, more than 9,000 software packages are available in BioConda alone,
85
Docker and Conda have been integrated into workflow systems such as Snakemake [73], Galaxy [1] and Nextflow [39], meaning a downloaded workflow definition can now be executed on a “blank” machine (except for the workflow engine) with the underlying analytical tools installed on demand. Even with using containers there is a reproducibility challenge, for instance Docker Hub’s retention policy will expire container images after six months,
87
These container and package systems only capture small amounts of metadata.
88
Docker and Conda can use
From this we see that computational workflows are themselves complex digital objects that need to be recorded not just as files, but in the context of their execution environment, dependencies and analytical purpose in research – as well as other metadata (e.g. version, license, attribution and identifiers).
It is important to note that having all these computational details in order to represent them in an RO-Crate is an ideal scenario – in practice there will always be gaps of knowledge, and exposing all provenance details automatically would require improvements to the data sources, workflow, workflow engine and its dependencies. RO-Crate can be seen as a flexible annotation mechanism for augmenting automatic workflow provenance. Additional metadata can be added manually, e.g. for sensitive clinical data that cannot be publicly exposed
89
FAIR principle A2:
RO-Crate has been established as an approach to packaging digital research artefacts with structured metadata. This approach assists developers and researchers to produce and consume FAIR archives of their research.
RO-Crate is formed by a set of best practice recommendations, developed by an open and broad community. These guidelines show how to use “just enough” standards in a consistent way. The use of structured metadata with a rich base vocabulary can cover general-purpose contextual relations, with a Linked Data foundation that ensures extensibility to domain- and application-specific uses. We can therefore consider an RO-Crate not just as a structured data archive, but as a multimodal scholarly knowledge graph that can help “FAIRify” and combine metadata of existing resources.
The adoption of simple Web technologies in the RO-Crate specification has helped a rapid development of a wide variety of supporting open source tools and libraries. RO-Crate fits into the larger landscape of open scholarly communication and FAIR Digital Object infrastructure, and can be integrated into data repository platforms. RO-Crate can be applied as a data/metadata exchange mechanism, assist in long-term archival preservation of metadata and data, or simply used at a small scale by individual researchers. Thanks to its strong community support, new and improved profiles and tools are being continuously added to the RO-Crate landscape, making it easier for adopters to find examples and support for their own use case.
Strictness vs flexibility
There is always a tradeoff between flexibility and strictness [116] when deciding on semantics of metadata models. Strict requirements make it easier for users and code to consume and populate a model, by reducing choices and having mandated “slots” to fill in. But such rigidity can also restrict richness and applicability of the model, as it in turn enforce the initial assumptions about what can be described.
RO-Crate attempts to strike a balance between these tensions, and provides a common metadata framework that encourages extensions. However, just like the RO-Crate specification can be thought of as a
Future work
The direction of future RO-Crate work is determined by the community around it as a collaborative effort. We currently plan on further outreach, building training material (including a comprehensive entry-level tutorial) and maturing the reference implementation libraries. We will also collect and build examples of RO-Crate
Furthermore, we want to better understand how the community uses RO-Crate in practice and how it contrasts with other related efforts; this will help us to improve our specification and tools. By discovering commonalities in emerging usage (e.g. additional Schema.org types), the community helps to reduce divergence that could otherwise occur with proliferation of further RO-Crate profiles. We plan to gather feedback via user studies, with the Linked Open Data community or as part of EOSC Bring-your-own-Data training events.
We operate in an open community where future and potential users of RO-Crate are actively welcomed to participate and contribute feedback and requirements. In addition, we are targeting a wider audience through extensive outreach activities
90
The main addition in the upcoming 1.2 release of the RO-Crate specifications will be the formalization of profiles
93
Footnotes
Acknowledgements
This work has received funding from the European Commission’s Horizon 2020 research and innovation programme for projects BioExcel-2
99
![]() " href="#fn-104" id="a-320">
104
" href="#fn-104" id="a-320">
104
Contributions
Author contributions to this article and the RO-Crate project according to the Contributor Roles Taxonomy CASRAI CrEDiT
107
![]() ]:
]:
We would also like to acknowledge contributions from:
Formalizing RO-Crate in First Order Logic
Below is a formalization of the concept of RO-Crate as a set of relations using First Order Logic.
RO-Crate Community
As of 2021-10-04, the
