Abstract
BACKGROUND:
FAM170B-AS1 is usually expressed low in all organs except for testicular tissues. No study was performed to explore its role in breast cancer (BC). Contradictory results were reported about hsa-miR-1202 and hsa-miR-146a-5p in BC.
OBJECTIVE:
The present study aimed to explore the involvement of FAM170B-AS1 in BC using bioinformatics predictive tools, followed by a practical validation besides exploring the impact of hsa-miR-1202 and hsa-miR-146a-5p in BC.
METHODS:
This study enrolled 96 female patients with BC, 30 patients with benign breast diseases (BBD), and 25 control subjects. The expressions of circulating FAM170B-AS1, hsa-miR-1202, and hsa-miR-146a-5p were quantified using qRT-PCR. These ncRNAs’ associations, predictive, and diagnostic roles in BC were statistically tested. The underlying miRNA/mRNA targets of FAM170B-AS1 in BC were bioinformatically predicted followed by confirmation based on the GEPIA and TCGA databases.
RESULTS:
The expression of FAM170B-AS1 was upregulated in sera of BC patients and hsa-miR-1202 was upregulated in sera of BBD and BC patients while that of hsa-miR-146a-5p was downregulated in BC. These FAM170B-AS1 was significantly associated with BC when compared to BBD. FAM170B-AS1 and hsa-miR-1202 were statistically associated with the BC’s stage, grade, and LN metastasis. FAM170B-AS1 and hsa-miR-146a-5p gave the highest specificity and sensitivity for BC. KRAS and EGFR were predicted to be targeted by FAM170B-AS1 through interaction with hsa-miR-143-3p and hsa-miR-7-5p, respectively. Based on the TCGA database, cancer patients having mutations in FAM170B show good overall survival.
CONCLUSIONS:
The present study reported that for the first time, FAM170B-AS1 may be a potential risk factor, predictive, and diagnostic marker for BC. In addition, FAM170B-AS1 might be involved in BC by interacting with hsa-miR-143-3p/KRAS and hsa-miR-7-5p/EGFR through enhancement or repression that may present a new therapeutic option for BC.
Abbreviations
Introduction
Being the most common cancer globally among women makes breast cancer (BC) the first cause of cancer-related deaths [1]. About 2.2 million new cases are diagnosed with BC. Therefore, BC accounts for 11.7% of all cancer incidence [2]. In 2019, about 59% of women diagnosed with BC were
The presence of a variety of genetic aberrations against BC makes it a very heterogeneous disease [7]. Hence, BC has five main molecular subtypes identified depending on the level of expression of receptors for estrogen (ER), progesterone (PR), and human epidermal growth factor receptor-2 (HER-2) on the surface of BC cells [7, 8]. Some of these subtypes are marked by significantly aggressive clinical course, proliferative activity and growth rate, poor prognosis, early metastasis, different targeted therapies, and risk of disease recurrence [9, 10, 11, 12, 13]. Hence, early screening and diagnosing the more aggressive subtypes of BC and identifying suspicious breast tumors are vital to reduce morbidity and mortality [14, 15].
Long noncoding RNAs (lncRNAs) and microRNAs (miRNAs) are reported to be associated with different types of cancer [16, 17]. LncRNAs can suppress mRNA expression by recruiting histone modification complexes, controlling the alternative splicing, and nuclear import [18]. LncRNAs can interact with miRNAs [19, 20, 21] to de-repress gene expression by competing with miRNAs for interaction with shared target mRNAs. miRNA binds to its complementary sequences in the 3’-UTR of their target mRNAs and post-transcriptionally represses their translation into protein [22]. Therefore, predicting and validating the lncRNA/miRNA crosstalk can modulate gene expression patterns that drive physiological and pathological processes [23, 24, 25].
Little is known about the Family with sequence similarity 170 member B-antisense RNA1 (FAM170B-AS1; ENSG00000234736). It is located on chromosome 10: 49,121,839-49,151,547 forward strand with four transcripts (
hsa-miRNA-1202 and hsa-miRNA-146a were reported to be associated with cancer progression, positively or negatively. hsa-miRNA-1202 is transcriped by MIR1202 gene on chromosome 6: 155,946,797-155,946,879 forward strand. It is reported to be upregulated in breast cancer [34] endometrial cancer [35], and lung cancer [36] while it is revealed to be downregulated in glioma cells [37] and HCC [38]. hsa-miR-1202 was reported to inhibit the proliferation of glioma cells and induce endoplasmic reticulum stress and apoptosis via regulating Rab1A in glioma [37]. Another study reported the inhibitory role of hsa-miR-1202 on HCC cell migration and invasion by targeting cyclin-dependent kinase 14 [38].
hsa-miR-146a is transcribed by the MIR146A gene on chromosome 5: 160, 485, 352-160, 485, 450 forward strand. Wu et al. [39] indicated that hsa-miR-146a-5p is downregulated in non-small cell lung cancer (NSCLC) tissues, whereas Wang et al. [40] reported an up-regulation of hsa-miR-146a-5p in NSCLC patients. Liu et al. [41] revealed the oncogenic mechanism of hsa-miR-146a-5p in NSCLC cell survival and migration by suppressing its target gene, tumor necrosis factor receptor-associated factor 6. In BC, several studies indicated that hsa-miR-146a can function as oncomiR [42, 43, 44, 45]. Other studies have demonstrated hsa-miR-146a as a tumor suppressor miRNA [46, 47]. The reports of Park et al. [48] and Kumaraswamy et al. [49] illustrated the tumor-suppressive effect of hsa-miR-146a via inhibiting epithelial-to-mesenchymal transition (EMT) and the metastatic potential in many cancer types, including BC. It has been proposed that hsa-miR-146a can sustain an epithelial phenotype in triple negative breast cancer (TNBC) via direct targeting of the TGF
To the best of our knowledge, the current study aimed to predict and validate the role of FAM170B-AS1 for the first time in breast cancer using the bioinformatics tool and QPCR, respectively, to construct FAM170B-AS1/miRNA/mRNA network. Another goal of the present study is to explore the roles of both hsa-miR-1202 and hsa-miR-146a-5p in BC.
Subjects and methods
Subjects
A total of 151 subjects were enrolled in the current study and divided into three groups: a control group (
Patients were chosen from those admitted to the National Cancer Institute, Cairo University, and Clinical Oncology Department, Faculty of Medicine, Ain Shams University, Cairo, Egypt. Full history and the pathological examinations of the breast mass from patients either with BBD or BC was obtained from their clinical sheets. The stages of BC, performed according to the tumor-node-metastasis classification system (TNM), were obtained from the clinical sheets. Furthermore, the expression of hormonal receptors (PR, ER, and HER-2) on the breast cancer tissue was examined using immunohistochemistry to identify the molecular subtype of BC.
The study was approved by the Scientific Research Ethics Committee, National Research Center, Dokki, Giza, Egypt (ID: 15209). Informed consent for using the samples for research purposes was obtained from all participants before beginning the study. The research was carried out under the Declaration of Helsinki, printed in the British Medical Journal (18 July 1964).
Blood sample collection
Three milliliters blood samples were withdrawn from individuals in tubes with polymer gel and clot activator (Greiner bio-one, GmbH, Australia). The blood was left to clot at 37∘C for 30 minutes, and all samples were centrifuged at 10,000
Tumor markers assay
The serum venous levels of carcinoembryonic antigen (CEA) and carbohydrate antigen 15.3 (CA 15.3) were estimated using commercial enzyme-linked immunosorbent assay kits provided by Immunospeccorporation, Netherlands.
Prediction of FAM170B-AS1 association with breast cancer
Predicting the association of FAM170B-AS1 in different diseases was performed using miRWalk v.2 (
Quantitative RT-PCR for hsa-miR-146a-5p and hsa-miR-1202 assays and FAM170B-AS1 validation
MiRNA was extracted from the sera of all the participants using an RNeasy mini kit (Qiagen, USA). The purity of the extracted RNA was detected at 260/280 nm using a nano-drop spectrophotometer (Quawell, Q-500, Scribner, USA). The extracted miRNA was reverse transcribed into cDNA using a MiScript II reverse transcription kit provided by Qiagen, USA. cDNA synthesis was performed using a thermal cycler (SureCycler 8800, Agilent, USA).
Real-time quantitative PCR was performed to analyse the expression of hsa-miR-1202 (HS_hsa-miR-1202 miScript Primer Assay, Cat no.MP00000259), hsa-miR-146-5p (HS_hsa-miR-146-5p miScript Primer Assay, Cat no.MS00003535), and FAM170B-AS1 (RT2 lncRNA qPCR Assay for Human, cat no. LPH11337A) using SYBR Green/ROX Master Mix (Qiagen, USA). Ready-made primers for the studied ncRNAs as well as their housekeeping genes [RNU6 for miRNA (HS_RNU6-2-11_ miScript Primer Assay, Cat no. MS00033740) and GAPDH for lncRNA (Hs_GPDH_1_SG QuantiTect Primer Assay, Cat no. QT00079247)] were provided from Qiagen, USA. The thermal reaction conditions were 95∘C for 15 min followed by 40 cycles at 94∘C for 15 s, 55∘C for 30 s, and 70∘C for 34 s using Max3005P QPCR system, Stratagene, Agilent biotechnology, USA. Expression for investigated miRNAs and lncRNA was assessed using the
Targeted prediction and functional analysis
LncBase predicted v2 database (
To further consider the potential functions, pathways, and networks of these target genes, the DAVID database (
In addition, the human protein atlas (HPA) (
Correlation to GEPIA and TCGA data
Gene expression profiling integrative analysis (GEPIA) was used to investigate the expression level of the selected target genes in normal breast tissue and BRCA tumors. Furthermore, GEPIA was utilized to analyse the correlation between the FAM170B-AS1 gene and the target genes in the BRCA tumor. In addition, the genetic alteration of FAM170B in primary cancers was investigated based on TCGA with their overall survival analysis.
Statistical analysis
The power of the current study was calculated utilizing G∗Power 3.1.9.7 software (Franz Faul, Universität Düsseldorf, Germany) (
The Statistical Package for Social Science version 26 for Windows (SPSS software package, Chicago, USA) was used to perform all statistical analyses. The data distribution was statistically determined using the Kolmogorov-Smirnov test with Lilliefors significance correction. The categorical variables were expressed as frequencies (percentages), and non-parametric data were expressed as median (interquartile range). The non-parametric variables were statistically analysed using the Kruskal-Wallis one-way ANOVA (all pairwise) test for multiple comparisons.
Binary logistic regression analyses were used to investigate the strength of the association of the expression of FAM170B-AS1, hsa-miR-1202, and hsa-miR-146a-5p with the susceptibility to breast tumors between either benign breast lesion versus control group, malignant tumor versus control group, and malignant tumor versus benign breast lesion. The strength of associations was measured by the odds ratio (OR) with a 95% confidence interval (CI) adjusted for age and menopause. Multiple linear regression analysis was used to assess the association of the independent variables relative to the dependent variables after transforming non-parametric variables into the logistic scale. The regression models were constructed using the “enter” analysis.
The receiver operating characteristic curve (ROC) was performed using SPSS vs. 26 to detect the area under the curve (AUC), cutoff points, sensitivity, and specificity for the tested miRNA-1202, miRNA-146a-5P, and FAM10B-AS1 between BC, BBD, and control groups, compared to the traditional tumor markers. All
Results
The clinical characteristics of all individuals in the studied groups are listed in Table 1. Concerning patients with BBD, 36.7% were identified to have follicular hyperplasia, 26.7% had fibrocytic changes, and 36.7% had intraductal papillomatosis. Of women with BC, 38.5, 34.4, 35.4, and 47.9% were diagnosed to be at the early stage with low-grade, ductal carcinoma in situ (DCI) and no lymph node (LN) metastasis, respectively,
Demographic characteristics of subjects in the different studied groups
Demographic characteristics of subjects in the different studied groups
Data are expressed as Median (INTER QUARTILE RANGE) for non-parametric variables and frequencies (percentages) for categorical variables. BBD: benign breast diseases, BC: breast cancer, DCI: ductal carcinoma in situ, IDC: invasive ductal carcinoma, ER: estrogen receptor, PR: progesterone receptor, HER-2: human epidermal growth factor receptor-2, LN: lymph node. The mean difference is significant at
Comparison of the levels of tumor markers as well as FAM170B-AS1, hsa-miR-1202, and hsa-miR-146a-5p expression in the different studied groups
Data are expressed as median (interquartile range). BBD: benign breast diseases, BC: breast cancer, CEA: carcinoembryonic antigen, CA 15.3: carbohydrate antigen 15.3. a: significance versus control group, b: significance versus benign breast diseases group. ∗: fold change versus control group, # : fold change versus benign breast diseases group. Significance had been adjusted by Bonferroni correction for multiple tests. The mean difference is significant at
Relationship between CEA and CA15.3 levels as well as ncRNAs expression in serum with clinicopathological parameters of patients with breast benign diseases
Data are expressed as a median. CEA: carcinoembryonic antigen, CA15.3: carbohydrate antigen 15.3. Significance had been adjusted by Bonferroni correction for multiple tests.
Relationship between CEA and CA15.3 levels in serum with clinicopathological parameters of patients with breast cancer
Data are expressed as a median. DCI: ductal carcinoma in situ, IDC: invasive ductal carcinoma, ER: estrogen receptor-positive, PR: progesterone receptor positive, HER-2: human epidermal growth factor receptor-2 positive, LN: lymph node. Significance had been adjusted by Bonferroni correction for multiple tests.
Relationship between FAM170B-AS1, hsa-miR-1202, and hsa-miR-146a-5p expression with clinicopathological parameters of patients with breast cancer
Data are expressed as a median. DCI: ductal carcinoma in situ, IDC: invasive ductal carcinoma, ER: estrogen receptor-positive, PR: progesterone receptor positive, HER-2: human epidermal growth factor receptor-2 positive, LN: lymph node. Significance had been adjusted by Bonferroni correction for multiple tests.
Association of FAM170B-AS1, hsa-miR-1202, and hsa-miR-146a-5p expression with breast cancer risk, compared with the traditional tumor markers
BC: breast cancer, BBD: benign breast diseases, OR: odds ratio, 95% CI: 95% confidence interval. # : Adjusted for age and menopause.
Multiple linear regression analysis for the association between FAM170B-AS1 and the studied miRNAs and the association of the tumor markers as well as the studied genes with the stage, grade, and LN metastasis
Data represented in Supplementary file Table S1 predicts the association of FAM170B-AS1 with different types of cancer in Homo Sapiens species. Using the lncRNA-disease association prediction (LDAP) method, FAM170B-AS1 was anticipated to be associated with breast cancer with a 0.1828 score.
Table 2 shows the expression levels of FAM170B-AS1, hsa-miR-1202, and hsa-miR-146a-5p in the sera of all the studied groups, compared to the serum levels of CEA and CA15.3 (traditional tumor markers). There were significant elevations in the serum level of CEA in both BBD patients (1.78-fold change,
Table 3 shows no significant difference (
Both the serum levels of CEA and CA15.3 were found to be similar (
Data in Table 5 illustrate the difference between the expression of the studied ncRNAs among BC patients with their age, menopause status, stage, grade, molecular subtypes, and LN metastasis. No significant difference was recorded in the expression of hsa-miR-1202 in BC patients with
The associations of the tumor markers and the studied ncRNAs with BBD and BC are illustrated in Table 6 using binary logistic regression analysis. The serum level of CA15.3 and the expression level of hsa-miR-146a-5p were not significantly associated with BBD (
Multiple linear regression analysis (Table 7) reported a significantly positive correlation between FAM170B-AS1 expression and the expression of hsa-miR-1202
Table 8 and Fig. 1 demonstrate the results of the ROC curve of the traditional tumor markers and the studied ncRNAs when comparing BC patients with non-BC patients (Control group
The ROC curve plot of CEA, CA15.3, FAM170B-AS1, hsa-miR-1202, and hsa-miR-146a-5P in BC group, compared to non-BC individuals (control group
AUC: area under the curve, CEA: carcinoembryonic antigen, CA 15.3: carbohydrate antigen 15.3.
The receiver operating characteristic curve of all tested markers in BC group compared to the groups of control 
A: Venn diagram identifies the overlapped miRNAs targeted by FAM170B-AS1 and associated with breast cancer identified. B: FAM170B-AS1-overlapped miRNA-mRNA network in breast cancer and constructed by Cytoscape.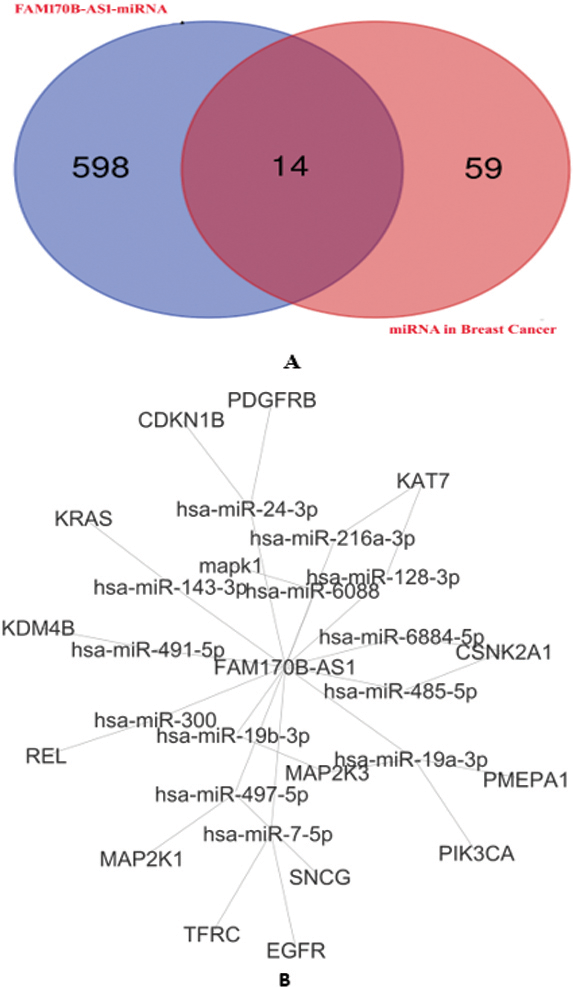
Correlation between the gene expression of FAM170B-AS1 with MAPK1 (A), EGFR (B), MAP2K1 (C), and KRAS (D) in BRCA tumor based on the GEPIA database.
The PPI network of the target genes.
Protein expression of the targeted genes in both normal breast and breast cancer tissues based on the HPA database. MAPK1 was moderately stained in glandular and myoepithelial cells of the normal breast (A-1) and was strongly stained in breast duct carcinoma (A-2). EGFR was moderately stained in glandular and myoepithelial cells of the normal breast (B-1) and was negatively stained in breast duct carcinoma (B-2). MAP2K1 was weakly stained in glandular cells of the normal breast (C-1) and was moderately stained in breast duct carcinoma (C-2). KRAS was negatively stained in glandular and myoepithelial cells of the normal breast (D-1) and was moderately stained in breast duct carcinoma (D-2).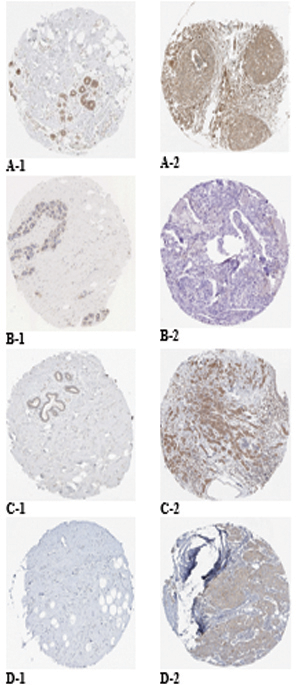
Overall survival of FAM170B genetic alterations in different types of primary cancer extracted from TCGA.
Supplementary file Table S2 illustrates the prediction of the interaction between FAM170B-AS1 and 710 miRNAs with the scores of predictions and binding besides the binding sites of miRNAs on FAM170B-AS1 lncRNA. Among these miRNAs, hsa-miR-1202 and hsa-miR-146a-5p were predicted to interact with FAM170B-AS1 with prediction scores of 0.611 and 0.539, respectively. The binding scores of hsa-miR-1202 and hsa-miR-146a-5p with FAM170B-AS1 were 0.018 and 0.007, respectively. One of the miRNAs predicted to interact with FAM170B-AS1 is hsa-miR-203a-3p with a prediction score of 0.555 and a binding score of 0.007. This miRNA is predicted by TargetScan 7.2 to interact with FAM170B gene (ENST00000311787.5) with 8mer which needs further practical validation. FAM170B is located on chromosome 10: 49, 131, 154-49, 134, 021 reverse strand. This gene codes for a protein with 283 amino acids.
In the current study, four miRNA target prediction algorithms were used to predict the potential target genes of miRNAs in breast cancer (Supplementary file Table S3). Several miRNA interactions with genes (259 interactions) were anticipated. The binding scores of miRNAs with genes were
The 16 genes were underwent for GO and pathways analyses. The GO analysis (Supplementary file Table S4) indicated that the target genes were involved in complex cellular pathways (BP), such as signal transduction, positive regulation of transcription, and cell proliferation. In addition, the target genes were located (CC) in the cytosol, nucleus, and plasma membrane. Furthermore, The GO molecular function of the target genes showed their involvement in numerous MFs, such as protein binding, ATP binding, protein kinase binding, and protein tyrosine kinase activity. The KEGG pathway analysis (Supplementary file Table S5) revealed that PD-L1 expression and PD-1 checkpoint pathway in cancer, PI3K-Akt signaling pathway, Ras signaling pathway, Rap1 signaling pathway, EGFR tyrosine kinase inhibitor resistance, ErbB signaling pathway, Estrogen signaling pathway, and MAPK signaling pathway are associated with the regulation of the 14 overlapped miRNAs.
In addition, the genes associated with PD-L1 expression and PD-1 checkpoint pathway in cancer, PI3K-Akt signaling pathway, Ras signaling pathway, Rap1 signaling pathway, EGFR tyrosine kinase inhibitor resistance, ErbB signaling pathway, Estrogen signaling pathway, and MAPK signaling pathway were selected based on the KEGG pathway analysis. Four genes (MAPK1, EGFR, KRAS, and MAP2K1) overlapped based on the Venn diagrams. Next, the current study investigated the preliminary expression level of the four genes based on GEPIA. Thus, MAPK1 and KRAS were upregulated, whereas MAP2K1 and EGFR were downregulated in BRCA tumors compared to normal BRCA tissues. Furthermore, positive correlations were found between the expression of FAM170B-AS1 and the genes expression of MAPK1 (
The String database was applied to construct the PPI network, and 66 PPI pairs with a combined score of
Moreover, based on the HPA database, strong staining in breast duct carcinoma was observed for MAPK1 (Fig. 5A-2), whereas moderate staining was observed for MAP2K1 (Fig. 5C-2) and KRAS (Fig. 5D-2) and negatively staining was observed for EGFR (Fig. 5B-2). Negative staining in normal breast tissues was observed for KRAS (Fig. 5D-1), while moderate staining was observed for MAPK1 (Fig. 5A-1) and EGFR (Fig. 5B-1), and weak staining was observed for MAP2K1 (Fig. 5C-1). Based on these results, EGFR and KRAS were selected. Using bioinformatics tool, it has been hypothesized that FAM170B-AS1 may dysregulate the expression of KRAS and EGFR genes in the PD-L1 expression and PD-1 checkpoint pathway in cancer, PI3K-Akt signaling pathway, Ras signaling pathway, Rap1 signaling pathway, EGFR tyrosine kinase inhibitor resistance, ErbB signaling pathway, Estrogen signaling pathway, and MAPK signaling pathway to participate in the different biological processes of BC.
Data collected from TCGA report 28 cases with ten types of primary cancers with somatic mutations in the FAM170B gene with different severity (six patients with the hematopoietic and reticuloendothelial system, five individuals with colon, four cases with bronchus and lung, three subjects with skin, two patients with breast, two individuals with other and ill-defined sites, two samples with the pancreas, and one sample for each of kidney, rectum, small intestine, and uterus). The types of mutations are listed in the Supplementary file Table S6. The high severity of mutations in FAM170B was observed in ductal & lobular neoplasm of breast cancer via deletion causing frameshift mutation and in ductal & lobular neoplasm of pancreatic cancer via substitution causing stop gained mutation. TCGA demonstrated that cancer patients with mutations in FAM170B show improved overall survival (Fig. 6).
Since the most diagnosed cancer in women is BC, more efficient diagnostic and therapeutic methods for BC are required. In the last decades, discovering specific and sensitive molecular biomarkers with statistically significant association with BC has attracted important attention for prediction, prognosis, and diagnosis [53, 54]. Among molecular biomarkers, the lncRNAs and miRNAs circulating in the blood are of great importance because of their stability [47, 48]. lncRNAs are proven to be associated with numerous types of cancer via many pathways, including cell proliferation, invasion, and metastasis [57, 58, 59]. lncRNAs can control these pathways through lncRNAs/miRNAs/mRNA interaction networks either by acting as oncogenic or tumor suppressor lncRNAs directly or indirectly [60, 61, 62]. Therefore, bioinformatic prediction of lncRNAs followed by experimental validation becomes one of the most important research fields in cancer [63].
Discovering novel biomarkers that are more specific and sensitive to BC diagnosis is still mandatory because of the limited sensitivity and specificity of CEA and CA15.3 for BC, especially at the early stage [64, 65]. The recent study by Jintao et al. [66] reported 85.71% sensitivity, 63.49% specificity of CEA, 64.29% sensitivity, and 80.95% specificity of CA15.3 for BC. In addition, measuring the levels of CEA and CA15.3 in sera of BC patients could not significantly predict the stage, grade, and LN metastasis of BC [67]. As far as we know, the current study bioinformatically anticipated and practically validated the involvement of FAM170B-AS1 in breast tumors for the first time. The upregulation of FAM170B-AS1 gene expression in the serum of BC patients and its significant associations with BC could make FAM170B-AS1 a potential predictive marker for BC among BBD patients. In addition, the present study revealed the probable diagnostic role of FAM170B-AS1 in BC by giving higher specificity (100%) and sensitivity (97%) than that of CEA and CA15.3. Multiple linear regression analysis performed in the present study reports that serum FAM170B-AS1 expression was incorporated as an independent predictor of stage, grade, and LN metastasis of BC, while CEA and CA15.3 did not. Previous studies demonstrated that increased CEA and CA15.3 levels in the serum of BC patients at diagnosis are associated only with higher stage, tumor size, and positive axillary lymph nodes [68, 69, 70, 71, 72, 73]. All of these make serum FAM170B-AS1 a promising biomarker for BC.
MiRNAs function as regulators for multiple genes by binding to their 3’-UTRs and, therefore, up or down-regulate their expression [74]. Concerning hsa-miR-1202, various reports suggest its valuable role in predicting the prognosis of cancers [35, 75]. Chen et al. found that silenced hsa-miR-1202 in endometrial cancer could enhance the apoptotic level of tumor cells, arrest tumor cells in the G1 phase, and decline the migratory and invasive abilities of the tumor cells [76]. Wang et al. illustrated the association of the upregulated hsa-miR-1202 with lymph nodes in papillary thyroid carcinoma [75]. Quan et al. demonstrated that hsa-miR-1202 inhibits tumor cell proliferation by targeting Rab1A in glioma cells [37]. Du et al. indicated that hsa-miR-1202 is down-regulated in HCC, which was associated with metastasis, clinicopathologic features, and worse prognosis of HCC via direct binding to the CDK14 gene to exert its anti-tumor effects [38]. These reports suggested that hsa-miR-1202 may play an oncogene or tumor suppressor role in cancers by controlling cell proliferation, apoptosis, migration, and invasion.
The current study documented an upregulation in hsa-miR-1202 gene in both BBD and BC patients. This upregulation varied among BC patients with stage, grade, pathological types, molecular subtypes, and LN metastasis of BC. Also, hsa-miR-1202 showed significant associations with both BBD and BC, compared to the control group, making hsa-miR-1202 a probable risk factor for both BBD and BC. The multiple linear regression analysis performed in the present study reveals that the upregulation of hsa-miR-1202 in sera is incorporated as an independent predictor of stage, grade, and LN metastasis of BC, compared to the traditional tumor markers (CEA and CA15.3). Measuring the expression level of hsa-miR-1202 in sera gave high specificity but low sensitivity towards BC. It has been previously shown that hsa-miR-1202 is an oncomiR in BC [34]. The study of Hamam et al. [34] showed an upregulation of hsa-miR-1202 in BC patients with stages I, II, and III, compared to stage IV, with potential utilization as a tumor biomarker for early detection. Furthermore, its expression was slightly higher in the HER2 and TN compared to patients with luminal subtypes.
Regarding hsa-miR-146a role in BC, a controversial role has been illustrated. Particularly, hsa-miR-146a has been revealed to downregulate BRCA1 directly [77], where another study showed a binding between BRCA1 and the promoter of MIR146A, which activates the transcription of MIR146A leading to attenuation of EGFR expression in a hsa-miR-146a-dependent manner [49]. Interestingly, a negative association between hsa-miR-146a and EGFR, NOTCH2, and SOX2 genes was practically reported and data extracted from TCGA dataset confirmed the reported negative association of hsa-miR-146a with EGFR gene which is upregulated in TNBC and its protein product enhances tumorigenesis and immune escape via glycolysis stimulation [78] and controls TNBC aggressiveness and stemness through the management of E2/E
In the present study, hsa-miR-146a-5p was downregulated in BC patients only. Their downregulation varied among BC patients with menopause status, stage, grade, pathological types, molecular subtypes, and LN metastasis of BC. Also, hsa-miR-146a-5p showed significant association with BC, compared to the control and BBD groups, making hsa-miR-146a-5p a probable risk factor for BC. The multiple linear regression analysis performed in the present study reveals that the down regulation of hsa-miR-146a-5p in sera is not incorporated as an independent predictor of stage, grade, and LN metastasis of BC. Measuring the expression level of hsa-miR-146a-5p in sera gave high specificity and sensitivity towards BC. Iacona and Lutz explained the repression in hsa-miR-146a is mainly due to hypermethylation of promoter, methylation, and deacetylation of the histone [80]. Recently, repression of hsa-miR-146a in TNBC cells was reported, which could drive EGFR abundance and signaling, leading to tumor progression and poor disease outcome [81].
One of the functions of lncRNAs is their counteract with miRNAs, making miRNA sponges [82, 83]. Multiple linear regression analyses carried out in the present research display significant correlations of the expressions of FAM170B-AS1 with both hsa-miR-1202 and hsa-miR-146a-5p. This statistical evidence confirmed the bioinformatic prediction of FAM170B-AS1 interaction with both miRNAs using LncBase Predicted v2 database. Also, the current study revealed that FAM170B-AS1 and hsa-miR-146a-5p, other than hsa-miR-1202, may be considered excellent diagnostic markers because of their higher specificity, sensitivity, and AUC.
Cross-talks between LncRNAs/miRNAs form complex regulatory networks of post-transcriptional gene regulation. Relying on the specific lncRNA/miRNA interaction, the lncRNA/miRNA axis can have tumor suppressor or oncogenic effects [84]. The present study used the bioinformatic analysis tool to construct FAM170B-AS1/miRNA interaction network in BC for the first time, followed by identifying their target mRNAs. According to GO analysis, the target genes are involved in complex cellular pathways, such as signal transduction, transcription, and cell proliferation. The KEGG pathway analysis revealed that the PD-L1 expression and PD-1 checkpoint pathway in cancer, PI3K-Akt signaling pathway, Ras signaling pathway, Rap1 signaling pathway, EGFR tyrosine kinase inhibitor resistance, ErbB signaling pathway, Estrogen signaling pathway, and MAPK signaling pathway are potentially correlated with the regulation of the 14 predicted miRNAs. As reported, these pathways are associated with the escape of tumor cells from immune surveillance [85], resistance to endocrine therapy [86], proliferation, apoptosis, differentiation, and survival [86, 87, 88, 89].
The detailed roles for FAM170B-AS1 in BC need more explanation. The current research work combined FAM170B-AS1 analysis with TCGA and GEPIA databases to explore the potential biological functions of FAM170B-AS1 in BC. Data collected from the TCGA database display that having mutations in FAM170B showed improved OS of patients with different types of cancer, including breast cancer. The present study also bioinformatically predicted the genes from the KEGG signaling pathways and the hub genes from PPI and hypothesized that FAM170B-AS1 may affect both KRAS expression through interaction with hsa-miR-143-3p and EGFR expression by interaction with hsa-miR-7-5p to contribute to different biological processes of BC.
KRAS is involved in the tumorigenesis and progression of breast cancers via activating AKT/MEK/ERK pathways signaling [90]. In addition, overactivation of the KRAS pathway occurs in TNBCs, leading to chemoresistance [91]. Studies targeting the KRAS gene instead of the KRAS protein have attracted increasing attention in recent decades. miRNAs targeting the UTR of KRAS were reported to reduce the level of KRAS and thus inhibit the progression of TNBCs [92]. It was reported that hsa-miR-143-3p is downregulated in TNBC patients, which is associated with EMT, prognosis, and resistance to cancer therapy [93, 94]. This therapeutic resistance could be due to cytokine-induced apoptosis inhibitor-1 protein activation, which may repress hsa-miR-143-3p expression [95]. Hence, FAM170B-AS1 may affect KRAS expression through hsa-miR-143-3p according to BC subtype, which needs further investigation.
The epidermal growth factor receptor (EGFR), a member of the erbB receptor family, controls proliferation, differentiation, and development in several human tissues, including the breast [96]. The upregulation of EGFR is observed in several solid tumors causing cancer progression, poor prognosis, and resistance to chemotherapy & radiation therapy [97]. The expression of EGFR mRNA varies among different BC subtypes, either by upregulation or downregulation. Consequently, understanding the way by which EGFR is dysregulated attracted great attention [98]. hsa-miR-7 has a tumor suppressor effect in breast cancers via suppressing proliferation and enhancing apoptosis [99, 100, 101] as well as inhibiting invasion & metastasis and mediating cytotoxic T-lymphocyte-mediated lysis of breast cancer cells [102, 103]. It was reported that hsa-miR-7 suppresses EGFR mRNA in lung, breast, and glioblastoma by binding to sites in its 3’-UTR [104]. Thus, FAM170B-AS1 may affect EGFR expression through hsa-miR-7-5p according to BC subtype, which needs further investigation.
Conclusions and future perspectives
The current study made a spot for the first time on the involvement of FAM170B-AS1 in BC. The study experimentally validated the bioinformatic prediction of FAM170B-AS1 association with BC. The associations of FAM170B-AS1, hsa-miR-1202, and hsa-miR-146a-5p had been reported in BC, making them probable risk factors for BC. The statistically significant correlations of FAM170B-AS1 with hsa-miR-1202 and hsa-miR-146a-5p verified the bioinformatic prediction of their interactions. FAM170B-AS1 and hsa-miR-1202 might be considered predictors of BC stage, grade, and LN metastasis. In addition, FAM170B-AS1 and hsa-miR-146a-5p might be appraised as excellent diagnostic markers for BC. The current study could bioinformatically predict FAM170B-AS1/hsa-miR-143-3p/KRAS and FAM170B-AS1/hsa-miR-7-5p/EGFR network in BC. This network needs further practical study in breast cancer cells to be confirmed. The mechanism by which FAM170B-AS1 is upregulated in breast tumors needs to be elucidated by studying the FAM170B-AS1 promoter region and histone. The present work opened a new era to design a therapeutic approach against BC by targeting FAM170B-AS1 that might lead to novel therapeutic strategies. In addition, The link between FAM170B-AS1 and FAM170B in BC needs to be practically explored.
Declaration
The authors declare that there is no conflict of interest.
Funding
Not applicable.
Availability of data and materials
The datasets used and/or analysed during the current study are available from the corresponding author upon reasonable request.
Authors contribution
Conception: Hala Mostafa Ghanem, Menha Swellam, and AlShaimaa Mohamed Taha.
Interpretation or analysis of data: AlShaimaa Mohamed Taha and Ahmed Saeed Abd ELhafeez.
Preparation of the manuscript: AlShaimaa Mohamed Taha and Ahmed Saeed Abd ELhafeez.
Revision for important intellectual content: Hala Mostafa Ghanem, Menha Swellam, and AlShaimaa Mohamed Taha.
Supervision: Hala Mostafa Ghanem, Menha Swellam, and AlShaimaa Mohamed Taha.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-230396.
Footnotes
Acknowledgments
The instruments listed in the current study were purchased through a grant from Science The Science, Technology & Innovation Funding Authority (STDF) through Capacity Building Grant Fund (CBG) [No. 4940]. This work was partially supported by a grant from STDF through Basic and Applied Research Support Grant Project (BARG) [No.15089], Egypt.
