Abstract
BACKGROUND:
STEAP3 is a metal reductase located on the plasma membrane close to the nucleus and vesicles. Despite numerous studies indicating the involvement of STEAP3 in tumor advancement, the prognostic value of STEAP3 in glioma and the related mechanisms have not been fully investigated.
METHODS:
Initially, we examined the correlation between STEAP3 expression and the survival rate in various glioma datasets. To assess the prognostic capability of STEAP3 for one-year, three-year, and five-year survival, we created receiver operating characteristic (ROC) curves and nomograms. Additionally, an investigation was carried out to examine the mechanisms that contribute to the involvement of STEAP3 in gliomas, including immune and enrichment analysis. To confirm the expression of STEAP3 in LGG and GBM, tumor tissue samples were gathered, and cell experiments were conducted to explore the impacts of STEAP3. The function of STEAP3 in the tumor immune microenvironment was assessed using the M2 macrophage infiltration assay.
RESULTS:
We found that STEAP3 expressed differently in group with different age, tumor grade IDH and 1p19q status. The analysis of survival illustrated that glioma patients with high level of STEAP3 experienced shorter survival durations, especially for IDH-mutant astrocytoma. Cox analysis demonstrated that STEAP3 had potential to act as an independent prognostic factor for glioma. The predictive value of STEAP3 for glioma prognosis was demonstrated by ROC curves and nomogram. Immune analysis showed that STEAP3 may lead to a suppressive immune microenvironment through the control of immunosuppressive cell infiltration and Cancer-Immunity Cycle. Combining enrichment analysis and cell experiments, we discovered that STEAP3 can promote glioma progression through regulation of PI3K-AKT pathway and M2 macrophage infiltration.
CONCLUSION:
STEAP3 plays significant roles in the advancement of glioma by regulating immune microenvironment and PI3K-AKT pathway. It has the potential to serve as a therapy target for glioma.
Introduction
In adults, glioma is a prevalent form of malignant brain tumor [1]. Gliomas are associated with poor prognosis, which poses a huge burden on families and society, especially for glioblastoma. The occurrence rate of glioblastoma rises with age, peaking in individuals between the ages of 75 and 84 [2]. A retrospective study conducted in China at a single center involved 1,285 patients diagnosed with glioma, in which 254 were identified as glioblastoma. The one-year and five-year survival rates were 61% and 9% respectively, basically consistent with the observations made in other countries [3].
The fifth edition of the WHO Classification of central nervous system (CNS) tumors made significant changes in the classification of gliomas by incorporating the used of molecular diagnostics in the classification system. As per the 2021 classification by WHO, adult-type diffuse gliomas are categorized into just 3 types: Astrocytoma, IDH-mutant (encompassing grade 2 to grade 4; comprising diffuse astrocytoma, anaplastic astrocytoma, and glioblastoma with IDH mutation as per the 2016 WHO classification); Oligodendroglioma, IDH-mutant and 1p/19q-codeleted; and Glioblastoma, IDH-wildtype [4]. Grading no longer depends on tumor histology entirely, as molecular events, such as EGFR gene amplification and TERT promoter mutation, can be used in the diagnosis of Glioblastoma without histological evidence [4, 5, 6]. Identifying novel therapeutic targets and formulating innovative treatment strategies are imperative to enhance the prognosis of individuals with glioma.
Six-transmembrane epithelial antigen of prostate (STEAP) are cell surface proteins consisting of four members, STEAP1-4. STEAP3 is a metal reductase that is involved in the physiological absorption and reduction of iron and copper [7, 8, 9], and has demonstrated significant involvement in various forms of cancer. STEAP3 can enhance sensitivity of renal cell carcinomas to ferroptosis through the regulation of p53/xCT pathway [10]. Additionally, it can stimulate the advancement of hepatocellular carcinoma by modulating the RAC1-ERK-STAT3 signaling [11]. It was reported that STEAP3 promoter includes a p53-binding site and can increase p53 expression to promote cell death in breast cancer [12]. In our prior investigation, we proved that elevated STEAP3 levels in gliomas were linked to unfavorable prognosis and developed a risk model based on the expression of the STEAP family [13]. During the study, we investigated the involvement of the STEAP protein family in glioma but we were unable to identify the specific contributions of individual members in STEAP family. Since previous studies on prognostic analysis of gliomas were based on the old WHO classification system, we examined the prognostic significance of STEAP3 in gliomas using the updated WHO classification system We also investigated the role of STEAP3 in the recruitment of M2 macrophages, which had not been studied previously.
Material and methods
Data source
To examine the expression of STEAP3 in various normal tissues and single cell types, the HPA (human protein atlas,
Tissue samples
The Ethics Committee approved the acquisition of tumor tissues from 10 glioma patients at The Second Hospital of Hebei Medical University. All samples were surgically resected gliomas and classified by senior physicians in pathology department of our hospital according to WHO standards (WHO II: 3 samples, WHO III: 3 samples, WHO IV: 4 samples). The inclusion criteria of glioma samples include: (1) Adult; (2) First diagnosis; (3) No tumor stroke; (4) No cerebral hernia; (5) No preoperative radiotherapy chemotherapy and other treatments.
Prognostic analysis
The Kaplan-Meier analysis was conducted using the R package Survminer. Univariate and multivariate Cox regression analysis were employed to assess whether STEAP3 served as a standalone prognostic factor for overall survival (OS). Using the SurvivorROC package, we drew the ROC curve to investigate the prognostic significance of STEAP3 in glioma [13]. In order to predict the OS in gliomas, nomograms were established using various prognostic factors including clinical features and STEAP3 expression in TCGA. We then confirmed whether the actual and predicted OS were consistent over 1, 3 and 5 years using calibration curves.
Immune microenvironment estimation
To determine the relative levels of infiltration of 22 immune cell types, the CIBERSORT algorithm was employed [17]. Next, we examined the correlation between STEAP3 expression and 22 different types of immune cells in TCGA dataset. The relative levels of the 22 immune cell types were ranked according to their scores.
In the year 2013, the Cancer-Immunity Cycle concept was introduced by Chen and Mellman [18]. and could be divided into 7 steps: (1) release of tumor cell antigen; (2) presentation of tumor antigen; (3) priming and activation; (4) recruitment of T cells into tumor; (5) infiltration of T cells into tumor; (6) recognition of tumor cells by T cells; (7) killing of tumor cells [19]. In this way, the immune system can efficiently eliminate cancer cells by means of the Cancer-Immunity Cycle. Cancerous cells elude the immune system by obliterating multiple stages within the Cancer-Immunity Cycle. In contrast, tumor immunotherapy enhances the immune system-mediated killing of tumor cells by amplifying or activating the Cancer-Immunity Cycle. Based on this point, we estimated the associations between expression of STEAP3 and the 7 steps of Cancer-Immunity Cycle using TCGA data.
Prediction of immunotherapy response
Genes involved in immune checkpoint, like CTLA-4 and PD-1, have a crucial function in the evasion of tumors from the immune system [20]. As a result, we acquired the immunophenotypic score (IPS) in TCGA from The Cancer Immunome Atlas (https//tcia.at TCIA) in order to anticipate the reaction to immune checkpoint inhibitors (ICI) [21]. A higher IPS score corresponds to increased immunogenicity and improved response to ICI. Furthermore, the TIDE [21] and ImmuCellAI [22] platforms were employed to forecast the efficacy of immunotherapy.
Enrichment analysis
Based on the median expression of STEAP3, the TCGA cohort of glioma patients was categorized into two groups: high-STEAP3 and low-STEAP3 patients. In TCGA, the identification of differentially expressed genes (DEGs) between high-STEAP3 and low-STEAP3 patients was performed using the R package called ’limma’, based on adj
Cell culture and transfection
Cell lines derived from glioma (U251, U87, A172, and LN229) and human astrocytes (HA) were acquired from the Cell bank of Chinese Academy of Sciences. The cells were cultivated in DMEM (Biological, Salem, MA, US) containing 10% FBS (Gibco, 10099141), and were kept at a temperature of 37
RT-qPCR
We extracted Total RNA from HA and 4 different glioma cell lines by utilizing the Superbrilliant test (Zhongshi, Tianjin, China). The Takara kit (Bori Medical, Beijing, China) was utilized for performing reverse transcription from mRNA to cDNA, whereas SYBR qPCR Mix kit (Vazyme, Nanjing, China) was employed for carrying out RT-qPCR. The RT-qPCR procedure included an initial pre-denaturation phase at 95
Immunohistochemical analysis
Paraffin-embedded slices underwent deparaffinization in xylene, then went through a sequence of graded alcohol solutions for rehydration. To retrieve the antigen, a water bath at 95
Western blot
Protein extraction reagent (K1015, APExBIO) was utilized to extract cellular proteins. The BCA method was utilized to ascertain the concentration of protein. Next, we utilized denaturing polyacrylamide gel electrophoresis to isolate protein samples, and subsequently moved the protein bands from the gel onto polyvinylidene fluoride membranes. The membranes were immersed in a solution to block them for 15 minutes, the solution was then removed and the membranes were left to incubate in the primary antibody overnight at a temperature of 4 degrees Celsius. Abcam and Themofisher provided the primary antibodies, which were diluted in the following ratios: anti-STEAP3 (1:1,500, ab151566), anti-AKT (1:1,500, ab38449), anti-phospho-AKT (1:1,500, ab38449), anti-mTOR (1:1,200, ab134903), anti-phospho-mTOR (1:1,000, ab109268), anti-P70S6K (1:1,200, A300-510A), anti-phospho-P70S6K (1:1,000, MA5-15202), and anti-GAPDH (1:10,000, ab8245). Following three washes, the membranes were subsequently exposed to goat anti-rabbit secondary antibody (1:10,000, ab216773, Abcam) for a duration of 2 hours in a light-free environment. To identify membranes, infrared imaging scanning devices (Odyssey LI-COR, USA) were employed.
CCK8 assay
U251 cells in logarithmic growth stage were planted into 96-well plates with a volume of 100
Colony formation assay
Tumor cells in the exponential growth stage were gathered and placed in the six-well dish with a density of 1000 cells per well, followed by a two-week incubation period. Afterwards, the cell cultures were treated with paraformaldehyde (4%) and then stained using crystal violet (0.1%). The numbers of colonies were analyzed using the Image J (version1.52 2p).
Transwell experiment
To assess cell invasion ability, the experiment was split into two groups: 1. a group using a coated matrix gel; 2. uncoated matrix gel to detect cell migration ability. U251 cells were placed in the upper chamber of Transwell plates at a density of 1
Wound healing experiment
After being placed in a six-well plate, the U251 cells were cultured for a duration of 24 hours. Next, a pipette tip was utilized to make a cut on the cells, forming a narrow opening. Photographs of the marks were captured using a microscope equipped with a camera (Olympus, Japan) at the identical location after 0, 24, and 48 hours. The assessment of the ability to heal was done by measuring the width of the scratches at various time intervals using Image J.
M2 macrophage infiltration assay
Expression of STEAP3 in tumor and normal tissues. (a) Analysis of STEAP3 expression in normal tissues. (b) Expression analysis of STEAP3 in different single cell types. (c) Analysis of STEAP3 expression in pan-cancer.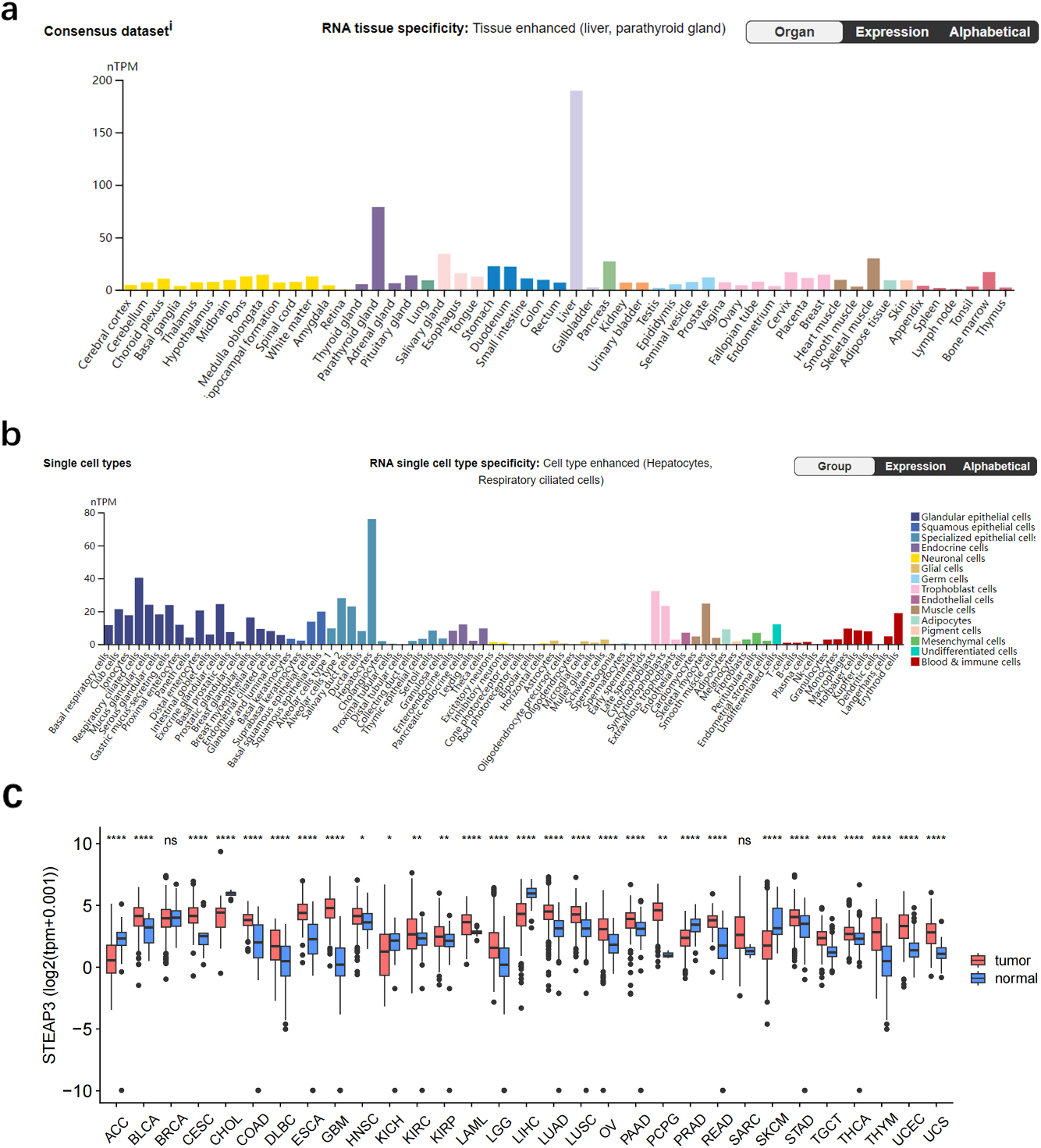
To induce differentiation into macrophages, THP-1 cells were subjected to treatment with 100 ng/ml PMA (Beyotime; shanghai, China) for a duration of 24 hours, in preparation for polarization of M2 macrophages. PCR was used to verify the expression of M0 markers, CD68. Polarization of the macrophages was induced by treating the cells with 20 ng/ml IL-4 (Nearshore proteins; Suzhou; China) and 20 ng/ml IL-13 (Nearshore proteins; Suzhou; China) for 48 hours in the presence of PMA. PCR was employed for the detection of the presence of M2 macrophage markers, namely CD163 and CD206.
Single cell sequencing analysis. (a) Cells from four glioma tissues were divided into seven cell types. (b) STEAP3 mainly expressed in immune cells and neoplastic cells.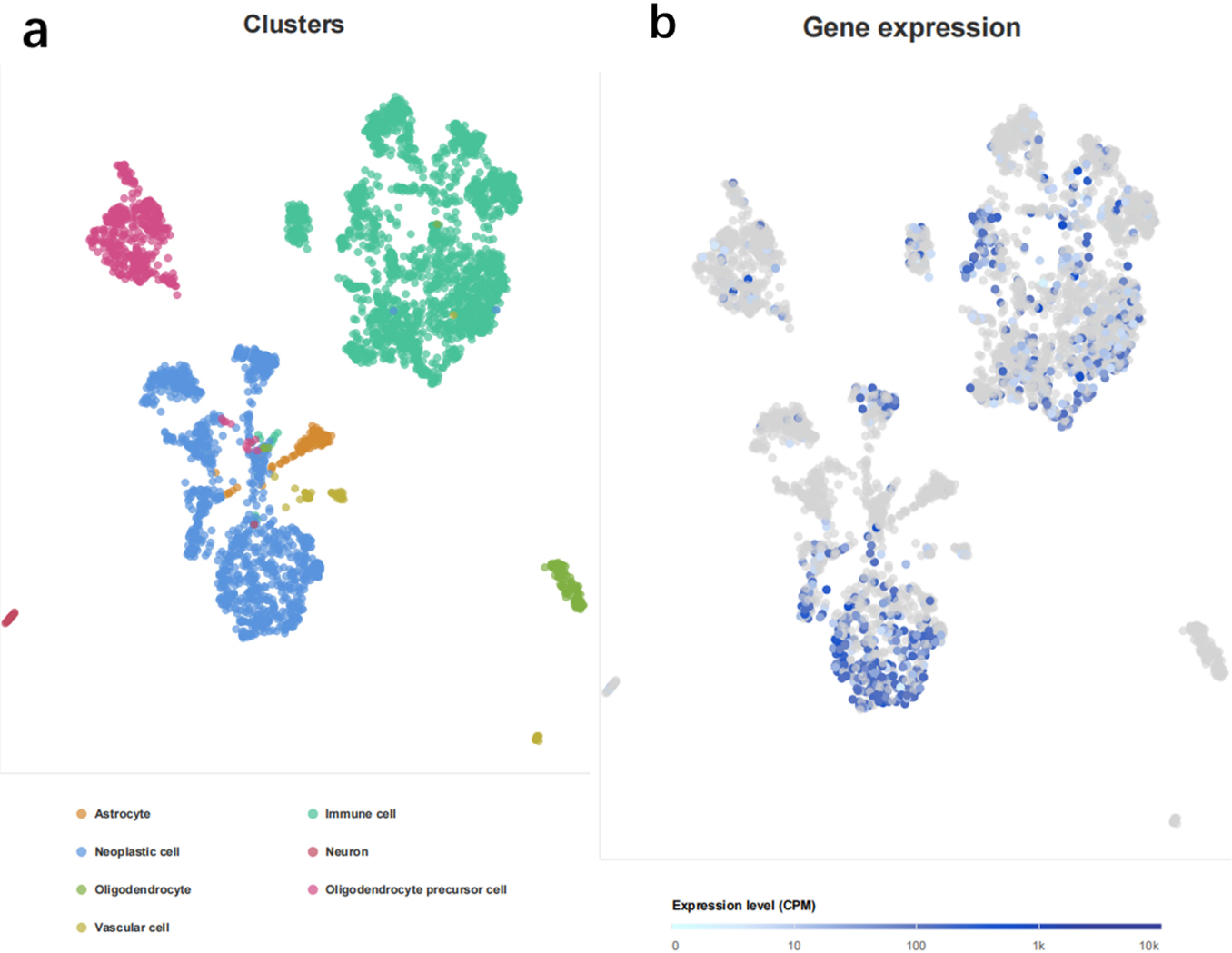
Infiltration of M2 macrophages was analyzed by seeding 5.0
Cell experiments in this study were repeated triplicate. Statistical analysis was performed through R software (version 4.2.3) or Prism 8 (GraphPad Inc, USA). Data are expressed as mean
Increased STEAP3 expression predicted poor prognosis for glioma. (a–i) Survival analysis in nine glioma populations. (j) The total HR value of STEAP3 was gathered using meta-analysis.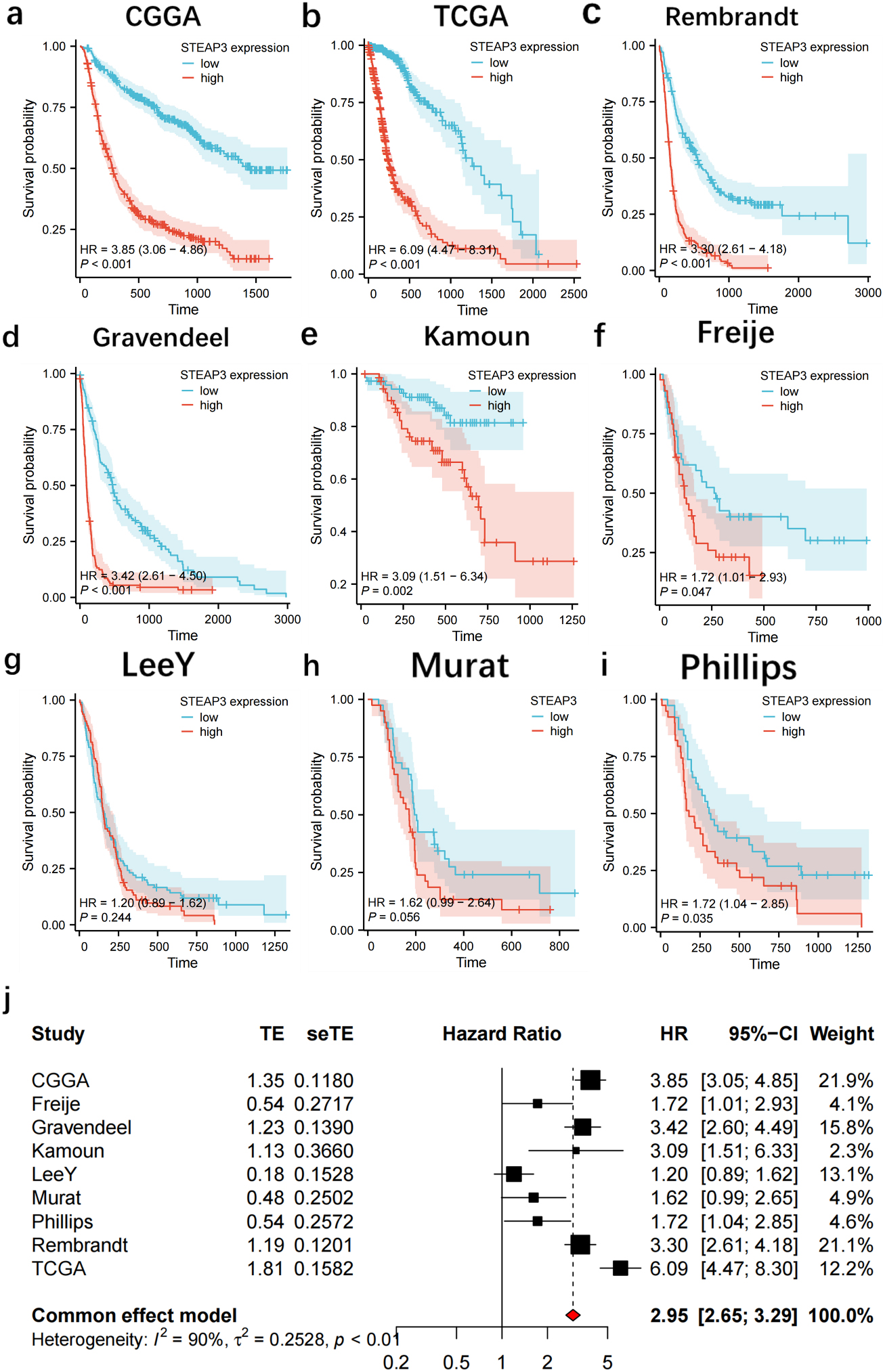
STEAP3 expression in normal and tumor tissues
STEAP3 expression was found to be elevated in the parathyroid gland, liver, skeletal muscle, and pancreas among various normal tissues (Fig. 1a) In different single cell types, STEAP3 expressed highly in glandular and specialized epithelial cells, while expressed lowly in neuronal cells (Fig. 1b). Pan-cancer analysis in TCGA showed that STEAP3 expression was up-regulated in UCS, UCEC, COAD, KIRP, OV, THYM READ, STAD, GBM, KIRC, LUAD, PAAD, HNSC, ESCA LAML, TGCT, LUSC, THCA, LGG, DLBC CESC and BLCA while down-regulated in ACC, CHOL, KICH, LIHC PRAD SKCM (Fig. 1c) Sample size of each dataset in pan-cancer analysis is as shown in Supplemental Table. Analysis of single-cell sequencing demonstrated that cells from four glioma tissues were classified into seven cell types (Fig. 2a), with STEAP3 expression being relatively high in immune cells and tumor cells (Fig. 2b).
Relationship between STEAP3 expression and clinical characteristics. (a) WHO grades, (b–c) IDH status and 1p19q codeletion, (d)age, (e) primary therapy outcome, (f) histological categories.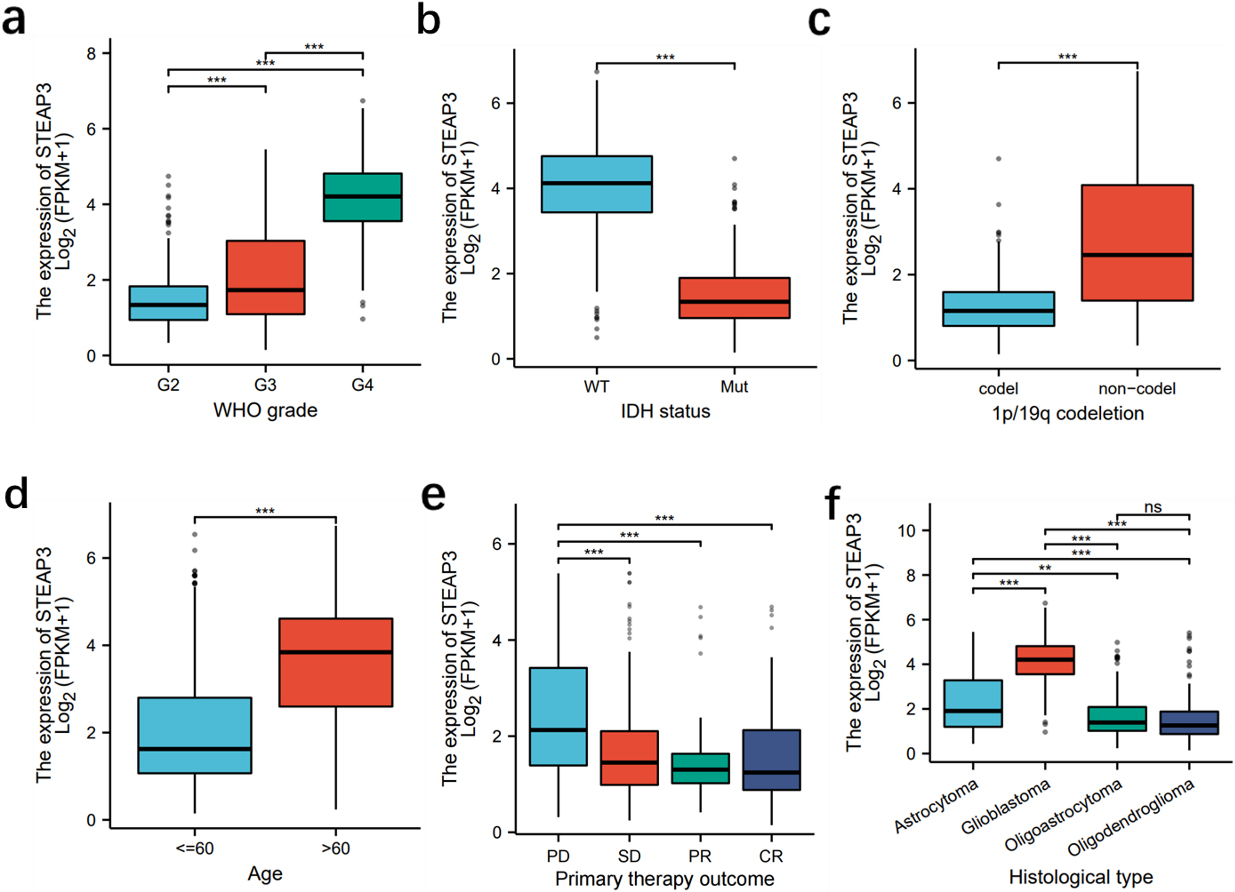
Survival analysis showed that patients in the high-STEAP3 group from CGGA (HR:3.85
Cox analysis and ROC curves. (a–c) Univariate and multivariate Cox analysis in CGGA, TCGA and Gravendeel. (d–f) ROC curves of STEAP3 in CGGA, TCGA and Gravendeel from 1 to 7 years.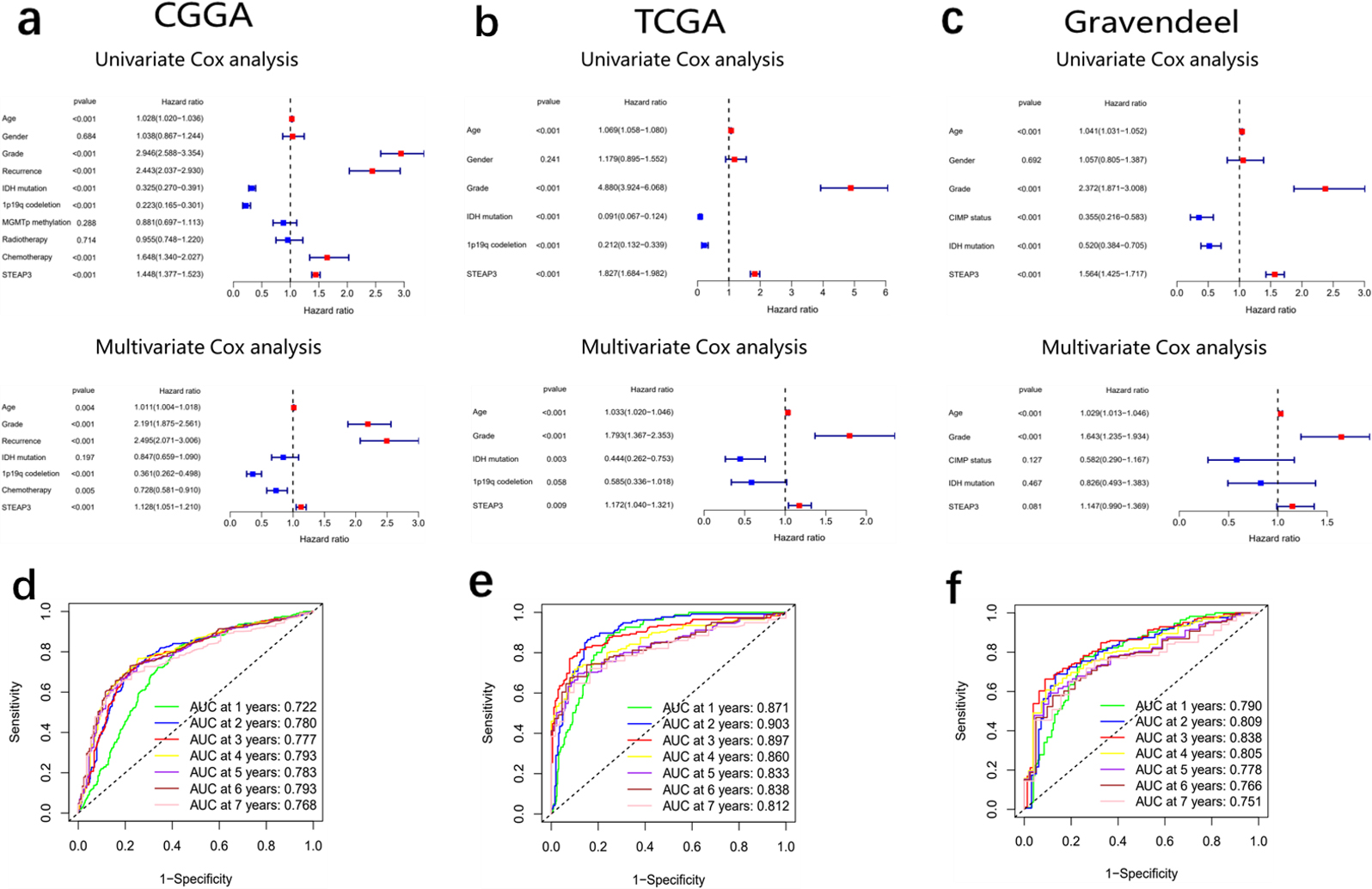
The proposal of 2021 WHO Classification of Tumors of the CNS improved our comprehension of glioma. To address the limitations in previous studies on STEAP3, we performed survival analysis in different glioma subtypes based on the new WHO Classification.
The results of survival analysis were showed as follows: (1) high-STEAP3 patients had poorer prognosis than low-STEAP3 patients in astrocytoma IDH-mutant (TCGA HR:167,
Examining the correlation between STEAP3 and clinical characteristics through analysis of the TCGA database, notable discrepancies in STEAP3 expression were observed among groups with distinct WHO grades, IDH status, 1P19q status, age, primary therapy outcome, and histological types (Fig. 4a–f).
Cox analysis in CGGA, TCGA and Gravendeel cohorts indicated that STEAP3 may serve as a standalone prognostic indicator in glioma (Fig. 5a–c). ROC curves demonstrated that STEAP3 had good ability in predicting glioma prognosis from 1 to 7 years (Fig. 5d–f).
We then developed a personalized nomogram that combined STEAP3 and other clinical indicators to quantitatively assess OS in glioma patients (Supplemental Figure 4a). We found the calibration curves of the nomogram were consistent with the standard curves in 1-, 3- and 5-year OS (Supplemental Figure 4b). We then compared STEAP3 with other clinical indicators using multivariate ROC curves and found the good predictive value of STEAP3 (Supplemental Figure 4c).
Immune cell infiltration analysis
The analysis of immune cell infiltration results showed that the STEAP3 expression exhibited a positive correlation with the scores of immunosuppressive cells, including Tregs, neutrophils, and M2 macrophages (Tregs, neutrophils and M2 macrophages) (Fig. 6a). In addition, our findings indicate a positive association between STEAP3 and the markers of TAM (CD68, IL10, and CCL2), M2 macrophage (MS4A4A, VSIG4 and CD163), neutrophil (ITGAM CEACAM8 and CCR7), as well as Treg (FOXP3, TGFB1, and CCR8) (Fig. 6b). These results suggested that STEAP3 might induce immunosuppression by enhancing the infiltration of immunosuppressive cells.
Roles of STEAP3 in immune microenvironment. (a) Heatmap showed the relationship between expression of STEAP3 and various immune cell types. (b) Correlation between expression of STEAP3 and cell markers of immune inhibitory cells.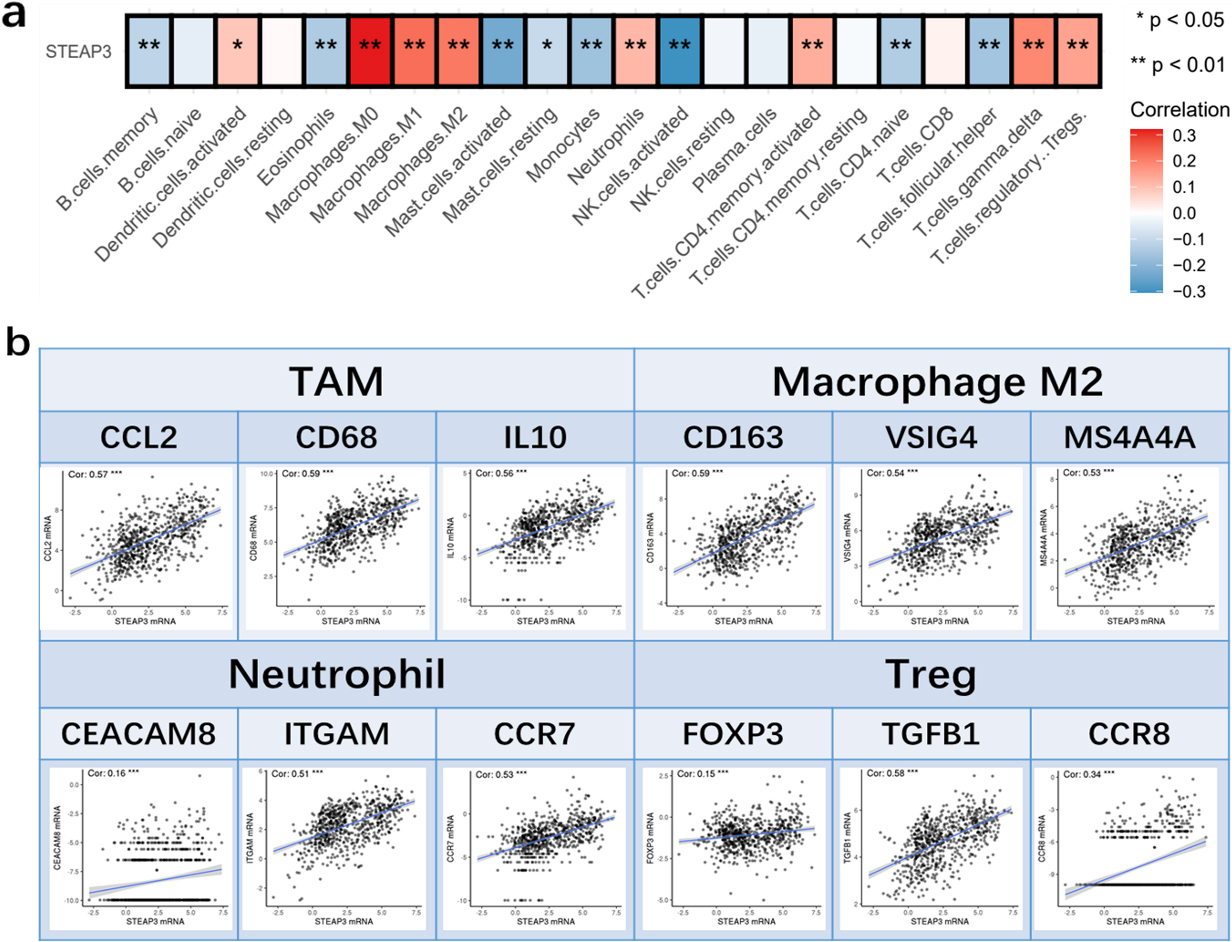
In our study, increased expression of multiple immune checkpoint genes was observed in the high-STEAP3 group (Fig. 7a). We then investigated the relationship between STEAP3 and Cancer-Immunity Cycle Our findings revealed a positive association between STEAP3 and the scores of Step1 and Step4, while indicating a negative correlation with the scores of Step7, Step5, Step3, and Step2 of the Cancer-Immunity Cycle (Fig. 7b–c). The findings suggest that STEAP3 may play a role in suppressing the immune system by controlling various stages of the Cancer-Immunity Cycle. Furthermore, it was observed that patients exhibiting elevated STEAP3 expression had significantly elevated TIDE scores (Fig. 7d), whereas the low-STEAP3 group demonstrated higher IPS scores in comparison to the high-STEAP3 group (Fig. 7e). The results indicated that individuals belonging to the low-STEAP3 group exhibited a more favorable reaction to immune checkpoint inhibitors (ICI). A previous study demonstrated that CD73 was a specific indicator for predicting the response of GBM to immunotherapy, and that CD73 knockdown enhanced the anti-tumor effect of ICI in a mouse model [23]. Our research revealed that the high-STEAP3 group exhibited elevated levels of CD73 expression (Fig. 7f), potentially accounting for their unfavorable reaction to ICI treatment. According to the results from TIDE and ImmuCellAI, the prediction of immunotherapy response indicated that the low-STEAP3 group had a higher response rate (54% in TIDE and 28% in ImmuCellAI) compared to the high-STEAP3 group (42% in TIDE and 7% in ImmuCellAI) (Fig. 7g–h).
Prediction of immunotherapy response in TCGA. (a) Relationship between immune checkpoint gene expression and STEAP3 expression. (b-c) Roles of STEAP3 in Cancer-Immunity Cycle. (d) The association between TIDE scores and STEAP3 expression. (e) The IPS in high- and low-STEAP3 groups. (f) CD73 expression in high- and low-STEAP3 groups. (g) TIDE online platform was used to predict response to immunotherapy. (h) ImmuCellAI online platform was used to predict response to immunotherapy.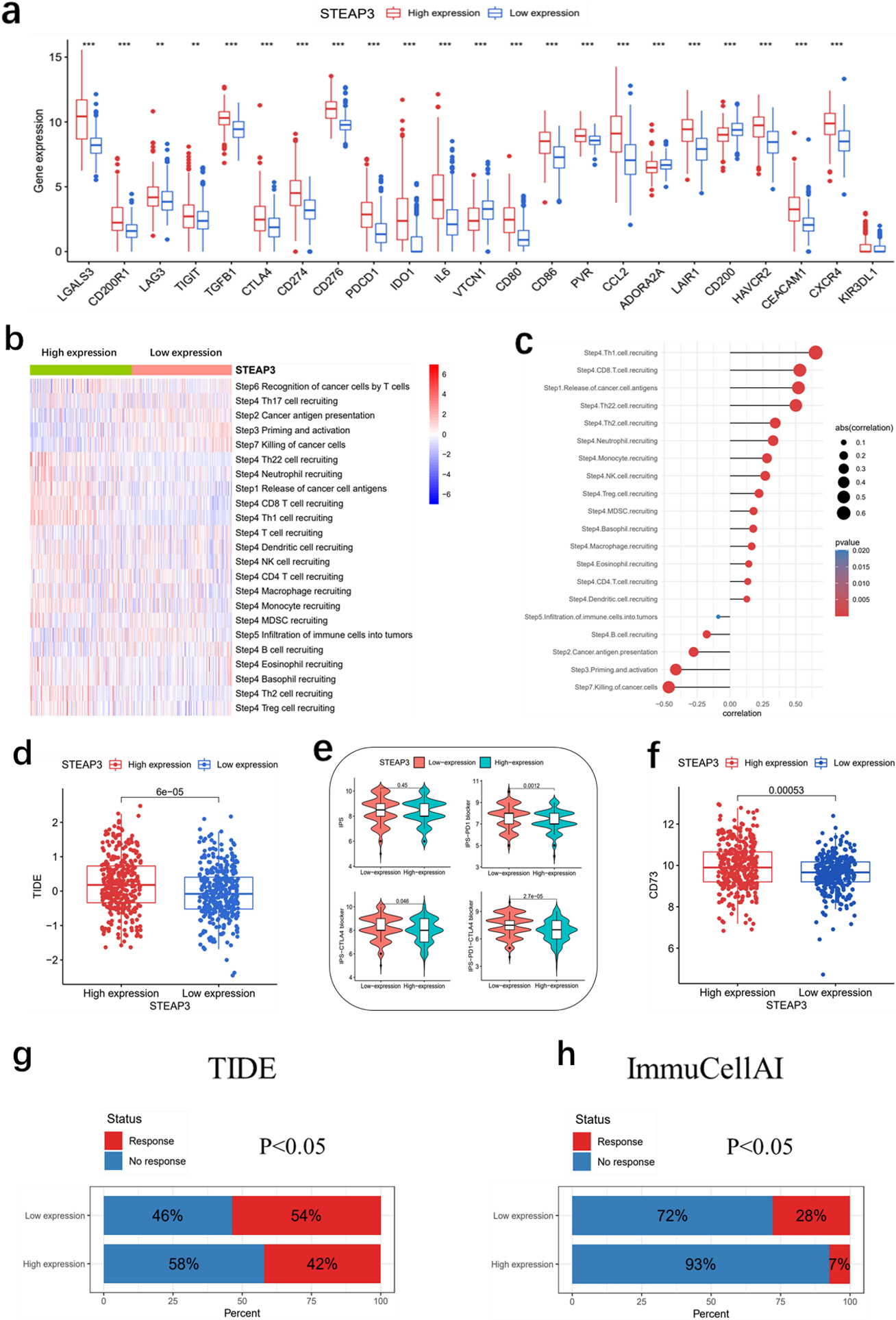
To determine the potential roles of STEAP3 in gliomas, we explored the DEGs between low-STEAP3 and high-STEAP3 patients in TCGA. We identified 1,436 DEGs, consisting of 450 downregulated genes and 986 upregulated genes (Fig. 8a). GO and KEGG enrichment analysis showed that the DEGs were enriched in neutrophil migration, JAK-STAT signaling pathway, PI3K-Akt signaling pathway, TNF signaling pathway, antigen processing and presentation and enzyme inhibitor activity (Fig. 8b–c). GSEA analysis illustrated that STEAP3 was involved in hypoxia, apoptosis and PI3K-Akt signaling pathway (Fig. 8d–e).
Enrichment analysis. (a) The DEGs between the high- and low-STEAP3 groups. (b) GO enrichment analysis. (c) KEGG enrichment analysis. (d–e) GSEA analysis. 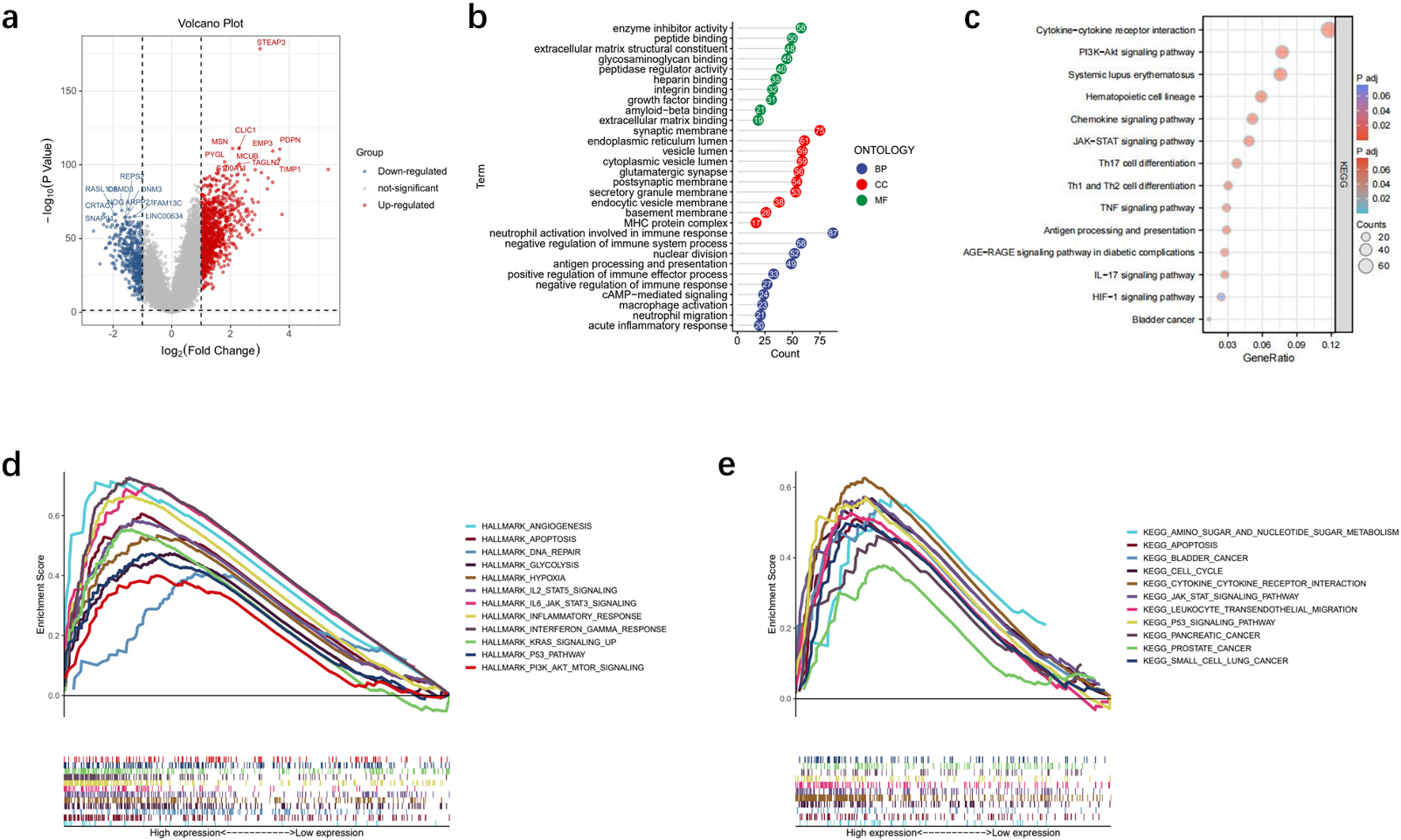
STEAP3 expression was higher in U251, U87, A172 and LN229 cell lines compared to HA cell line (Fig. 9a, c). In GBM, the immunohistochemical analysis revealed a greater expression of STEAP3 compared to LGG (Fig. 9b).
Expression Level of STEAP3 in cell lines and glioma tissues. Knockdown of STEAP3 significantly reduced GBM cell progression. (a) RT-qPCR showed the levels of STEAP3 in HA and glioma cell lines. (b) Immunohistochemical images showed the expression of STEAP3 in LGG and GBM. (c) Western blot showed the levels of STEAP3 in HA and glioma cell lines. (d) STEAP3 knockdown in U251 cells. (e-f) CCK-8 and colony formation assays demonstrated that the U251 cells proliferation was significantly reduced after knocking down STEAP3. (g) Transwell assay showed that knockdown of STEAP3 can inhibit invasion and migration of U251 cells. (h) Wound healing illustrated that the relative wound width was reduced after knocking down STEAP3. 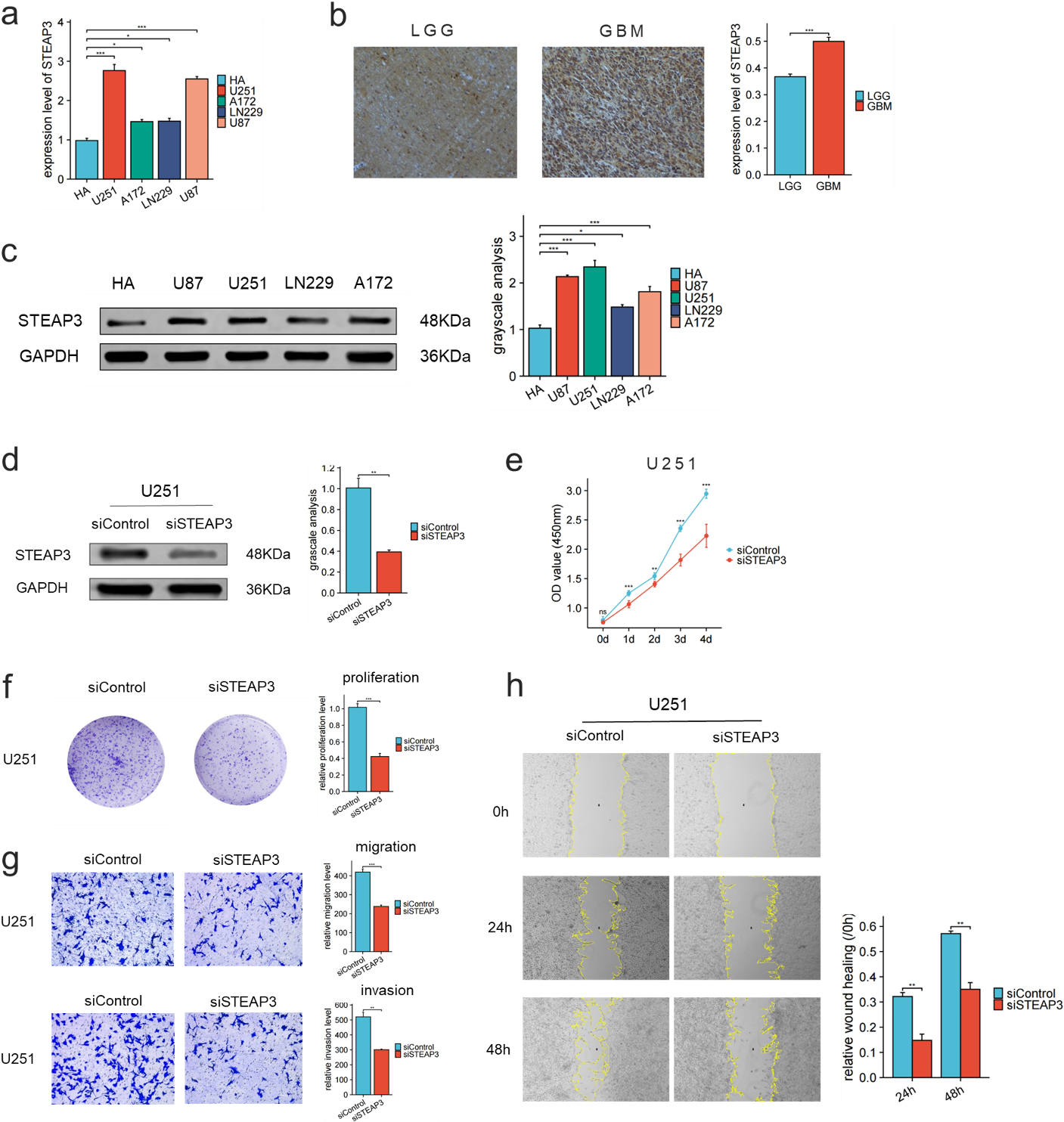
We then investigated the roles of STEAP3 in glioma using siRNA-mediated knockdown of STEAP3. Western blot analysis confirmed the knockdown of STEAP3 expression in U251 cells transfected with siRNA (Fig. 9d). CCK-8, colony formation and wound healing assays indicated that knockdown of STEAP3 significantly inhibited proliferation, invasion and migration ability of U251 cells (Fig. 9e–h).
M2 macrophage infiltration assay was then conducted after STEAP3 knockdown to determine the relationship between STEAP3 and M2 macrophages. After using classical inducing methods, we observed the transformation from THP-1 cells to M2 macrophages (Fig. 10a–b). The findings from our study indicate that the infiltration capacity of M2 macrophages was notably diminished following the suppression of STEAP3 (Fig. 10c).
STEAP3 knockdown significantly decreased recruitment of M2 macrophages in U251 cells. (a) Morphological changes from THP-1 to M2 macrophages. (b)RT-qPCR results showed the relative levels of CD68, CD163 and CD206 in the transformation from THP-1 cells to M2 macrophages. (c) M2 macrophage infiltration was significantly reduced after STEAP3 knockdown in U251. (d) PI3K-AKT pathway was suppressed after STEAP3 knockdown. 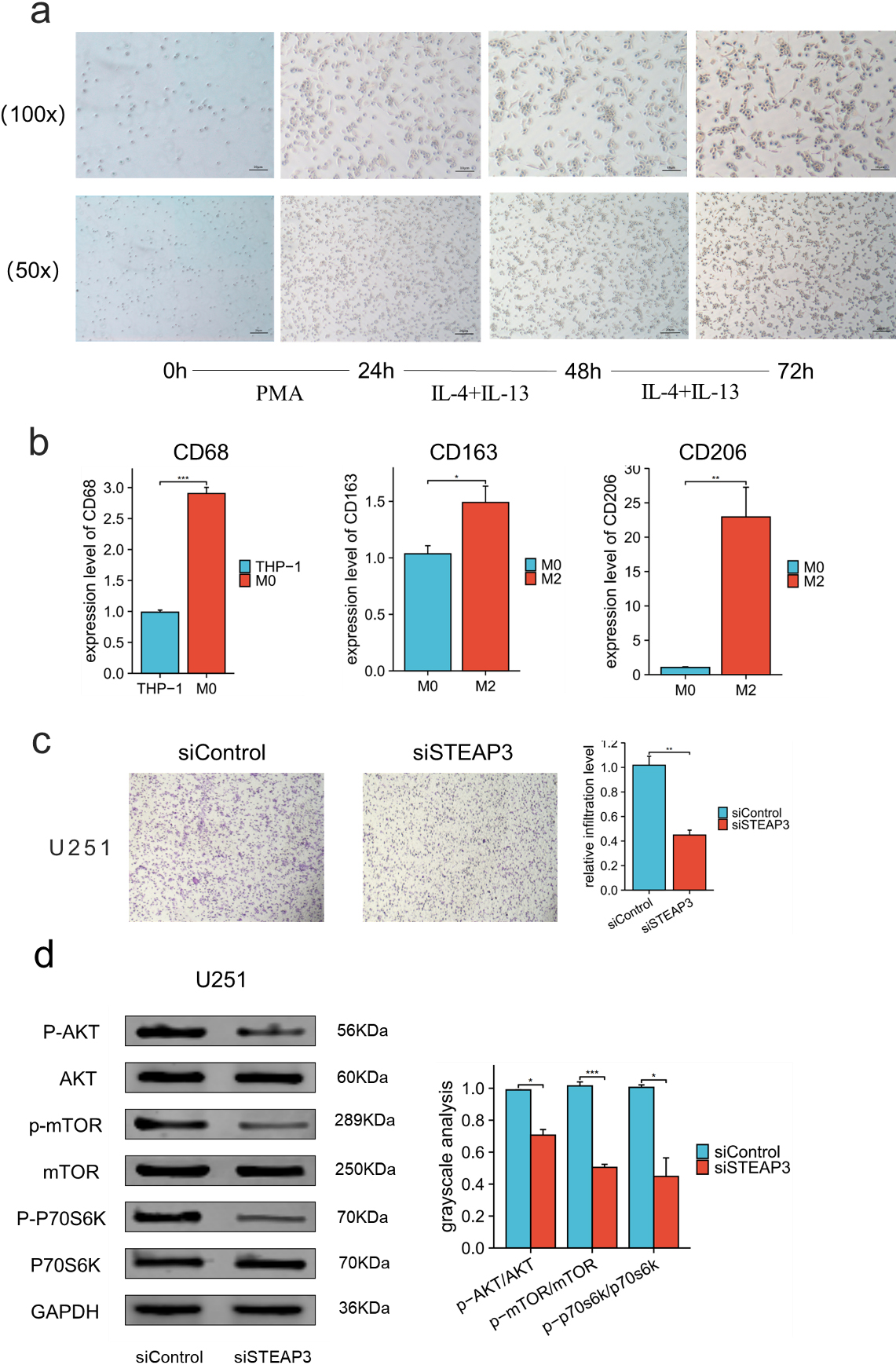
Western blot analysis showed that STEAP3 knockdown decreased the ratio of p-Akt/Akt, p-mTOR/mTOR and p-P70S6K /P70S6K in U251 cell line. This suggests that STEAP3 may promote glioma progression through PI3K-AKT pathway (Fig. 10d).
Discussion
Glioma represents the prevalent and highly malignant type of primary brain neoplasm. The aggressive expansion of gliomas, which is marked by indistinct borders between the tumor and healthy tissue, poses great challenges in achieving complete removal of the tumors [24, 25] Prostate six transmembrane epithelial antigen 3 (STEAP3) is a metal reductase located on the plasma membrane close to the nucleus and vesicles. Research had shown that STEAP3 is highly expressed in colorectal cancer, prostate cancer, breast cancer, glioma, and various other diseases [26, 27]. Short survival time in various types of cancers was also linked to elevated STEAP3 expression [28] Although the prognostic value of STEAP3 for glioma has been previously demonstrated, most of the studies were conducted based on the old WHO classification which did not account for the glioma subtypes. In our study, survival analysis in different glioma subtypes showed that STEAP3 had good prognostic value in IDH-mutant astrocytoma while the prognostic value in other glioma subtypes was limited or unclear. It was suggested that therapy targeting STEAP3 might achieve better response in IDH-mutant astrocytoma.
There is some evidence in the past research showing that STEAP3 was closely associated with tumor development and progression in glioma. STEAP3 was reported to be related to EMT process in glioma and STEAP3 knockdown could inhibit the migration and invasion of T98G cells [29]. Han reported that STEAP3 could predict short-term survival and promote glioblastoma progression by regulating TfR-STAT3 pathway and mesenchymal transition [30]. Our previous work revealed the critical roles of STEAP family in glioma and found that STEAP3 knockdown reduced the proliferation ability of U87 and U251 cells [13]. However, the potential mechanism of STEAP3 had not been fully verified in our past research. Herein, we found that STEAP3 promoted the malignant phenotype by regulating PI3K-AKT pathway and infiltration of M2 macrophages.
Immune microenvironment analysis suggested that STEAP3 may regulate the immune phenotype and tumor immune cycle and lead to a suppressive immune microenvironment. Treatment predictions showed that ICI is more effective in patients with low expression of STEAP3. Earlier research has suggested that the immune system cells and supportive cells have a crucial part in the immune response against tumors [31]. Several types of immune cells, including M2 macrophages, neutrophils, Treg cells, and Th2 cells, have a crucial function in creating an immunosuppressive environment [32]. Our results showed a positive correlation between STEAP3 and the quantity of immunosuppressive cells, suggesting that STEAP3 participated in multiple steps of Cancer-Immunity Cycle. These findings illustrated that STEAP3 played a critical role in glioma immune microenvironment. M2 macrophages promote tumorigenesis and progression probably by activating Th2-type immune response [33]. M2 macrophages may promote tumor progression and poor prognosis by inhibiting CD8+ T cell function [34]. We investigated the relationship between STEAP3 and M2 macrophages and found that the infiltration of M2 macrophages in U251 cells was significantly diminished after knockdown of STEAP3.
The development of immunotherapy has made great progress in increasing survival rate of malignant tumors [35]. Although T-cell therapies have shown excellent efficacy in a variety of cancers and can significantly improve overall survival rates, glioma patients do not benefit from these treatments [36, 37]. This could be attributed to high expression of CD73. CD73 can be used as a specific target to predict GBM response to immunotherapy, and CD73 knockout has been shown to enhance the anti-cancer impact of ICI treatment [23]. Through the utilization of TIDE and ImmuCellAI [22], it was discovered that individuals exhibiting diminished levels of STEAP3 expression experienced greater advantages from ICI treatment in comparison to those with elevated STEAP3 expression. Although high-STEAP3 patients had elevated expression of immune checkpoint genes, they did not benefit from ICI treatment due to upregulation of CD73. In addition, the enrichment analysis showed that DEGs are enriched in antigen recognition and delivery, immune response, cell cycle and PI3K-Akt-mTOR pathway. Our in vitro experiment showed that STEAP3 knockdown inhibits glioma progression via PI3K-AKT pathway.
Our study investigated the predictive significance of STEAP3 in nine glioma cohorts and performed subgroup prognostic analysis based on the new WHO Classification. While Cox analysis revealed that STEAP3 is an independent prognostic factor for glioma, its prognostic significance remains uncertain or restricted in glioblastoma (IDH-wildtype) and oligodendroglioma. We then used bioinformatics analysis to investigate the underlying mechanisms of STEAP3 in glioma especially in the immune microenvironment. In vitro experiments showed that STEAP3 can promote glioma cell progression through PI3K-AKT pathway and regulate M2 macrophage infiltration. Our results suggested that targeting STEAP3 could be a promising approach to inhibit tumor growth and improve immune microenvironment in glioma. However, there are still some shortcomings in our study. There is a need to validate our results using prospective and multicenter clinical cohort studies. Further investigation, including in vivo and in vitro experiments is necessary to elaborate the exact functions of STEAP3 in glioma.
Conclusions
We identified STEAP3 as a prognostic factor for glioma patients, particularly for IDH-mutant astrocytoma. Our findings indicated that STEAP3 contributed to the advancement of glioma through its regulation of immune microenvironment and PI3K-AKT pathway. Our research provides new understandings about the molecular mechanisms and target therapy in glioma.
Abbreviations
Ethical approval
This study was approved by the Research Ethics Committee of the Second Hospital of Hebei Medical University. Approval letter No. 2021 – R519
Funding
This research was supported by National Natural Science Foundation of China (81870984); National Key R & D Program Intergovernmental Cooperation on International Scientific and Technological Innovation of the Ministry of Science and Technology of China (2017YFE0110400); Hebei Natural Science Foundation General Project – Beijing-Tianjin-Hebei Basic Research Cooperation Project (H2018206675); Special Project for the Construction of Hebei Province International Science and Technology Cooperation Base (193977143D); 2017 Hebei Provincial Outstanding Clinical Medical Personnel Training and Basic Research Funding Project; National funded project of Hebei Provincial Excellent Clinical Medical Personnel Training and Basic Research Project in 2019. 2023 Hebei Province Introducing Foreign Intelligence Program. 2023 Government Funding for Excellence in Clinical Medicine Program.
Authors’ contributions
Conception: Zongmao Zhao.
Interpretation or analysis of data: ZihanSong, Zijun Zhao.
Preparation of the manuscript: Zihan Song, Siyu Zhu, Qianxu Jin, Revision for important intellectual content: Yunpeng Shi, Shiyang Zhang, Zairan Wang, Yizheng Wang.
Supervision: Zongmao Zhao.
Data availability
The datasets generated during the current study are available in the data repository: 1. UCSCXenaShiny (
Disclosure
The author reports no conflicts of interest in this work.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-230217.
sj-docx-1-cbm-10.3233_CBM-230217.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-230217.docx
