Abstract
BACKGROUND:
Hepatocellular carcinoma (HCC) is one of the leading causes of cancer related deaths world over. Early diagnosis and effective treatment monitoring significantly improves patients’ outcomes. FKBP11 gene is highly expressed in HCC and could play a role in its development, early diagnosis and treatment.
OBJECTIVE:
This study aimed to evaluate the expression of FKBP11 in HCC, its correlation with patients’ clinical characteristics and potential role in HCC development.
METHODS:
Expression was determined by bioinformatics analysis, quantitative real-time PCR, western blot, and immunohistochemistry. CCK-8, Transwell and wound healing assays were used to investigate involvement in HCC development.
RESULTS:
FKBP11 was significantly upregulated in HCC cells, tissues and blood (all
CONCLUSION:
This study thus found that FKBP11 is upregulated in HCC, and the upregulation promotes HCC development. FKBP11 levels are significantly reduced post-surgery and could be a potential diagnostic and prognostic marker for HCC.
Introduction
World over, liver cancer is the second highest cause of cancer related deaths [1, 2], with hepatocellular carcinoma (HCC) making up about 80% of the cases [3]. It is estimated that there will be approximately 1 million newly diagnosed HCC cases by 2030 [4]. Major risk factors for HCC include hepatitis B and C virus infection, liver cirrhosis and alcoholic liver disease. Chronic hepatitis B virus infection remains the major cause of HCC to date [5, 6, 14, 7]. The 5-year-survival rate of HCC patients is very low (
FK506 binding protein 11 (FKBP11) is an enzymatic protein that catalyzes the folding of proline-containing polypeptides. It’s C-terminal contains a putative transmembrane domain and a motif found within the endoplasmic reticulum (ER) membrane proteins, where it functions in protein folding and secretion [10, 11]. It belongs to the FKBP family of proteins, consisting of 16 numbers that are collectively called immunophilins [10]. It modulates various kinases and cellular factors and is associated with T-cell activation, cell metabolism, cellular homeostasis, tumor carcinogenesis, and tumor progression [12]. FKBP11 mRNA is abundant in secretory tissues such as the liver and pancreas. Its gene encodes a 22 kDa pre-protein containing a 25 residue leucine-rich N-terminal leader sequence, whose cleavage leaves a 19 kDa mature protein, giving the protein its alternative name FKBP19 [13].
Recent evidences increasingly suggest that FKBP proteins regulate cell cycle, survival and apoptotic signaling pathways and influence tumorigenesis and the response to chemotherapies and radiotherapies [14, 15]. Related studies in cancers other than HCC have suggested that FKBP11 promotes cell proliferation and tumorigenesis in oral squamous cell carcinoma via p53-related pathways [16], and is associated with poor prognosis in Clear-cell renal cell carcinoma [17]. In HCC, it has been found that FKBP11 is highly expressed and that it could play a significant role in HCC development [13]. In this study, we evaluated the expression level of FKBP11 in HCC, its correlation with clinicopathological characteristics of the HCC patients, and its role in the pathogenesis of HCC.
Materials and methods
Bioinformatics analysis of FKBP11 expression in hepatocellular carcinoma
We explored the expression of FKBP11 gene and its importance in hepatocellular carcinoma using different bioinformatics tools. We analyzed the TCGA publicly available datasets for differential expression of FKBP11 in HCC and normal liver samples, its relative expression in different stages of HCC, race, histological types of HCC and association with hepatitis B virus infection. We then analyzed its involvement in prognosis of HCC patients using survival analysis.
Sample collection
A total of 46 HCC tissues (30 males and 16 females, mean age 57.7
RNA extraction, cDNA synthesis and quantitative teal-time Polymerase Chain Reaction (qRT-PCR)
Total RNA was extracted from whole blood, tissues and cell cultures using the RNAeazy kits for whole blood, and tissues/cells; (Cat: R6814-02; Cat: R6812-01, Biotek Co., Ltd., China), according to the manufacturer’s instructions. RNA concentration was determined using the Nano-drop 2000 spectrophotometer (Thermo Scientific Inc., Waltham, MA, USA), and then reverse transcribed into cDNA using ReverTra Ace
Cells culture
Hepatocellular carcinoma (HCC) cell lines; HepG2, Huh7, and Hep3B and the normal liver cell line LO2 were all purchased from Procell Inc. (Wuhan, China). All cells were cultured in DMEM medium (Gibco, USA) supplemented with 10% fetal bovine serum (FBS, Gibco, USA), and 1% Penicillin-streptomycin antibiotics.
Lentivirus cell transfection
Two FKBP11 siRNAs were designed and synthesized by HanBio, Beijing China; siRNA1: GGGCAATCATTCCTTCTCA; and siRNA2: GAGAAGCGAAGGGCAATCA. The siRNAs were constructed into a plasmid and cloned into a pHBLV lentiviral vector to generate pHBLV-FKBP11. The pHBLV-FKBP11 was then transfected into HEK 293 T cells and virus particles harvested 48 h post transfection, with the packaging plasmid psPAX2 and the envelope plasmid pMD2.G using Lipofectamine 2000 reagent (Invitrogen). Huh-7, and Hep3B cells were then infected with the recombinant lentivirus-transducing units using 1
Cell proliferation assay (CCK-8)
To determine cell proliferation, the Cell Counting Kit-8 (CCK-8; Dojindo, Kumamoto, Japan) assay was conducted. Briefly; the cells were seeded at 3
Cell migration assays (Transwell and scratch)
To determine cell migration ability, Transwell and scratch assays were conducted. Briefly, cells were first suspended in 200
Western blot analysis
Total proteins from the cells were extracted using RIPA lysis buffer (Cat: P0013; Beyotime, Shanghai, China) mixed with a protease inhibitor; PMSF (Cat: G2008; Thermo Fisher Scientific, Waltham, MA, USA). The protein concentration was measured using BCA proteins assay kit (Cat: P0010, Beyotime, Shanghai, China). Appropriate volumes equaling to 10 ug of protein were then loaded and separated by electrophoresis on 10% SDS-PAGE gel, then transferred onto PVDF membranes (Millipore, Sigma). The membranes were blocked with 5% skimmed milk, and then incubated overnight with primary antibodies against FKBP11 (Cat: bs-13176; dilution: 1:2,000; Bioss, China), and GAPDH (Cat: K200057M; dilution: 1:5000; Proteintech, China). The blots were then incubated with HRP secondary antibodies (Cat: AS1106; dilution: 1:6000, Aspen, China) for 1 hour at room temperature and the bands visualized using the ECL machine (Thermo Fisher Scientific, Inc.).
Immunohistochemistry analysis
Three pairs of tissues (HCC:
Statistical analysis
Normally distributed data were presented as mean
Results
Bioinformatics analysis of FKBP11 expression in Hepatocellular carcinoma
Expression
First, we explored the mRNA expression of FKBP1 in hepatocellular carcinoma (HCC) and normal liver samples using the publicly available TGCA data sets. GEPIA2 (
Bioinformatics analysis of FKBP11 expression using the TCGA datasets GEPIA2 and OncoDB. A: FKBP11 is a pan cancer gene whose expression is upregulated in various cancers, including HCC. B: differential expression of FKBP11 mRNA in HCC versus normal samples (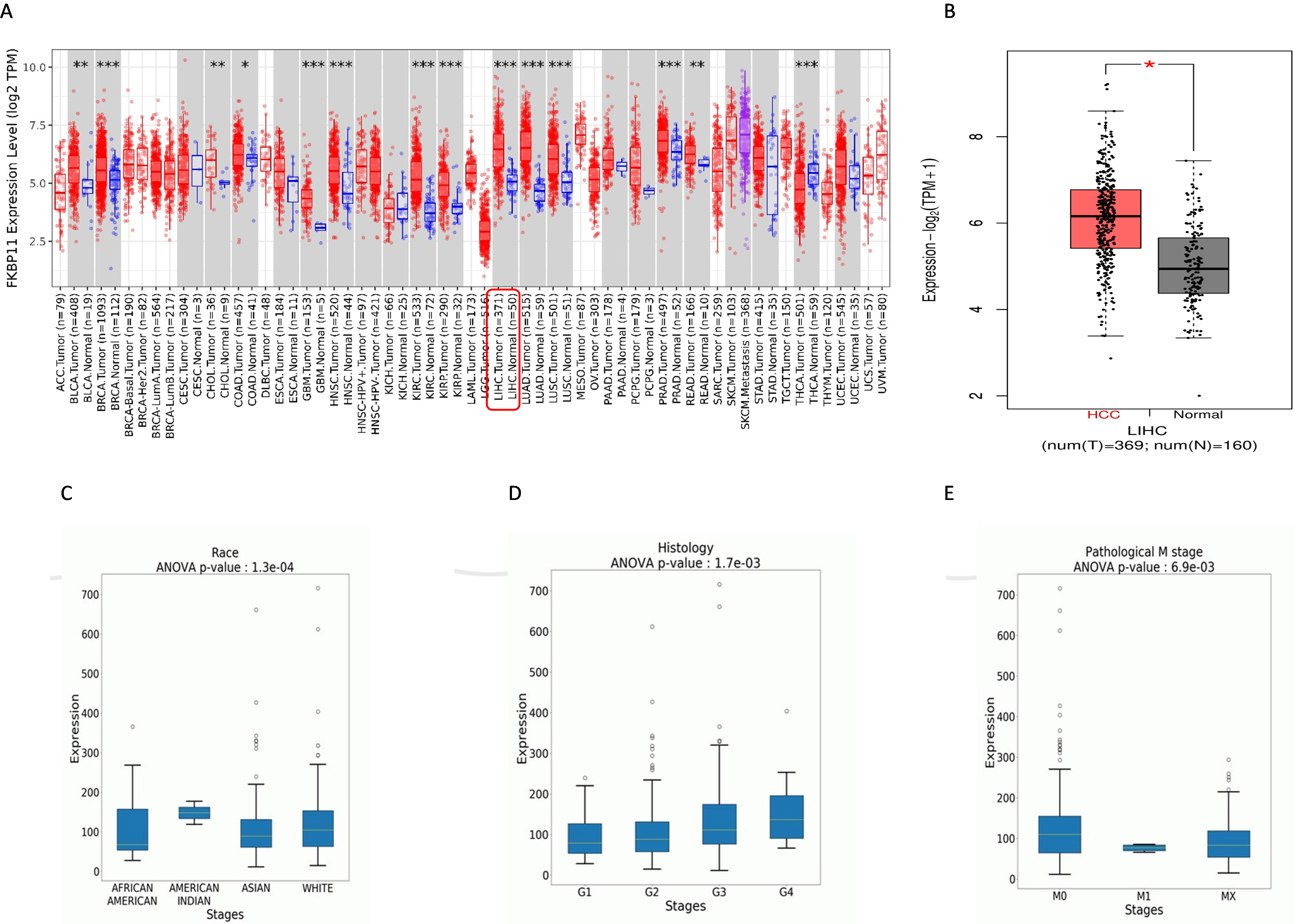
Survival analysis revealed that there is no statistically significant difference in overall survival (OS) between HCC patients with high FKBP11 expression and those with low expression, however, recurrence free survival (RFS), progression free survival (PFS) and disease specific survival (DSS) were significantly different among the different clinical characteristics of patient’s with high and low FKBP11 expression as follows: White people with low FKBP11 expression had better RFS (
Kaplan-Meiyer survival analysis for race, alcohol consumption, and gender. A: Recurrence free survival (RFS) among white people. Those with low FKBP11 expression had better outcomes than those with high expression (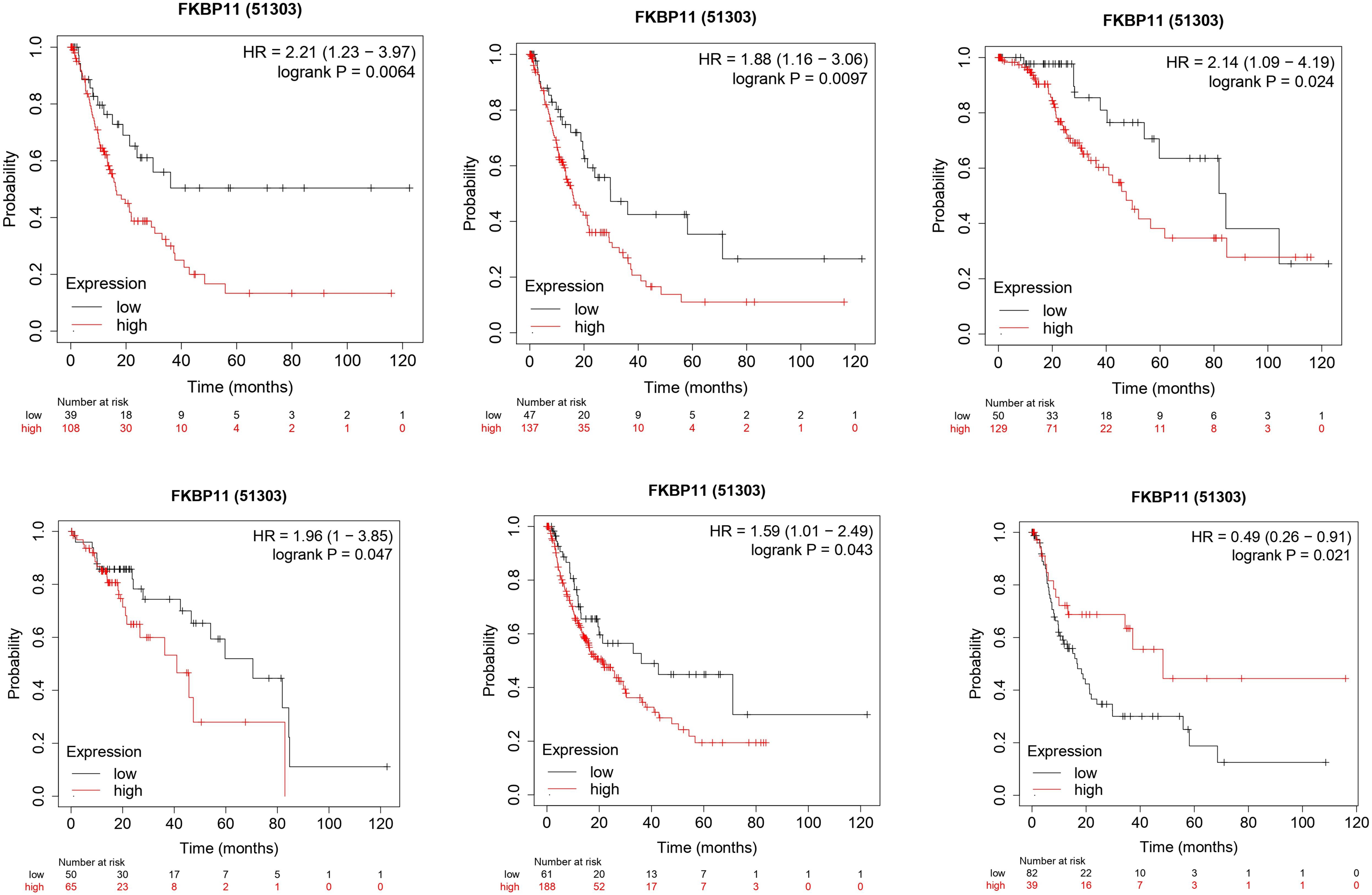
We then explored the association between FKBP11 expression and hepatitis B virus (HepBV) infection in HCC patients with HepBV. Here, the analysis showed that FKBP11 was significantly upregulated in HCC patients with HepBV infection than those without (Fig. 3A). Survival analysis showed that although HCC patients with HepBV and low expression of FKBP11 had better OS, the difference was not statistically significant (
Expression of FKBP11 in hepatitis B positive and negative HCC patients and their survival analysis graphs. A: FKBP11 was upregulated in HCC patients with hepatitis B virus infection than in those who were negative for the virus (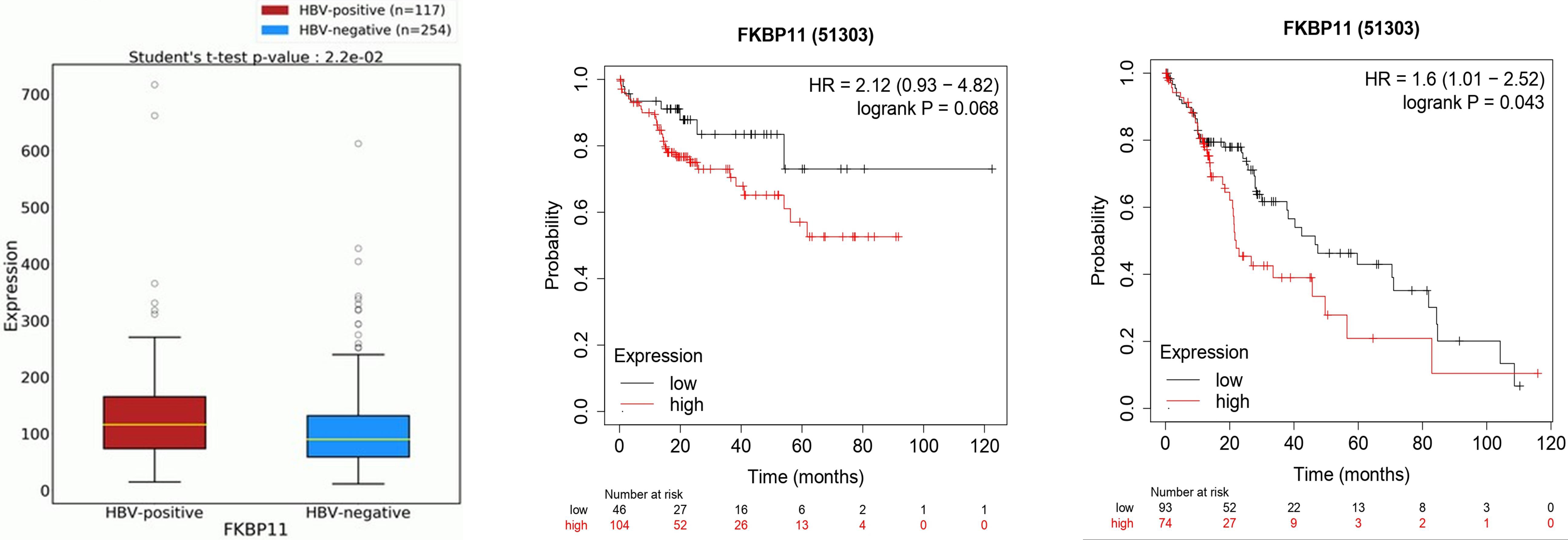
To confirm the bioinformatics results, quantitative real time PCR, western blot and immunohistochemistry were used to determine the differential expression of FKBP11 in HCC tissues, cell lines, and their corresponding normal controls. Here, FKBP 11 exhibited significantly high expression levels in HCC tissues compared to their adjacent normal controls, (Fig. 4A;
Expression of FKBP11 gene in HCC. A: FKPB11 is upregulated in HCC tissues compared to normal adjacent cancer tissues, dependent 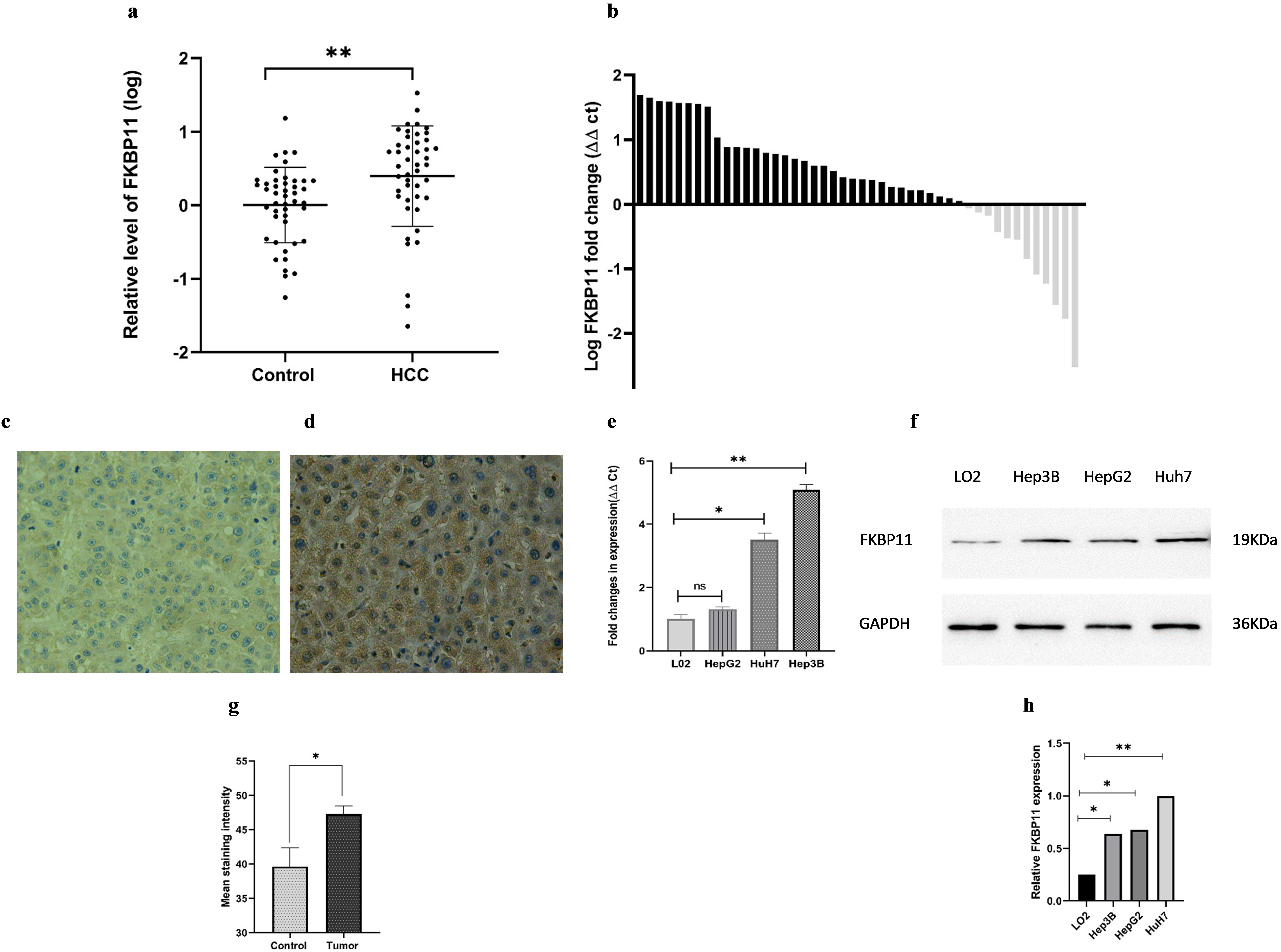
Clinicopathological relevance analysis of FKBP11 in HCC patients
Key. Data presented as mean and corresponding standard deviations. AST: Aspartate aminotransferase; ALT: Alanine aminotransferase; GGT: Gammaglutamyl transferase. AFP: Alpha-feto protein.
Demographic and clinical characteristics of the patients
Data are presented as Median (25 Percentiles, 75 Percentiles).
Details of the patients’ associated clinical parameters of all the 46 HCC tissues were analyzed and summarized in Table 1. Of the 46 patients 30 were male and 16 female. 28 patients were above the age of 55. In terms of tumor characteristics, 44 belonged to TNM I
FKBP11 level in peripheral blood
We then explored the level of expression of FKBP11 in peripheral blood of 62 HCC patients, 40 liver cirrhosis patients, and 65 healthy control cases. Demographic and clinical characteristics of the patients are presented in Table 2. No significant difference was observed in gender as a risk factor, however, significant differences were found in age, ALT, AST, GGT, TBIL, DBIL and TP. As shown in Fig. 5A, the expression levels of FKBP11 differed significantly between HCC and cirrhosis (
FKBP11 expression was downregulated post-surgery
Next, we evaluated whether peripheral blood expression level of FKBP11 could be used to monitor treatment dynamics. We examined the expression of FKBP11 in 14 pairs of plasma samples collected from the same patients before and approximately two weeks after surgery. Interesting, FKBP11 was downregulated in 71.4% (10/14) of the samples, indicating that FKBP11 could be useful in monitoring treatment progress in HCC (Fig. 5C).
FKBP11 knockdown inhibits HCC cell proliferation, and migration
FKBP11 in peripheral blood. A: Expression levels in HCC and liver cirrhosis vs control, Kruskal-Wallis test, and Dunn’s post-hoc analysis (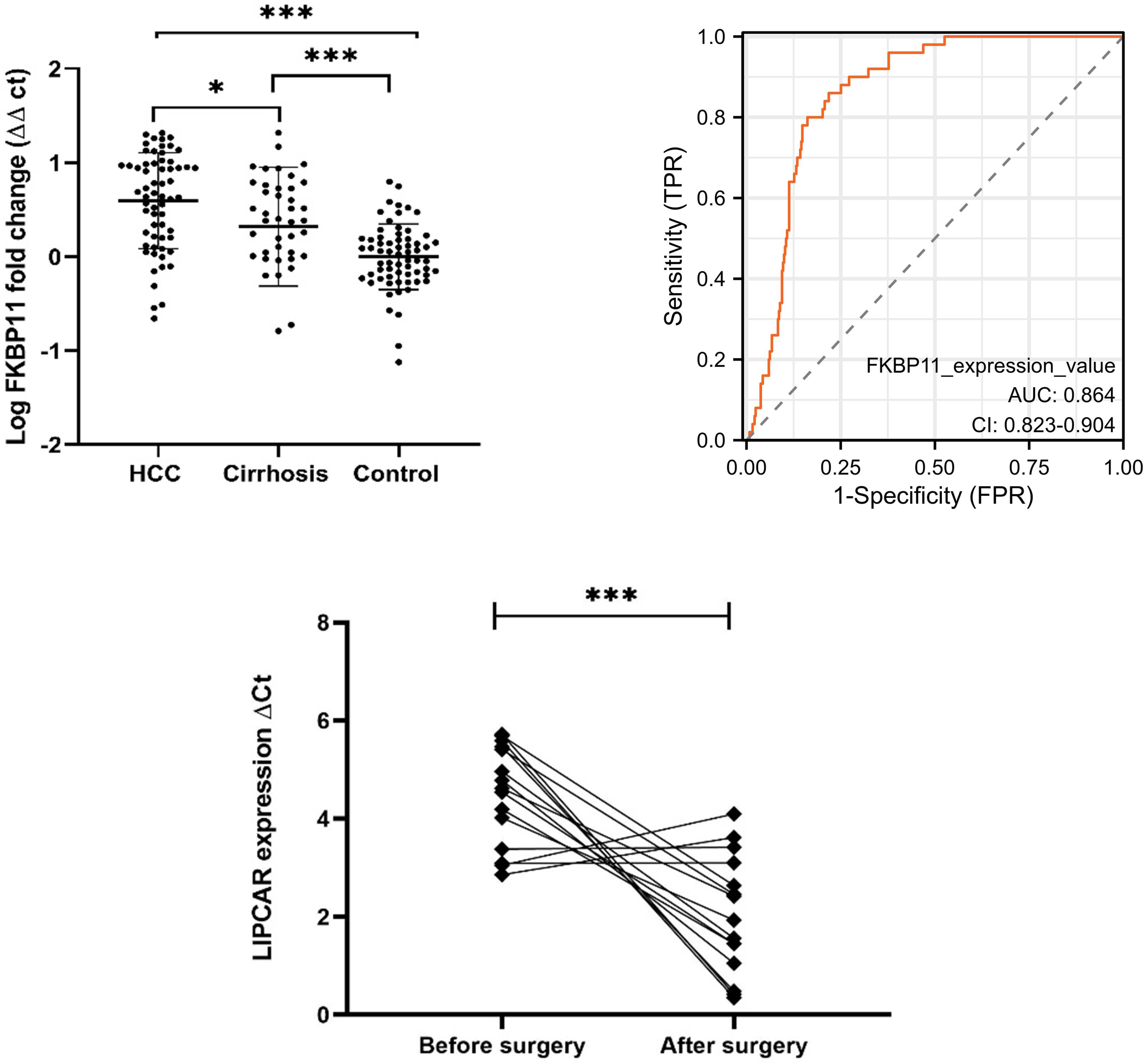
Proliferation and migration assays. A: siRNA knockdown of FKBP11 in Hep3B and Huh-7 cells. B and C: CCK-8 assay results. D–F: Transwell assay results in Huh-7 cells, (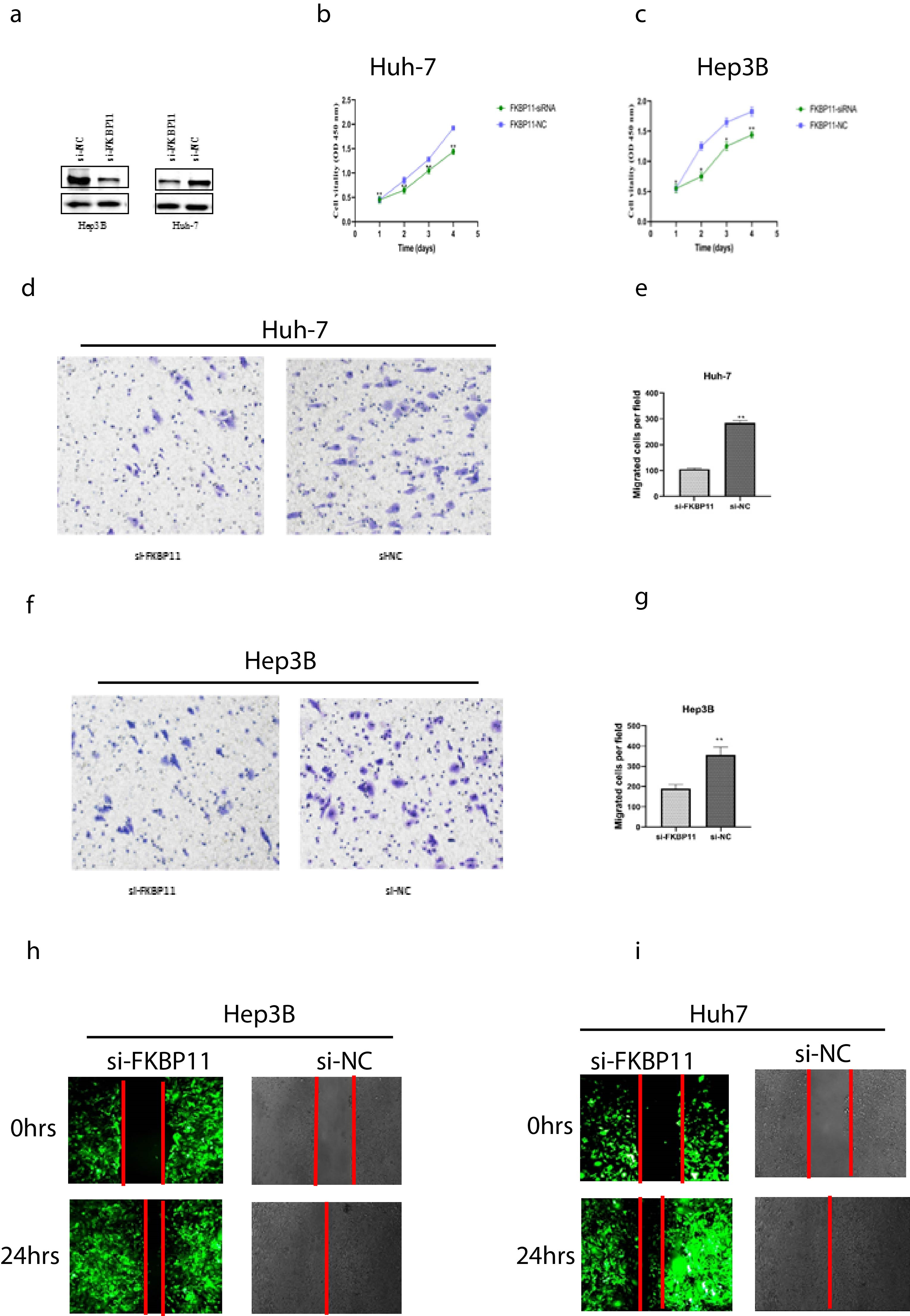
After demonstrating cellular expression, we then determined the role of FKBP11 in HCC progression. We used siRNA to knock down FKBP11 in both Hep3B and HuH7 cells. Transfection efficacy was confirmed by western blot analysis (Fig. 6A). CCK-8 assay showed that compared to the control group, proliferation of the si-FKBP11 transfected cells was significantly inhibited. The degree of inhibition became pronounced as culture time extended, stagnating at around 72 h (Fig. 6B and C). Similarly, si-FKBP11 transfected cells manifested significantly reduced ability to migrate compared to si-NC cells in Transwell and wound healing assays, (Fig. 6D–K, respectively. All
Hepatocellular carcinoma remains a major public health concern to date, owing to its high incidence, often late diagnosis and poor prognosis [19, 20]. According to reports, persistent hepatitis B infection (HBV) is directly responsible for more than 50% of the cases, making HCC a major burden in HBV endemic regions of the world [21]. To improve early diagnosis and prognosis, it’s vital that new biomarkers are identified, since AFP that is the most widely used biomarker to date shows limited sensitivity and specificity [22, 23].
FK506 binding protein 11 (FKBP11) is an enzymatic protein that catalyzes the folding of proline-containing polypeptides [10]. Recent evidence suggest that it regulates cell cycle, survival and apoptotic signaling pathways and influence tumorigenesis and the response to chemotherapies and radiotherapies [12]. Furthermore, it is highly expressed in hepatocellular carcinoma, suggesting that it could play a specific role in the development of hepatocellular carcinoma [13].
In this study, we assessed the expression of FKBP11 in hepatocellular carcinoma and its potential utility as a marker of prognosis and treatment monitoring. The study demonstrated that FKBP11 is significantly upregulated in hepatocellular carcinoma patients compared to controls. Analysis of the association between its upregulation in HCC tissues and clinical characteristics of the tissue providers showed that its expression was significantly associated with HepBV infection. None of the patients in this study had information on hepatitis C virus infection, so we couldn’t examine its association with hepatitis C. From the bioinformatics analysis, HCC patients with HepBV infection and high expression of FKBP11 had worse survival outcomes than those with low expression. According to Peneau et al. [24], HepBV integration into liver cells led to the distant alteration of cancer driver genes such as TERT, TP53 and MYC resulting into HCC with very poor prognosis. Whether this scenario could be true for FKBP11 needs to be studied.
From the peripheral blood samples, there was no difference in gender, smoking and drinking as risk factors among the subject groups, however age, (
Our study is in agreement with a study by Lin et al. [13] who showed that FKBP11 was upregulated in hepatocellular carcinoma. However, Lin et al. only demonstrated expression in cancer tissues unlike ours which also involved cultured cells and peripheral blood leukocytes. Other cancer related studies have shown that FKBP11 promotes cell proliferation and tumorigenesis in oral squamous cell carcinoma via p53-related pathways [16], and is associated with poor prognosis in Clear-cell renal cell carcinoma [17]. In this regard, we explored the potential role of FKBP11 in HCC pathogenesis. Knock down of FKBP11 in HCC cell lines (Hep3B and HuH-7) significantly reduced proliferation and migration of the cells in vitro. These results suggest that FKBP11 could be involved in promoting HCC development. Whether it is involved in HCC metastasis or not, and its molecular mechanism of involvement are areas of future studies.
Limitations
Our study had the following limitations;
First, although we demonstrated the upregulation of FKBP11 in HBV associated hepatocellular carcinoma, and its promotion of HCC, the exact mechanism by which it promotes the cancer is still unknown. This is a likely area of research in our future studies.
Second, downregulated expression after surgery does not necessarily mean it’s applicable for prognosis prediction and treatment monitoring. To confirm this a long follow-up study is required where the reduction in expression can be correlated with patients’ survival outcomes and other factors associated with HCC prognosis.
Conclusion
In summary, FKBP11 expression was upregulated in peripheral blood, cultured cells and hepatocellular carcinoma tissues compared to controls, and was significantly associated with hepatitis B virus infection. Its differential expression in HCC patients versus normal individuals has potential for diagnostic application in HCC, and monitoring of treatment progress. Lastly, Knockdown of FKBP11 in HCC cells significantly reduced cell proliferation and migration, suggesting a possible role in promoting HCC development.
Author contributions
Conception: Erick Thokerunga; Huang Fangfang.
Interpretation or analysis of data: Erick Thokerunga.
Preparation of the manuscript: Erick Thokerunga.
Revision for important intellectual content: Christian Cedric Bongolo, Gilbert Akankwatsa, Simon Peter Rugera, and JianCheng Tu.
Supervision: JianCheng Tu.
Funding
This study was supported by grants from Zhongnan Hospital of Wuhan University Science and Technology Innovation and Education Fund (Project cxpy2018067) and National Basic Research Program of China (973 program) (2012CB720605).
Institutional review board statement
Ethical approval was sought from the Zhongnan Hospital of Wuhan University Research Ethics Committee, number 2018063. All participants provided signed informed consent to participate.
Data availability statement
All the data involved in this study are with the corresponding author and shall be made readily available upon request.
Footnotes
Acknowledgments
The authors acknowledge the contributions of Ms. Bena Binoga for her grammatical review of this manuscript.
Conflict of interest
The authors declare no conflict of interest.
