Abstract
BACKGROUND:
It is of great clinical significance to discover novel biomarkers for neck squamous cell carcinoma (HNSCC) treatments. We discovered a potential cancer-related gene, Cornichon Family AMPA Receptor Auxiliary Protein 4 (CNIH4), that can be a biomarker for HNSCC.
METHODS:
We access multiple open databases and analyzed bulk mRNA-sequencing, protein staining, and single-cell mRNA-sequencing data of HNSCC and investigated the diagnostic and prognostic value of CNIH4 in HNSCC. The potential association between CNIH4 and the immune microenvironment of HNSCC was also estimated.
RESULTS:
CNIH4 was significantly up-regulated in HNSCC compared with non-cancer tissues. Higher CNIH4 resulted in a shorter overall survival time and we further constructed a survival nomogram for clinical applications. 2012 and 421 genes were identified as positive and negative differentially expressed genes of CNIH4 in HNSCC respectively. These genes were mostly mapped to “Cell cycle”, “DNA replicate”, “Cytokine-cytokine receptor interaction” KEGG pathways. Functions associated with CNIH4 were “stemness”, “cell cycle”, and “DNA repair” in single-cell data. CNIH4 potentially affected immune cell infiltration levels and cancer immune therapy.
CONCLUSION:
CNIH4 is a potential diagnostic and prognostic biomarker associated with cancer stemness and immunity in HNSCC.
List of abbreviation
CNIH4: Cornichon Family AMPA Receptor Auxiliary Protein 4
HNSCC: head and neck squamous cell carcinoma
TCGA: The Cancer Genome Atlas
TIDE: Tumor Immune Dysfunction and Exclusion
ICB: immune checkpoint blockade
OCLR: one-class logistic regression machine learning algorithm
TISCH: Tumor Immune Single-cell Hub
UMAP: Dimensionality reduction via Uniform Mani- fold Approximation and Projection
Introduction
As one of the most influential cancer types in the world, head and neck squamous cell carcinoma (HNSCC) results in five hundred thousand new cancer cases each year and causes approximately three hundred thousand deaths [1, 2]. The most common clinical management of HNSCC includes chemotherapy, radiotherapy, and surgery [3]. Over the past ten years, although several novel therapeutic methods for HNSCC patients have been developed, the HNSCC-related death number is still high. The challenges in the early diagnosis and precise prognosis of HNSCC patients prevented the treatments of this disease. The delay of an in-time therapy and the following lymph node metastasis lead to a much higher death rate [4, 5]. Therefore, the discovery of novel biomarkers of HNSCC for clinical HNSCC diagnosis and prognosis is urgent in the field.
Our preliminary screening revealed a potential cancer-related gene, Cornichon Family AMPA Receptor Auxiliary Protein 4 (CNIH4), which encoded a protein interacting with newly synthesized G protein-coupled receptors (GPCR) and regulates their export from the endoplasmic reticulum [6]. CNIH4 has been reported to play potential roles in cancer. A study revealed that CNIH4 could promote colon cancer metastases [7], while in breast cancer, CNIH4 was found to be one of the cancer marker genes [8]. CNIH4 was also identified as one of the genes in the five gene signatures for prognostic of hepatocellular carcinoma patients [9]. However, the clinical values of CNIH4 in many other cancer types remain largely unknown and the roles of CNIH4 in HNSCC required further investigation.
Here, we investigated the value of CNIH4 as a diagnostic and prognostic biomarker for HNSCC. We also proposed that CNIH4 is a regulator for the cancer cell biology and tumor immune microenvironment in HNSCC. This study provided a clinical molecular marker for the improvement of clinical HNSCC treatment.
Methods
RNA-seq and clinical data acquisition
Tissue samples expression data were downloaded from TCGA (The Cancer Genome Atlas) HNSCC cohort in January 2020 following the rules and policies.
RNA-seq data analysis
R foundation for statistical computing (2020) version 4.0.3 and the ggplot (v3.3.2) was used for the expression and survival analysis and the plotting.
Oncomine analysis
The expression and copy number of CNIH4 were also analyzed using the ONCOMINE [10], a cancer microarray database and integrated data-mining platform. All the data mining and differential analyses were done with the ONCOMINE platform. Detailed analysis methods can be found in this paper [10].
Immune cell analysis
The immune cell infiltration levels were calculated using the xCell algorithms [11]. R foundation for statistical computing (2020) version 4.0.3 and the ggplot (v3.3.2) was used for the plotting.
Stemness analysis
The one-class logistic regression machine learning algorithm (OCLR) [12] algorithm was used to calculate the mRNAsi (stemness index based on mRNA expression) score for the evaluation of cancer stemness.
Immune therapy analysis
Potential immune checkpoint blockade (ICB) response prediction was calculated using the Tumor Immune Dysfunction and Exclusion (TIDE) [13] algorithm with the TCGA HNSCC cohort.
Immunohistochemistry staining
The representative images of CNIH4 were accessed from HPA (Human Protein Atlas) [14]. HPA044268 antibody was used for protein staining. The sample details and the general pathological annotations and results were provided by HPA.
Single-cell sequencing data acquisition and analysis
The CancerSEA [15] was used to analyze the functional correlation to the CNIH4 expression. Single-cell datasets GSE103322 [16] (5902 single cells from 18 patients with oral cavity tumors, including five matched pairs of primary tumors and lymph node metastases) were analyzed. The TISCH (Tumor Immune Single-cell Hub) [17] was used to analyze the expression of CNIH4 in immune cells and malignant cells. Data sets GSE103322 [16] and GSE139324 [18] (130,721 single cells from peripheral and intra-tumoral CD45
Statistical significance
Results were considered statistically significant at
Results
The expression of CNIH4 in HNSCC
In this study, we compared the CNIH4 expression in HNSCC and non-cancer tissues. Data suggested that the level of CNIH4 in cancer was up-regulated compared with non-cancer tissues (Fig. 1A1). In high-grade HNSCC, tissues had a significantly higher expression of CNIH4 than low-grade tissues (Fig. 1A2). HNSCC had a significantly higher expression of CNIH4 in male over female patients (Fig. 1A3). The pTNM IV had significantly higher CNIH4 than pTNM I, but the other comparison had no significant difference (Fig. 1A4). Results showed no significance among pT stagings or pN stagings (Fig. 1A5–6). HNSCC had a slightly higher expression of CNIH4 in white over black (Fig. 1A7) and in smoke patients over non-smoke patients (Fig. 1A8). The limit in pack years for smokers is 20 pack years. In addition, the cancer-noncancer paired comparison also supported the conclusion that CNIH4 was up-regulated in cancer over non-cancer (Fig. 1B). The clinical relevance of the high (50–100%, red) and low (0–50%, blue) CNIH4 groups were also analyzed and displayed in a table (Table 1). To observe the up-regulating protein level of CNIH4 in HNSCC, we accessed and observed the protein expression of CNIH4 in tissue staining. Representative images revealed that HNSCC tissues had a higher staining signal of CNIH4 over normal oral mucosa (Fig. S1). However, these are just representative images of the staining for future reference only, thus, we do not have any statistical analysis and quantitative analysis to show the difference. Moreover, to further validated the up-regulation of CNIH4 in HNSCC, we explored the expression and copy number of CNIH4 in HNSCC and normal tissues across 27 data sets. The expression or copy number of CNIH4 in HNSCC was up-regulated in normal tissues in 20 of the 27 data sets included. The overall statistical analysis demonstrated that the mRNA expression or copy number of CNIH4 in HNSCC was up-regulated compared with normal tissues. (Fig. S2).
The expression of CNIH4 in HNSCC. (A1) The mRNA expression of CNIH4 in HNSCC and normal head and neck tissues. TCGA and GETx data were plotted. (A2–A8) The mRNA expression of CNIH4 in HNSCC of different clinicopathological groups. (B) The mRNA expression of CNIH4 in HNSCC and normal head and neck tissues. TCGA paired data were plotted, and cancer and non-cancer samples were linked. Paired 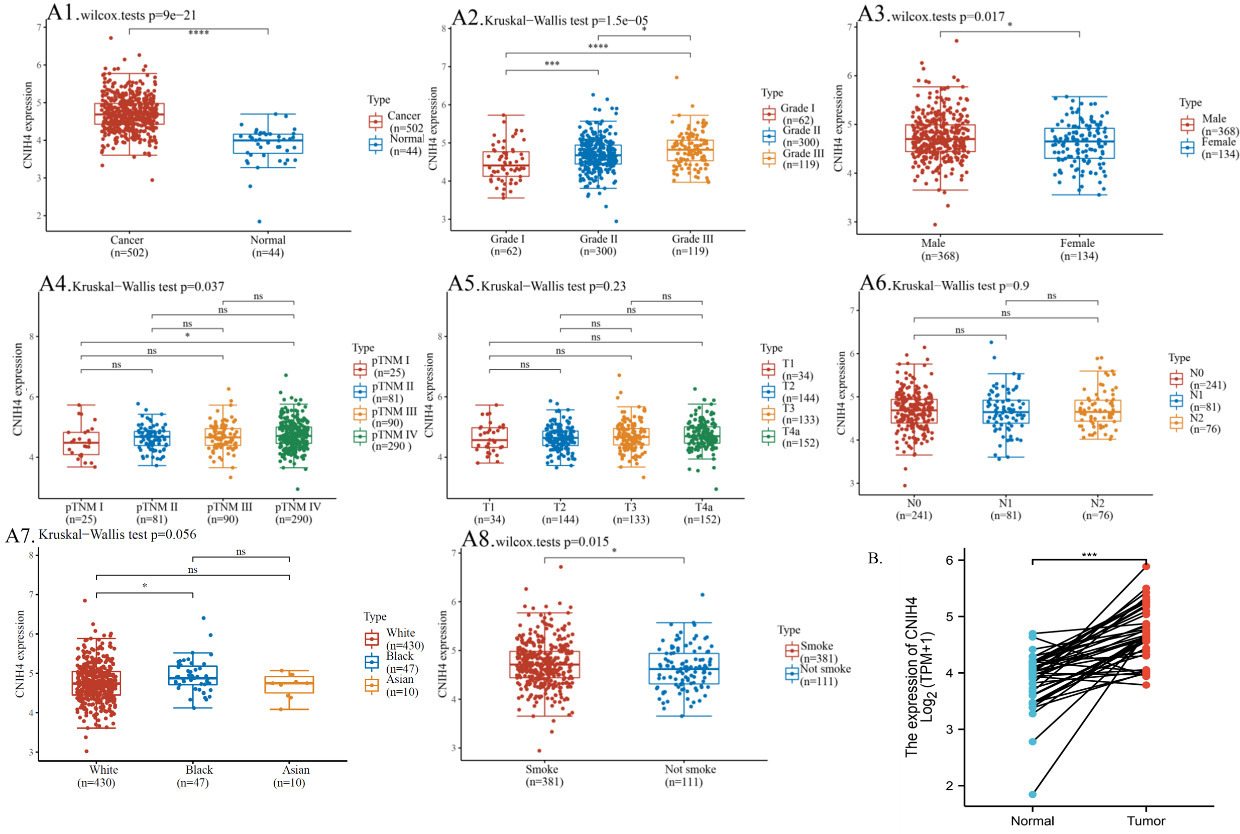
Distribution of HNSCC patients with different clinicopathological variables in CNIH4 high and low groups
To investigate the association between CNIH4 and the survival of HNSCC patients, we conducted a Kaplan-Meier plotting and log-rank test to compare the overall survival of the CNIH4 high (50–100%, red) and low (0–50%, blue) patients. Results showed that the Hazard ratio (HR) of CNIH4 high was 1.5 (95% CI
The prognostic value of the CNIH4 in HNSCC. (A) Kaplan-Meier plot of the high (50–100%, red) and low (0–50%, blue) CNIH4 groups. Log-rank analysis results were shown. (B) The overall survival univariate Cox regression of HNSCC. (C) The overall survival multivariate Cox regression of HNSCC. (D) Nomogram for the overall survival of HNSCC. (E) Calibration plots of the nomogram. 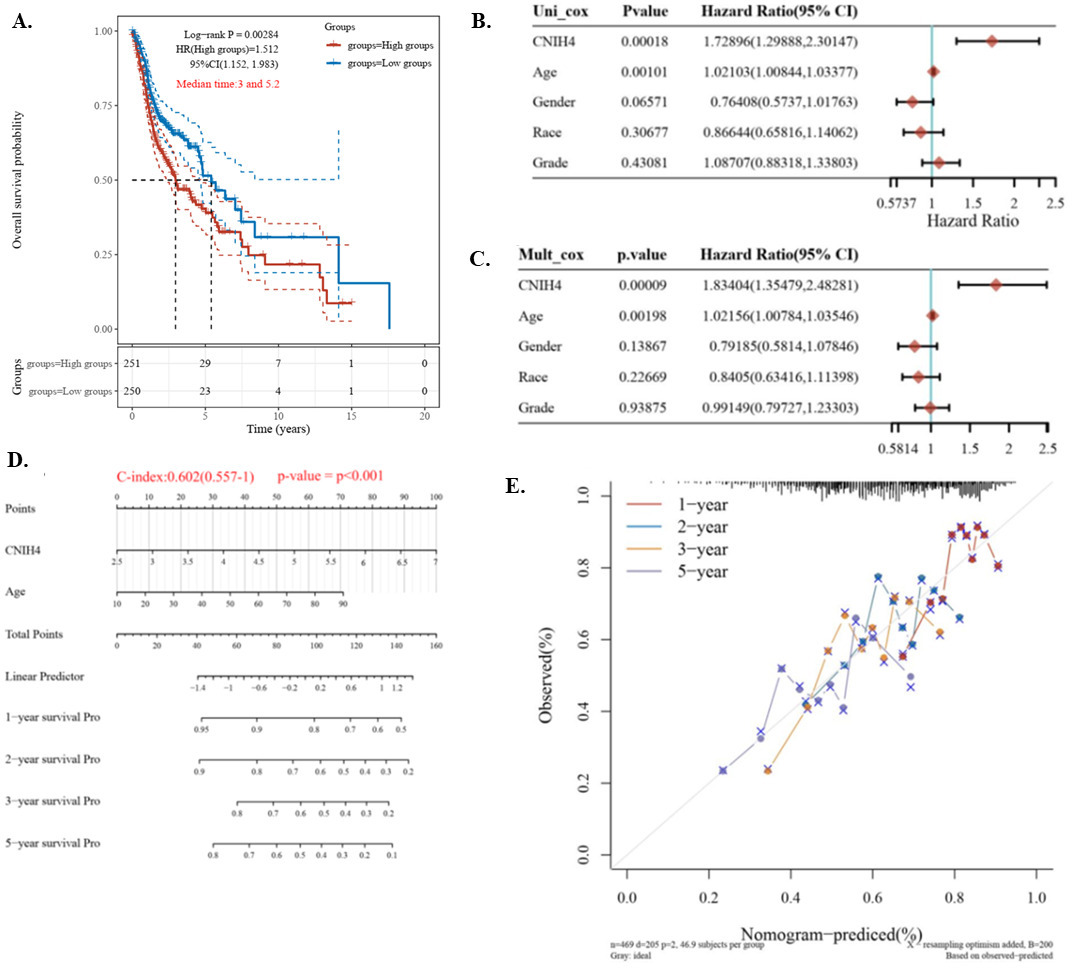
To investigate the potential functional association of CNIH4 in HNSCC, we analyzed differentially expressed genes (DEGs) between CNIH4 high (75–100%) and low (0–25%) groups. The analysis identified 2012 positive DEGs and 421 negative DEGs of CNIH4 in HNSCC (Fig. 3A–B). These DEGs were further investigated in the GO terminologies and the KEGG pathways enrichment analysis. The positive DEGs were found associated with the “Cell cycle”, while the negative DEGs were associated with the “Cytokine-cytokine receptor interaction” KEGG pathways. In GO enrichment, positive DEGs were associated with “organelle fission”, “nuclear division”, “mitotic nuclear division”, “chromosome segregation”, and “DNA replication”, while negative DEGs were associated with “epidermis development”, “skin development”, “epidermal cell differentiation”, “keratinization”, and “keratinocyte differentiation” (Fig. 3C). Other enriched terms were immune-associated, such as “T cell activity”. Generally, these DEGs were found associated with cancer growth and immune cells, indicating that CNIH4 might affect HNSCC stemness and immunity.
The differentially expressed genes (DEGs) enrichment analysis. (A) Volcano plots of CNIH4 DEGs. (B) Heat map and hierarchical clustering analysis of the DEGs. (C) The GO and the KEGG pathways enrichment analysis of CNIH4 DEGs in HNSCC.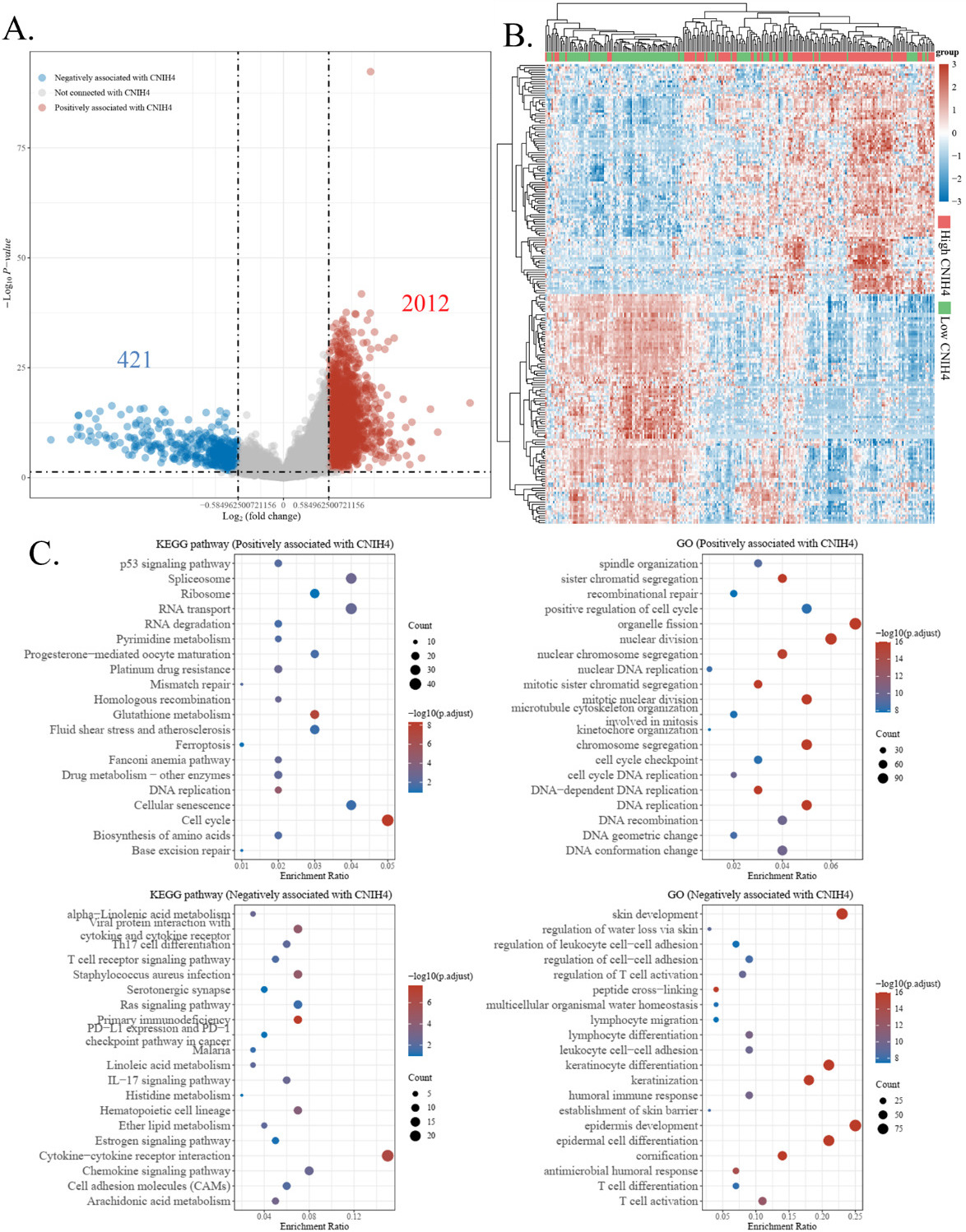
Association of CNIH4 and stemness. (A) Correlations between CNIH4 expression and functional scores in HNSCC single-cell datasets. The GSE103322 data (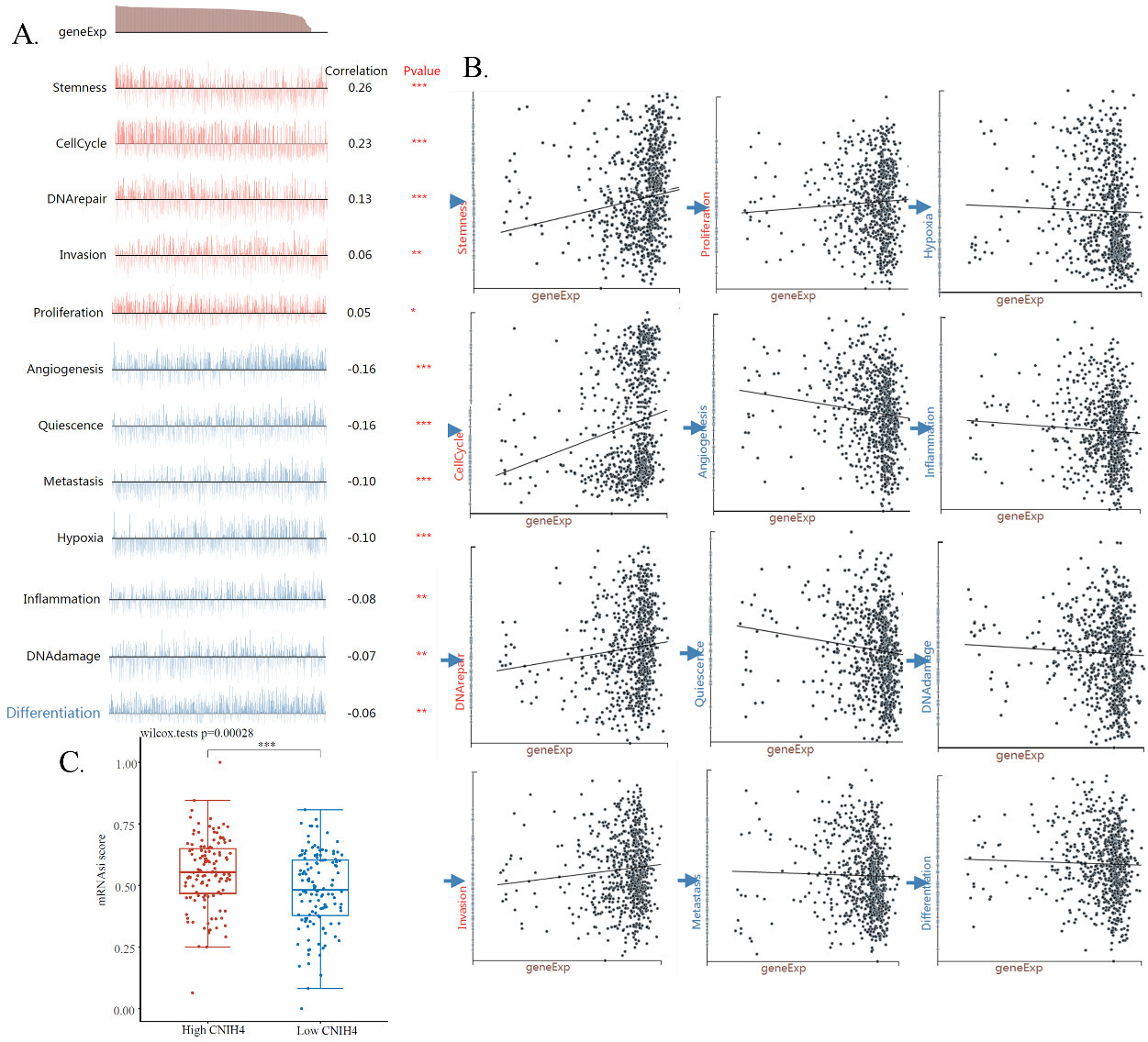
To further study the functional association of CNIH4 in HNSCC, we accessed and analyzed an HNSCC single-cell mRNA-sequencing data set, which included 2150 single HNSCC cells. The correlations of CNIH4 and cancer functional scores were calculated. The calculation showed that 12 functional scores were significantly correlated with the expression of CNIH4. Specifically, the expression of CNIH4 was positively correlated with scores of stemness, cell cycle, DNA repair, invasion, and proliferation with coefficients of 0.26, 0.23, 0.13, 0.06, and 0.05 respectively, and was negatively correlated with angiogenesis, quiescence, metastasis, hypoxia, inflammation, DNAdamage, and differentiation with coefficients of
The expression of CNIH4 in different types of cells in HNSCC. The data were accessed and analyzed using the TISCH. (A) The average expression of CNIH4 in different cell types across datasets (Heatmap). (B1–B2) HNSCC_GSE103322. (C1–C2) HNSCC_GSE139324. The gene expression level was displayed using the UMAP method. 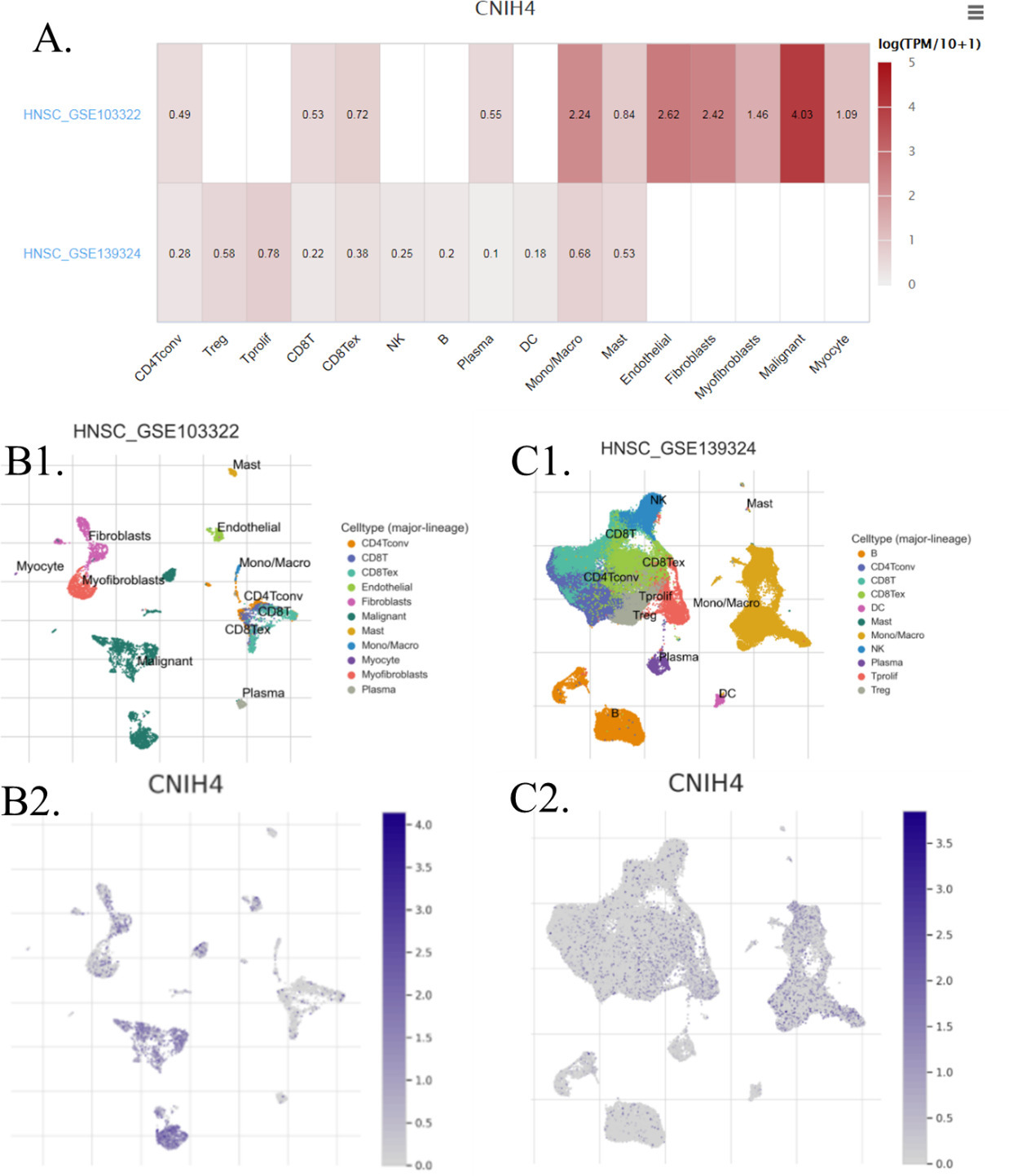
The effect of CNIH4 on the immunity in HNSCC. (A) Comparison of immune cell infiltration levels of high CNIH4 (75–100%) vs. low CNIH4 (0–25%) groups. The xCell algorism was used to calculate the levels of immune cell infiltration levels. The significance was calculated using the Wilcox test. 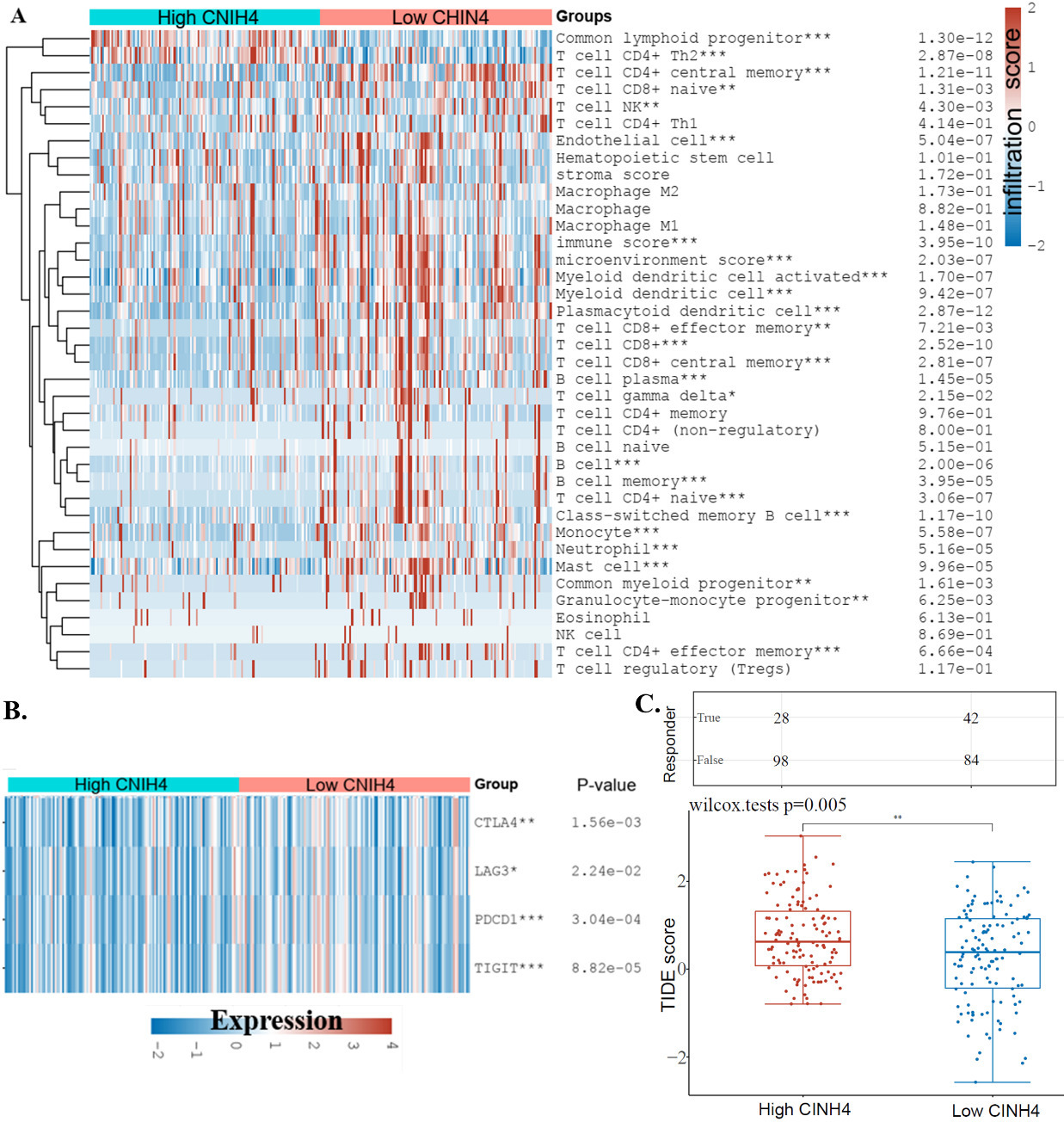
Another point found by the enrichment study was the potential association of CNIH4 and immune cells in HNSCC. To estimate this association, we analyzed the distribution of CNIH4 in different cell types in HNSCC. Single-cell seq data suggested that, in HNSCC, malignant cells expressed much higher CNIH4 than immune cells (Fig. 5), indicating that the expression of CNIH4 in HNSCC bulk data was mainly dependent on the expression of CNIH4 in malignant cells. In addition, we analyzed the immune cell infiltration levels using TCGA bulk data. The immune cell infiltration levels between the CNIH4 high (75–100%) and low (0–25%) groups were compared. Results showed that immune cells up-regulated in the CNIH4 high group included common lymphoid progenitor and T cell CD4
This study reported that CNIH4 was overexpressed in HNSCC compared to normal tissues. Data revealed that this overexpression might result from the increased gene copy number in HNSCC, but other factors such as transcriptional factors or methylation can also contribute to the overexpression of CNIH4 in cancer. In addition, we found that the expression of CNIH4 was higher in the higher grade HNSCC, hence, we proposed that CNIH4 was a potential prognostic biomolecule in HNSCC. A previous study reported that CNIH4 was upregulated in colon cancer [7], which was consistent with our initial proposal that cancer tissues might express higher CNIH4 than their corresponding normal tissues. Therefore, we believe that CNIH4 has a similar cancer-noncancer expression pattern and role in different tumor types. However, to date, the expression cancer-noncancer pattern of CNIH4 in most other cancer types has not been studied.
Figure 1 and Table 1 are different in comparison and statistical analysis. Figure 1 compared expression levels in different groups and used a
One of the most striking findings of this study was that CNIH4 expression potentially affects the overall survival of HNSCC patients. The Cox regression analysis indicated that CNIH4 was a risky factor for HNSCC patients and was independent of patients’ age, gender, grade, and race. Although a previous study suggested that CNIH4 was valuable in the prognosis of hepatocellular carcinoma patients [9], the prognostic value of CNIH4 for HNSCC patients has never been reported. We suggested that most of the poor survival was caused by stemness or immune alteration or other aspects rather than by metastasis. Based on Table 1 M0-M1 analysis, most of the patients analyzed were at M0 which meant no distant cancer spread has been found. In this study, we constructed a nomogram to demonstrate the potential clinical practice of CNIH4 for HNSCC prognosis. Hence, our analysis provided a biomarker candidate for clinical HNSCC diagnosis and prognosis.
Although our results showed that a higher expression of CNIH4 was associated with worse survival of HNSCC patients, it was not clear what roles CNIH4 played in HNSCC. Previously, a study reported that the protein secretion modulator TMED9 drives CNIH4 signaling to inhibit TMED3-WNT-TCF, thereby promoting the migration of colon cancer [7]. Yet, our results showed a potential role of CNIH4 in cancer growth. In the present study, the enrichment results showed that the CNIH4 level was positively associated with stemness-related terms, such as “cell cycle” and “DNA replication”. Cancer proliferation is largely dependent on cancer stem cells [20]. The OCLR score also demonstrated that a higher CNIH4 expression might result in a higher stemness of HNSCC. In addition, in the single-cell analysis, stemness, cell cycle, and DNA replicate were the three most CNIH4-correlated states with considerable large coefficients. The single-cell sequencing gets about 20,000 gene expressions. CNIH4 does not have the highest correlation. But this study focused on CNIH4, not the other genes, hence, although other genes might have an even higher correlation, we did not mention them. In the correlation analysis, we only look at one gene, CNIH4. These single-cell data further supported CNIH4 as a biomarker for cancer stemness in HNSCC. The association between CNIH4 and cancer stemness might account for the association between the high expression of CNIH4 and severe HNSCC. Further functional validation of the stemness association of CNIH4 in HNSCC cells is required in the future.
Another interesting finding in the enrichment study was that CNIH4 might be involved in the immunity in HNSCC. These results raised our interest in the effect of CNIH4 in tumor immune therapy. Immunotherapies such as immune checkpoint therapy (ICT), tumor vaccines, immune adaptive therapy, and immunomodulators have been applied for the treatment of many cancer types [21]. However, for HNSCC, more than 70% of patients who received programmed death 1 (PD-1) or programmed death-ligand 1 (PD-L1) blockade did not benefit from immunotherapy [22, 23]. These disappointing results urged a better understanding of the complex biological mechanism underlying the immune response within the HNSCC microenvironment. Therefore, it is of great significance to explore novel immunotherapy targets or biomarkers for the development of immunotherapy regimens in the clinic. CNIH4, as demonstrated in this study, is a potential predictive biomarker for the ICB response. Therefore, we are looking forward to the development of CNIH4 as a clinically applicable index with a clinical referential significance for HNSCC immunotherapy.
The levels of CNIH4 in single-cell data were analyzed in this study. The data set GSE103322 included expression data of 5902 single cells from 18 patients with oral HNSCC, including cells of primary tumors and lymph node metastases tumors, while GSE139324 included 130,721 single cells from peripheral and intra-tumoral CD45
Our results showed that HNSCC tumor cells expressed a much higher level of CNIH4 than immune cells. A previous study has reported that CNIH4 interacted with newly synthesized G protein-coupled receptors (GPCR) and controlled their export from the endoplasmic reticulum. The role of GPCRs in HNSCC and their clinical relevance to the disease have been extensively explored [24]. GPCR was found to activate the epidermal growth factor receptor (EGFR) and mediate EGFR-independent signaling pathways to promote the growth of HNSCC [25]. We proposed that the potential cancer-promoting effect of CNIH4 in HNSCC might be associated with GPCR and its relative signaling pathways, including pathways that potentially regulated cancer stemness. While in immune cells, regarding the low expression level of CNIH4, we suggested that CNIH4 might not be functional in most immune cells.
In addition, for the interpretation of our immune infiltration analysis, these single-cell expression analysis results provided evidence that a higher level of CNIH4 in HNSCC resulted from higher levels of CNIH4 in tumor cells rather than in immune cells. Therefore, the immune infiltration analysis revealed the effect of CNIH4 expression in cancer cells on the immune microenvironment. Results showed that CNIH4 was negatively associated with most of the immune cell infiltration levels. The only two positive associated immune cell types were common lymphoid progenitor and T cell CD4
In conclusion, this study demonstrated that CNIH4 is a diagnostic and prognostic biomarker candidate for HNSCC patients and can potentially affect the cancer stemness and tumor immune microenvironment of HNSCC cells.
Ethics approval and consent to participate
The authors confirm that all methods were performed following the relevant guidelines and regulations.
Consent for publication
All the authors have given consent for publication.
Availability of data and materials
The datasets analyzed during the current study are available in the TCGA and GEO repositories (GSE103322 and GSE139324).
Author contributions
Conception: Hengrui Liu.
Interpretation or analysis of data: Hengrui Liu and Yixue Li.
Preparation of the manuscript: Hengrui Liu and Yixue Li.
Revision for important intellectual content: Hengrui Liu.
Supervision: Hengrui Liu.
Author information
The corresponding author of this study, Hengrui Liu, is a Principal Investigator at Tianjin Yinuo Biomedical Co., Ltd.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-220143.
sj-docx-1-cbm-10.3233_CBM-220143.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-220143.docx
Footnotes
Acknowledgments
We thank Yi Liu for the technical support in the bioinformatics study. We thank Weifen Chen, Zongxiong Liu, Yaqi Yang, Xinming Li, Ruihua Wang, Yinghua Wang, Siying Wang, Pinghui Zeng, and Shengjie Wang for their help in the study.
Conflict of interest
The authors claimed that there is no conflict of interest.
