Abstract
BACKGROUND:
Acute myeloid leukemia (AML) is a significantly heterogeneous malignancy of the blood. Cytogenetic abnormalities are crucial for the prognosis of AML. However, since more than half of patients with AML are cytogenetically normal AML (CN-AML), predictive prognostic indicators need to be further refined. In recent years, gene abnormalities are considered to be strong prognostic factors of CN-AML, already having clinical significance for treatment. In addition, the relationship of methylation in some genes and AML prognosis predicting has been discovered. RASGEF1A is a guanine nucleotide exchange factors of Ras and widely expressed in brain tissue, bone marrow and 17 other tissues. RASGEF1A has been reported to be associated with a variety of malignant tumors, examples include Hirschsprung disease, renal cell carcinoma, breast cancer, diffuse large B cell lymphoma, intrahepatic cholangiocarcinoma and so on [1, 2]. However, the relationship between the RASGEF1A gene and CN-AML has not been reported.
METHODS:
By integrating the Cancer Genome Atlas (TCGA) database 75 patients with CN-AML and 240 Gene Expression Omnibus (GEO) database CN-AML samples, we examined the association between RASGEF1A’s RNA expression level and DNA methylation of and AML patients’ prognosis. Then, we investigated the RASGEF1A RNA expression and DNA methylation’s prognostic value in 77 patients with AML after allogeneic hematopoietic stem cell transplantation (Allo-HSCT) as well as 101 AML patients after chemotherapy respectively. We investigated the association between sensitivity to Crenolanib and expression level of RASGED1A in patients by integrating 191 CN-AML patients from BeatAML dadataset. We integrated the expression and methylation of RASGEF1A to predict the CN-AML patients’ prognosis and investigated the relationship between prognostic of AML patients with different risk classification and expression levels or methylation levels of RASGEF1A.
RESULTS:
We found that RASGEF1A gene high expression group predicted poorer event-free survival (EFS) (
CONCLUSION:
Higher RASGEF1A RNA expression and lower DNA methylation predicts CN-AML patients’ poorer prognosis. The RASGEF1A high expression level from patients with CN-AML have better sensitivity to Crenolanib. The integrative analysis of RASGEF1A RNA expression and DNA methylation can provide a more accurate classification for prognosis. Lower RASGEF1A expression is a favorable prognostic factor for AML patients receiving chemotherapy or Allo-HSCT. 2010 European Leukemia Net’s risk classification can be improved by RASGEF1A expression levels or methylation levels.
Introduction
Acute myeloid leukemia (AML) is a highly heterogeneous malignancy of the blood in genetic basis and prognosis. The main feature of AML is the clonal proliferation of myeloid blasts in bone marrow, peripheral blood and other tissues, which may cause impaired hematopoiesis and bone marrow failure, with symptoms like frequent infections, easy bruising or bleeding, anemia, bone or abdominal pain, etc. [3, 4, 5]. The incidence of AML is approximately 4.3 per 100,000 persons in the United States and 3.7 per 100,000 persons in Europe [6]. As a disease commonly incident in elderly people, the median diagnostic age of AML is 67 years old, and nearly one-third of AML patients are diagnosed at the age of 75 or more [3]. For a long time, the common chemotherapy of AML has been “3
Cytogenetic abnormalities have long been considered one of the most significent factors in the prognosis of AML, which have been affirmed in clinical practice, for instance patients with inv(16), t(15; 17) as well as t(8; 21) have better estimate [10, 11]. However, nearly one-half of the AML patients are cytogenetically normal (CN), cannot predicting prognosis by cytogenetic markers [11, 12]. In recent year, several genetic mutations have been described, providing more possibility for predicting prognosis [11, 13]. The 4-years survival rate of AML patients with CEBPA mutations is nearly 60%, and AML patients with NPM1 mutation have a 50% 4-year survival rate approximately. Patients with mutations of the CEBPA or NPM1 genes get a favorable prognosis, and BAALC, MLL-PTD and CEBPA can help to predict the response of patients obtaining allogeneic hematopoietic stem cell transplantation (ALLO-HSCT). FLT3–Internal Tandem Duplication (FLT3-ITD) patients own an adverse prognosis, whose 4-years survival rate is only 20% to 25% [14, 15, 16]. Mutations such as RUNX1, DNMT3A and ASXL1 are cytogenetically normal AML (CN-AML) patients’ adverse prognostic factors [17, 18, 19, 20]. Above findings illustrated that gene abnormalities are extreme valuable prognostic factors for CN-AML, providing research directions for us to find precise risk stratifications.
DNA methylation is a direct chemical modification of DNA that form a 5-methylcytosine through transferring a methyl group to the cytosine’s C5 position. DNA methylation controls gene expression mostly through recruiting proteins can inhibit expression of gene or impairing the transcriptional activators’ binding, mainly to reduce gene expression [21]. In addition, methylation also exerts its effects by other mechanisms, including splicing variants, down regulating microRNA, fostering DNA rearrangements, etc. [22, 23, 24]. Previous studies have demonstrated that DNA methylation levels are closely related to the AML patients’ prognosis. For example, high GADD45A methylation level can predict poor survival of AML patients, and the decrease of SCIN expression due to promoter methylation results in adverse prognosis of AML patients, while high HOXA5 methylation levels are valuable biomarkers for prognosis in AML patients [25, 26, 27]. Methylation further affects the prognosis of disease through several mechanisms, indicating that gene methylation also has important research significance in AML patients.
RasGEF1A belongs to RasGEF family and is capable of regulating the transformation and activation of Rap2A, a member of Ras family [28]. Studies showed that Rap2A is a direct target of p53, and the over expression of Rap2A may promote the invasion and migration of many cancers, including lung cancer, renal cell carcinoma, osteosarcoma, nasopharyngeal carcinoma, and gliomas of the central nervous system and was associated with prognosis [29, 30, 31, 32, 33, 34]. A study exhibited that the expression of RASGEF1A is significant for intrahepatic cholangiocarcinoma cells’ survival and migration, while the suppression of RASGEF1A inhibits the growth of ICC cells [35]. It has been reported that RASGEF1A is highly expressed in brain, moderately expressed in bone marrow, spleen, lymph node, testis and gall bladder, weakly expressed in other 21 examined tissues [36]. But the prognostic significance of the expression and methylation of RASGEF1A genes in CN-AML has not been reported so far. Our study integrated CN-AML patients’ two independent groups to studied the prognosis value of RASGEF1A RNA expression level and DNA methylation level in CN-AML patients.
Methods
Data source
We integrated RNA expression profiles of 75 the Cancer Genome Atlas (TCGA) CN-AML samples (
The expression of RASGEF1A was investigated in normal hematopoietic cells from GSE42519 (
All AML samples measured are come from bone marrow or peripheral blood mononuclear cells and the expression and methylation level of AML samples are measured before receiving chemotherapy or Allo-HSCT. TCGA AML patients were diagnosis including white cell counts of peripheral blood, blast percentages of blood and bone marrow, FAB subtypes, cytogenetic risk classification, immunophenotypes and and the frequencies of recurrently mutated genes discovered (e.g., FLT3, DNMT3A, NPM1, TP53, etc.). We collected a total of 164 TCGA AML samples including 67 patients only received Allo-HSCT treatment including 33 CN-AML patients, 92 patients only received chemotherapy including 37 CN-AML patients, and 5 CN-AML patients were not received chemotherapy or Allo-HSCT treatment. In 75 CN-AML patients including 38 female and 37 male and they were divided two group including 10 RASGEF1A high expression and 65 RASGEF1A low expression. Their gene expression was measured using RNA-seq. The 162 AML of GSE12417 U133B dataset and the 78 AML of GSE12417 U133 Plus dataset were normal karyotype and received chemotherapy or transplant treatment. Their gene expression was measured using microarray. The 191 CN-AML patients from BeatAML were normal karyotype received chemotherapy or transplant treatment. Their gene expression was measured using RNA-seq (Fig. S1).
RNA expression profiles and DNA methylation profiles analysis
RNA expression microarray of each CN-AML sample from GEO database was calculated through the method of robust multiarray averaging method (RMA). The expression levels of each probe were transformed with log2. RNA expression data (RNA-seq) and DNA methylation data (HumanMethylation450 chip) were obtained from TCGA database. RNA expression levels of each gene were displayed with RPKM (Reads Per Kilobase per Million mapped reads). It was transformed with log2(FPKM
Statistics
The statistical results were analyed through ggplot2 and survminer packages of R software v3.1.3. Survival curves were designed by Kaplan-Meier estimation. Survival analysis was performed by the Log-rank test. The unpaired
Results
The 75 TCGA CN-AML samples’ baseline characteristics
Baseline CN-AML patient characteristics in the RASGEF1A-high expression and RASGEF1A-low expression group from TCGA database
Baseline CN-AML patient characteristics in the RASGEF1A-high expression and RASGEF1A-low expression group from TCGA database
Compare the survival levels of the RASGEF1A-high expression group and the RASGEF1A-low group in CN-AML patients. (A) EFS and OS in 75 CN-AML patients from the TCGA database (EFS, 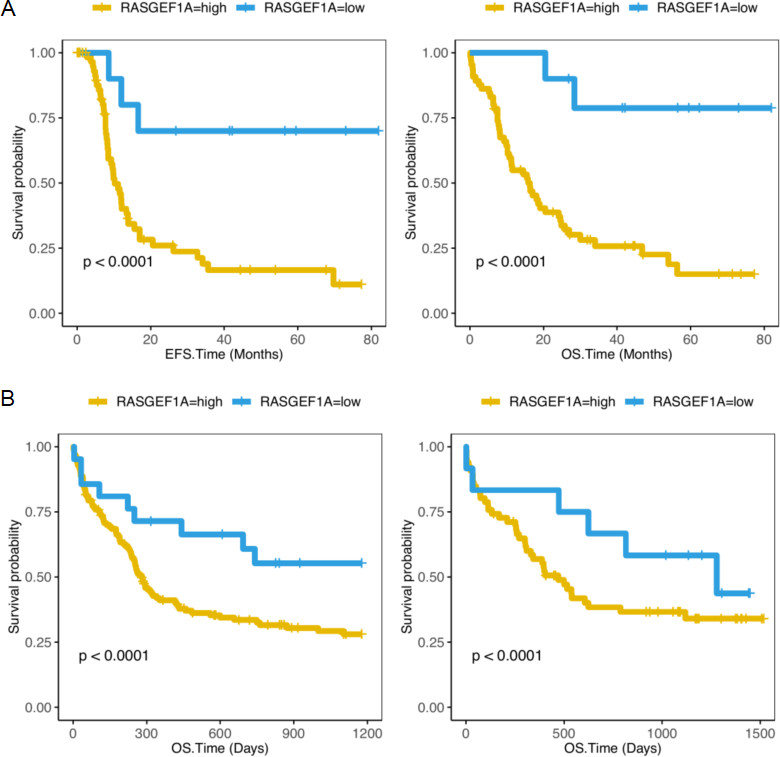
We examine the baseline characteristics of the 75 TCGA CN-AML including 10 CN-AML patients with RASGEF1A-low expression and 65 patients with RASGEF1A-high expression and 240 GSE12417 CN-AML patients including 162 from GSE12417 U133B and 78 from GSE12417 U133 Plus. In 75 TCGA CN-AML patients, we found that the baseline features of the two groups matched. Except for recurrence (
To address the correlation between RASGEF1A expression level and survival rate of AML samples, we conducted a study. We examine RASGEF1A expression level and prognosis in 75 TCGA CN-AML patients and 240 GSE12417 CN-AML patients. We found 10 patients with low RASGEF1A expression from TCGA CN-AML has a better EFS and OS (
The 75 TCGA CN-AML samples’ multivariate analysis
Table 2 clearly illustrates that the expression level of RASGEF1A is a robust independent risk factor for event-free survival (HR
Multivariate analysis for EFS and OS in the TCGA CN-AML
Multivariate analysis for EFS and OS in the TCGA CN-AML
BM_BLAST, bone marrow blast cell; WBC, peripheral blood WBC; PB_BLAST, peripheral blood blast cell; RASGEF1A, RASGEF1A RNA expression; EFS, event-free survival time; OS, overall survival time.
To further understand the correlation between RASGEF1A expression and the prognosis of samples with AML received chemotherapy or Allo-HSCT, we compared the survival rates of patients with low and high RASGEF1A expression received treatment. In 67 AML patients after Allo-HSCT, RASGEF1A-low expression group had more favorable prognosis than RASGEF1A-high expression group (EPS:
Compare the survival levels of the RASGEF1A-high expression group and the RASGEF1A-low expression group in AML patients receiving chemotherapy or Allo-HSCT. (A) EFS and OS in 67 AML patients receiving Allo-HSCT from the TCGA database (EFS, 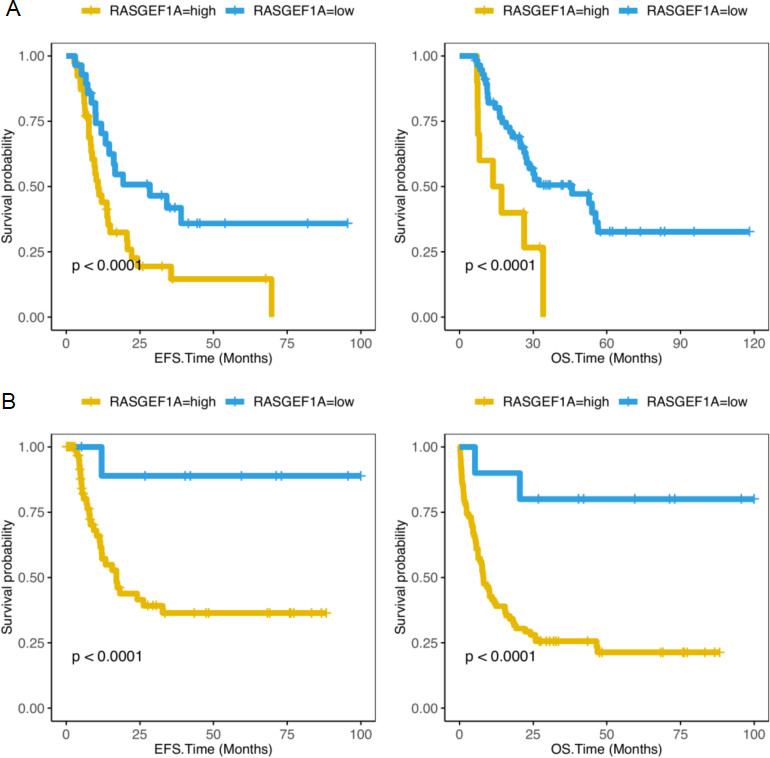
Compare the survival levels of RASGEF1A-high methylation group and RASGEF1A-low DNA methylation group in CN-AML patients, and AML patients receiving chemotherapy or Allo-HSCT. (A) EFS and OS in 85 CN-AML patients from the TCGA database (EFS, 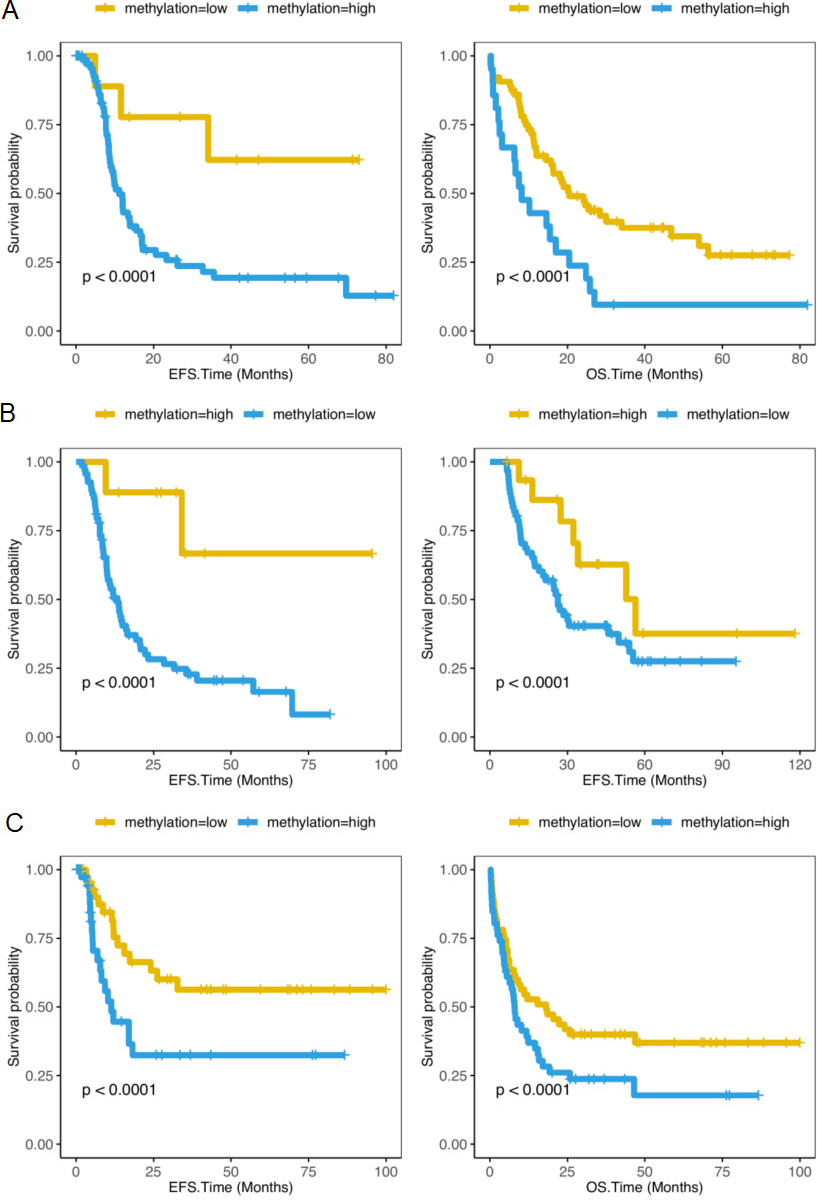
We detected the methylation levels of RASGEF1A at the same time. In 85 TCGA CN-AML patients, the survival rate was elevated in RASGEF1A-high methylation group (EPS:
Combination of expression and methylation levels of RASGEF1A provide a accurate prognostic classification for patients with CN-AML. (A) EFS and OS of G1, G2 and G3. G1 represents RASGEF1A-high expression and RASGEF1A-low methylation group, G2 represents RASGEF1A-high expression and RASGEF1A-high methylation and RASGEF1A-low expression and RASGEF1A-low methylation, G3 represents RASGEF1A-low expression and RASGEF1A-high methylation group. A Log-rank test was used to compare the survival curves of high and low gene expression. CN-AML, cytogenetically normal acute myeloid leukemia. Event-free survival time (months); OS, Overall survival time (months). Left side: the x-axis represents the EFS time (months); the y-axis represents the survival probability; Right side: the x-axis represents the OS time (months); the y-axis represents the survival probability.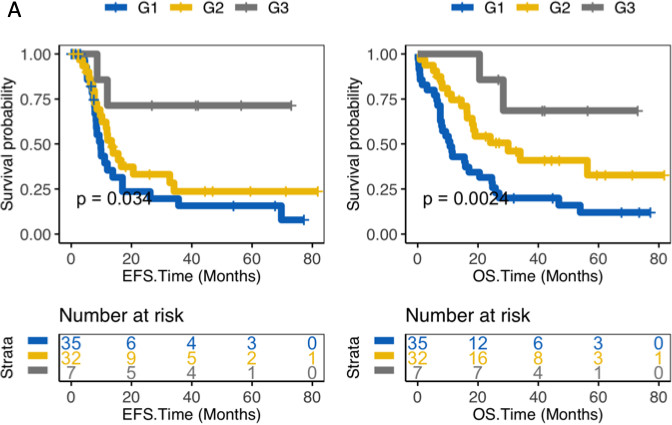
We comprehensively examine expression and methylation levels of RASGEF1A of 74 CN-AML patients and obtained a more accurate method for prognosis classification. As shown in Fig. 4, 74 patients were divided into 3 groups, which are RASGEF1A-high expression and RASGEF1A-low methylation group (G1), RASGEF1A-high expression and RASGEF1A-high methylation or RASGEF1A-low expression and RASGEF1A-low methylation (G2), RASGEF1A-low expression and RASGEF1A-high methylation group (G3). In EFS, G3 had the best prognosis, G2 had moderate prognosis and G1 had the worst prognosis among the 3 groups (EFS:
To investigate the relationship between prognostic of AML patients with different risk classification and expression levels or methylation levels of RASGEF1A, we compare prognostic of 4 groups including AML patients with good risk (
Dependence of RASGEF1A on AML
We examined RASGEF1A expression of hematopoietic cells from GSE42519 datasets. Band cells, which are immature neutrophils, had a significantly higher expression level of RASGEF1A than other hematopoietic cells (
The BeatAML CN-AML samples with RASGEF1A high expression level with better sensitivity to Crenolanib
To investigate the relationship between RASGEF1A expression level and sensitivity to Crenolanib in CN-AML patients, we integeted 191 CN-AML patients from BeatAML including 50 RASGEF1A low expression and 141 RASGEF1A high expression. We found the IC50 (half maximal inhibitory concentration) of Crenolanib of CN-AML patients with RASGEF1A high expression level is lower, which suggested RASGEF1A-high had better sensitive to Crenolanib (Fig. S1). We found high FLT3 or high PDGFRA expression level group had better sensitive to Crenolanib, while higher PDGFRB expression level had lower sensitive to Crenolanib (Fig. S7).
Discussion
With the gradual development of research on AML, the heterogeneity in the disease is becoming more common, and cytogenetics is increasingly recognized as an independent predictor of prognosis in AML patients. However, though many prognostic factors have been identified, there is more than 50% of patients do not have cytogenetic marker, which are cytogenetically normal AML patients [11, 12, 13]. The good news is many gene abnormalities that escape cytogenetic detection have been discovered, like CEBPA, NPM1, KIT, FLT3-ITD, having great significance in predicting prognosis [17, 18, 40]. In addition, the role of gene methylation in prognosis has also been discovered, like high GADD45A methylation predicting low survival of AML patients [25]. Therefore, new gene abnormalities are still required for further discovery, in order to provide clues for predicting prognosis, studying the pathogenesis and exploring new therapeutic targets. In this study, we integrated survival of 75 TCGA samples with CN-AML and 240 GEO CN-AML samples, examined the effect of RASGEF1A expression on CN-AML patients’ prognosis, including patients received chemotherapy or Allo-HSCT, and the influence of RASGEF1A methylation levels on CN-AML samples’ prognosis.
In this study, we found that RASGEF1A can be used as a CN-AML patients’ prognostic factor. We compared the OS and EFS of 75 TCGA samples with CN-AML and 240 CN-AML patients from GEO database with or without high RASGEF1A expression, including CN-AML patients’ overall prognosis and respective prognosis of AML patients received chemotherapy or Allo-HSCT. The study showed that the prognosis of patients in RASGEF1A-high expression group of RASGEF1A was poorer than the RASGEF1A-low expression group (
In addition, we studied the association between the methylation of RASGEF1A gene and patients survival rates. As can be seen from Fig. 3A, the EFS and OS of patients are much better in RASGEF1A-high methylation group than RASGEF1A-low methylation group (
Furthermore, we found that the integrative analysis of gene expression and methylation can provide a more accurate method of prognostic classification. In Fig. 4, we can see that CN-AML samples can be separated into 4 groups by RNA level and methylation level, and the difference in EFS and OS among the 4 groups were statistically significant (
RASGEF1A is an important guanine nucleotide exchange factors of Ras. Previous studies have exposed that RASGEF1A high expression is related to the survival and migration of intrahepatic cholangiocarcinoma cells, and our study has shown a correlation with the prognosis of patients with AML [35]. This may suggests that this gene further contribute to the pathogenesis of AML and is associated with some malignant diseases. At the same time, this gene was detected in the bone marrow and other 26 detected tissues, especially in brain, lymph nodes, spleen and testis [29, 30, 31, 32, 35, 36]. We can guess that there are also potential correlations with RASGEF1A gene and diseases in those tissues, which requires further studies.
What is regrettable is that this experiment does not involve molecular mechanisms. To date, no prospective studies have proved the clinical significance of RASGEF1A in the treatment of CN-AML.
In conclusion, the high RASGEF1A expression level and low methylation level suggest poor survival and adverse prognosis for CN-AML patients. The integrative analysis of RNA and methylation level can provide a more accurate classification for prognosis. The CN-AML patients with RASGEF1A high expression level have better sensitivity to Crenolanib. Low RASGEF1A expression is a favorable prognostic factor for AML patients receiving chemotherapy or Allo-HSCT. RASGEF1A expression levels or methylation levels is help to improve risk classification of 2010 ELN.
Funding
This work was funded by National Natural Science Foundation of China (81800195 and 81460315), Key Clinical Projects of Peking University Third Hospital (BYSYZD2019026, BYSYDL2021006), Beijing Natural Science Foundation (7132183 and 7182178), China Health Promotion Foundation (CHPF-zlkysx-001), Wu Jieping Medical Foundation (320.6750), Scientific Research Foundation (20141114) from Health Commission of Jiangxi Province, and Science and Technology Research Foundation (GJJ14676) from Educational Commission of Jiangxi Province, China.
Contributions
Conception: HMJ, ZPZ, YLZ and CJY.
Interpretation or analysis of data: XH, WLZ and WF.
Preparation of the manuscript: XH, WLZ, WF, XNL, PY, JW, MXZ, SXL, WZ, XRZ, GHD, HMJ, ZPZ, YLZ and CJY.
Revision for important intellectual content: XH, WLZ, WF, XNL, PY, JW, MXZ, SXL, WZ, XRZ, GHD, HMJ, ZPZ, YLZ and CJY.
Supervision: HMJ, ZPZ, YLZ and CJY.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210407.
sj-docx-1-cbm-10.3233_CBM-210407.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210407.docx
Sources of support
The sources of data resources are The Cancer Genome Atlas (TCGA) database (
Footnotes
Acknowledgments
The authors thank TCGA database, GEO database, BeatAML dataset and DepMap Portal database which are all available online.
Conflict of interest
The authors declare that they have no conflict of interest.
