Abstract
BACKGROUND:
MicroRNAs (miRs) are frequently altered in colorectal cancer (CRC) and can be used as prognostic factors.
OBJECTIVE:
To confirm in stage III CRC patients a reported miR signature that was associated to the presence of metastatic disease. To correlate miR expression with microsatellite instability (MSI) and mutations in RAS and BRAF.
METHODS:
miR-21, miR-135a, miR-206, miR-335 and miR-Let-7a expression was analyzed by RT-qPCR in 150 patients out of the 329 patients used to analyze MSI and RAS and BRAF mutations. Association with disease free survival (DFS) and overall survival (OS) was analyzed. Data was confirmed by a multivariate analysis.
RESULTS:
MiR-21 high expression (
CONCLUSIONS:
Association of increased expression of miR-21 and better prognosis in the poor prognostic group may be of interest and could be explored in future prospective clinical trials.
Keywords
Introduction
The CRC represents an important health problem, being the third cause of death in developed countries [1]. Around 75% of patients are diagnosed with localized disease and almost half of them are in stage III. 5-year disease-free survival of colon cancer stage III patients, treated only with surgery, varies widely depending on T-N sub-stages, from 79.6% for T1N1a to 13.9% for T4N2b. Adjuvant chemotherapy is generally recommended for all stage III subgroups, but the magnitude of benefit increases with the higher risk of relapse [2]. Apart from TN sub-stage, few data derived from clinical or molecular studies have provided robust enough results to be implemented in clinical practice. In this scenario, most patients with low risk stage III are overtreated with the adjuvant chemotherapy treatment while patients in the very high risk group will relapse after the current recommended oxaliplatin-based adjuvant therapy. Molecular alterations such as high microsatellite instability (MSI-H), RAS or BRAF mutations, correlate with histopathological characteristics and clinical different colorectal cancer types and they have prognostic impact in advanced disease [3, 4]. However, none of these molecular factors have a great prognostic impact in stage III, and therapeutic options are not driven by them.
It is essential in clinical practice to identify prognostic and predictive factors that permit identification of patients with increased risk of relapse, those who benefit from chemotherapy, and those for whom this treatment is unnecessary and could even be harmful given its pronounced toxicity.
MicroRNAs (miRs) regulate gene activity post-transcriptionally by inhibition of protein translation. In cancer, they can function as oncogenes or as tumor suppressors, and miR signatures can serve as promising biomarkers for diagnosis, prognosis, and treatment efficacy monitoring over the disease course [5]. In this context, A miR expression signature containing miR-21, miR-135a, miR-335, miR-206 and miR-let-7a that was associated with the presence of metastatic disease was reported [6]. In this study, we have analyzed the expression of this miR signature in patients with stage III CRC to confirm its correlation with clinical outcome. In the absence of literature data, we explore potential associations with the presence of BRAF and RAS mutations as well as Microsatellite Instability (MSI), in order to correlate miRNA expression signatures with well-known patterns of colorectal cancer subtypes.
Materials and methods
Patients
We analyzed 329 samples out of a total of 472 samples of patients with CRC stage III treated with adjuvant therapy included in a previous study of our group [7]. Tumor samples were taken from the Biobanks of the centers associated to the Cooperative Spanish Group for the Treatment of Digestive Tumors (TTD) and of the Biobank of the Hospital Clinico San Carlos B.0000725 (PT20/00074), integrated in the Spanish National Biobanks Network and they were processed following standard operating procedures with the appropriate approval of the Ethical and Scientific Committees. The study was approved by the Institutional Review Board of the Hospital. All samples collected included the corresponding informed consent following The Code of Ethics of the World Medical Association (Declaration of Helsinki, British Medical Journal 18 July 1964). Unanalyzed samples were excluded from the study due to lack of tumor blocks.
RNA and DNA extraction
RNA and DNA were extracted from 4–5 cuts (7
DNA mutational analysis
RAS KRAS (12Val, 12Asp, 13Asp, 12Cys, 12Ser, 12Ala, A146T, K117N, A146V, A183C); NRAS (Q61L, Q61R, Q61K, G12D) and BRAF (600Glu) mutations were detected by End-point Multiplex PCR amplification, followed by hybridization in a low-density microarray, using the CLART
Analysis of microsatellite instability
MSI Analysis System, Version 1.2 (Promega) was used to detect microsatellite instability in DNA from FFPE tumor tissue. Following the instructions of the company, PCR products were separated by capillary electrophoresis using an Applied Biosystems
Analysis of miRNA expression
We analyzed the expression of the miR-21, miR-135a, miR-206, miR-335, miR-let-7a and miR-103 (miR-Control [8]) by RT-qPCR in 150 samples out of 329 initial samples. This selection was based on the quantity (
TaqMan qPCR was carried out using miRs specific primers: miR-21 (ref:000397), miR-135a (ref:000460), miR-206 (ref:000510), miR-335 (ref:000546), miR-Let-7a (ref:000377) and miR-103 (ref:000439) (Thermo Fisher Scientific). The initial volume for each miR was: 1
Data was processed using the LinRegPCR software [9] through which the N
Cutoff finder
Cutoff points for survival analysis were determined using Cutoff Finder software [10]. Among the five methods available, fit of mixture model procedure was chosen to obtain the cutoff point. This procedure fits two Gaussian distributions to the histogram of the biomarker (in our case, miRNA expression values). Optimal cutoff is established as the point where the probability density functions of both distributions coincide.
Statistical analysis
Categorical variables were summarized with absolute numbers and the percentage of each group regarding the total population under investigation. Chi-Square-test (or Fisher’s exact test) were used to determine statistical differences in categorical variables distribution between groups. Continuous variables were summarized, and within-group variation was compared by dispersion analysis using pairwise Wilcoxon-Mann-Whitney test. Overall survival and progression free survival was represented using the Kaplan-Meier model. We performed univariate analysis using the Cox proportional hazards regression model to determine the significance of those observed differences. Those variables that presented a significative effect in survival were included in a multiple regression model (along with diagnosis age, due to the well-known impact in prognosis) in order to obtain the effect of each variable in term of hazard ratios (HRs).
All statistical tests were two-tailed, and a
Results
Patients
The main characteristics of the study cohorts are shown in Table 1.
Patient characteristics (
329)
Patient characteristics (
T, tumor; N, node; TNM, tumor-node-metastasis; ECOG PS, Eastern Cooperative Oncology Group performance status; 5FU, 5-fluorouracil; FOLFOX, 5-fluorouracil plus leucovorin plus oxaliplatin; XELOX, capecitabine plus oxaliplatin; Mut, mutated; Wt, wildtype; N/A, unknown; MS, microsatellite; MSI-H, microsatellite instability High; MSS/MSI-L, microsatellite stability.
Association between miR-21, miR-135a, miR-206, miR-335 and miR-Let-7a expression levels and mutations in RAS, BRAF and MSI were analyzed. Our results show a significant association of higher expression levels of miR-21 to MSI-H tumors (
Box plot of the association of miR-21 and miR-335 with MSI and of miR-135a with mutations in RAS. 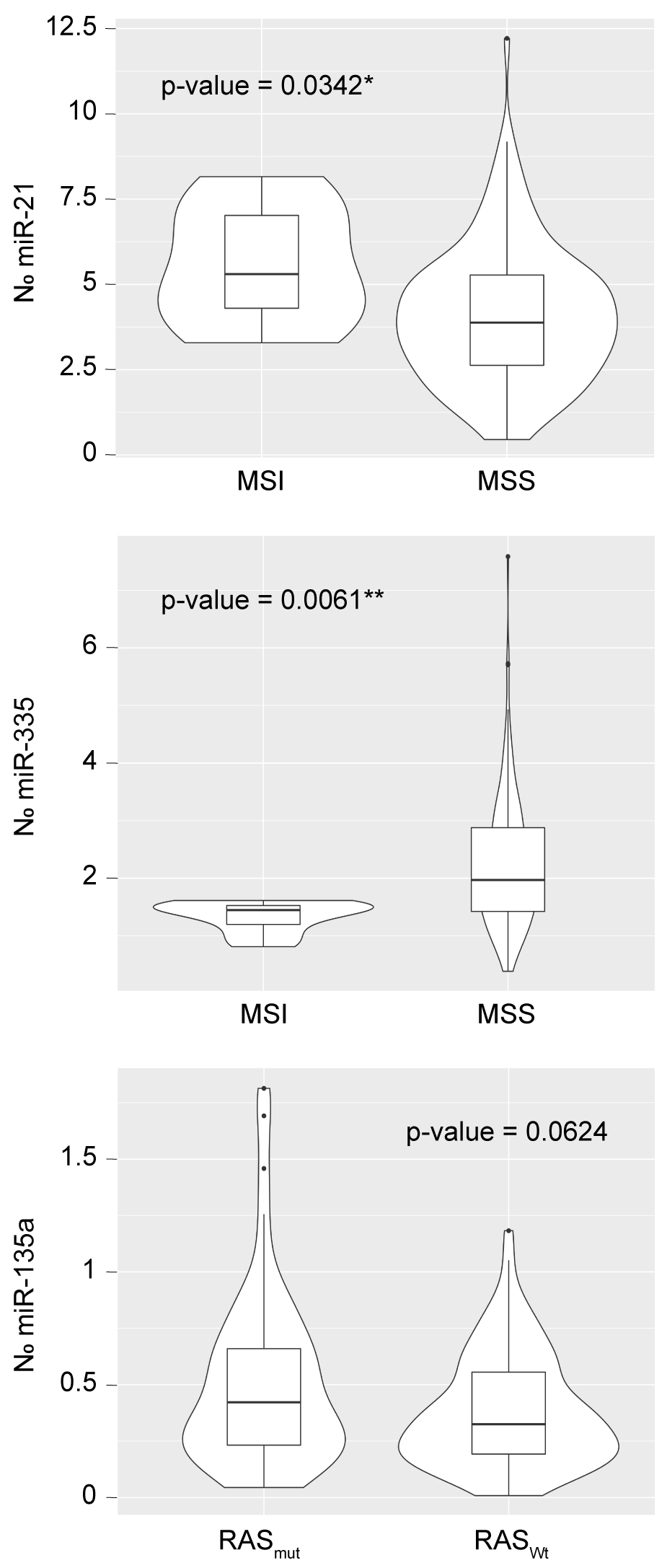
DFS Kaplan-Meier curves according to miR expression. 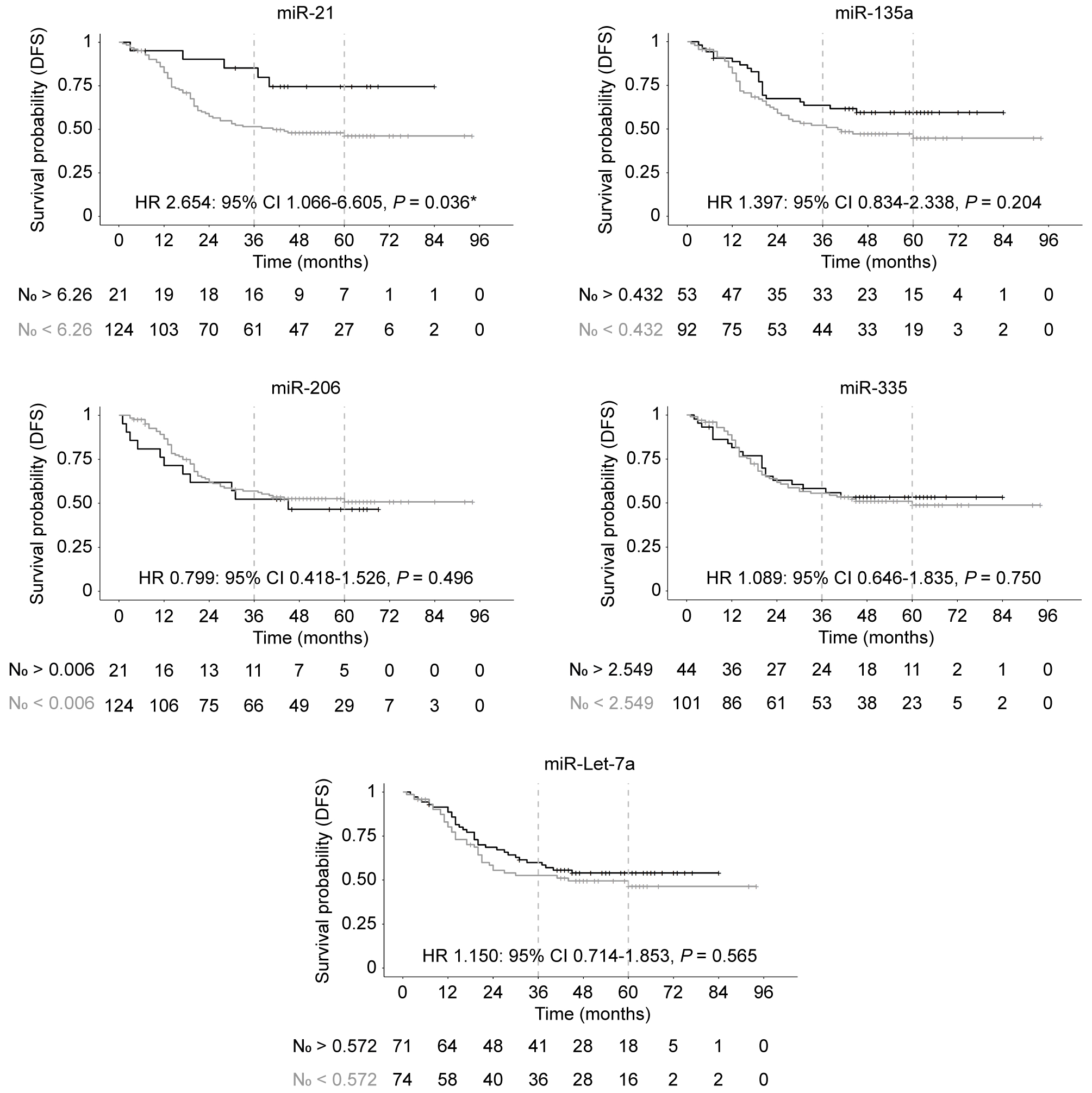
OS Kaplan-Meier curves according to miR expression. In all cases, Grey line: N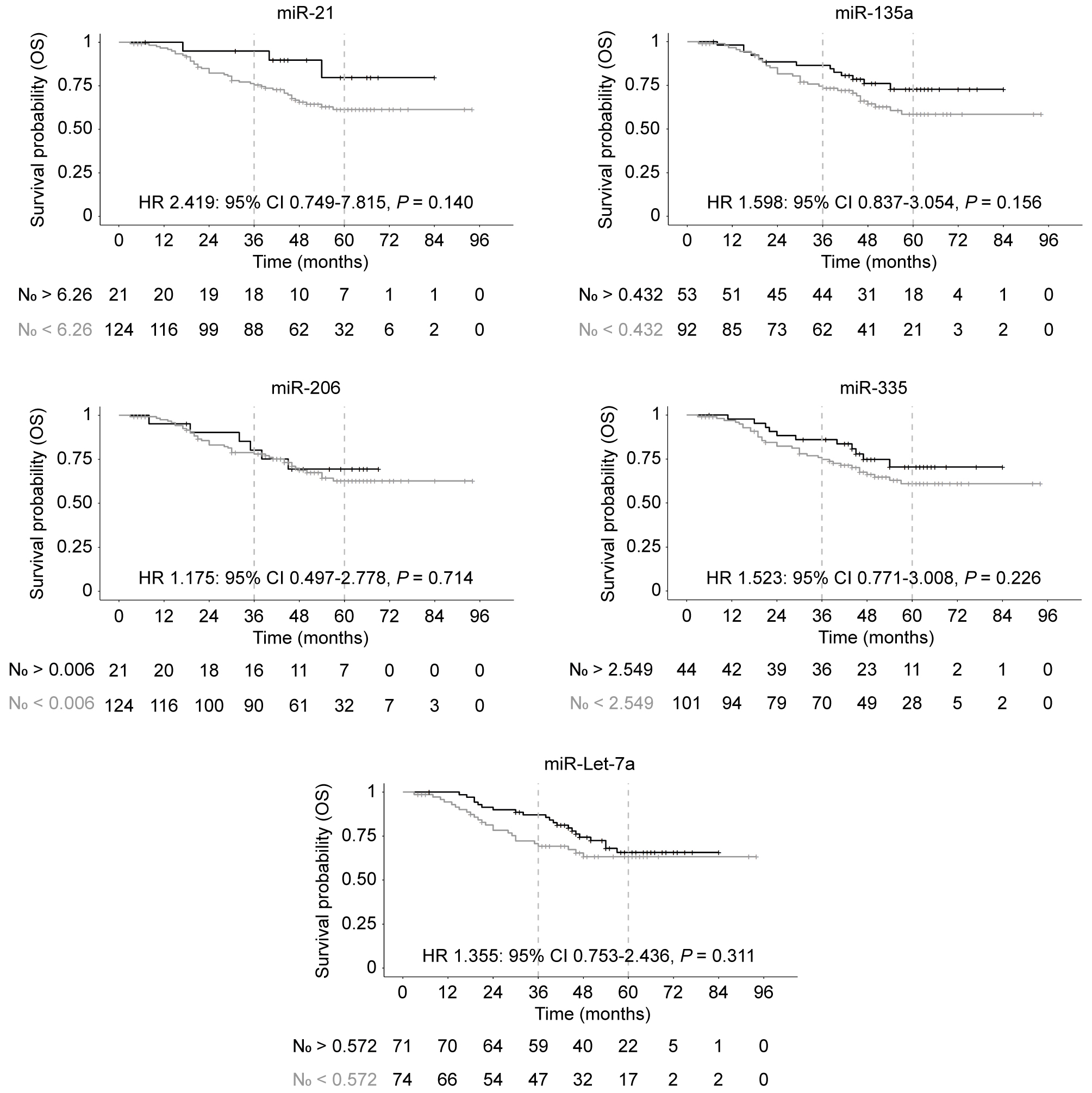
DFS and OS Kaplan-Meier curves according to MS, RAS and BRAF mutations. Grey line: MSS/MSI-L and Wild Type; black line: MSI-H and Mutations in RAS and BRAF. At the bottom of each graph, the number of patients contained in each group that remain throughout the follow up period.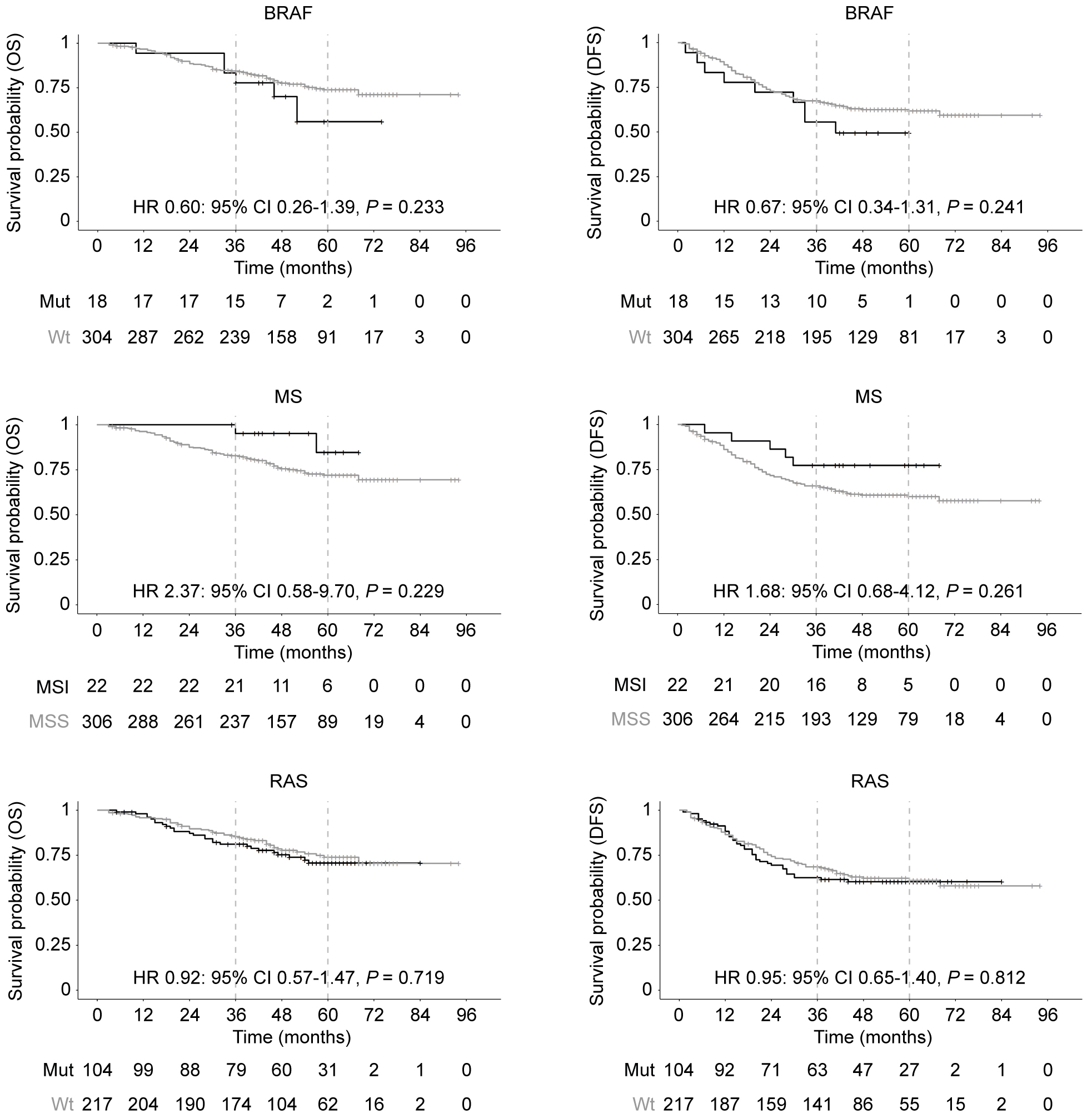
DFS Kaplan-Meier curves according to miR-21 expression and prognostic statement. A) DFS of the good prognostic group (T1-3/N1): grey line; and the poor prognostic group (T4 or N2): black line. B) DFS of the effect miR-21 levels in the good prognostic group (T1-3/N1). C) DFS of the effect miR-21 levels in the poor prognostic group (T4 or N2). Grey line: N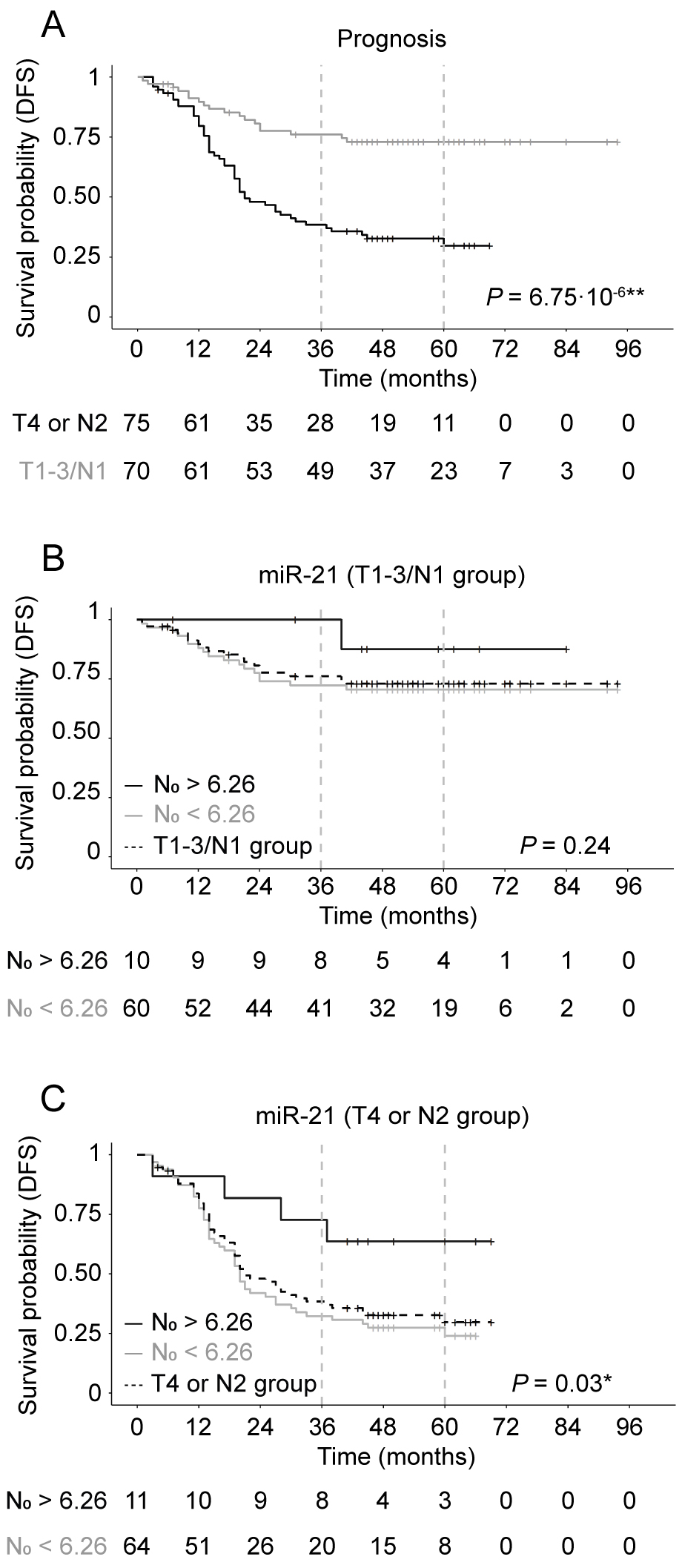
No significant associations were found between the expression levels of any of the analyzed miRs and the presence of RAS or BRAF mutations with the exception of a trend (
To analyze the association of miR expression levels with Disease Free Survival (DFS) and Overall Survival (OS) a cutoff point for each of the analyzed miRs was generated.
The expression levels of the studied miRNAs vary from miR-21 showing the highest expression levels to miR-206 which showed the lowest expression levels. MicroRNA-21 N
Association of MSI and mutations in RAS and BRAF with recurrence and survival
Our results showed that there are no significant differences in DFS or OS between wild type and mutated RAS or BRAF and between MSS/MSI-L or MSI-H tumors (Fig. 4).
Impact of “prognostic grouping” (low risk T1-3/N1) and (high risk T4 or N2) in DFS and association with miR-21 expression
Stage III CRC shows a heterogeneous clinical behavior. Two main groups with different survival, T1-3/N1 and T4 or N2 groups, have been described [2]. Our results shows that there is a significant difference in DFS (
Since we found that miR-21 is associated with DFS we analyzed the effect of prognostic grouping and miR-21 expression (Fig. 5B and C). miR-21 expression levels higher than the cutoff point (N
The impact in DFS of miR-21 expression levels, prognosis group and age of diagnosis was confirmed by a multivariate analysis. The obtained results were HR 2.981: 95% CI 1.196–7.432,
Discussion
In the present study we have analyzed in patients with stage III CRC, the expression of a panel of miRNAs that was associated to the presence of metastatic disease in CRC [6]. Our results did not show any significant association between miR-135a, miR-206, miR-335 or miR-Let-7a expression levels and DFS or OS. However, we found that high levels of miR-21 correlated with a better DFS. This is conflictive, since a general consensus is that high level of tissue miR-21 expression is associated with poor DFS in patients with CRC [11, 12]. However, in many of these studies individual TNM stages are not considered, stages I to IV are pooled together, stage II is mixed with stage III. Only stage II is analyzed individually in some of these studies. Particularly stage III is the least represented and none of these studies analyzed stage III by itself [11, 12]. This may not be a reliable way to assess OS or DFS, since it is based on a population that is too heterogeneous. In this context, other reports described differences among stages of the effects of high miR-21 expression on DFS or OS. Bovell et al. [13] showed that increased expression of miR-21 correlated with shorter OS of only stage IV patients. Likewise, Kang et al. [14] reported that high miR-21 expression was mainly localized in the stroma and was associated with shorter DFS in stage II but not in stage III CRC. These studies are in accordance with our results. Furthermore, an interesting study in hepatocellular carcinoma shows that deletion of miR-21 in mice promotes the development of hepatocellular carcinoma that is associated with increased expression of oncogenes like Cdc25A [15]. Overexpression of Cdc25A correlates with poor prognosis in cancer and Cdc25A has been validated as a miR-21 target in colon cancer cell lines [16]. These results are in agreement with our data and question the general consensus of the association of high expression of miR-21 and poor clinical outcome.
Another factor to take into account is MSI, the presence of MSI-H is a good prognostic factor [17]. The combination of MSI-H and high levels of miR-21 could contribute to the better DFS we found in our population of stage III CRC patients. Interestingly, we found a significant correlation of MSI-H with high levels of miR-21, to our knowledge this has not been reported previously. In sporadic CRC, MSI-H is predominantly caused by the inhibition of the transcription of the mismatch repair genes, generated by promotor methylation. A different mechanism to decrease mismatch repair genes, could be triggered by miR-21, since it has been reported that miR-21 binds to the 3’-UTRs of MSH2 and MSH6 mRNA to repress protein translation of these mismatch repair genes [18]. This mechanism could explain the association that we found between high levels of miR-21 and MSI-H.
Our results showed a significant association of lower levels of miR-335 and MSI-H tumors, coinciding with Mjelle et al. [19] results, but we did not find an association of miR-335 levels and survival. However, the effect of miR-335 levels in CRC is unclear since some studies indicate miR-335 expression is higher in tumor tissues compared with their matched normal tissue and that miR-335 promotes tumorigenicity of CRC cells by inhibition of RASA1 and activation of the RAS/ERK pathway [20]. Other authors indicate that miR-335 expression levels are higher in tumors that metastasized compared with those that do not [6]. In contrast, other reports suggest that miR-335 inhibit invasion and metastasis in CRC and that miR-335 expression is lower in tumor tissues than in adjacent normal mucosa [21, 22] or report that serum miR-335 levels are lower in stage III and IV compared to stage I and II [23].
Our results did not show any significant association between miR-21, miR-206, miR-335 or miR-Let-7a expression levels and the presence of RAS or BRAF mutations but we found a trend with higher levels of miR-135a and the presence of mutations in RAS.
Post-hoc analysis of the PETACC-8 revealed that MSI and BRAF mutations were not prognostic factors. KRAS mutation was significantly associated with shorter DFS and OS. When the analysis was performed according to MSS/MSI-L or MSI-H, both KRAS and BRAF mutations were independent poor prognostic factors for MSS/MSI-L tumors, while in the MSI-H subgroup, KRAS was not prognostic and BRAF mutation was associated with longer DFS [17, 24]. We did not find significant differences in DFS or OS between wild type and mutated RAS or BRAF and between MSS/MSI-L or MSI-H tumors. A recent report of an ACCENT pooled analysis of 12 adjuvant trials focused on MSI stage III colon cancer showed that MSI-H tumors had better prognosis for N1 stage but not for N2, and the addition of oxaliplatin improved DFS and OS in stage III [25].
Nowadays, the most promising predictive factor for DFS in stage III colon cancer is the analysis of ctDNA after surgery and at the end of adjuvant chemotherapy [26], but it needs to be confirmed in large prospective trials.
Our study has several limitations including the small sample size with few patients in most of the analyzed subgroups and the heterogeneity of this colon cancer population. Only 22 patients had MSI-H tumors and 18 BRAF mutation. Therefore, the conclusions related to those groups need to be taken with caution.
In the future, exploratory studies like this need to be conducted in localized colorectal cancer in order to find prognostic and predictive factors that allow the clinicians to take individualized therapeutic decisions. Patients with stage II MSI-H tumors can be selected to surveillance avoiding adjuvant chemotherapy.
Conclusions
Our panel of miRNA expression tested in this study does not allow to select specific subgroups of patients in which changes in clinical practice can be done. Nevertheless, some data such as the association of high expression levels of miR-21 and better prognosis in the poor prognostic group may be of interest and could be explored in future prospective clinical trials.
Footnotes
Acknowledgments
This work was partially funded by: Fondo de Investigación Sanitaria (FIS) Instituto de Salud Carlos III Pi12/00172; IMMUNOTHERCAN Comunidad de Madrid S2017/BMD-3733; Fundacion Mutua Madrileña; Bayer Healthcare; Fundacion 2000 Merck-Serono.
We thank Inmaculada Ruiz from the Cooperative Spanish Group for the Treatment of Digestive Tumors (TTD) and Elena Molina from the Biobank of the Hospital Clinico San Carlos.
Author contributions
Conception: Javier Sastre, Eduardo Diaz-Rubio and Beatriz Perez-Villamil.
Interpretation or analysis of data: Tania Calvo-López, Mateo Paz-Cabezas, Patricia Llovet and Maria Dolores Ibañez.
For the interpretation of clinical data: Vicente Alonso-Orduña, J.M
Preparation of the manuscript: Tania Calvo-López and Beatriz Pérez-Villamil.
Revision for important intellectual content: Javier Sastre, Eduardo Diaz-Rubio and Beatriz Perez-Villamil.
Supervision: Javier Sastre, Eduardo Diaz-Rubio and Beatriz Perez-Villamil.
