Abstract
Uveal melanoma (UM) is the most common primary intraocular malignancy in adults with high metastasis rates. The O6-methylguanine DNA methyl transferase (MGMT) is involved in chemoresistance of Dacarbazine (DTIC) treatment. Our previous study found that the combination of oncolytic adenovirus H101 and DTIC in the treatment of UM cells shows a synergistic antitumor effect mainly though down-regulation of MGMT. MGMT knockdown by shRNAs increases the sensitivity of uveal melanoma cells to DTIC treatment. The protein hemostasis of MGMT is important for the antitumor effect of DTIC. Tripartite motif-containing protein 72 (TRIM72) belongs to the tripartite motif (TRIM) proteins family and was identified as a novel E3 ligase for MGMT, which interacts with and mediates the ubiquitination of MGMT. TRIM72 knockdown increases the protein levels of MGMT, while reduces the ubiquitination of MGMT. Further study indicated that MGMT is highly expressed in UM cells, and the protein levels of MGMT and TRIM72 shows a negative correlation. UM cells that ectopically expressing TRIM72 shows increased sensitivity to DTIC treatment, which is consistent with the antitumor affect exhibited by H101. These results suggest that TRIM72 is a promising therapeutic target for UM treatment.
Abbreviation
Introduction
Uveal melanoma (UM) is the most common primary intraocular malignancy in adults, and gives rise to metastases in 50% of cases [1, 2]. For the highly metastatic nature, it is an intractable disease with a low survival rates, despite adequate local treatment [3, 4]. Risk factors include light-coloured eyes, fair skin, congenital ocular melanocytosis, ocular melanocytoma and the BAP1-tumour predisposition syndrome [1]. Ocular treatment aims at preserving the eye and useful vision and, if possible, preventing metastases. Recently, surgery combined with chemotherapy, immune-therapy, gene therapy and other methods have been used to the treatment of UM [5].
Dacarbazine or DTIC (dimethyl-triazeno-imidazol carboxamide) is an alkylating agent, which results in DNA methylation, is commonly used to treat melanoma [6]. O6-methylguanine DNA methyl transferase (MGMT) is a DNA repair protein which transfers the methyl group from guanine to a cysteine residue, repairing the cytotoxic lesion induced by alkylating agents, such as DTIC [7]. The overexpression of MGMT is a key mechanism of cancers resist to alkylating agents [8]. In our previous studies, we found that combination of oncolytic adenovirus H101 and dacarbazine enhances antitumor ability against uveal melanoma cells through down-regulation of MGMT [9, 10].
Sequences of the shRNAs for MGMT and TRIM72
Sequences of the shRNAs for MGMT and TRIM72
Protein homeostasis is fundamental in normal cellular function and cell survival, and ubiquitination is an evolutionary conserved highly-orchestrated enzymatic cascade essential for normal cellular functions and homeostasis maintenance [11, 12, 13, 14]. The ubiquitination system relies on a defined set of cellular enzymes, among them, substrate-specific E3 ubiquitin ligases (E3s) are the most critical players [12]. Ubiquitination is involved in the regulation of many eukaryotic signaling pathways and aberrant ubiquitin signaling is currently an active area in cancer research [15]. So far, several mechanisms for the down-regulation of MGMT gene expression have been used in new strategies to decrease chemoresistance and enhance anti-tumor ability [16]. MGMT serves as an important target for cancer treatment. The regulation of MGMT protein homeostasis at post-translational level might be a promising field for uveal melanoma therapy.
In this study, TRIM72 was identified as a novel E3 ligase for MGMT, which mediates the ubiquitination and degradation of MGMT, thus favors the antitumor effect of DTIC. Therefore, TRIM72 may act as a potential therapeutic target for UM.
Cell culture, transfection and treatment
The normal human pigment epithelial cell line ARPE-19 and three human uveal melanoma (UM) cell lines 92.1, MuM2B and OMM1 were kindly gifted from Professor Xianqun Fan (Shanghai Jiao Tong University, China). All these cells were cultured in DMEM medium, supplemented with 10% FBS and 100 U/mL penicillin and 100mg/mL streptomycin (all from Gibco, USA), in a 37
Plasmids construction
The shRNAs for MGMT and TRIM72 were synthesized as oligos (Sangon, China), annealed and inserted into the pLKO.1 vector that was digested with EcoRI and AgeI (NEB, USA), the specific sequences for shRNAs seen in Table 1. The plasmids containing TRIM72 and MGMT were amplified from the cDNA of human HEK293T cells, and inserted into pCDH, pCADNA3.0, pGBKT7, pET22B or pGEX4T-1 vector. The plasmids of MGMT mutants were generated using the QuikChange Site-Directed Mutagenesis Kit (Stratagene, USA). Plasmids that expressing UBA1, UBCH5A, Ubiquitin and Usp2cc were kindly provided by Professor Yuexiang Wang (Chinese Academy of Sciences, Shanghai, China).
Cell viability assay
MuM2B or 92.1 cells were seeded into 96-well plates, and treated with DITC, H101, or both agents in combination. Cell viability was determined using Cell Counting Kit-8 (CCK8, Jiangsu, China) solution according to the manufacturer’s instructions at different time points, and the product was quantified spectrophotometrically at a wavelength of 450 nm using a microplate reader (Bio-Rad, USA). The absorbance is directly proportional to number of viable cells. The 0 h time point was defined as 6 h after the cells were seeded. All experiments were conducted with six replicates and repeated three times.
Yeast two-hybrid screening
The MGMT ORF (open reading frame) was cloned into pGBKT7, generating the bait plasmid, named pGBKT7-MGMT, which contained the in-frame fusion of GAL4 DNA binding domain. Yeast two-hybrid (Y2H) screening was performed by transforming yeast strain (Mav203 strain) that harbored bait vector, with the pACT2 prey vectors for human E3 ubiquitin-protein ligases cDNA expression library. Yeast transformants were first grown on to the plate on SD-2 (deficient in Leu, Trp) for selection of yeast cells containing both bait and prey vectors, and then transferred to SD-4 (deficient in Leu, Trp, His and Ura) plates to screen for E3 ligase proteins that potentially interact with human MGMT. The interaction was confirmed by transforming yeast Mav203 cells with the indicated bait and prey vectors, and allowing the transformants to grow on the SD-2 or SD-4 agar plates for approximately 3 days at 30
Co-immunoprecipitation (Co-IP), immunoprecipitation and immunoblotting
For Co-IP, cells were lysed in 500
Expression and purification of recombinant proteins
Recombinant proteins were purified as previously described [17]. Briefly, His6- or GST-tagged proteins were expressed in BL21 E.coli cells. After 400 mM isopropyl-
GST pull-down assay
Purified His6 tagged proteins (25
In vitro ubiquitination assay
In vitro ubiquitination assays were performed as previously described [18]. Briefly, recombinant 100 ng His6-Ub, 100 ng UBA1-His6 (E1), 100 ng UBCH5A-His6 (E2), 300 ng Gst-TRIM72 (E3) and 300 ng MGMT-His6 proteins were added to ubiquitination buffer (25 mM Tris-Cl, 100 mM NaCl, 1 mM DTT, 5 mM MgCl
Statistical analysis
The data were analyzed by one-way ANOVA with Tukey’s post-hoc test using GraphPad Prism 5 (GraphPad Software Inc., San Diego, USA). A
Results
Down-regulation of MGMT by oncolytic adenovirus (H101) or shRNAs enhances the sensitivity of uveal melanoma (UM) cells to Dacarbazine (DTIC) treatment
Our previous studies found that combination of oncolytic adenovirus H101 and dacarbazine (DTIC) enhances antitumor ability against uveal melanoma cells through downregulation of MGMT [9, 10]. This phenomenon was confirmed in two other uveal melanoma cell lines, as showed in Fig. 1A, combination of H101 and DTIC enhanced the antitumor ability against 92.1 and MuM2B cells. The protein levels of MGMT mainly down-regulated by H101 but not DTIC (Fig. 1B), and we also found that down-regulation of MGMT expression by H101 in a dose dependent manner (Fig. 1C). To further investigate the role of MGMT in uveal melanoma, three shRNAs targeting MGMT were designed and tested in 92.1 and MuM2B cells. Immunoblotting analysis indicated that MGMT was significantly decreased in cells stably transfected with shMGMT-1 and shMGMT-3 (Fig. 1D), and these two shRNAs were selected for further study. The cell viability of 92.1 and MuM2B cells that stably expressing MGMT shRNAs, treated with DTIC or not, was detected by the CCK-8 assay at different time points, and the results indicated that MGMT knockdown increased the sensitivity of uveal melanoma cells to DTIC treatment (Fig. 1E).
Down-regulation of MGMT by oncolytic adenovirus (H101) or shRNAs enhances the sensitivity of uveal melanoma (UM) cells to Dacarbazine (DTIC) treatment. (A) Combination of H101 and DTIC enhanced the antitumor ability against UM cells. Cell viability of PBS (Control), DTIC (5 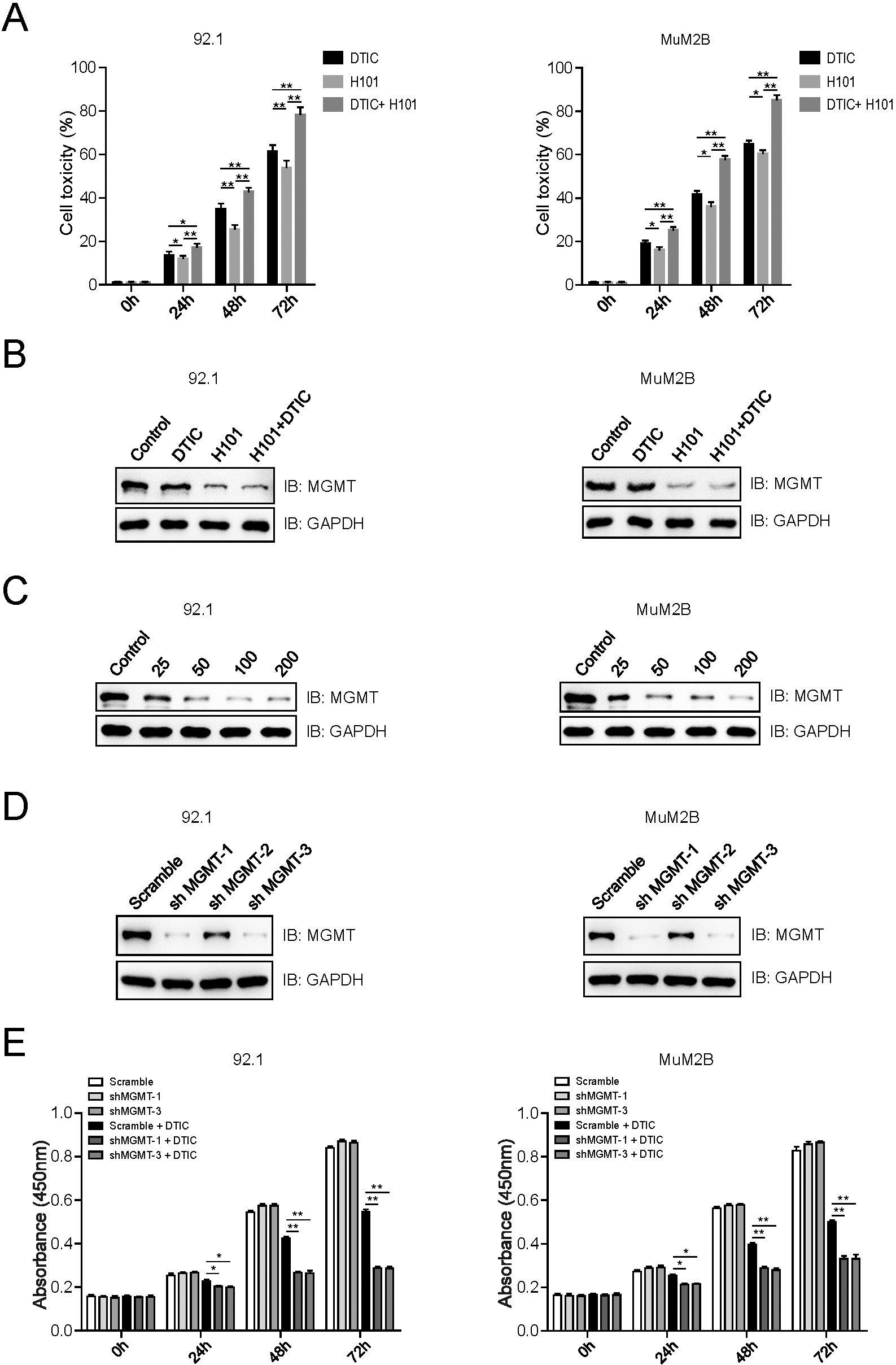
TRIM72 interacts with MGMT. (A) Yeast two-hybrid (Y2H) screening identified TRIM72 as an interacting partner for MGMT. MGMT was used as a bait. SD-2, deficient in Leu and Trp; SD-4, deficient in Ura, His, Leu and Trp. (B) Endogenous TRIM72 interacted with MGMT in UM cells. The lysates of 92.1 cells were immunoprecipitated with anti-IgG or anti-MGMT antibody, and subjected to immunoblotting analysis. (C) TRIM72 interacted with MGMT in vitro. Recombinant GST-tagged TRIM72 and His6-tagged MGMT were purified, and then GST pull-down assays were performed and subjected to immunoblotting analysis. PD, GST pull-down. (D) The N-terminal region of MGMT directly interacted with the RING domain of TRIM72. GST pull-down assays were performed with indicted recombinant GST tagged TRIM72, MGMT or their fragments, and His6 tagged TRIM72 or MGMT. RING, RING (short for Really Interesting New Gene) domain; B-BOX, B-Box-type zinc finger domain; PRYPSRY, the conserved B30.2/SPRY domain for TRIM proteins.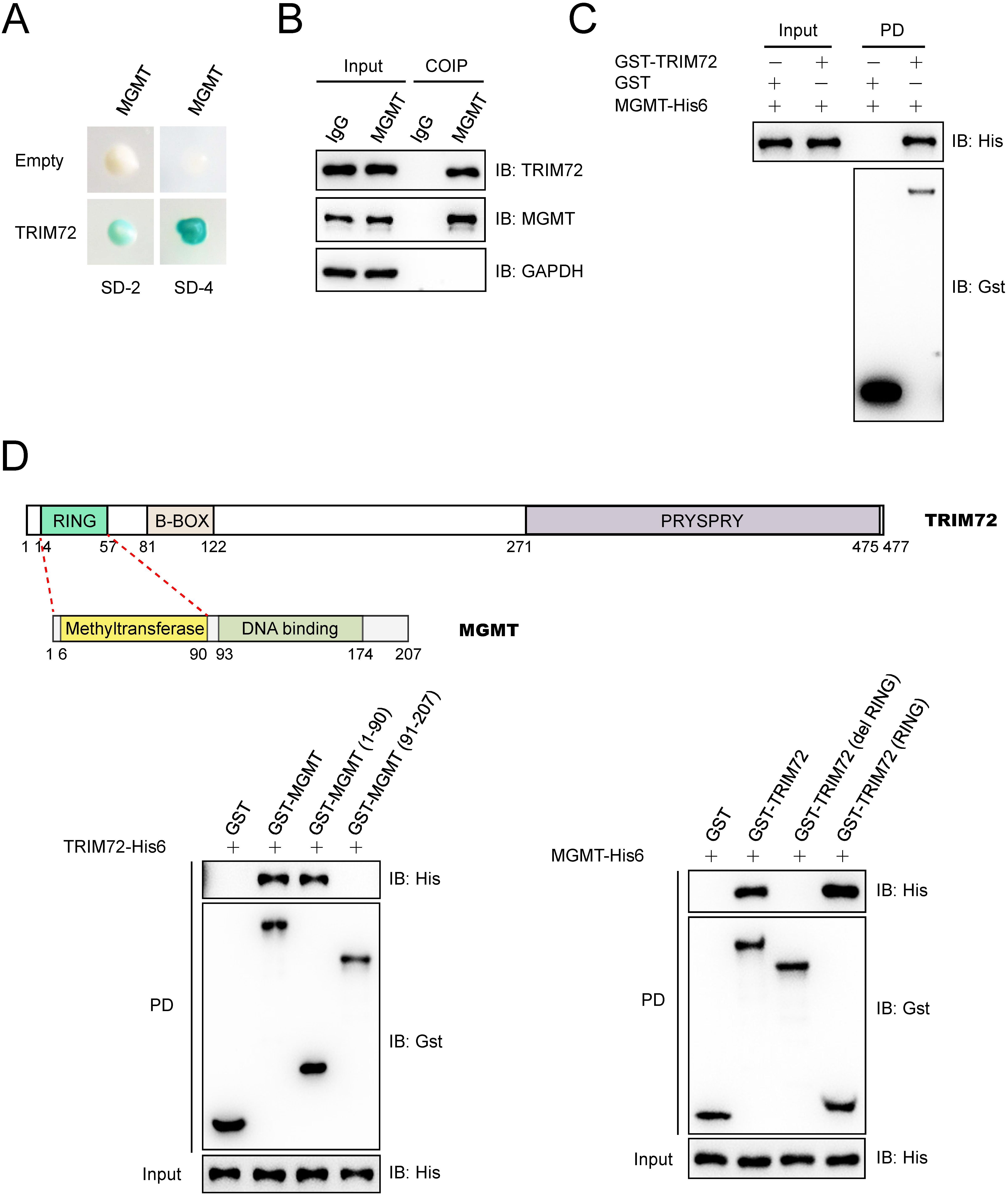
TRIM72 mediates the ubiquitination and degradation of MGMT. (A) The knockdown efficiency of shRNAs targeting TRIM72 was tested by immunoblotting analysis. The shRNAs for TRIM72 were transfected into 92.1 cells, and puromycin selection was used to establish stable expressing cell lines. The levels of TRIM72 were detected by immunoblotting analysis. (B) TRIM72 knockdown increased the protein levels of MGMT. The lysates of 92.1 cells that stably expressing with TRIM72 shRNAs were subjected to immunoblotting analysis with indicated antibodies. (C) TRIM72 promoted the degradation of MGMT in a dose-dependent manner. 92.1 cells that transiently transfected with different amounts of pCDNA3.0-TRIM72-Myc or empty vectors were subjected to immunoblotting analysis using indicated antibodies after treatment with 100 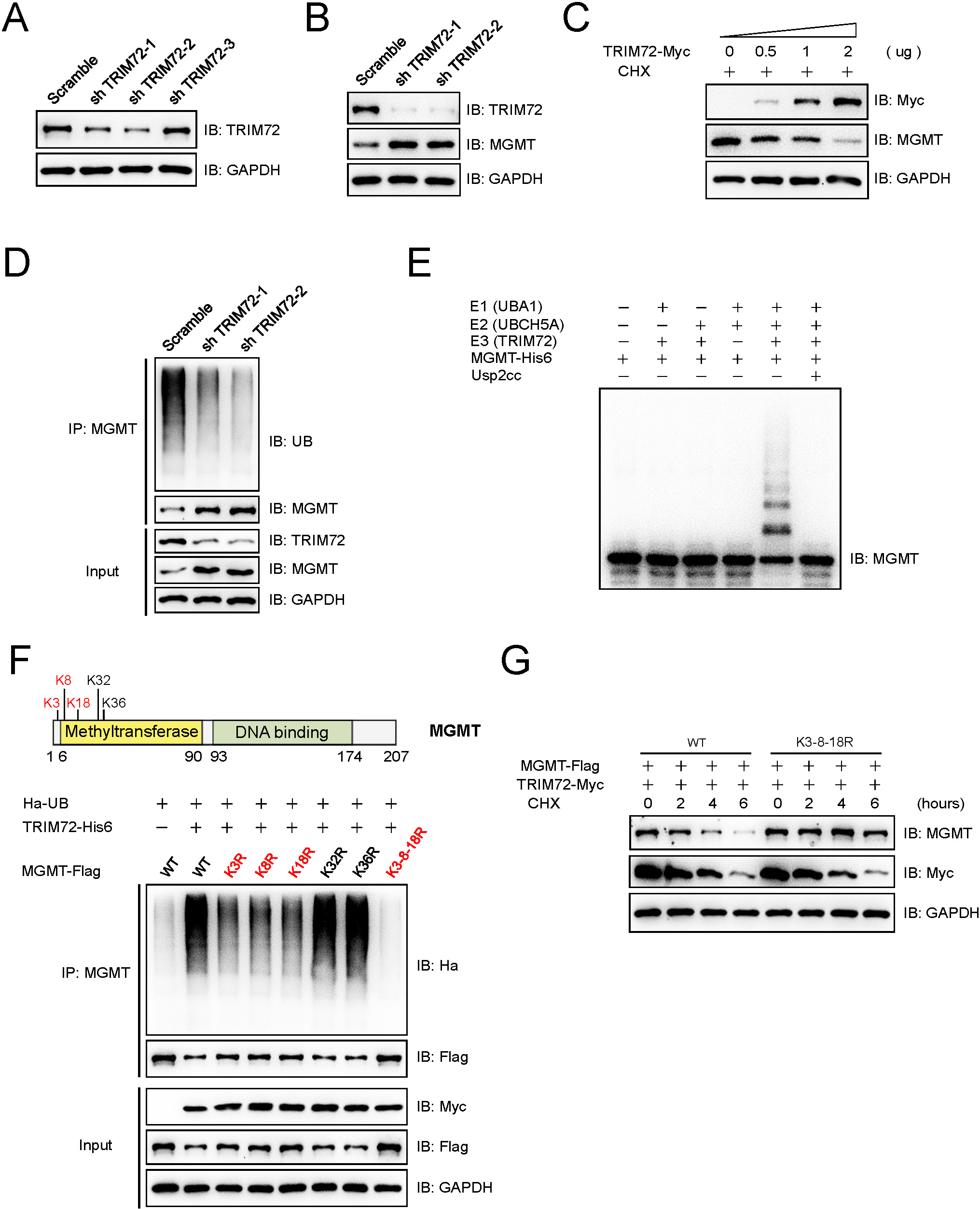
Ectopically expressing TRIM72 in UM cells increases their sensitivity to DTIC treatment. (A) Protein profile of TRIM72 and MGMT in human pigment epithelial cell line (ARPE-19) and uveal melanoma cell lines (92.1, MuM2B and OMM1) detected by immunoblotting analysis. (B) The protein levels of TRIM72 and MGMT in UM cells stably expressing TRIM72 were detected by immunoblotting analysis. 92.1 and MuM2B were transfected with empty vectors (PCDH) or vectors that expressing TRIM72 (PCDH-TRIM72), and puromycin selection was used to establish stable expressing cell lines. (C) Ectopically expressing TRIM72 in UM cells increases their sensitivity to DTIC treatment. 92.1 and MUM2B cells stably expressing TRIM72 were seeded into 96-well plates, treated with DTIC (5 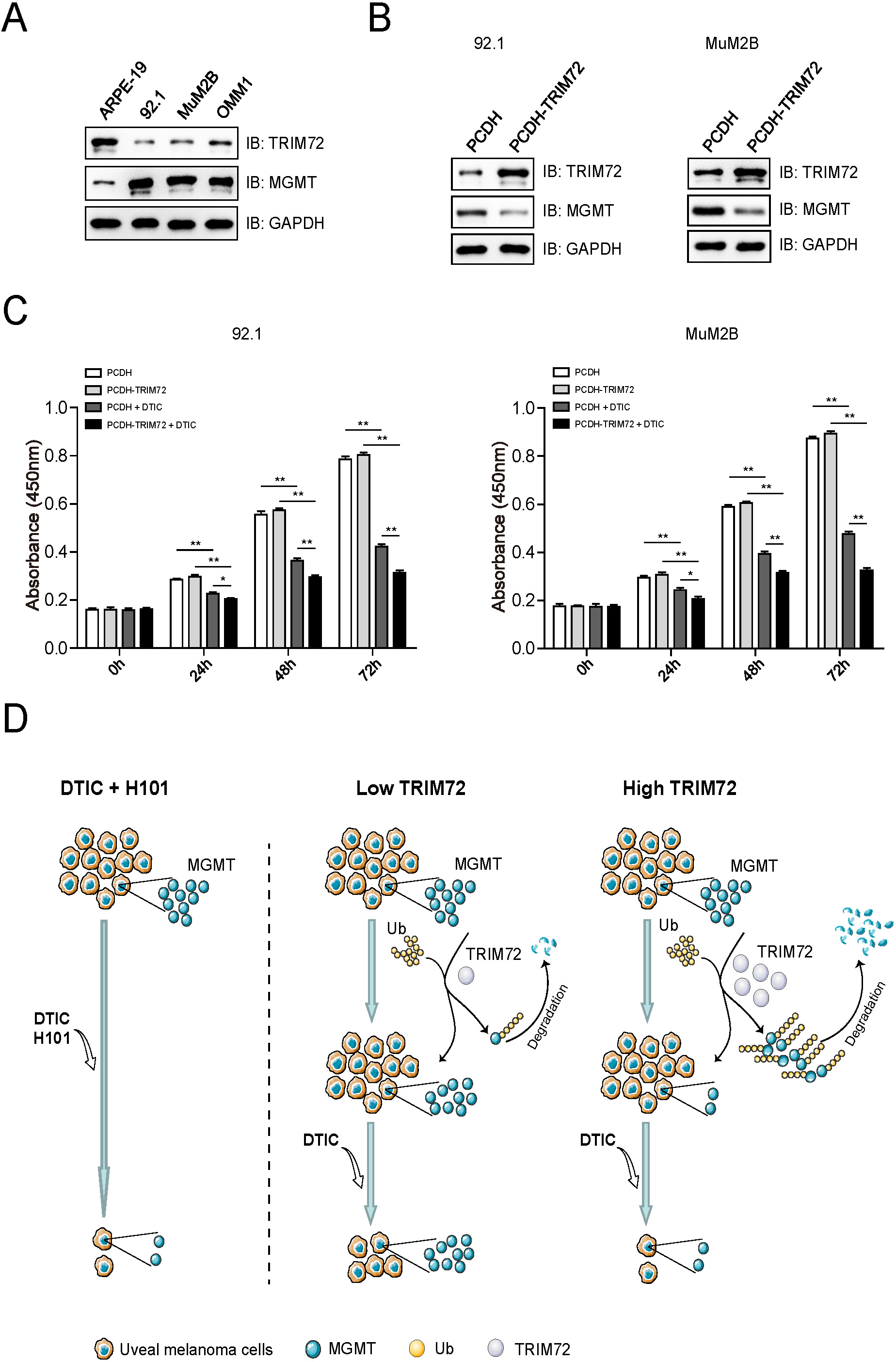
To explore how the homeostasis of MGMT protein is regulated in UM cells, yeast two-hybrid (Y2H) screening was performed to identify the potential E3 ligase for MGMT. Three proteins that interacted with MGMT were identified, and they are HACE1, MARCH2 and TRIM72. HACE1 and SYVN1 proteins are mainly located in the cytoplasm, while TRIM72 and MGMT mainly located in the nucleus of cells (The human Protein Atlas database), so TRIM72 was selected for further study. TRIM72 was identified as an interacting partner for MGMT and validated in yeast cells (Fig. 2A). Co-immunoprecipitation (Co-IP) assay showed that endogenous TRIM72 formed complexes with MGMT in 92.1 cells (Fig. 2B), and GST pull-down assay indicated that TRIM72 directly interacted with MGMT in vitro (Fig. 2C). Further study found that the N-terminal region of MGMT directly interacted with the RING domain of TRIM72 (Fig. 2D).
TRIM72 mediates the ubiquitination and degradation of MGMT
To explore the effect of TRIM72 on MGMT protein homeostasis, three shRNAs targeting TRIM72 were designed and tested in 92.1 cells. As showed in Fig. 3A, TRIM72 was significantly decreased in cells stably transfected with shTRIM72-1 and shTRIM72-2, and these two shRNAs were selected for further study. The protein levels of MGMT were obviously increased in TRIM72 knockdown 92.1 cells compared with the scramble group (Fig. 3B), and further study found that TRIM72 promoted the degradation of MGMT in a dose-dependent manner (Fig. 3C). Ubiquitination assay indicated that TRIM72 knockdown reduced the ubiquitination of endogenous MGMT in 92.1 cells (Fig. 3D). An in vitro ubiquitination assay was performed with indicated recombinant proteins, UBA1 as the ubiquitin-activating enzyme (E1), UBCH5A as the ubiquitin-conjugating enzyme (E2) and TRIM72 as the ubiquitin-protein ligase (E3), together with other components. Only when E1, E2 and E3 were all present, the ubiquitination of MGMT could be detected, and this signal efficiently eliminated by Usp2cc which is the catalytic core of human deubiquitinase Usp2 (Fig. 3E). Ubiquitin is usually covalently attached to the lysine (K) residues of targeting proteins, and there are five lysine (K) residues in the N-terminal region of MGMT which is essential for TRIM72 binding. Three Lys residues (K3, K8 and K18, shown in red color) in MGMT turned out to be the major sites for TRIM72 mediated ubiquitination in 92.1 cells, as the MGMT mutants that bears these individual Lys-to-Arg (K-to-R) substitutions showed attenuated ubiquitination, while the mutant bearing simultaneous K-to-R substitutions (K3-8-18R) at the three Lys residues almost totally abolished TRIM72 mediated ubiquitination on MGMT (Fig. 3F). Further study indicated that TRIM72 mediated the degradation of wild type MGMT but not K3-8-18R mutant in 92.1 cells (Fig. 3G), which again demonstrated the authenticity of these three sites.
Ectopically expressing TRIM72 in UM cells increases their sensitivity to DTIC treatment
Protein profile indicated that the expression of MGMT was higher in the three UM cell lines (92.1, MuM2B and OMM1) compared to the normal human pigment epithelial cell line (ARPE-19), while TRIM72 showed lower expression levels in UM cells compared to ARPE-19 (Fig. 4A). 92.1 and MuM2B cells that stably expressing TRIM72 were constructed, and the protein levels of TRIM72 and MGMT showed negative correlation (Fig. 4B). The cell viability of 92.1 and MuM2B cells that stably expressing TRIM72, treated with DTIC or not, was detected by the CCK-8 assay at different time points, and showed that ectopically expressing TRIM72 in UM cells increases their sensitivity to DTIC treatment (Fig. 4C).
Discussion
In this study, we found that combination of oncolytic adenovirus H101 and dacarbazine (DITC) enhances antitumor ability against 92.1 and MuM2B cells (Fig. 1A), which was consistent with our previous studies [9, 10]. H101 decreased the expression of MGMT was again verified (Fig. 1B), which enhanced the antitumor effect of DITC. The down-regulation of MGMT expression by H101 in a dose dependent manner and 50 MOI seems to be the most cost-effective concentration for UM treatments in this study (Fig. 1C).
H101, an oncolytic adenovirus with no E1B and portions of E3 regions, is designed to replicate in tumor cells that have inactivated p53 [9]. The p53 gene rarely has mutation in patients with UM and UM cell lines including 92.1 cells [19] which were involved in our study, while MuM2B cells carrying mutated p53 [20], but H101 showed antitumor effect on both of them, which was consistent with other studies that the replication of H101 may be independent of p53 [21, 22].
MGMT knockdown showed the enhanced sensitivity of UM cells to DTIC treatment (Fig. 1E), which was similar to the antitumor effect of H101 combined with DTIC. The down-regulation of MGMT serves as an important method for UM therapy. TRIM72 belongs to the tripartite motif (TRIM) proteins family, and accumulating studies indicate that these proteins regulate various biological processes [23], which was identified as a new E3 ligase for MGMT ubiquitination and degradation (Figs 2 and 3). TRIM72 interacts with and ubiquitinates the N-terminal region of MGMT that contains methyltransferase domain (Fig. 2D and F), so we speculate that the ubiquitination modification will also affect its catalytic activity, which need to be further study.
The expression of MGMT shows a significant association with DTIC chemoresistance. According to previous studies, high MGMT expression levels in tumor cells usually means pronounced resistance to DTIC [24, 25]. The expression of MGMT was higher in the three UM cell lines compared to the normal human pigment epithelial cell line (ARPE-19), while TRIM72 showed lower expression levels in UM cells compared to ARPE-19 (Fig. 4A). In UM cells, TRIM72 knockdown increased the protein levels of MGMT, while TRIM72 overexpression reduced the protein levels of MGMT (Figs 3B and 4B). UM cells that ectopically expressing TRIM72 showed increased sensitivity to DTIC treatment (Fig. 4C), which was consistent with the antitumor affect exhibited by H101. Thus TRIM72 acts as a promising therapeutic target for UM treatment, and an ongoing effort is to test the function of TRIM72-MGMT axis in animal models and patient sample.
Conclusions
The combination of oncolytic adenovirus H101 and DTIC in the treatment of UM cells shows a synergistic antitumor effect mainly though down-regulation of MGMT. TRIM72 was identified as a novel E3 ligase for MGMT, which interacts with and mediates the ubiquitination of MGMT. Ectopically expressing TRIM72 shows enhanced sensitivity to DTIC treatment, which was consistent with the antitumor affect exhibited by H101 (Fig. 4D).
Author contributions
Conception: Ping Chen and Biyun Cun.
Interpretation or analysis of data: Xun Li, Cong Yang, Ning Luo, Yunzhi Yang and Yan Guo.
Preparation of the manuscript: Ping Chen and Biyun Cun.
Revision for important intellectual content: Xun Li, Cong Yang, Ping Chen and Biyun Cun.
Supervision: Ping Chen and Biyun Cun.
Data availability
All the data generated or analyzed during this study are included in this published article and materials are available for other researchers after the manuscript is published.
Footnotes
Acknowledgments
This work was funded by the Joint Project of Yunnan Provincial Science and Technology Department and Kunming Medical University (2019FE001(-022)) and the National Natural Science Foundation of China (81301952).
Conflict of interest
The authors declare that they have no conflict of interest.
