Abstract
BACKGROUND:
Hepatocellular carcinoma (HCC) is one of the most common types of cancer in the world and a reason behind different oncogenes activation and tumor suppressor genes inactivation. Hyper-methylation of tumor suppressor genes including RASSF1a, GSTP1, p16, and APC cause gene silencing as well as tumor cell invasion. STAT 1 gene is a part of signaling cascade of JAK/STAT and any dysregulation in signaling has been implicated in tumor formation.
OBJECTIVE:
The current investigation focus on the methylation role of STAT1 gene as a non-invasive biomarker in the progression and diagnosis of hepatocellular carcinoma.
METHODS:
STAT1 gene methylation status in 46 HCV induced hepatocellular carcinoma patients and 40 non-HCC controls were examined by methylation specific PCR. STAT1 gene expression was examined by real time PCR and further validated by various bioinformatics tools.
RESULTS:
STAT1 methylation in HCV-induced HCC (67.4%) was significantly higher compared to the non-HCC controls (
CONCLUSION:
After using peripheral blood samples we observed that STAT1 silencing caused by aberrant methylation could be used as potential non-invasive biomarker for the diagnosis of HCV induced hepatocellular carcinoma. We conclude that blood as a sample source could be used instead of biopsy for early detection of HCC.
Schematic flow of the recruitment of study participants: All the cases were assessed for HCV and HBV viral profile by real time PCR; all selected HCC cases were positive for HCV and negative for HBV. All the subjects were recruited from Bait-ul-Sukon cancer hospital (Karachi, Pakistan). Samples were taken in EDTA tubes and stored at 4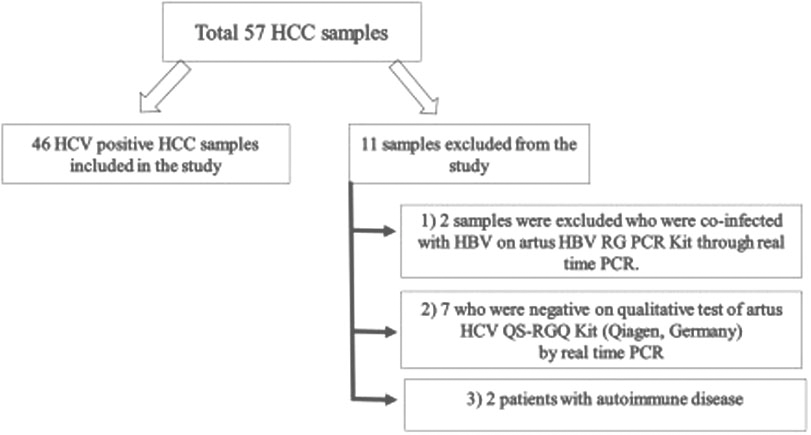
Hepatocellular carcinoma (HCC) is one of the most common types of cancer in the world and a reason behind different oncogenes activation and tumor suppressor genes inactivation [1, 2]. Various oncogenes like cyclin D1, c-myc and
Currently alpha fetoprotein (AFP) assay, tumor biopsy and ultrasonography (US) are considered standard methods for the diagnosis of HCC [14]. But due to low sensitivity, these both are not satisfactory methods for early detection of HCC [15]. Moreover, biopsy is considered invasive technique with a high risk of seeding tumor along the biopsy tract therefore; there is urgent need for the discovery of new effective sample source for the detection of HCC.
STAT 1 gene is a part of signaling cascade of JAK/ STAT [16] and any dysregulation in signaling has been implicated in tumor formation. STAT1 controls several cellular processes, such as the activity of antimicrobials, cell proliferation and has vital role in immune responses [17]. Previous Study shows that promotor hyper-methylation of STAT1 was observed in cervical cancer and gastric cancer patients [18, 19]. STAT1 function is still mystery in cancer biology, most of the research reveals tumor promoter effects of STAT1 while other evidence shows tumor suppressor role in cancer [16]. Another study revealed that STAT1 methylation in tumor tissue cause ovarian cancer [20]. Most of the studies were conducted on tissue samples however no study has been reported to find the correlation between STAT1 hypermethylation and expression in peripheral blood samples of HCV induced HCC. To the best of our knowledge present study is first to demonstrate the STAT1 gene silencing by hypermethylation in peripheral blood samples. That could be used as potential noninvasive biomarker for the diagnosis of HCV induced HCC.
The present study aimed to investigate the methylation status of STAT1 in peripheral blood of HCV induced HCC patients and cancer free controls by methylation specific polymerase chain reaction (MSPCR). Moreover, explore the correlation between STAT1 methylation and mRNA expression through real time RT PCR and the bioinformatics tool human protein atlas.
Materials and methods
Study subjects
A total of 57 HCC samples were used in this study. Among them, 11 patients were excluded under the exclusion criteria (Fig. 1). Remaining 46 HCV-induced HCC (28 males, 18 females; mean age 51.2
Primer sequences and conditions of STAT1 genes for MSP-PCR
Primer sequences and conditions of STAT1 genes for MSP-PCR
Genomic DNA was extracted using QIAamp DNA Mini Kit (Qiagen, Germany) as per manufacturer protocol. The quantity and purity of extracted DNA checked by NanoDropTM 1000 spectrophotometer (Thermo Fisher Scientific, Copenhagen, Denmark) and agarose gel electrophoresis. Extracted DNA (1
Methylation-specific PCR (MSP)
The bisulfite converted DNA is used as a template using primer specific for either methylated or unmethylated sequences and amplified by using EpiTect MSP kit (Qiagen, Germany) [20] Table 1. Negative and EpiTect positive control DNA (Qiagen, Germany) were included in each set of reaction. Amplified product was loaded on 2% agarose gels and visualized by gel documentation system (Bio-Rad, USA).
RNA extraction and RT PCR
RNA was extracted using QIAamp RNA blood mini Kit (Qiagen, Germany) as per manufacturer protocol. Further DNA digestion was performed using RNase-Free DNase set (Qiagen, Germany), that provides efficient on-column digestion of DNA. Total RNA was reverse transcribed by Revert Aid First Strand cDNA synthesis kit (Thermo Scientic, USA). STAT1 mRNA levels were assessed by real time RT-PCR using syber-green chemistry. After that, cDNA was added to primers and Ultra-Fast SYBR Green (Agilent, USA) and amplified using CFX96 thermal cycler (Bio Rad, USA). In each case cyclic threshold were obtained and expression levels were then evaluated by the
Insilico analysis of STAT1 gene
STAT-1 genes interacting partners were investigated using String 11 database (
RNA and protein expression of STAT1 gene in different types of cancer including liver cancer and STAT1 gene expression in different types of liver cells were evaluated by the human protein Atlas (
(a) The clinical history of HCC and healthy individuals: Elevated and suppressed levels of aminotransferase (ALT), alkaline Phosphatase (ALP), gama GT (GGT), total bilirubin, direct bilirubin, prothrombin time (PT), hemoglobulin (HG), platelets and albumin were expressed in mean 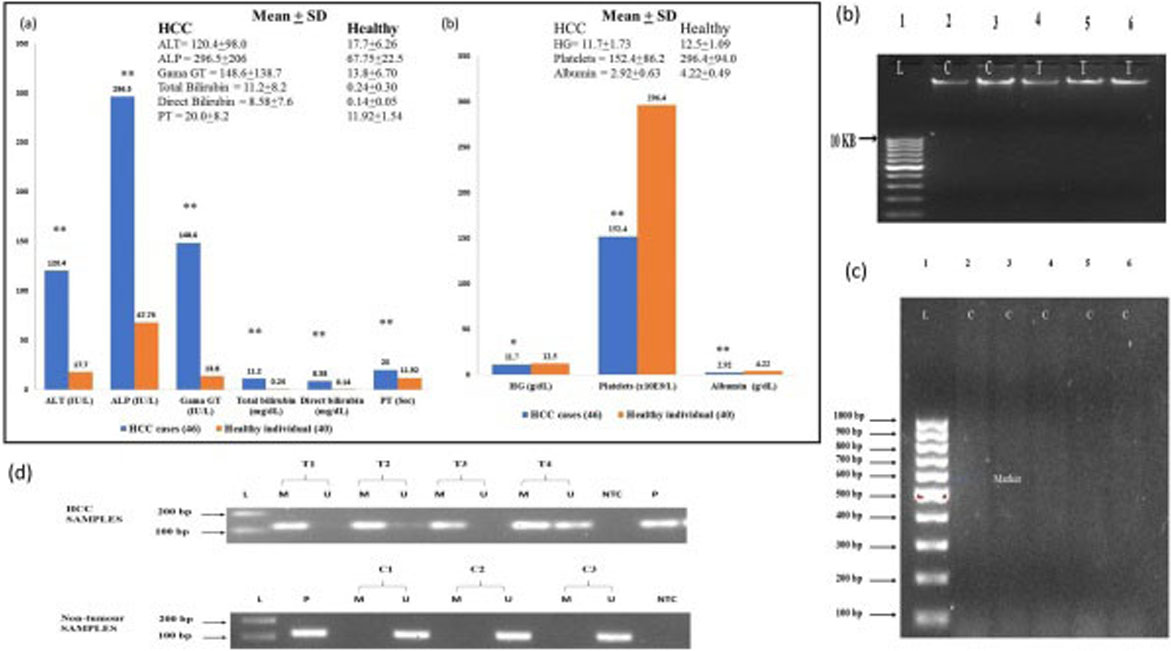
Data were illustrated as mean
Results
Clinico-pathological study
A total of 46 HCV induced HCC cases and 40 controls were included in this study (Fig. 2a). They underwent clinically indicated blood test and liver function test. Values of alanine aminotransferase (ALT), alkaline phosphatase (ALP), gama GT (GGT), total bilirubin, direct bilirubin and prothrombin time (PT) in HCC patients was higher than that in the control group (
Methylation results of STAT1 gene in disease and normal condition
Methylation results of STAT1 gene in disease and normal condition
STAT1 methylation, Co-evolution and alignment: Schematic diagram showed the domain of STAT1 and its various PTMs.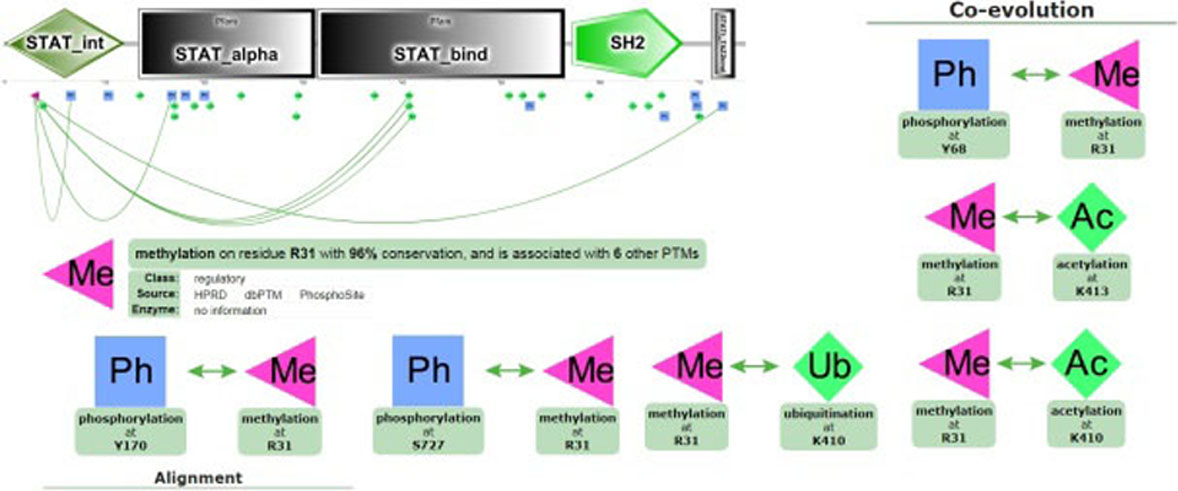
All patients’ characteristics were collected from the patients’ record (Fig. 2a) and the DNA was extracted from peripheral blood of HCC patients and healthy controls (Fig. 2b). Extracted genomic DNA was then subjected to bisulfite conversion and run on 2% agarose gel with a 100 bp marker. Bisulfite-converted DNA showed smearing pattern from 1,500 bp down to 100 bp shown in (Fig. 2c). To determine the STAT1 methylation, bisulfite-converted DNA was amplified with primers specific for CpG island promoter region of the STAT1 gene (Fig. 2d). According to our result aberrant DNA methylation in STAT1 was detected in 31 of 46 (67.4%) cases. In contrast 36 (90%) healthy control show no methylation signal Table 2. Result shows that STAT1 methylation was significantly higher in HCV-induced HCC as compared to healthy controls (
STAT1 methylation by PTMcode2
STAT1 methylation was examined by PTMcode2 on residue R31 with 96% conservation and its association with 6 other PTM. R31 methylation are associated with phosphorylation on residue Y68, S727, Y170; acetylation on residue K410, K413 and ubiquitination on K410 (Fig. 3).
STAT1 protein expressions in different types of cancer including liver cancer through the human protein Atlas (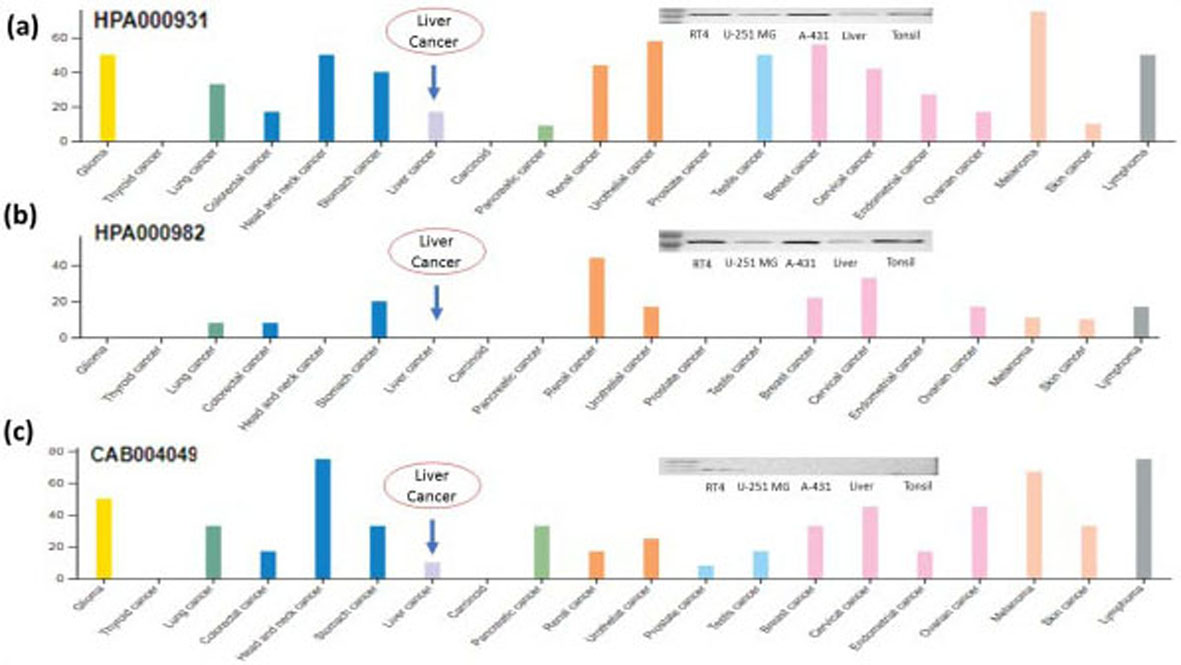
(a) Expression of STAT1 RNA in several cancers including liver cancer through the human protein Atlas (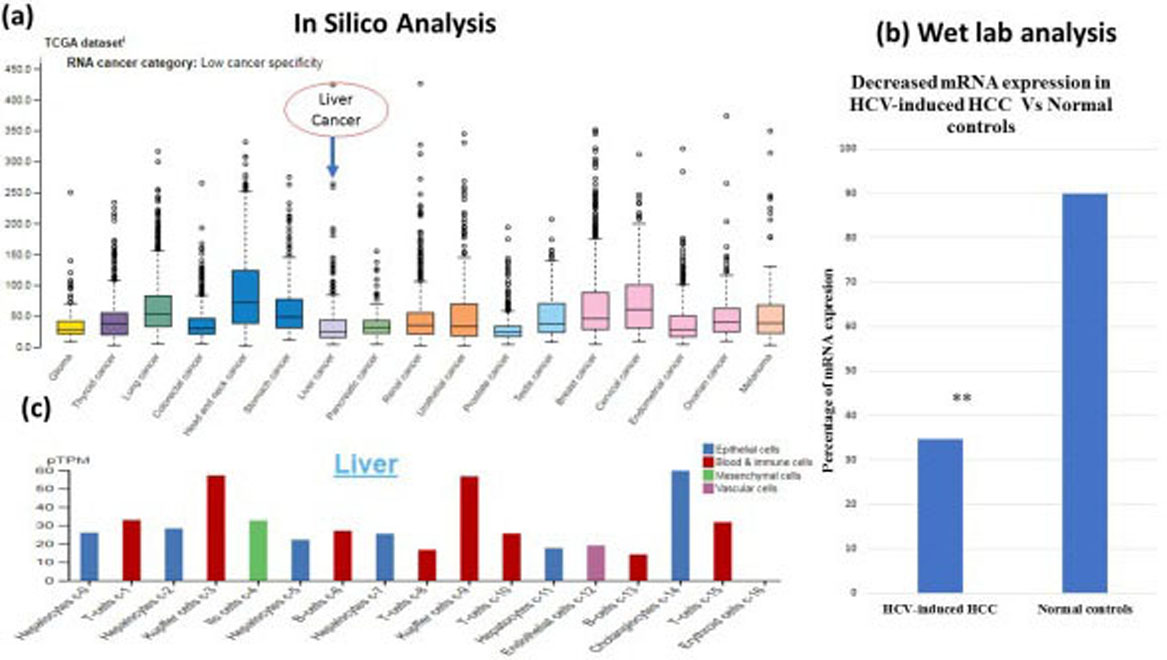
(a) Interaction of STAT1 protein with other interacting proteins by STRING 11 (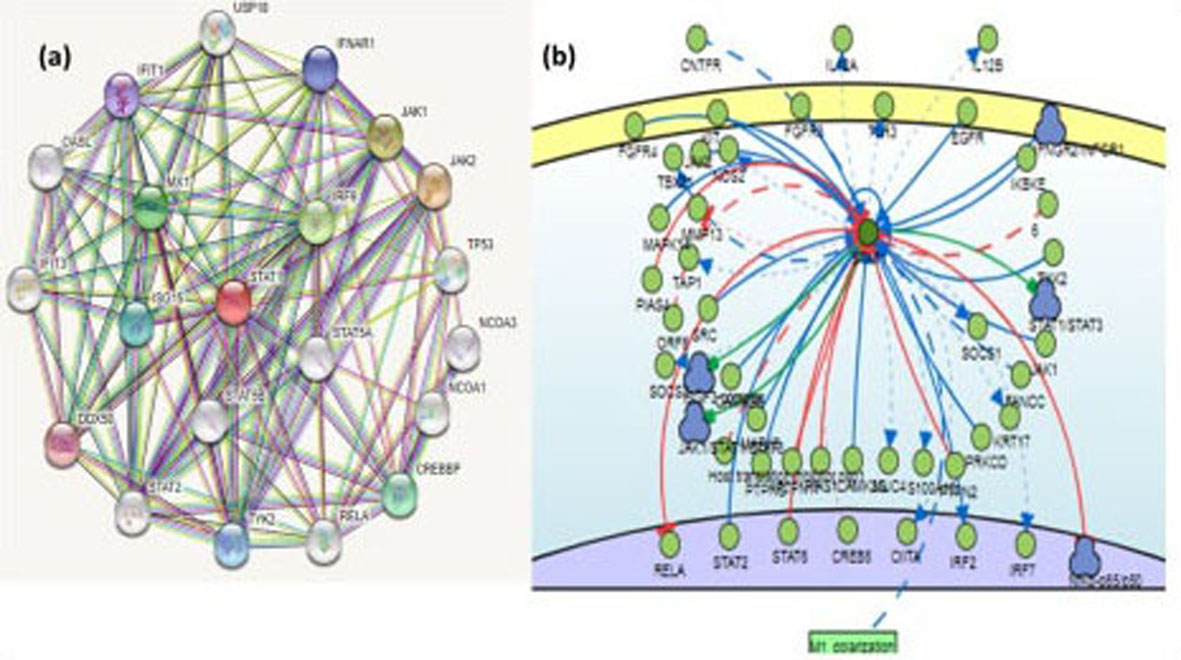
STAT1 RNA expression was examined by real time PCR. Our result shows that STAT1 mRNA expression level in HCV-induced HCC was significantly lower than that in normal individuals (
Insilico study of STAT1 protein and RNA expression
RNA and protein expression of STAT1 was evaluated by the human protein atlas. Protein expression in lymphomas, gliomas, melanomas, testis, urothelial, renal, stomach, cervical and breast cancers showed moderate to strong cytoplasmic and nuclear anti-STAT1 rabbit antibody HPA000931 positivity (Fig. 4a and b). Selected cases of most cancer tissues showed moderate to strong cytoplasmic positivity against another STAT1 antibody CAB004049 (Fig. 4c). Prostate, endometrial, thyroid, liver cancer and carcinoid were mainly negative against CAB004049 (Fig. 4c). Over all three Anti-STAT1 antibodies HPA000931, HPA000982, CAB004049 shows decrease expression of STAT1 protein in liver cancer and most abundant in lymphoid tissues (Fig. 4a–c).
STAT1 RNA was highly expressed in head and neck cancer with the median 71.8 (
Identification of the target gene of STAT1 constructing its regulatory network
Functional association network of STAT-1 with neighboring partner was developed through STRING 11 and Signor 2.0 interaction network. STAT1 gene interacts with activated signaling proteins of the tyrosine signaling including JAK family kinases. The predicted functional partners of STAT1 gene with confidence score are JAK2-1 (0.998), IRF9 (0.997), MX1 (0.996), CREBBP (0.995), ISG15 (0.997), TYK2 (0.994), IFNAR1 (0.993), IFIT1 (0.992), DDX58 (0.992) (Fig. 6a), FGFR4 (0.1354), FGFR3 (0.1364), KIT (0.5622), MAPK14 (0.2197), PIAS4 (0.3534), SRC (0.3586), CAMK2B (0.2197), ISGF3 COMPLEX (0.2197), RELA (0.783), STAT2 (0.971), STAT6 (0.2197), JAK1/STAT1/ STAT3 (0.1364), MAPK8 (0.1364), PTPRG (0.1364), PTPN11 (0.3586), PIAS1 (0.5653), CAMK2G (0.2197), IRF2 (0.1364), PTPN2 (0.4576), PRKCD (0.2197), STAT1/STAT3 (0.1364), NfKb-p65/p50 (0.1364), EGFR (0.2817), IKBKE (0.337) and IFNGR2/IFNGR1 (0.2197). Indirect interactions were observed with CNTFR (0.1364), TBX21 (01364.), ORF6 (0.1364), Host translation inhibitor nsp1 (0.2197), M1 polymerization (0.1364), FANCC (0.379), non-structure protein 6 (0.4203) (Fig. 6b).
Discussion
Hypermethylation of tumor suppressor genes such as RASSF1a [9], SMPD3 [22], DOK1 [9] and P15-INK4b [23] causes gene silencing, which leads to the induction of HCC [24, 25]. STAT1 is major gene in the Janus Kinase/signal transducer and activator of transcription signaling pathway [26]. JAK/STAT pathway has potential role in different tumors [27, 28]. Studies have shown that STAT1 expression is decreased in numerous cancers including head and neck cancer, breast cancer, leukemia and multiple myeloma [29]. In addition, STAT1 has been reported to prevent the growth of neoplastic and benign cells by regulating the expression of anti-proliferative genes, for instance BCL-xL and caspases [30].
Our data showed that, promoter methylation in HCV induced HCC individuals was significantly higher (
We divided our study in two steps (1) wet lab analysis and (2) insilico analysis. Firstly, we compile clinicopathological parameters data of HCV associated HCC patients and healthy individuals. Total bilirubin, ALT, ALP, GGT, direct bilirubin and PT was significantly higher in HCV induced HCC patient (
After that we performed major part of research by extracting genomic DNA from HCV associated HCC patients and healthy controls. Then bisulfite conversion of DNA leads to DNA fragmentation and limited base pairing that results in appearance of smearing pattern from 1500 bp to 100 bp on agarose gel shown in Fig. 2a–c. Finally, methylation in promoter region of STAT1 gene and its expression was detected through MSP PCR and real time PCR respectively. Aberrant methylation in STAT1 gene was detected in 31 (67.4%) cases in contrast to healthy controls 4 (10%), (
Furthermore, it is evaluated that STAT1 protein expression in liver cancer was lower with HPA000931, HPA000982 and CAB004049 antibodies (Fig. 4a–c). According to the results of human protein atlas, 2 of 12 patients with liver cancer have medium expression of STAT1 protein, 1 patient has low expression while 9 patients have no STAT1 protein expression (Fig. 4a–c).
Among all liver cells expression of STAT1 gene was higher in cholangiocytes c-14 with
Moreover, the predicted functional partner of STAT1 gene as by String 11 and Singor 2.0 with high confidence score were JAK2, JAK1, IRF9, MX1, CREBBP, ISG15, TYK2, IFNAR1, IFIT1 and DDX58., FGFR4, FGFR3, KIT, MAPK14, PIAS4, SRC, CAMK2B, ISGF3 COMPLEX, RELA, STAT2, STAT6, JAK1/STAT1/STAT3, MAPK8, PTPRG, PTPN11, PIAS1, CAMK2G, IRF2, PTPN2, PRKCD, STAT1/STAT3, NfKb-p65/p50, TYK2, EGFR, IKBKE and IFNGR2/IFNGR1 (Fig. 6a and b). Cellular responses to cytokine (KITLG/SCF), interferon and other growth factors are mediated by the STAT1 signal transducer and transcription activator. Interferon binding to cell surface receptors activates protein kinases, resulting in the activation of Jak kinases and the phosphorylation of tyrosine to form a complex called ISGF3 transcription factor complex (Fig. 6). This complex reaches the nucleus and the phosphorylated STATs dimerize and interact with ISGF3G/IRF-9. ISGF3 binds to the IFN Stimulated Response Element (ISRE) allowing an antiviral transcription of IFN-stimulated (ISG) genes that drive the cell. STAT1 is phosphorylated on tyrosine and serine residues in response to type II IFN (IFN-gamma). It then forms an IFN-gamma-activated factor (GAF) homodimer, moves into the nucleus and links to the IFN gamma-activated sequence (GAS). This accelerates target gene expression, inducing an antiviral cellular state and making it responsive towards KIT and KITLG/SCF signaling. Furthermore, it mediates cellular responses to stimulated FGFR gene family that directly belongs to the STAT family transcription factors [32]. The ERK/MAPK pathway generally involves a series of adaptors (Shc, GRB2, Crk, etc.) that connect the receptor to a guanine nucleotide exchange factor (SOS, C3G, etc.). This transmits the signal to small proteins that attach to GTP (Ras, Rap1), which in turn activates the central cascade unit composed of MAPK family (Raf). They enter the cytosol and relocate in nucleus may cause phosphorylation of numerous transcription elements, resulting in variable gene expression [33]. Furthermore, in many cytokine receptor systems, JAKs and STATs are critical components; they regulate development, survival, differentiation, and pathogen resistance [34, 35]. Our finding demonstrates that aberrant methylation status of STAT1 may results in transcriptionally silencing of gene and STAT1 promoter methylation may be a potential biomarker for HCC diagnosis.
Conclusion
Present study provided strong evidence that aberrant promoter methylation is the primary mechanism of STAT1 silencing in HCV induced HCC. Analysis of STAT1 methylation could be a potent noninvasive biomarker for HCC diagnosis. For future studies, it is suggested that the probability of tumor formation in liver might be reduced by using demethylating reagents for reducing the STAT1 hypermethylation.
Abbreviations
Hepatocellular carcinoma (HCC), Methylation specific PCR (MSP PCR)
Acknowledgments
We thank the staff of the Molecular Genetics Laboratory, Dr. Ziauddin hospital for helpful discussions and support.
Author contributions
RK and FN Conceived and designed the experiments, UZ and NNS performed the experiment. UZ and FN analyzed and interpreted the data, RK and NNS revised for important intellectual content and contributed reagents, material, analysis tools or data, UZ and RK prepare the manuscript.
Footnotes
Conflict of interest
None.
