Abstract
BACKGROUND:
Colorectal cancer (CRC), one of the most common human malignancies, is a leading cause of the cancer-related mortality. 5-FU is a first-line chemotherapeutic agent against CRC. Although CRC patients responded to 5-FU therapy initially, a part of patients succumbed to CRC due to the acquired drug resistance. Thus, investigating molecular mechanisms underlying chemoresistance will contribute to developing novel strategies against colorectal cancer.
OBJECTIVE:
Accumulation evidence revealed pivotal roles of long non-coding RNAs (lncRNAs) in tumorigenesis and chemoresistance of CRC. However, the precise roles and molecular mechanisms of lncRNA-HCG11 in CRC remain unclear. This study aimed to investigate the biological roles and underlying mechanisms of HCG11 as well as its molecular targets in regulating the cellular metabolism processes, which facilitate the chemoresistance of CRC.
METHODS AND RESULTS:
This study uncovers that HCG11 was significantly upregulated in CRC tumors tissues and cell lines. Moreover, HCG11 was elevated in 5-FU resistant CRC tumors. Silencing HCG11 inhibited colon cancer cell proliferation, migration, invasion and glucose metabolism and sensitized CRC cells to 5-FU. In addition, we detected increased HCG11 expression level and glucose metabolism in the established 5-FU resistant CRC cell line (DLD-1 5-FU Res). Furthermore, microRNA-microArray, RNA pull-down and luciferase assays demonstrated that HCG11 inhibited miR-144-3p which displays suppressive roles in colon cancer via sponging it to form a ceRNA network. We identified pyruvate dehydrogenase kinase 4 (PDK4), which is a glucose metabolism key enzyme, was directly targeted by miR-144-3p in CRC cells. Rescue studies validated that the miR-144-3p-inhibited glucose metabolism and 5-FU sensitization were through targeting PDK4. Finally, restoration of miR-144-3p in HCG11-overexpressing DLD-1 5-FU resistant cells successfully overcame the HCG11-faciliated 5-FU resistance via targeting PDK4.
CONCLUSION:
In summary, this study reveals critical roles and molecular mechanisms of the HCG11-mediated 5-FU resistance through modulating the miR-144-3p-PDK4-glucose metabolism pathway in CRC.
Introduction
Colorectal cancer, one of the most diagnosed malignancies, is associated with poor prognosis and high mortality rate [1]. Currently, surgical resection is the primary approach for CRC patients at early stage [2]. In addition, combination therapy with chemotherapy and/or radiotherapy reduced the risk of recurrence and improved overall survival of CRC patients at metastatic or advanced stages [3]. 5-fluorouracil (5-FU) is a traditionally first-line anti-cancer drug for CRC patients through interfering DNA synthesis, resulting in DNA damage and cancer cell death [4]. Over the last decades, 5-FU has been applied as an effectively chemotherapeutic agent for colorectal cancer [5]. However, development of intrinsic and acquired 5-FU resistance remains a major barrier to its clinical applications, leading to severe side-effects due to intolerable dosages [6, 7]. Therefore, understanding the precise mechanism of 5-FU resistance is critical to developing therapeutic strategies for colon cancer patients.
Long non-coding RNAs (lncRNAs) are a clan of endogenous RNAs with relatively large size (
The “Warburg effect” describes an aberrant glucose metabolism feature of cancer cells that they demand elevated glucose supply and metabolize the majority of glucose into lactate [19]. Moreover, reprogramming of glucose metabolism renders cancer cells advantages for survival under chemotherapeutic pressures [20]. Thus, targeting the dysregulated glucose metabolism could be a potential strategy to effectively treat cancer patients [20]. It was known that the pyruvate dehydrogenase kinase 4 (PDK4), which negatively regulates the convert of pyruvate to acetyl-CoA [21], plays important roles in glucose metabolism and cancer progressions. In this study, the roles of HCG11 in glucose metabolism and 5-FU resistance of colon cancer will be evaluated. Targets of HCG11 will be explored. These results manifested important roles of the HCG11-miR-144-3p-PDK4 axis in 5-FU resistance, offering novel insights for CRC patient treatment.
Materials and methods
Collection of CRC patient specimens
A total of forty paired human colon tumor specimens and adjacent normal tissues were obtained from CRC patients during surgery in the Department of Chinese PLA General Hospital (Beijing, China) from July 2016 to August 2018. This study was approved by the Institutional Review Board of Chinese PLA General Hospital. Written informed consent from patients was provided. CRC patients did not receive other local or systemic therapies prior to surgery. Patient samples were examined by three independent pathologists for the diagnosis of CRC cancers. After dissection, tissues were frozen immediately in liquid nitrogen and stored at
Cell culture and reagents
Human colorectal cancer cell lines DLD-1, LoVo, Caco2, HT-29, HT-116 and SW480, as well as normal colon epithelial cell FHC were obtained from the American Type Culture Collection (ATCC). Cells were maintained in RPMI-1640 medium (Invitrogen, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS) (Invitrogen, Waltham, MA, USA), 50 U/ml Penicillin and 50
Bioinformatics analysis
Bindings between HCG11 and miR-144-3p, miR-144-3p and PDK4 3’UTR were predicted from the starBase 2.0 of ENCORI:
Transfections of miRNA, siRNA and plasmid DNA
Colon cancer cells were seeded into 6-well plates at density of 5
RNA isolation and qRT-PCR
Total RNA was extracted from CRC tissues and cells using TRIzol Reagent (Invitrogen). Quality and quantity of RNA samples were assessed by Nanodrop. Complementary DNA (cDNA) was synthesized from extracted total RNA (1
RNA pull-down assay
The RNA pull-down assay was performed according to previous reports [25]. Briefly, HCG11 and its antisense RNA as well as a scramble control were biotin-labeled from RiboBio Co. Ltd (Guangzhou, China). Probs were incubated with CRC cell lysate for 2 hours. Mixture was incubated with Streptavidin-coupled agarose beads (Thermo Fisher Scientific, Shanghai, China) for 2 hours to down the RNA-RNA complexes. The amount of miR-144-3p in the RNA-RNA complexes was measured by qRT-PCR. Experiments were repeated three times.
Luciferase assay
Cells (5
MicroRNA microarray analysis
Total RNA was extracted using the RNeasy Kits (Qiagen) followed by purification using the RNeasy column (Qiagen) with a modified protocol. After quantification, RNAs were labeled by the Exiqon miRCURY LNATM arrays, Hy3/Hy5 Power Labeling Kit which enzymatically adds a fluorescent label to the 3’-end of the miRNAs, followed by hybridization onto an microRNA array in a SureHyb hybridization chamber (Agilent) according to the manufacturer’s protocols. Images were obtained using an Agilent microarray scanner. The Bioconductor package LIMMA and R program were performed to analyze the raw data and statistical analysis. The background-corrected intensity data were then normalized. MiRNAs that were up or down regulated more than 1.5-fold were selected as potential targets. Heatmap of the up or downregulated miRNAs was created using the GraphPad Prism 7 software (La Jolla, CA, USA).
Measurement of glucose metabolism
The glucose metabolism rate was assessed via glucose uptake assay, extracellular acidification rate (ECAR), and oxygen consumption (OCR). The glucose uptake was determined using a glucose test kit (Applygen Technologies, Beijing, China) according to the manufacturer’s instruction. The ECAR and OCR were measured using the Seahorse XF Glycolysis Stress Test Kit, XF Cell Mito Stress Test Kit and Seahorse XF96e analyzer (Agilent, Santa Clara, CA, USA) according to the manufacturer’s instructions. Results were normalized according to cell protein amount of each group. Experiments were performed in triplicate and repeated three time.
Wound healing assay
CRC cell motility capacity was determined by wound healing assay. Colon cancer cells (1
Transwell assay
Transwell chambers (Invitrogen, Waltham, MA, USA) were used to determine cell invasive capacity. Cell culture medium (0.5 ml) supplemented with 15% FBS was added to the center of chambers. Colon cancer cells (5
Cell viability assay
Cell viability in response to 5-FU treatment was determined by MTT assay (Sigma-Aldrich, Shanghai, China) according to the manufacturer’s instructions. Briefly, colon cancer cells (1
Cell apoptosis assay
Cell apoptosis/death rate of CRC cells in response to 5-FU treatment was determined using an Annexin V/FITC Apoptosis Detection Kit I (BD Biosciences, San Jose, CA, USA) according to the manufacturer’s instruction. After 5-FU treatments, cells were trypsinized and centrifuged at 2000 rpm for 3 minutes. Cells were washed by PBS then resuspended in binding buffer (300 ml). Cells were stained with 5 ml propidium iodide (PI) and 5 ml Annexin V FITC for 15 minutes in the dark. Data was obtained and analyzed using a FACScan flow cytometer and Flowjo software (BD Biosciences). Experiments were repeated three times.
HCG11 is upregulated in colon cancer. (A) Expression levels of HCG11 from CRC tumors (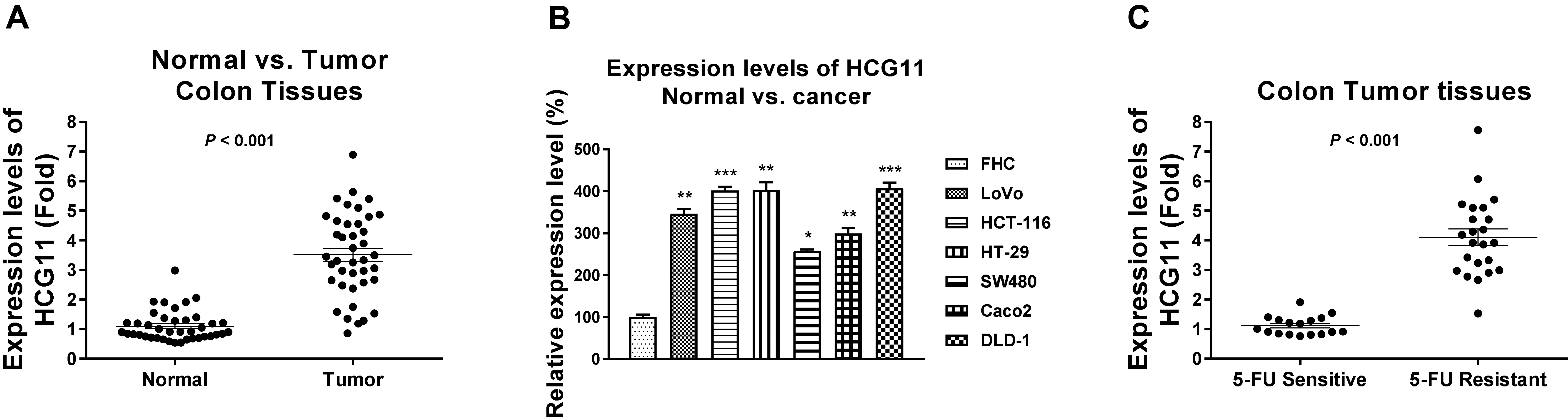
HCG11 is positively associated with CRC progresses and 5-FU resistance. (A, B) HCG11 was knocked down by siRNA in CRC cells. Cell proliferation rates were examined in 0, 24, 48 and 72 hours. (C) The above transfected cells were subjected to wound healing assay and (D) transwell assay. (E) The glucose uptake, lactate product and (F) oxygen consumption rate were examined in DLD-1 cells without or with HCG11 knockdown. (G) DLD-1 and (H) LoVo cells without or with HCG11 knockdown were treated with 5-FU at the indicated concentrations for 48 hours. Cell viability was determined by MTT assay. 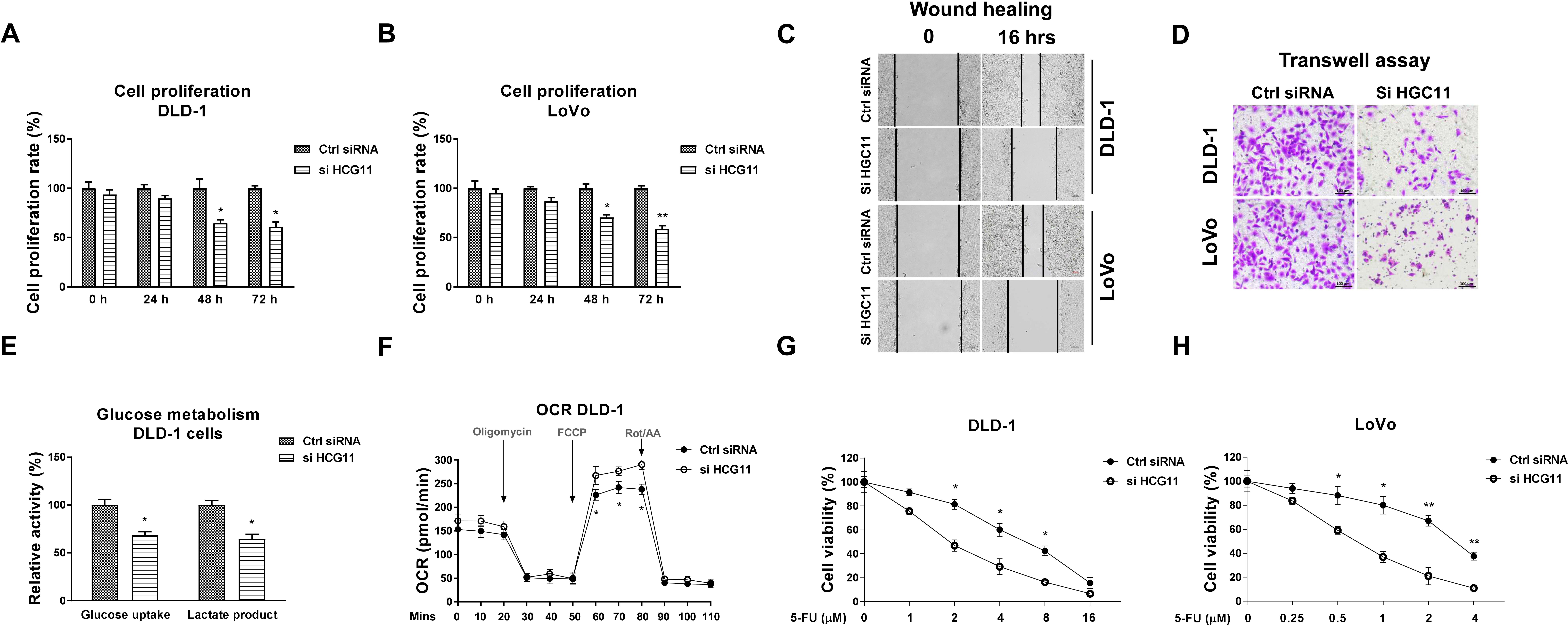
Total proteins were extracted from CRC cells using RIPA buffer (Thermofisher, Waltham, MA, USA) with 1
Statistical analysis
Data were analyzed using GraphPad Prism 7 software (La Jolla, CA, USA) and were displayed as mean
5-FU resistant colon cancer cells display upregulated glucose metabolism and HCG11. (A) The 5-FU sensitivities of DLD-1 parental and 5-FU resistant cells were determined by MTT assay and (B) clonogenic assay. (C) HCG11 expression levels were detected in DLD-1 parental and 5-FU resistant cells by qRT-PCR. (D) The glucose uptake, (E) extracellular acidification rate (ECAR) and (F) oxygen consumption rate (OCR) of DLD-1 parental and 5-FU resistant cells were measured. 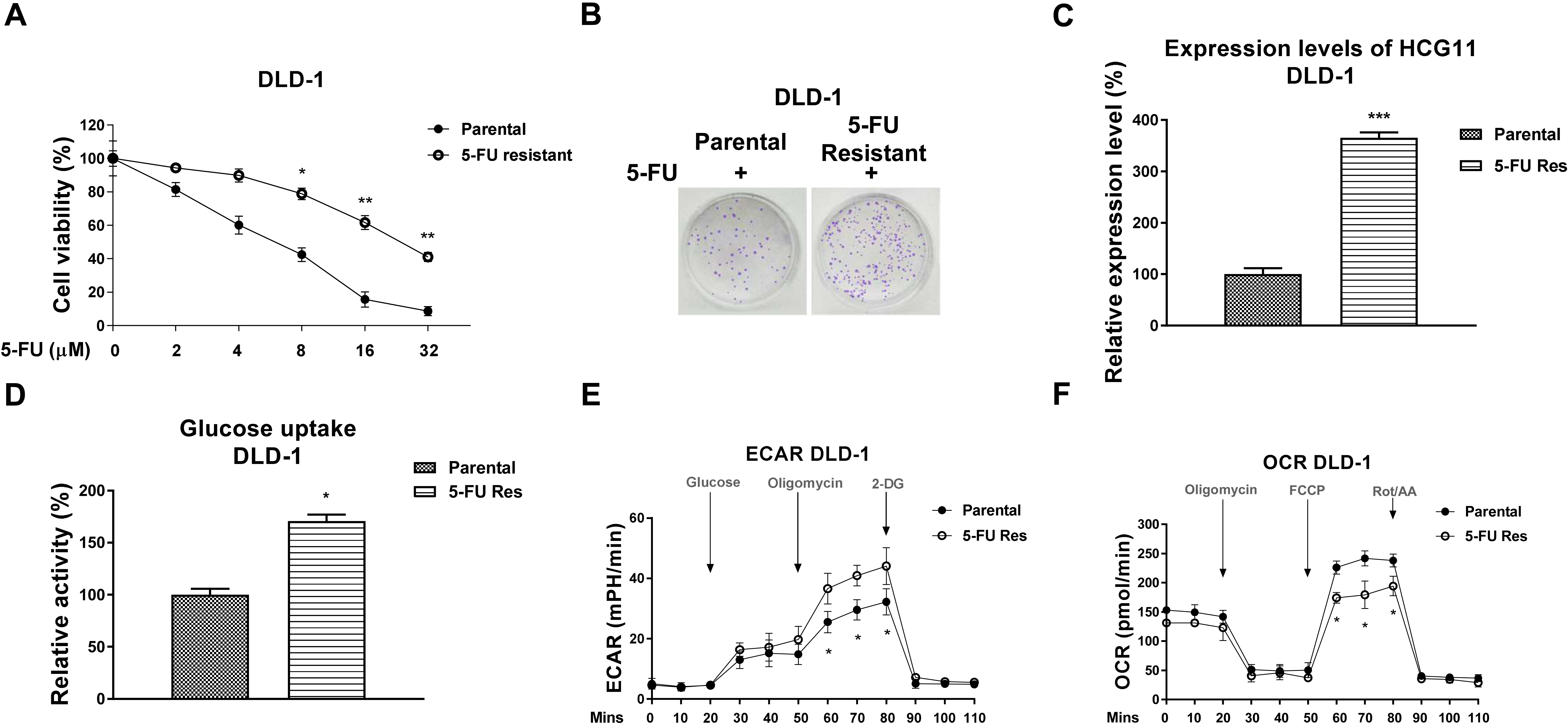
HCG11 is upregulated in colon cancer
Previous studies uncovered oncogenic roles of HCG11 in multiple cancers [10, 11, 12, 13, 14, 15, 16, 17, 18]. Yet, the precis functions of HCG11 in colon cancer have not been elucidated. To evaluate the biological roles of HCG11 in colon cancer, expression levels of HCG11 in 40 paired CRC and adjacently non-tumorous tissues were examined. Results from qRT-PCR demonstrated that HCG11 was significantly elevated in colon cancer tissues (Fig. 1A). Consistently, HCG11 was apparently upregulated in six CRC cancer cell lines compared with normal colon epithelial cell line, FHC (Fig. 1B). In addition, HCG11 was detected to be significantly upregulated in 5-FU resistant CRC tumor specimens compared with those from 5-FU sensitive CRC tumor tissues (Fig. 1C). Summarily, these results suggest that HCG11 plays oncogenic roles and is positively associated with 5-FU resistance of colon cancer.
Characteristics of CRC patients enrolled in this study
Characteristics of CRC patients enrolled in this study
HCG11 sponges miR-144-3p to downregulate its expression. (A) DLD-1 cells without or with HCG11 silencing were subjected to microRNA microArray analysis. (B) Results from microArray were validated by qRT-PCR in DLD-1 and LoVo cells. (C) Prediction of the binding of HCG11 with miR-144-3p from StarBase. (D) Pearson’s correlation coefficient analysis illustrated a negative correlation between HCG11 and miR-144-3p in CRC tumor specimens. (E) DLD-1 and LoVo cells were transfected with HCG11 siRNA for 48 hours, expression levels of miR-144-3p were detected by qRT-PCR. (F) Biotin pull-down assay was performed in DLD-1 and LoVo cells. Expression levels of miR-144-3p was assessed by qRT-PCR. (G) Dual-luciferase reporter assay was performed in DLD-1 and (F) LoVo cells with co-transfection of WT-HCG11 or Mut-HCG11 with control miRNA or miR-144-3p. 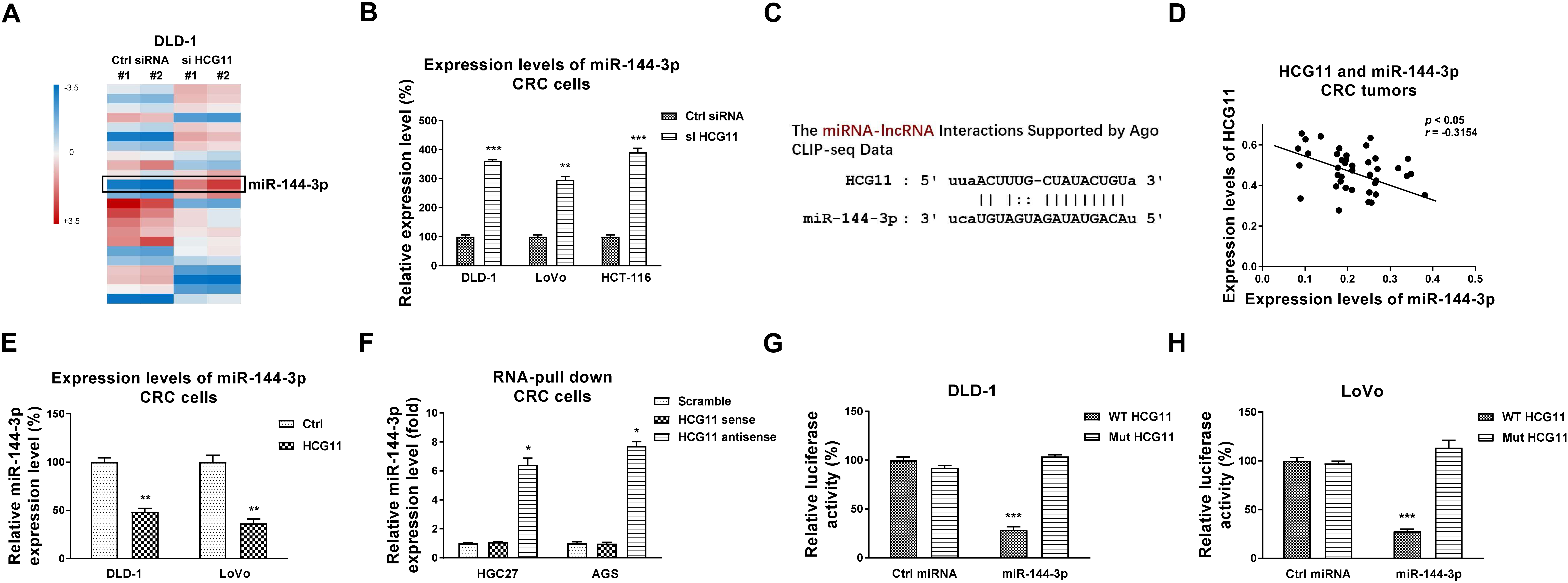
miR-144-3p is negatively associated with CRC progresses and 5-FU resistance. (A) Expression levels of miR-144-3p in normal or CRC specimens were analyzed from UALCAN. (B) Expression levels of miR-144-3p were detected in CRC tumor specimens (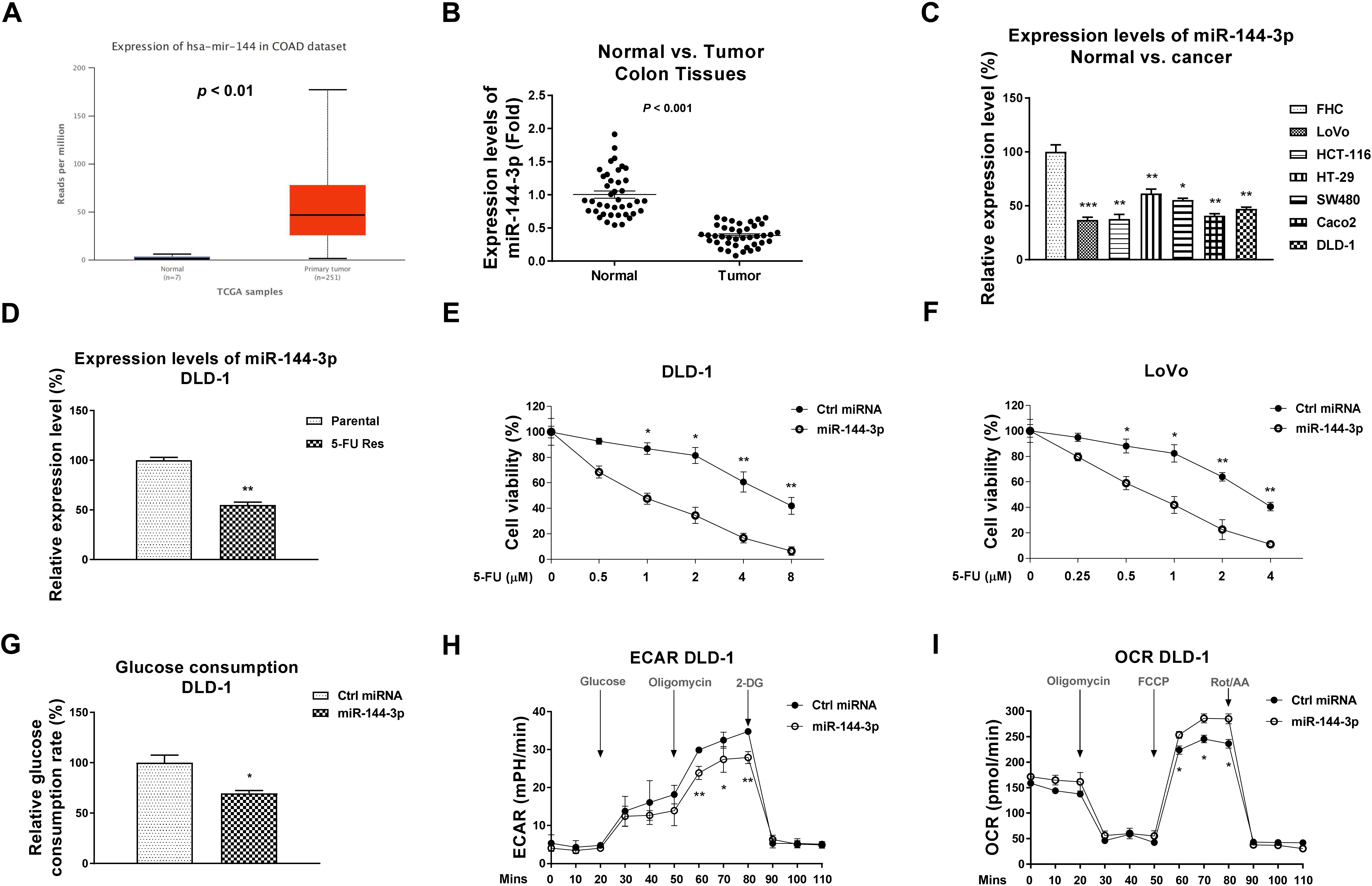
miR-144-3p directly targets PDK4 in CRC cells. (A) Prediction of miR-144-3p binding sites on 3’UTR of PDK4 from StarBase. (B) mRNA expression levels of PDK4 in CRC tumor specimens and adjacent normal colon tissues were detected by qRT-PCR. (C) DLD-1 and (D) LoVo cells were transfected with control siRNA or siPDK4 for 48 hours. Cell viabilities in response to 5-FU treatments were determined by MTT assay. (E) Pearson’s correlation coefficient analysis revealed a negative correlation between miR-144-3p and PDK4 mRNA in CRC tumor specimens. (F) DLD-1 and LoVo cells were transfected with control miRNA or miR-144-3p precursor for 48 hours, protein levels of PDK4 were determined by Western blot. (G, H) Dual-luciferase reporter assay was performed in CRC cells by co-transfection of luciferase vector containing 3’UTR of WT-PDK4 or Mut-PDK4 with control miRNA or miR-144-3p. 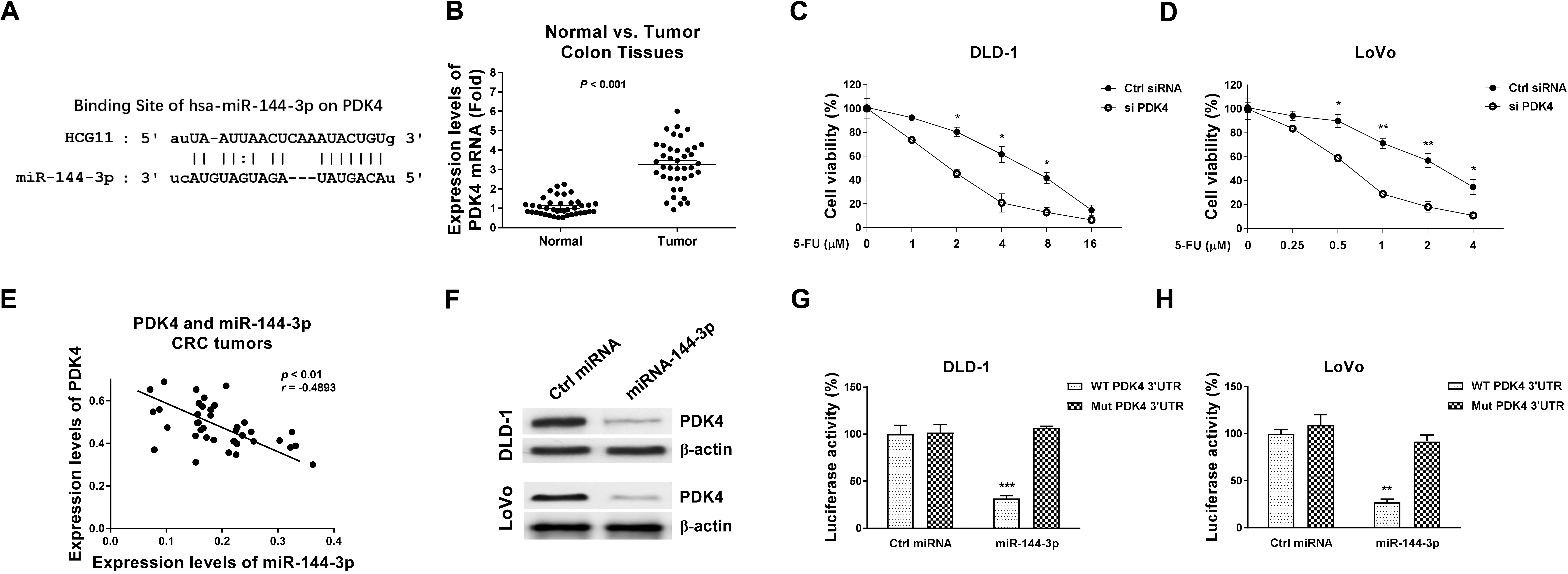
To investigate the direct functions of HCG11, LoVo and DLD-1 cells were transfected with siHCG11 to specifically knockdown its expressions (Fig. S1). Expectedly, CRC cells with HCG11 silencing displayed remarkedly inhibited cell proliferation rate (Fig. 2A and B), migration and invasion capacities (Fig. 2C and D). Given the dysregulated glucose metabolism affected the chemosensitivity of tumors [19], we investigated the roles of HCG11 in glucose metabolism of colon cancer cells. The glucose uptake and lactate production, two readouts for glycolysis reactions from HCG11-low expressing cells, were attenuated (Fig. 2E). Moreover, the oxygen consumption rate (OCR), which reflects the overall mitochondrial functions, was significantly increased in HCG11-silenced cells (Fig. 2F), suggesting HCG11 drives a glucose metabolic switch from oxidative phosphorylation to anerobic glycolysis in colon cancer cells. Given the acquired chemoresistance is a hallmark of cancer cells, we examined the effects of HCG11-knockdown in 5-FU sensitivity of CRC cells. Consistent with the above results, silencing HCG11 significantly sensitized CRC cells to 5-FU (Fig. 2G and H). The 5-FU IC50s of DLD-1 and LoVo control cells were 6.16
5-FU resistant CRC cells display upregulated HCG11 and glucose metabolism reprogramming phenotypes
We then explored the underlying mechanisms of the HCG11-promoted 5-FU resistance. A 5-FU resistant cell line (5-FU Res) was established from DLD-1 by selecting survival cells under stepwise increased concentrations of 5-FU treatments for 6 months. The DLD-1 5-Fu resistant cells displayed higher survival rate under 5-FU treatments compared with parental cells detected by cell viability assay and clonogenic assay (Fig. 3A and B). The 5-FU IC50 of DLD-1 5-FU Res cells was 24.4
HCG11 downregulates miR-144-3p in CRC cells via sponging it as a ceRNA
Considering that lncRNAs function through sponging miRNAs to block their expressions [8, 9], we investigated the molecular targets of HCG11 in 5-FU resistant colon cancer. A microRNA-microArray analysis was performed. Results in Fig. 4A demonstrated that among the miRNAs which were differentially regulated in HCG11-silencing DLD-1 cells, miR-144-3p was detected to be upregulated by HCG11 knockdown. The microArray results were further validated by qRT-PCR in DLD-1 and LoVo cells (Fig. 4B). Moreover, bioinformatics analysis showed miRNA-144-3p contains putative HCG11 binding sites (Fig. 4C), suggesting HCG11 might downregulate miR-144-3p via direct binding it to form a ceRNA network. To assess the clinical correlation between HCG11 and miR-144-3p, Pearson’s correlation coefficient analysis was performed. Expectedly, a significantly negative correlation between HCG11 and miR-144-3p was observed in CRC tumor specimens (Fig. 4D). Furthermore, DLD-1 and LoVo cells with lower HCG11 expression levels displayed significantly inhibited miR-144-3p expressions (Fig. 4E). To confirm the binding of HCG11 on miR-144-3p, an RNA-pull down assay was performed. qRT-PCR results clearly demonstrated that the biotin-labeled antisense HCG11 probe effectively pulled down enriched amount of miR-144-3p from DLD-1 and LoVo cells, compared with those from HCG11 sense RNA probe and scramble control (Fig. 4F). To validate whether HCG11 directly binds on miR-144-3p, luciferase assays were performed by co-transfection of CRC cells with luciferase vector containing wild type HCG11 (WT-HCG11) or binding site mutant HCG11 (Mut-HCG11) plus control miRNA or miR-144-3p. Expectedly, luciferase activities of CRC cells with co-transfection of WT-HCG11 plus miR-144-3p were significantly suppressed compared with that from the Mut-HCG11 plus miRNA-144-3p co-transfection (Fig. 4G and H). Taken together, the above results elucidated that HCG11 downregulates miR-144-3p in CRC cells through sponging it as a ceRNA.
Overexpression of miR-144-3p sensitizes colon cancer cells to 5-FU
To determine the effects of miR-144-3p on the 5-FU sensitivity in colorectal cancer, we examined miR-144-3p expression levels in CRC tumors specimens and normal colon tissues. As we expected, miR-144-3p was significantly downregulated in colon tumors (Fig. 5A and B). In addition, downregulated miR-144-3p was observed in six CRC cancer cell lines compared with normal colon epithelial cell line, FHC (Fig. 5C). MiR-144-3p was detected to be remarkedly low-expressed in DLD-1 5-FU resistant cells (Fig. 5D), suggesting miR-144-3p plays suppressive roles in 5-FU resistant colon cancer. Consequently, the effects of miR-144-3p overexpression on the 5-FU sensitivity were evaluated. CRC cells with miR-144-3p overexpression exhibited significantly increased 5-FU sensitivity (Fig. 5E and F). Moreover, overexpression of miR-144-3p effectively blocked the glucose uptake (Fig. 5G), ECAR (Fig. 5H) and OCR (Fig. 5I) in colon cancer cells. Summarily, the above results uncovered a tumor suppressive role of miR-144-3p in CRC cells. Moreover, overexpression of miR-144-3p contributes to sensitization of CRC cells to 5-FU.
Rescue of PDK4 overcomes the miR-144-3p-attenuated glucose metabolism and 5-FU sensitization in CRC cells. (A) DLD-1 cells were transfected with control, miR-144-3p alone or plus PDK4, expression levels of PDK4 were detected by Western blot. (B) Glucose uptake, (C) lactate product and (D) oxygen consumption were measured from the above transfected cells. (E) The above transfected cells were treated with 5-FU for 48 hours, 5-FU sensitivity was determined by MTT assay and (F) Annexin V apoptosis assay. 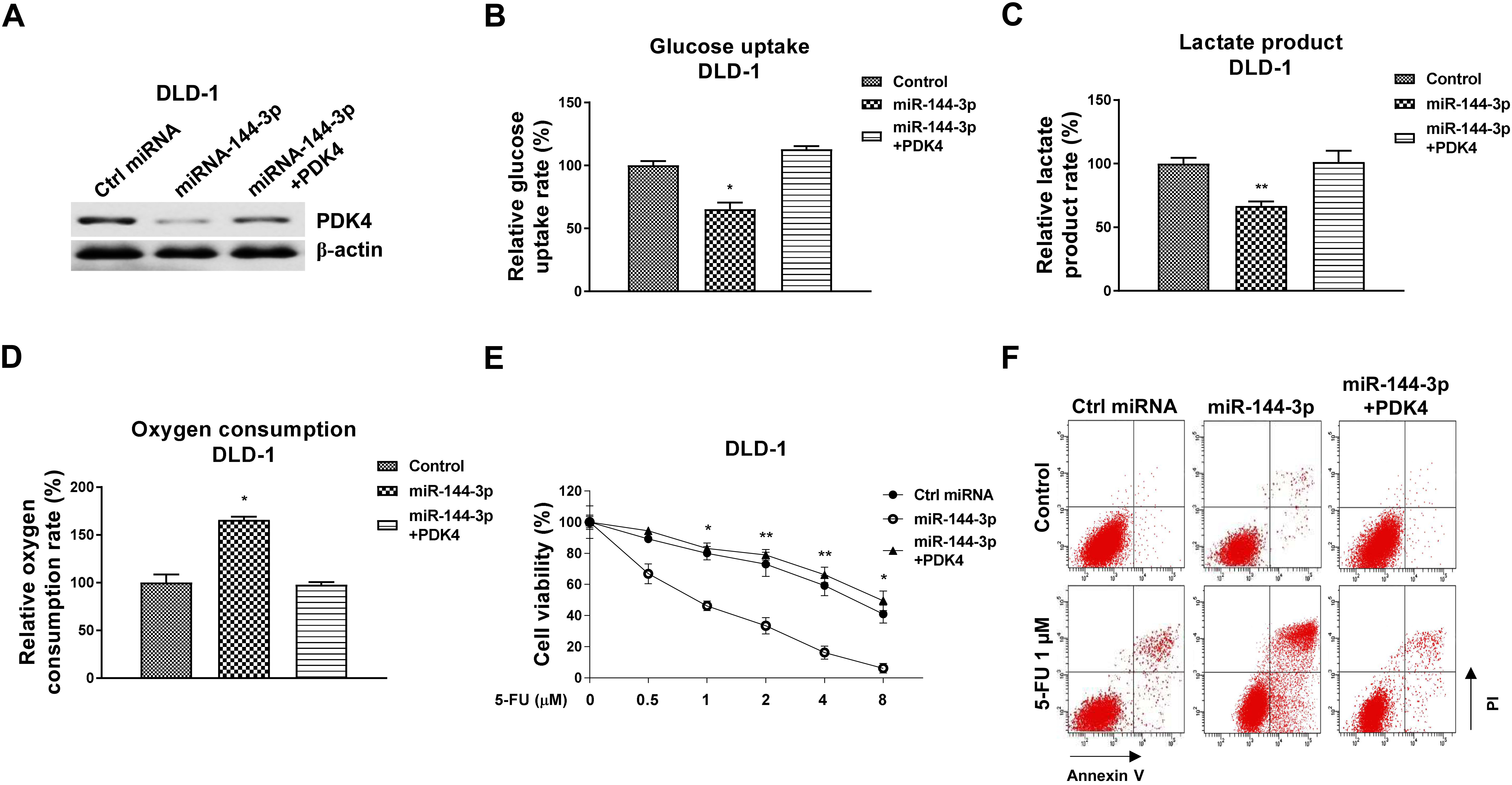
The HCG11-miR-144-3p-PDK4 axis in 5-FU resistance. (A) Correlation between HCG11 and PDK4 expression levels was analyzed from CRC tumor specimens and adjacent normal colon tissues. (B) DLD-1 5-FU Res cells were transfected with control, HCG11 alone or combined with miR-144-3p for 48 hours, expression levels of PDK4 were detected by Western blot. (C) Glucose uptake, (D) lactate product and (E) oxygen consumption from the above transfected cells were examined. (F) The above transfected cells were treated with 5-FU at the indicated concentrations, 5-FU sensitivity was determined by Annexin V apoptosis assay. 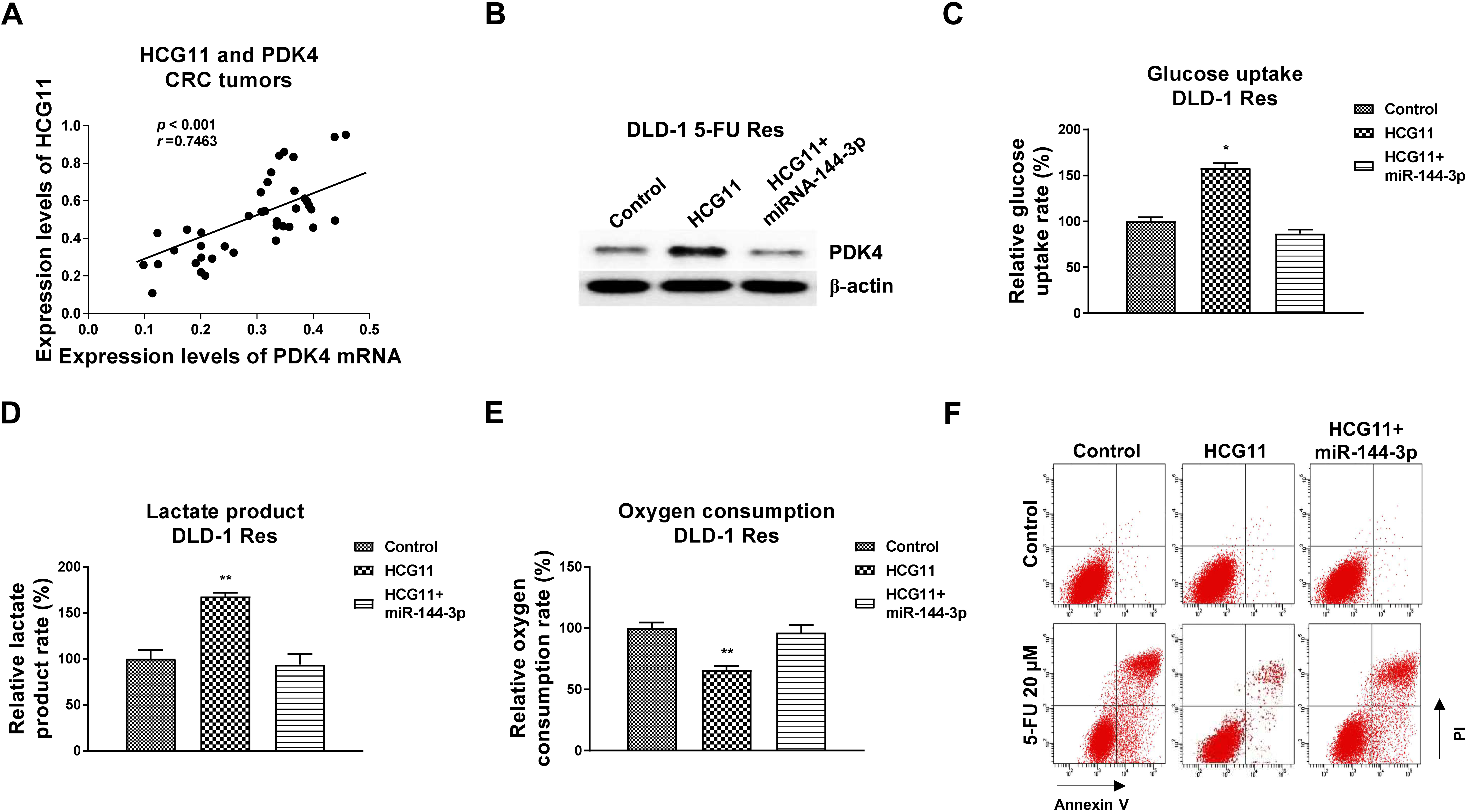
Accumulating studies reported that miRNA functions through binding to the 3’UTR of their target mRNAs, leading to blocking the mRNA transcriptions [26]. We then analyzed the mRNA targets of miR-144-3p from the non-coding RNA database, starBase2.0. Interestingly, we observed the 3’UTR of PDK4, which regulates the reaction of pyruvate to lactate or acetyl-CoA during the glucose metabolism [21], contains a putative miR-144-3p binding sites (Fig. 6A). We then assessed the clinical relevance of PDK4 in colon cancer. Expression level of PDK4 was significantly upregulated in CRC tumor specimens compared with normal colon tissues by qRT-PCR (Fig. 6B). To verify the biological roles of PDK4 in 5-FU sensitivity of CRC cells, PDK4 was silenced by siRNA in DLD-1 and LoVo cells. Expectedly, CRC cells with PDK4 silencing were more sensitive to 5-FU compared with control cells (Fig. 6C and D). We then evaluated the clinical correlation between miR-144-3p and PDK4. The Pearson’ correlation coefficient analysis demonstrated that higher PDK4 expression level was significantly associated with lower miR-144-3p expression level in CRC tumor specimens (Fig. 6E), supporting that miR-144-3p targets PDK4 in colon cancer. Consistently, overexpression of miR-144-3p effectively downregulated PDK4 protein expressions in CRC cells (Fig. 6F). To further validate whether miR-144-3p could target the 3’UTR of PDK4, luciferase reporter assays were performed by co-transfection of luciferase vector containing 3’UTR of WT-PDK4 or binding site mutant-PDK4 with control miRNA or miR-144-3p into CRC cells. Expected results demonstrated overexpression of miR-144-3p significantly blocked the luciferase activity of vector containing WT-PDK4 3’-UTR but not that of the binding site mutant PDK4 3’-UTR (Fig. 6G and H).
To investigate whether the miR-144-3p-modulated glucose metabolism switch and 5-FU sensitization were through targeting PDK4, rescue experiments were conducted. Transfection of PDK4 in miR-144-3p-overexpressing DLD-1 cells successfully rescued PDK4 protein expression level (Fig. 7A). Expectedly, restoration of PDK4 in miR-144-3p-overexpressing CRC cells significantly recovered the glucose uptake (Fig. 7B), lactate product (Fig. 7C) and oxygen consumption (7D). Furthermore, DLD-1 cells with PDK4 rescue showed decreased 5-FU sensitivity compared with those from cells with miR-144-3p overexpress alone by MTT assay and Annexin V apoptosis assay (Fig. 7E and F). Taken together, these data validated that miR-144-3p sensitizes CRC cells to 5-FU through targeting the PDK4-glucose metabolism pathway.
Inhibition of HCG11 sensitizes 5-FU resistant CRC cells via the miR-144-3p-PDK4 axis
We evaluated whether the HCG11-promoted 5-FU resistance was through the miR-144-3p-PDK4-glycolysis axis. Bioinformatics analysis from the lncRNA database, Starbase, indicated HCG11 expression level is positively associated with PDK4 in colon cancer (Fig. S2). Meanwhile, qRT-PCR results clearly verified that CRC patients with higher HCG11 expression level were associated with PDK4 upregulation (Fig. 8A). Thus, mechanism rescue experiments were performed by co-transfection of DLD-1 5-FU Res cells with control, HCG11 overexpression plasmid alone or plus miR-144-3p. Western blot results showed that overexpression of HCG11 significantly upregulated PDK4 expression level, which was further blocked by co-transfection of miR-144-3p (Fig. 8B). Expectedly, co-transfection of HCG11 with miR-144-3p recovered the glucose uptake (Fig. 8C), lactate product (Fig. 8D) and oxygen consumption (Fig. 8E). Consequently, under 5-FU treatments, we detected significant recovery of 5-FU sensitivity in miR-144-3p-rescued DLD-1 5-FU Res cells compared with that from cells with HCG11 overexpression alone (Fig. 8F). Summarily, the above results consistently demonstrated that the HCG11-promoted 5-FU resistance was through modifying the miR-144-3p-PDK4-glucose metabolism axis.
Discussion
Colorectal cancer is considered one of the most common human malignancies, leading to high death rate worldwide [1]. In the last two decades, the overall prognosis and survival rates of CRC patients have remarkedly improved, due to early screening and advanced therapeutic approaches [2, 3]. Despite 5-Fluoropyrimidine has been widely used in clinical treatment of CRC, development of intrinsic and acquired chemoresistance limits its clinical applications [4, 5]. Currently, while a number of studies described multiple molecules which are responsible for 5-FU resistance [6, 7], the precise mechanisms of 5-FU resistance in CRC are still under investigation. Therefore, understanding the precise mechanism of 5-FU resistance is critical for developing therapeutic strategies for colon cancer patients. This study uncovered non-coding RNA-modulated molecular targets and mechanisms of 5-FU resistance in colon cancer.
Accumulating evidence revealed that lncRNAs play vital roles in the occurrence and development of cancers [9]. However, the biological roles and mechanisms of HCG11 in colon cancer have not been investigated. Studies reported that HCG11 was dysregulated in variously malignant tumors [10, 11, 12, 13, 14, 15, 16, 17, 18]. Interestingly, HCG11 functioned as either oncogene or tumor suppressor in these studies due to tissue specific expressions [10, 11, 12, 13, 14, 15, 16, 17, 18]. Our present study for the first time showed that HCG11 was significantly upregulated in colon tumors and cell lines. Moreover, HCG11 expression level was positively associated with 5-FU resistance of CRC patients, suggesting HCG11 plays an oncogenic role in CRC. Silencing HCG11 effectively sensitized CRC cell to 5-FU. Consistent results demonstrated that HCG11 was significantly upregulated in 5-FU resistant colon cancer cells compared with parental CRC cells, indicating
HCG11 is a potential biomarker and therapeutic target of CRC.
Mounting studies reported that lncRNAs acted as sponges of miRNAs to downregulate their expressions [27]. Here we report miR-144-3p, which acts as a tumor suppressor in colon cancer cells, was a target of HCG11 in CRC cells. MicroRNA-microArray and RT-qPCR assay consistently demonstrated that overexpression of HCG11 significantly downregulated miR-144-3p expression levels. Subsequently, RNA-pull down and luciferase assay validated the direct bind of HCG11 with miR-144-3p in CRC cells. We further identified the glucose metabolism key enzyme, PDK4, as a direct mRNA target of miR-144-3p in CRC cells. Rescue experiments clearly verified the miR-144-3p-promoted 5-FU sensitization was through direct targeting PDK4.
Reprogramming of glucose metabolism characterized with increased anaerobic glycolysis and suppressed oxidative phosphorylation, a phenomenon called “Warburg effect”, was a new hallmark of cancer [19]. In addition, accumulation evidence manifested that dysregulated glucose metabolism is tightly correlated with chemoresistance [20], suggesting targeting glucose metabolism could be an effectively anti-cancer approach. From the established 5-FU resistant CRC cell line, we observed significantly upregulated glycolysis rate and decreased mitochondrial oxidative phosphorylation in 5-FU resistant cells. Consequently, HCG11 was detected to be upregulated and miR-144-3p was suppressed in 5-Fu resistant CRC cells compared with parental cells. Importantly, mechanisms rescue results showed silencing HCG11 suppressed glycolysis and recovered oxidative phosphorylation of CRC cells through regulating the miR-144-3p-PDK4 axis, suggesting targeting the HCG11-promoted glucose metabolism pathway could potentially overcome chemoresistance of CRC. This study still has limits that the current molecular mechanisms of the HCG11-promoted glucose metabolism and 5-FU resistance were assessed only by in vitro cell model. The in vivo xenograft mice experiments which could verify the in vitro results were lacking. In addition, since lncRNA and miRNA could target multiple molecules, we cannot exclude other signaling pathways which are potentially regulated by HCG11 or miR144-3p in the 5-FU sensitivity of CRC cells.
In summary, we demonstrate that lncRNA HCG11 plays oncogenic roles in 5-FU resistant CRC. Mechanically, HCG11 sponges miR-144-3p as a ceRNA, leading to de-repression of the miR-144-3p target, PDK4 in colon cancer. Our discovery will contribute to the development of novel approaches for treatment of chemoresistant CRC.
Authors’ contributions
Quan-Li Han: Investigation, Methodology, Writing – review& editing; Mu-Hong Deng and Zhi Cui: Investigation, Methodology; Qi Wang: Statistical Analysis.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210212.
sj-docx-1-cbm-10.3233_CBM-210212.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210212.docx
Footnotes
Acknowledgments
None.
Conflict of interest
The authors declare no competing financial and non-financial interests.
