Abstract
BACKGROUND:
Due to the heterogeneous nature of Diffuse Large B-cell Lymphoma (DLBCL), the mechanisms underlying tumor development and progression have not yet been fully elucidated.
OBJECTIVE:
This study aimed to compare the characteristics of plasma exosomes of DLBCL patients and healthy individuals and to evaluate the exosomal interactions between DLBCL cell lines and normal B-cells.
METHODS:
Exosome isolation was performed using an ultracentrifugation-based protocol from plasma of 20 patients with DLBCL and 20 controls. The expression of miRNAs from exosome samples was analyzed using a miRNA expression microarray. The presence of exosome-mediated communication between the lymphoma cells and normal B-cells was determined by the co-culture model.
RESULTS:
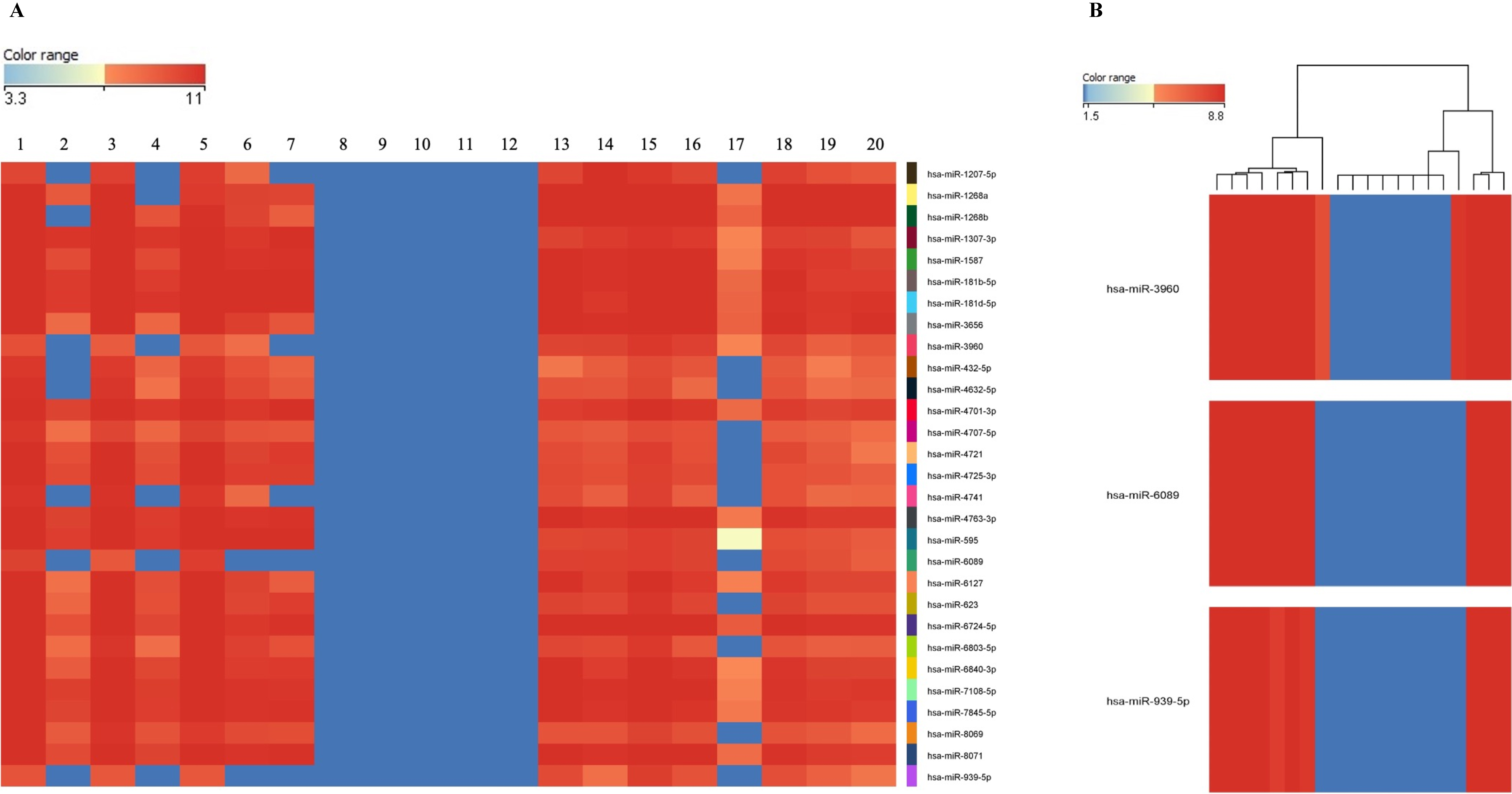
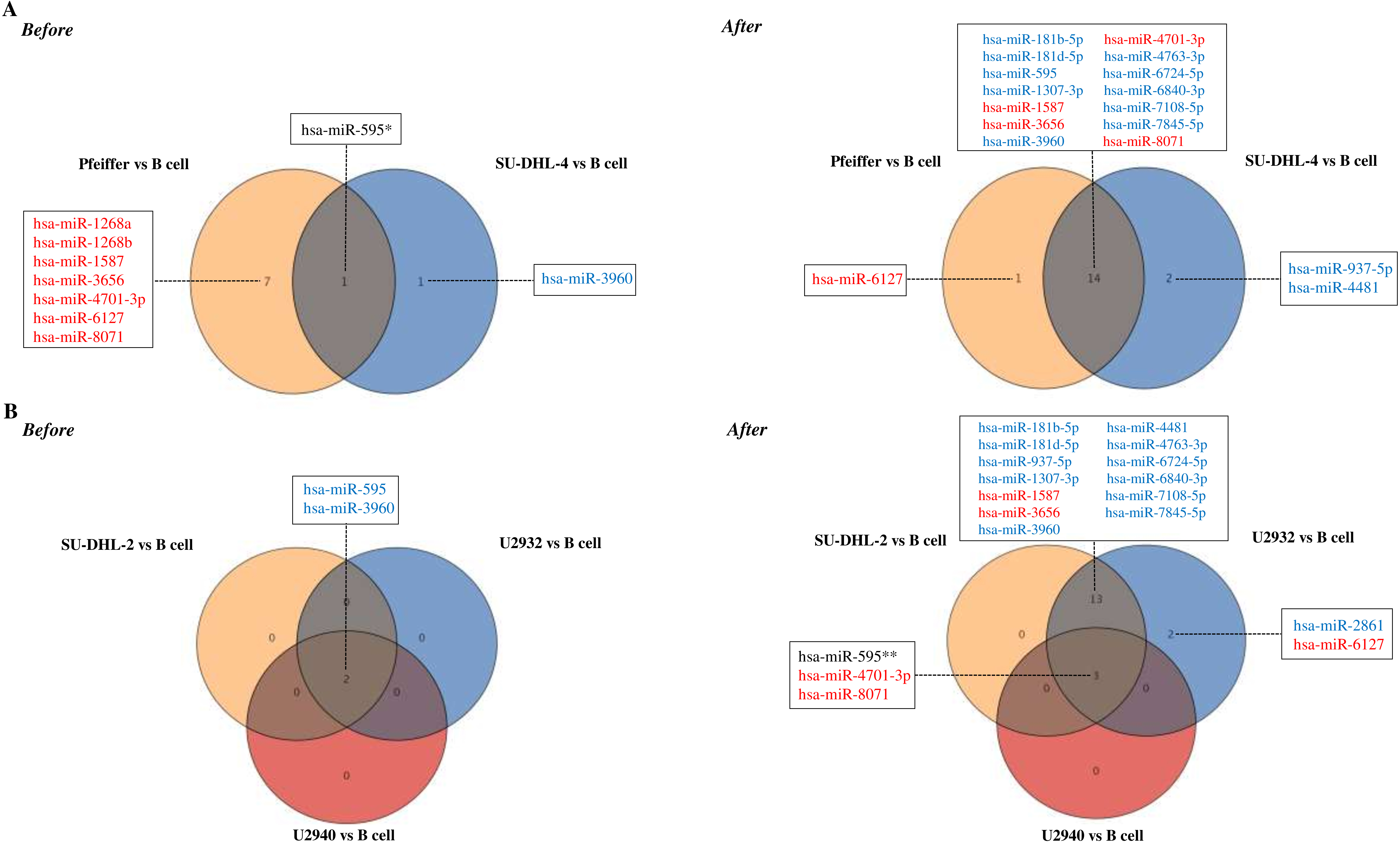
A significant increase in plasma exosome concentrations of DLBCL patients was observed. There was also a significant decrease in the expression of 33 miRNAs in plasma exosomes of DLBCL patients. It was determined that normal B-cells internalize DLBCL-derived exosomes and then miRNA expression differences observed in normal B-cells are specific to lymphoma-subtypes.
CONCLUSIONS:
MiR-3960, miR-6089 and miR-939-5p can be used as the miRNA signature in DLBCL diagnosis. We suppose that the exosomes changed the molecular signature of the target cells depending on the genomic characterization of the lymphoma cells they have originated.
Introduction
Diffuse Large B-cell Lymphoma (DLBCL) accounts for about 30–40% of Non-Hodgkin’s lymphomas (NHLs). Although there is no detailed data on the incidence of DLBCL in Turkey, NHL ranks the eighth and seventh most common cancer in women and man, respectively [1]. DLBCL is highly heterogeneous in terms of genetic, immunological, morphological, and clinical features [2, 3]. New biomarkers that will provide insight into disease diagnosis, prognosis, and response to therapy are needed since the analyzes which are used to classified into subtypes in routine clinical practice do not meet all the biological parameters.
Circulating miRNAs have many characteristics of ideal biological markers such as stability, being tightly conserved among species, and being different according to the tissue or clinical stage [4, 5]. In recent years, especially in the diagnosis and classification of cancer, the superiority of exosomal miRNA profiles as biomarkers has been emphasized [6, 7, 8]. There are limited data on exosomal miRNA profiles and their roles in the diagnosis and pathogenesis of DLBCL.
Exosomes that have changed the image of “cellular garbage collectors” to “information-carrying vesicles” over time, play an active role in intercellular communication. Exosomal cargo compositions are considered as a fingerprint of originating cells and transferred to the target cells [9, 10]. There are limited studies on the internalization of lymphoma-derived exosomes by peripheral blood cell populations [11, 12]. The common finding in these studies is that B-cells effectively internalize lymphoma-derived exosomes, but the effects of exosomes after internalization by target B cells are not yet known.
In this study, we aimed to (i) compare exosomal miRNA expression profiles in DLBCL patients and healthy individuals, and (ii) determine whether DLBCL-derived exosomes are mediators that transfer miRNA-specific information to normal B-cells.
Materials and methods
The study group
The minimum number of samples was calculated (
Since gender is not considered as a risk factor for DLBCL in the literature, 20 voluntary healthy individuals who matched the patients’ ages were selected as the control group. For the group, inclusion criteria included no prior cancer history, no chronic and autoimmune disease history, age between 18 and 80, and no previously receiving cytotoxic drug treatment.
This study was approved by the Institutional Ethics Committee, Pamukkale University (Approval no: 14, Approval date: 27.08.2013), and written informed consent was provided by all participants.
Exosome isolation from plasma samples
Peripheral blood sample collection was performed as previously described [13, 14]. Briefly, venous blood sample (10 ml) were taken through a
Exosome concentrations were estimated using a commercial kit based on the measurement of acetylcholinesterase enriched activity in exosomes (Exocet, SBI).
Characterization of isolated exosomes
Morphological characterization
To characterize and visualize exosome morphology, scanning transmission electron microscopy (STEM) and scanning electron microscopy (SEM) were used. Briefly, a 5-
Determination of total protein concentration in the exosome samples
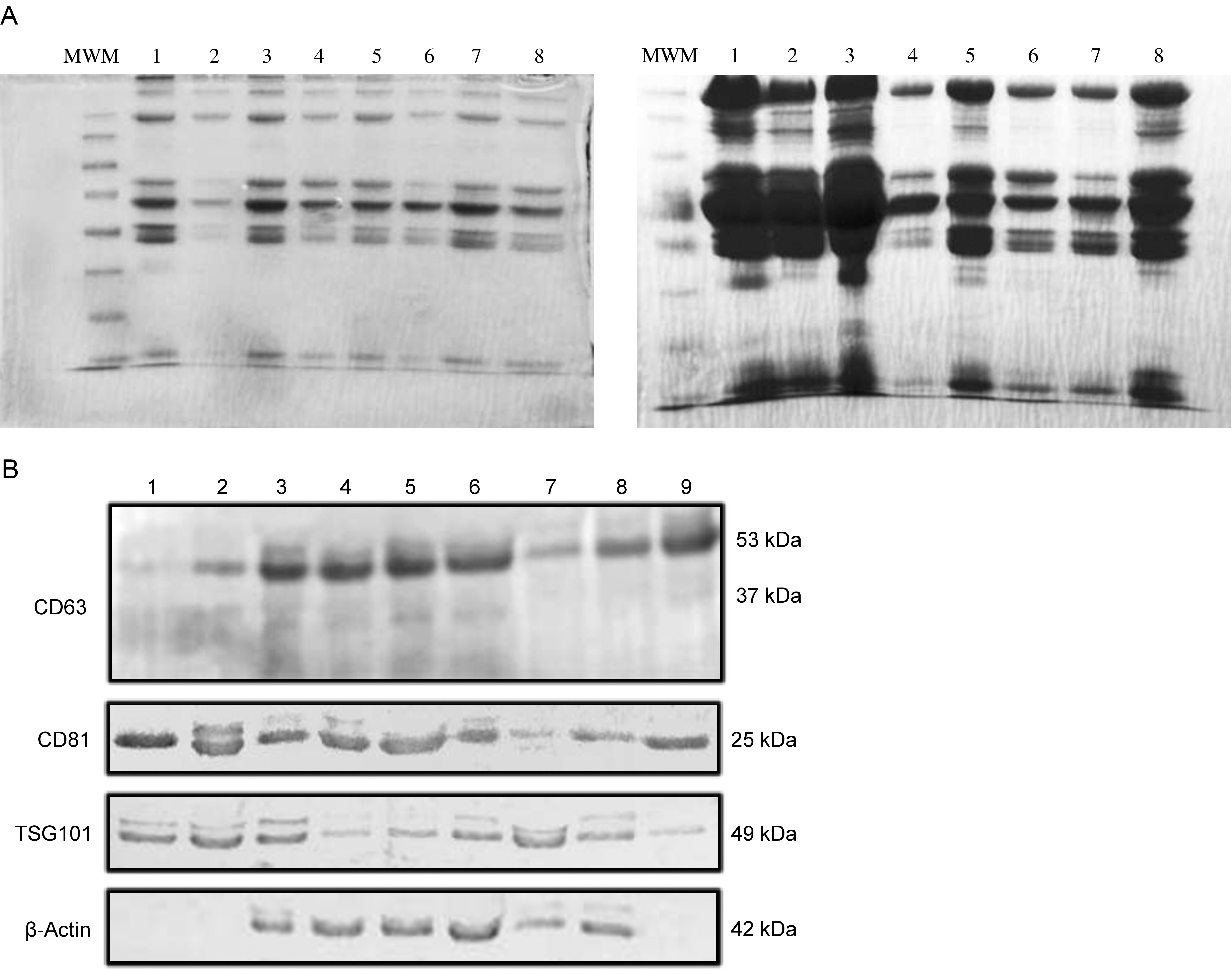
Exosomal protein lysates were prepared in the lysis buffer (Cell Signalling Tech.), including protease inhibitor cocktail (Cell Signalling Tech.) and EDTA (Sigma). The lysates were used in determining total protein amounts, imaging total proteins by SDS-PAGE, and Western Blot analysis.
Total protein amounts of exosomes were determined by the Bradford method (DC Protein Assay, BioRad, Protein Standard II, BSA, BioRad).
Detection of exosomal marker proteins
According to the MISEV2014 guidelines, we checked the expression of CD63, CD81, and TSG101 as exosomal protein markers by western blot [14]. Western blot analyzes were completed using anti-CD63 (ab68418-Abcam), anti-CD81 (ab155760-Abcam), anti-TSG101 (ab30871-Abcam), anti-
Exosomal total RNA extraction
Before total RNA isolation, exosome samples were treated with RNase A (Invitrogen) in order to avoid external RNA contamination. Total RNA isolation was completed using Trizol reagent (Invitrogen). Briefly, 100
Co-culture model
B-cell isolation from peripheral blood mononuclear cells
Due to the increased exosome internalization capacity in B-cells with age, a 5-ml peripheral blood sample was taken from a 62-year-old healthy individual [15]. B-cells were isolated by negative selection (MACS Human B Cell Isolation Kit II, Miltenyi Biotec) and analyzed on a flow cytometer (Beckman Coulter).
The B-cells were cultured in RPMI-1640 (Gibco, Life Technologies), supplemented with 10% exosome-depleted Fetal Bovine Serum (FBS) (Gibco, Life Technologies) and 1% penicillin/streptomycin (Gibco, Life Technologies) at 37
DLBCL cell lines and culture conditions
A total of 5 cell lines belonging to different DLBCL subtypes were used: Pfeiffer (CRL-2631, ATCC) and SU-DHL-4 (CRL-2957, ATCC) for the GC B cell-like (GCB), SU-DHL-2 (CRL-2956, ATCC) and U2932 (ACC 633, DSMZ) for the activated B cell-like (ABC), and U2940 (ACC 634, DSMZ) for the primary mediastinal B-cell lymphoma (PMBL) subtype. All cell lines were obtained from the Leibnitz Institute DSMZ-German Collection of Microorganisms and Cell Cultures (
Before the co-culture experiment, the exosome isolation was performed on normal B-cells and all of the lymphoma cell lines as previously described. In the co-culture model, 6-well cell culture dishes (Greiner Bio-One) were used, and the lymphoma cells were added to each well at a concentration of 1
STEM and TEM images of exosomes isolated from plasma samples of a healthy individual and a DLBCL patient. A-1: STEM (Magnification: 125.00 K X, Scale bars: 200 nm) and A-2: TEM image (Magnification: 67.00 K X, Scale bars: 200 nm) of exosomes in a healthy individual. B-1: STEM (Magnification: 100.00 K X, Scale bars: 200 nm) and B-2: TEM image (Magnification: 50.00 K X, Scale bars: 200 nm) of exosomes in a patient. This figure also shows that patient’s plasma contains more particles than a healthy individual. Pa: target particle, PaR: particle diameter by point to point measure. 
miRNA microarray analysis was performed using SurePrint G3 Human miRNA r21 Array Kit (Agilent Technologies). The labeling, hybridization, and washing procedures were performed according to the manufacturer’s instructions. Each sample was repeated three times and multiple correlation analyzes were completed by taking the mean values. For validation of the microarray data, differentially expressed miRNAs (down-regulated: hsa-miR-3960, hsa-miR-6089, hsa-miR-939-5p, hsa-miR-1207-5p, hsa-miR-595; up-regulated: miR-6803-5p) were investigated by real-time-PCR using the miRCURY LNA
Association between plasma exosome concentration, total exosomal protein concentration and clinicapathological variables in DLBCL patients
Association between plasma exosome concentration, total exosomal protein concentration and clinicapathological variables in DLBCL patients
Footnotes
Acknowledgments
This study was supported by a grant from The Scientific and Technological Research Council of Turkey, TUBITAK (Grant No. 114S442). The authors are grateful to Dr. Sevil Zencir for helpful discussions and Dr.Nazan Keskin for STEM and TEM assistance.
Conflict of interest
The authors involved in this study declare no conflict of interest.
Author contributions
Conception: V.C., and G.O.C.
Interpretation or analysis of data: V.C., G.O.C., S.H., I.C.B., E.T., N.S.T., and G.C.
Preparation of the manuscript: V.C., G.O.C., S.H., I.C.B., E.T., G.B., and K.Y.
Revision for important intellectual content: K.Y.
Supervision: V.C., and G.O.C.