Abstract
BACKGROUND:
The discovery of miRNA/mRNA interactions in several biological samples prompted the researchers to explore new biomarkers in tumors.
OBJECTIVE:
We aimed to investigate the interactions of miRNA/mRNA in response to radiotherapy in the plasma samples of rectal cancer patients.
METHODS:
Five microarray datasets related to cancerous and non-cancerous individuals were first used to construct networks. The databases of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) were applied to analyze pathway enrichment. The plasma samples were then collected from 55 patients with recently diagnosed rectal cancer and 10 healthy subjects. For radiotherapy courses, the patients have consecutively received 30 sessions of local radiation for six weeks. At last, the expression of selected genes and miRNAs was experimentally measured before and after radiotherapy by qPCR, and the protein levels of the target genes were measured by ELISA assay. We evaluated the therapeutic responses based on the tumor regression grade of the Dworak classification.
RESULTS:
We identified 5 up-regulated and 5 down-regulated miRNAs and 8 up-regulated and 3 down-regulated genes of the databases. There was a significant increase in tumor suppressor miRNAs, including miR-101-3p, miR-145-5p, miR-26a-5p, miR-34a-5p, and a significant decrease in oncomiRs, including miR-221-3p and miR-17-5p, after radiotherapy compared to the pre-treatment. Moreover, the up-regulated miR-17-5p and miR-221-5p and the down-regulated miR-101-3p and miR-145-5p were directly related to rectal cancer through the interaction with the Wnt, RAS, PI3K, and TGF-
CONCLUSIONS:
It seems that monitoring the miRNA/mRNA interactions during radiotherapy can be an appropriate diagnostic tool to track the recovery process and respond to standard therapies.
Introduction
Many studies have investigated radiation’s effects on the cells’ biological behavior, like cell death, chromosomal aberration, and mutagenesis. Observations have shown that the sensitivity of cells to radiation is different. The cells with a high proliferation or division are more sensitive to radiation than cells with a low proliferation or division [1]. Studies showed that some cells are not capable of DNA damage repair (DDR) caused by radiation, and the damage can directly or indirectly cause cell death. In this setting, microRNA (miRNA) may play an essential role in regulating DDR-related processes and altering tumors’ sensitivity to radiation [2]. Therefore, evaluating miRNA expression in radiated patients can give us helpful information on how tumors can resist or sensitize the radiation [3]. For example, miR-21 has been known to progress in many diseases that may increase malignancy. The miR-21 level increases in tumor cells after radiation [4]. Likewise, Yan et al. showed that the miR-101 expression is significantly associated with poor clinical outcomes in colorectal cancer patients and can inhibit the expression of ataxia telangiectasia mutated (ATM) and DNA-PKC genes. Therefore, overexpression of this miRNA in tumor cells may increase their sensitivity to radiation [5].
Moreover, He et al. showed that colorectal cancer (CRC) patients with a low serum miR-101 had a more reduced 5-year overall survival than patients with a high serum miR-101 level. Therefore, it might be a valuable marker for diagnosis and prognosis [6]. MiR-145 is another miRNA that exhibits tumor suppressor activity in several cancers, including colon cancer. The overexpression of miR-145 in SW620 and DLD1-SNAI1 cells could sensitize these cells to radiation therapy [7]. In this setting, miR-145 and miR-101 are associated with biological processes such as proliferation, growth, and apoptosis. Yang et al. also found that overexpression of miR-100 could increase the sensitivity of CCL244 cells to radiation. For the first time, they suggested that miR-100 may play an essential role in regulating colorectal tumor cells [8]. Likewise, several studies have shown that many crucial proteins in colorectal cancer signaling, such as WNT, PI3K, EGFR, P53, TIMP, and epithelial-mesenchymal transition (EMT), can control colorectal cancer via miRNA. Zheng et al. observed that miR-106b overexpression led to cell radioresistance through direct interaction with PTEN and P21 proteins, increasing tumor cells’ survival and proliferation. They detected that miR-106b could activate the PI3K/AKT signaling pathway via PTEN inhibition [9, 10].
Generally, the discovery of miRNA/mRNA interactions in several biological samples prompted the researchers to explore new biomarkers in tumors [11]. In this context, analysis of relevant miRNA/mRNA interactions may be a suitable strategy for the early detection, monitoring, and prognosis of different types of cancer, including colorectal cancer [12]. Therefore, it appears that changes in miRNA/target gene expression in response to radiotherapy can provide valuable information on how to use adjuvant therapies to improve radiation-based treatments. We aimed to develop novel predictive miRNA/target gene pathways for rectal cancer using a resourceful systematic approach.
Materials and methods
Microarray data analysis
Expression profiles of miRNAs (GSE125961 and GSE112955) and mRNAs (GSE44172, GSE123390: GPL17586, and GSE81986: GPL570) in rectal cancer patients were extracted from the Gene Expression Omnibus (GEO) database (
Predicted target genes of candidate miRNAs
The target genes of miRNAs were identified using the online predictive programs, including miRmap (
GO term and KEGG pathway analysis by the FunRich dataset
The pathway enrichment analyses of gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases were executed using the FunRich dataset, the software for the functional classification of genes.
Experimental design and sampling
Fifty-five patients diagnosed with rectal cancer and ten healthy individuals were recruited from October 2016 and over one year. All the included participants were informed about the study protocol, and the written consent was taken. This trial was registered in the Iranian Registry of Clinical Trial (Clinical trial number: IRCT2016072618745N9).
The eligible cases were recruited after reviewing the medical records of patients who had previously been diagnosed with rectal cancer based on pathologic reports. Inclusion criteria comprised the age between 30–70 years, non-metastatic stage II or III rectal cancer patients, Karofsky Performance Status
Plasma samples were collected from healthy volunteers and patients with rectal cancer before and after radiotherapy. First, approximately 10 ml blood samples were taken from all participants using Vacutainer disposable blood collection tubes. The blood was then centrifuged at 3000
ELISA assay
VEGF (ab100663, Sensitivity: 10 pg/ml, Range: 8.23–6000 pg/ml), SMAD4 (ab253211, Sensitivity: 52.31 pg/ml, Range: 125–8000 pg/ml), ZEB2 (LS-F13506, Sensitivity: 0.312, Range: 0.312–20 ng/ml), TGFBR2 (MBS7223133, Detection Range: 1.0–25 ng/ml, Sensitivity: 0.1 ng/ml), STAT3 (ab176655, Sensitivity: 15
The selected protein levels were determined by a sandwich ELISA as follows: aliquots of 100
Real-time PCR analysis
The quantitative PCR (qPCR) was used to evaluate the expression of miRNA and genes. First, the total RNA was extracted from the plasma and PBMC samples. Then, the plasma (250
A list of primers for the RT-PCR
A list of primers for the RT-PCR
All the participants underwent surgical tumor resection after neoadjuvant radiotherapy (30 sessions of local radiation for six weeks). The patients were classified based on TNM 8
Statistical analysis
The statistical analysis was carried out using GraphPad Prism 7.04 (San Diego, CA) and SPSS 18 (IBM, New York, USA) statistical analysis software. The one-sample K-S test was used to evaluate the normality of the data. The
Results
Identification of differentially expressed miRNAs (DEMs) and differentially expressed genes (DEGs)
The datasets of miRNAs (GSE125961 and GSE112 955) and mRNAs (GSE44172, GSE123390: GPL17586, and GSE81986: GPL570) have included 102 samples of rectal cancers. According to our analysis, 96 miRNAs were found to have differential expression in the samples. About 75 of 1067 DEGs were identified as novel genes. The top 5 up-regulated miRNAs, including miR-17, miR-20a, miR-221, miR-23a, and miR-200c, and the top 5 down-regulated miRNAs, including miR-34a, miR-141, miR-145, miR-26a, and miR-101, were represented in Table 2. Besides, the target genes of the selected miRNAs were represented in Table 3.
The candidate miRNAs in rectal cancer
The candidate miRNAs in rectal cancer
The candidate genes in rectal cancer
To examine the biological functions of 75 DEGs, GO and KEGG analyses were performed in the DAVID database. The enrichment analysis of miRNAs and their target genes included cell apoptosis, migration, and proliferation pathways (Fig. 1). We demonstrated the top 10 enriched pathways in Fig. 1. All of these genes were selected for the qPCR for further validation.
Pathways enrichment analysis related to deriver miRNAs and their target genes.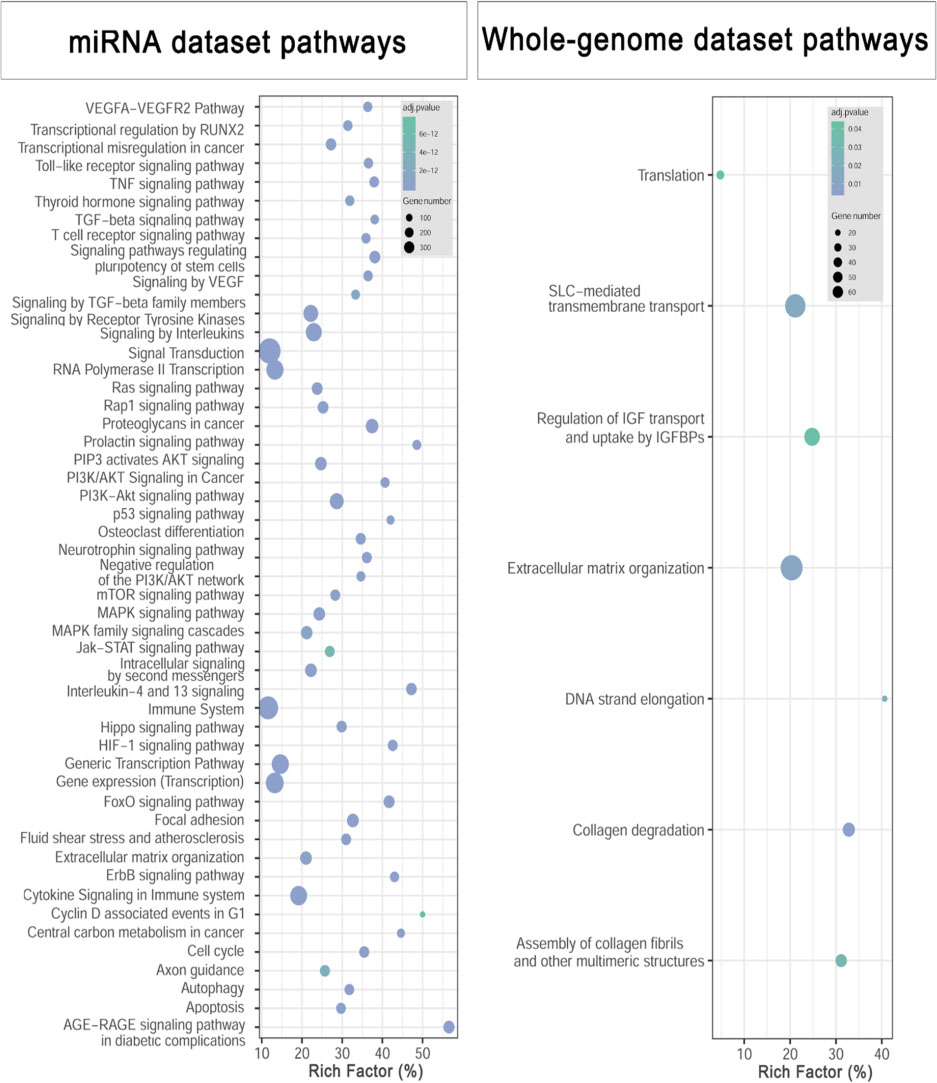
The mean age was 57.3
Demographic characteristics of the participants in the present study
Demographic characteristics of the participants in the present study
BMI: Body mass index.
Fifty-five patients with localized rectal cancer and ten healthy subjects were examined for evaluating the expression levels of miR-17, miR-200c-3p, miR-23a-3p, miR-20a-5p, and miR-221-3p before and after radiotherapy. Our results showed that the expression level of miR-17-5p (
The expression level of the selected miRNAs, miR-17-5p (a), miR-221-3p (b), miR-20a-5p (c), miR-200C-3p (d), and miR-23a-3p (e), before and after radiotherapy in rectal cancer patients compared to the healthy subjects. The relative expression of selected miRNAs was normalized using U6 as the reference RNA. 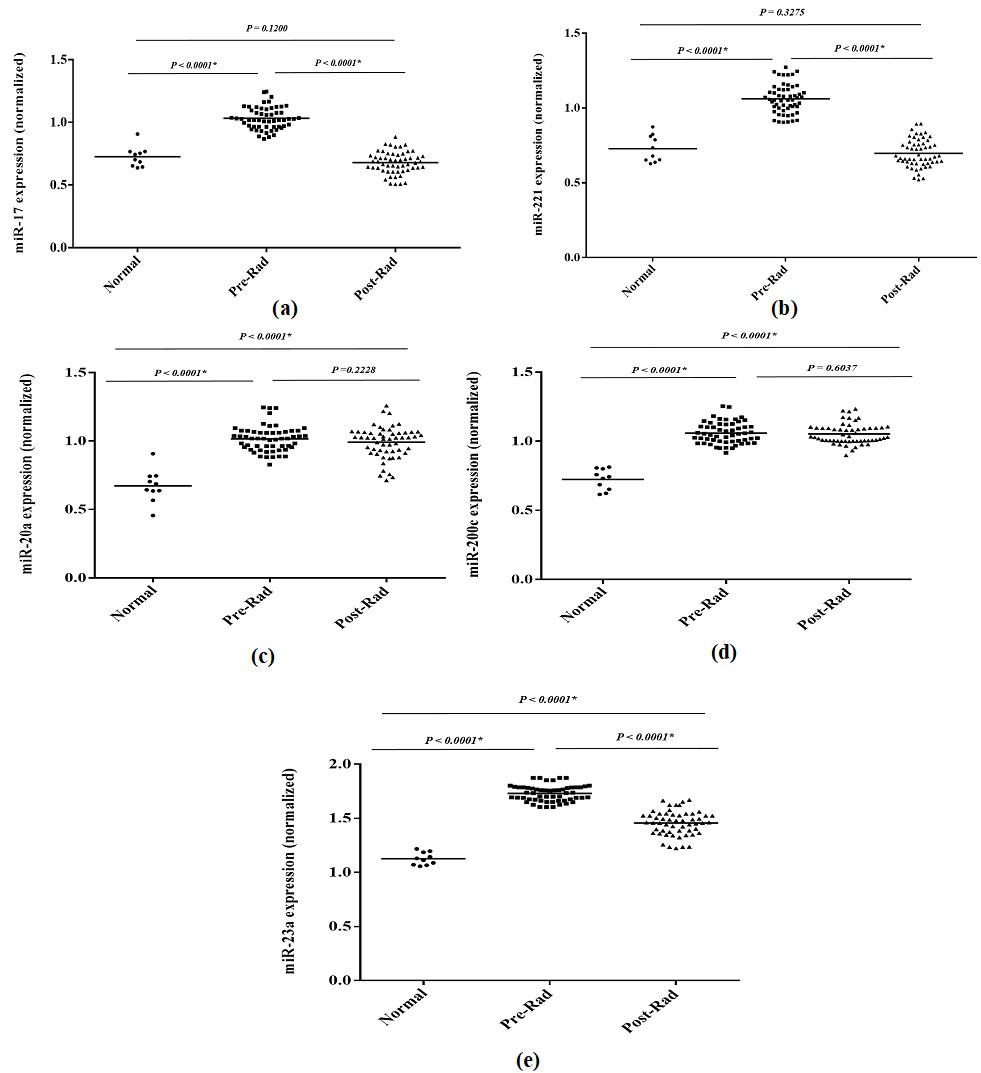
The expression levels of miR-26a-5p (
The selected miRNAs’ expression, miR-141-3p (a), miR-145-5p (b), miR-26a-5p (c), miR-101-3p (d), miR-34a-5p (e), before and after radiotherapy in rectal cancer patients compared to the healthy subjects. The relative expression of selected miRNAs was normalized using U6 as the reference RNA. 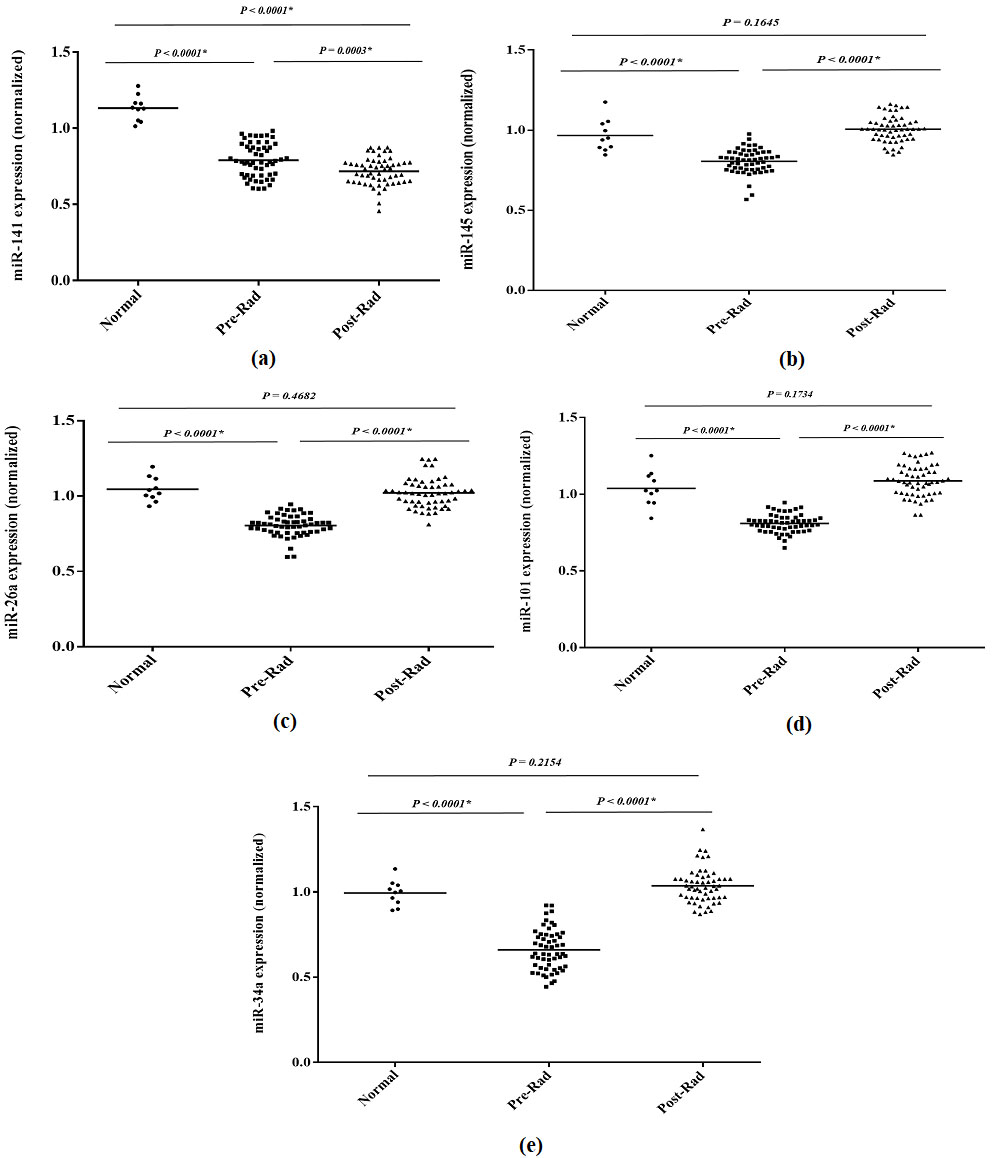
The results showed that the expression levels of oncoproteins, including VEGF, SMAD4, ZEB2, TGFBR2, STAT3, NOTCH1, TET2, and MYC, were significantly increased in the cancerous samples compared to the normal group. Likewise, a similar trend was consistent as the advanced stage of disease (Table 5). In contrast, the expression levels of tumor suppressor proteins, including PTEN, WEE1, and RB1, were significantly decreased in the advanced stages compared to the primary stages and the normal subjects (Table 5).
The expression of the selected oncogenes (a) and tumor suppressor genes (b) before and after radiotherapy in rectal cancer patients compared to the healthy subjects. The relative expression of genes was normalized by using b-actin as the internal control gene. 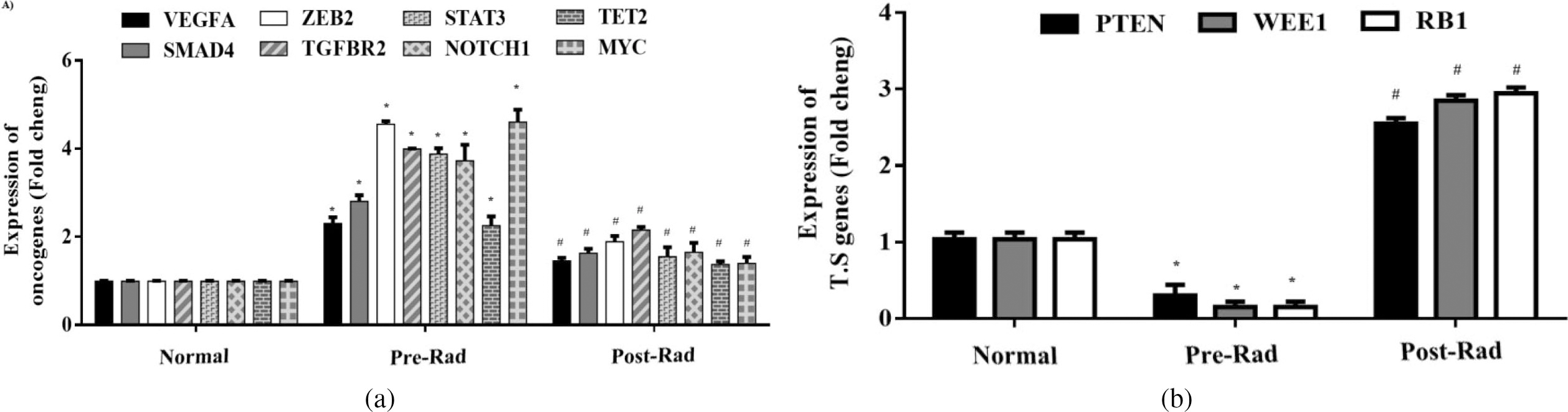
The protein levels of selected tumor suppressor and oncoproteins in rectal cancer patients in response to radiotherapy
We examined the expression levels of the VEGF, SMAD4, ZEB2, TGFBR2, STAT3, NOTCH1, TET2, and MYC oncogenes in the patients before and after radiotherapy compared to the healthy subjects (Fig. 4). Our results showed that the expression levels of these genes were significantly different before radiation from that of the normal subjects. Similarly, there was a significant difference in their expression levels after radiotherapy compared to the pre-treatment (Fig. 4a).
Receiver operating characteristic (ROC) curve to evaluate the potential efficacy of (a) miR-221, miR-17, and miR-23, (b) the combination of miR-221 and miR-17, and (c) miR-26 to differentiate patients with response to radiotherapy (TRG 3/4) from non-respondent (TRG 1/2). TRG: tumor regression grade.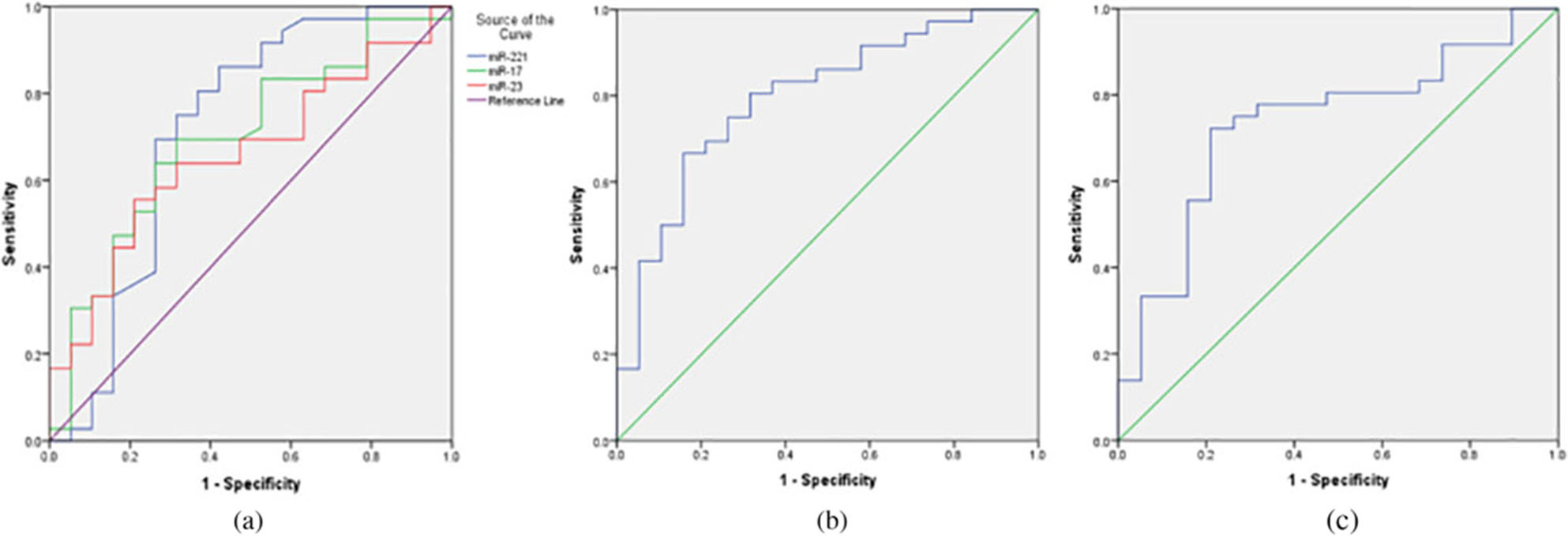
Finally, we have compromised the expression levels of PTEN, WEE1, and RB1 tumor suppressor genes before and after radiotherapy compared to the healthy subjects. The expression levels of these genes were significantly different before radiotherapy from that of the normal subjects. Their expression levels were very different after radiotherapy than the pre-treatment, indicating the positive effects of radiotherapy on these tumor suppressor genes (Fig. 4b).
The ten miRNAs were studied among the participants to validate their potential role as a predictor of response to radiotherapy. The patients were categorized into TRG3/4 and TRG1/2 groups as respondents and non-respondents, respectively. Interestingly, the patients who were in TRG3/4 (65.4%) group had significantly lower miR-221 (
ROC analysis was performed to investigate the potential utility of miR-221, miR-17, miR-23 and miR-26 as predictive biomarkers of response to radiotherapy. The AUC value for expression of miR-221, miR-17, and miR-23 was 0.717 (95% CI
Moreover, miR-221 and miR-17 provided a better predictive profile with an AUC value of 0.795 (95%CI: 0.651–0.920) (Fig. 5b). For the miR-26, the AUC was 0.735 (95% CI
Discussion
Our results showed the tumor suppressor miRNAs’ expression, including miR-101-3p, miR-145-5p, miR-26a-5p, and miR-34a-5p, and also the expression of oncomiRs, including miR-221-3p and miR-17-5p, changed significantly after radiotherapy compared to the pre-treatment in the rectal cancer patients. Moreover, there was a significant difference in the expression level of the oncoproteins and the tumor suppressor proteins after radiotherapy compared to the pre-treatment. The analyses also showed that the up-regulated miR-17-5p and miR-221-5p and the down-regulated miR-101-3p and miR-145-5p were directly related to rectal cancer via the Wnt, RAS, PI3K, and TGF-
Previous studies have shown that miRNAs could predict and determine treatment response to cancer treatments. They can also increase the sensitivity of tumors to radiation by inhibiting target genes. In this setting, miRNAs may modify current therapeutic strategies and make them more targeted and effective. Therefore, miRNAs are not only biomarkers involved in cancer prediction, prognosis, diagnosis, and monitoring, but also they can be used as therapeutic targets in many cancers [20, 21]. Our results showed that changes in miRNA expression in response to tumor radiation could provide helpful information on using adjuvant therapies to improve radiation-based treatments [22]. In this setting, four main pathways can be activated by the growth factor receptors in response to radiation.
The first pathway is PI3K/AKT, which modifies the expression of BAX, BIM, BCL2, and FOXO in response to radiation. PTEN is a tumor suppressor protein that plays a crucial role in regulating this pathway. Our results showed that the PTEN level was significantly different after radiotherapy compared to the pre-treatment. In this setting, the miR-21 has been shown to target and inhibit PTEN protein. This miRNA plays a central role in the occurrence or progression of cancer and may increase malignancy [23]. Moreover, Zheng et al. investigated the role of miR-106b in the sensitivity of human colorectal cells to radiotherapy. They observed that the increased expression level of miR-106b led to radiation resistance through direct interaction with PTEN and P21 proteins and improved cell survival and proliferation under radiation. They also observed that miR-106b could activate the PI3K/AKT signaling pathway by restricting PTEN protein and enhancing cell proliferation [9]. Similar to our results, Drebber et al. observed that the expression levels of miR-21 and miR-145 were increased and decreased, respectively, in tumor tissues. Their observations indicated reduced miR-21 and an increase in miR-145 after treatment in rectal tumors [4]. Thus, these miRNAs can be used to monitoring patients with radiotherapy.
The second pathway involves MAPK that promotes cell proliferation controlled by RAS and RAF activations. Our results showed that the expression levels of miR-17-5p, miR-200c-3p, miR-23a-3p, miR-20a-5p, and miR-221-3p were significantly different after radiotherapy than the pre-treatment. Moreover, high levels of miR-17-3p expression were related to a shorter disease-free survival [24]. Likewise, elevated plasma levels of miR-221 could be used as a potential biomarker to predict poor overall survival in CRC patients [25]. Similar to our results, Let-7 [26], miR-145 [27], and miR-143 have been identified as tumor suppressors that can inhibit RAS and RAF expressions [28]. Studies have also shown that the Let-7 family can increase tumor cell sensitivity to radiation by decreasing the RAS family genes [29]. According to our results, the up-regulation of miR-143 and miR-145 may improve the rectal cells’ radiosensitivity. Thus, these miRNAs may be used as biomarkers to predict therapy responses.
The third pathway is the transformation of epithelial-mesenchymal transition cells, leading to cancer cell metastasis. Several miRNAs are involved in EMT regulation, such as the miR-200 family. Our results showed that the expression level of miR-200c-3p was significantly different after radiotherapy compared to the pre-treatment. Sun et al. found that miR-429, a member of the miR-200 family, was significantly down-regulated in colon cancer [30]. According to our results, Hur et al. found that miR-200 was abnormally expressed in metastatic colon tumors correlated with reducing the expression of the target genes, such as ZEB1, ETS1, and FLT1 genes. This issue could up-regulate E-cadherin and down-regulate Vimentin sequentially, leading to the EMT signaling pathway [31]. Observations have also shown that factors such as ZEB1 have an active role in the EMT pathway through the interaction of miR-200 and miR-141 [10]. Therefore, the down-regulation of miR-200 might improve the radiosensitivity of the colorectal cells.
The fourth pathway is the P53, identified as a mutated tumor suppressor in 50–75% of all colorectal cancers. P53 can induce several miRNAs’ expression and maturation, including Let-7a, miR-133a, miR-34, and miR-16 in colon cancer cells [32]. Our results showed that the expression levels of miR-26a-5p, miR-101-3p, miR-145-5p, miR-34a-5p, and miR-141-3p were significantly different after radiotherapy compared to the pre-treatment. Similar to our results, the miR-34 family can play a central role in the cell cycle, proliferation, apoptosis, and angiogenesis, which targets the CDK4/6, cyclin E2, E2F2, BCL2, and SIRT proteins. Moreover, the miR-34 expression is decreased in colorectal cancers, which may be due to the deletion of 1p36 or miR-34 promoter’s methylation [32]. Therefore, the increased expression of miR-34 and Let-7a in tumor cells can increase the sensitivity to radiotherapy.
Overall, our results provided helpful information on inhibiting the expression of proteins involved in miRNAs’ cancer-related pathways. Although some of these interactions are only predicted, this dual computational approach can provide critical information for conducting new validation studies. We showed that over-expressed miRNAs, such as miR-221-3p, and down-expressed miRNAs, such as miR-101-3p and miR-26a-5p, could interact more genes in the signaling pathways. Generally, our observations may hypothesize that miRNAs can enhance proliferation and inhibit cell death. These results can support future studies that may determine the sensitivity, specificity, and efficacy of selected miRNAs, representing particular interactions with genes and molecular pathways in rectal cancer.
Additionally, the previous studies showed that patients with the complete response could undergo less invasive strategies such as “wait and watch” approaches [33]. The supporting literature indicated that such an approach could increase the quality of life in patients who were responsive to neoadjuvant chemoradiotherapy [33]. Contrarily, the patients with resistance to neoadjuvant chemoradiotherapy should manage properly based on their response rate to the treatments. This issue can result in modifying the irradiation dose or administration of different chemotherapeutic agents. Likewise, understanding the underlying mechanisms for resistance can improve the efficacy of radiation therapy by overcoming radioresistance. In the current work, we observed that from 10 selected miRNAs, four associated with the response to radiotherapy. In this respect, we observed that the miR-221, -17, and -23 were response-related miRNAs in patients with locally advanced rectal cancer. This profile has not been previously evaluated as a tool for treatment monitoring in locally advanced rectal cancer to the best of our knowledge. However, some other novel miRNAs were reported by which the response to chemoradiotherapy was predicted [34, 35]. In this respect, it was demonstrated that the miR-31 was associated with poor overall survival and a higher rate of resistance in locally advanced rectal cancer [34]. Besides, it was demonstrated that post-surgical expression of miR-345 was more elevated in rectal cancer patients resistant to chemoradiotherapy [36].
Moreover, Campayo et al. reported that the overexpression of miR-21, miR-99-b, and miR-375 was observed among the patients who showed a poor response rate to the chemoradiotherapy. Consistently, they provided similar findings regarding the possible role of let-7b in enhancing radiosensitivity in rectal cancer patients [37]. Interestingly, we observed that post-surgical higher expression of tumor suppressor miRNAs, miR-26, was associated with better radiosensitivity. This issue can imply that the target pathways of these miRNAs can be a possible target for targeted therapy for improving the efficacy of chemoradiotherapy.
Limitation and clinical application
Although we assessed a panel of the biomarkers to predict the response to treatment in locally advanced rectal cancer patients, our low sample size may diminish the validity of the findings. The clinical decision on the management of rectal cancer is substantially made based on the different variables before initiation of any clinical intervention. However, the current decision-making system is not accurate enough. As a result, around 30% of the patients showed no clinical response to performed neoadjuvant interventions [38]. About 70 miRNAs are associated with treatment resistance in rectal cancer patients [39, 40]. The possible valuable role of miRNAs in the clinic can be range from the biomarker to candidate for the targeted therapy. So far, multiple techniques developed to facilitate the clinical applications of miRNAs in patient management. These procedures can comprise antisense oligonucleotides or antagomirs, locked nucleic acids, peptide nucleic acids, the newest miRNA sponges and miRNA masking techniques, and an increased tumor suppressor miRNAs by miRNA mimics or viral vector-encoded miRNA replacement [40]. Despite their considerable impact on patient management, these techniques have been studied on a limited scale of rectal cancer patients. We could not find any active clinical trial regarding using these techniques on patients with locally advanced rectal cancer based on our search. Therefore, more studies should be conducted to assess the impact of these strategies in managing radioresistance patients.
Conclusion
We indicated that the interactions of selected miRNAs and target genes were associated with cell apoptosis, migration, and proliferation that can play crucial roles in rectal cancer. They may also be new candidate biomarkers to monitor conventional radiotherapy.
Abbreviation
DNA damage repair; microRNA; ataxia telangiectasia mutated; colorectal cancer; epithelial-mesenchymal transition; Gene Expression Omnibus, GO, gene ontology; Kyoto Encyclopedia of Genes and Genomes; peripheral blood mononuclear cells; Foetal Bovin Serum; Dimethyl sulfoxide; bovine serum albumin; quantitative PCR; tumor regression grade; receiver operating characteristic; differentially expressed miRNAs; differentially expressed genes; body mass index.
Author contributions
Conception: SK, MRK, VK, and AMA.
Interpretation or analysis of data: SK, MRK, ML, AN, and AMA.
Preparation of the manuscript: SK, MRK, and AMA.
Revision for important intellectual content: TA, AP, and EE.
Supervision: AMA.
Funding
None of the funding sources had any role in the study design, the collection, analysis, and interpretation of data, or the decision to submit the article for publication.
Compliance with ethical standards
Ethical approval
All procedures performed in the studies involving animal participants were under the ethical standards of the institutional and national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The study was also conducted under relevant national and international guidelines and approved by the Institutional Animal Care and Use Committee of Tehran University of Medical Sciences.
Footnotes
Acknowledgments
Research reported in this publication was supported by Elite Researcher Grant Committee under award number (Grant Number: 982679) from the National Institute for Medical Research Development (NIMAD), Tehran, Iran, and the Tehran University of Medical Sciences (Grant Number: 32359).
Conflict of interest
The manuscript authors have no conflicts of interest to declare and are responsible for the paper’s content.
