Abstract
BACKGROUND:
Recent studies demonstrated the involvement of mesenchymal stem/stromal cells (MSCs) in carcinogenesis, but the molecular mechanism behind this transformation is still obscured.
OBJECTIVE:
To screen both the expression levels of polycomb and trithorax epigenetic regulators and TrP53 mutations in early and late MSC culture passages in an attempt to decipher the mechanism of spontaneous transformation.
METHODS:
The study was conducted on early and late passages of MSC culture model from C57BL/6J mice. The expression profile of 84 epigenetic regulators was examined using RT2 profiler PCR array. TrP53 mutations in the DNA binding domain was screened. Codons, amino acids positions and the corresponding human variants were detected in P53 sequences.
RESULTS:
Sixty-two epigenetic regulators were dysregulated. Abnormalities were detected starting the third passage. Nine regulators were dysregulated in all passages. (C
CONCLUSION:
In vitro expanded MSCs undergo transformation through alteration of epigenetic regulators which results in genomic instability and frequent P53 mutations.
Introduction
Mesenchymal stem cells (MSCs) are heterogeneous cell populations that serve as reserves for tissue regeneration in the presence of disease or injury. Although MSCs are found in various tissues, bone marrow-derived stem/stromal cells (BMSCs) have been most thoroughly investigated [1]. These cells were first isolated from bone marrow by Friedenstein in 1967 and have been described as plastic adherent cells with a fibroblast-like morphology [2]. In 2006, the International Society for Cellular Therapy (ISCT) established criteria for identifying unique populations of MSCs. They defined MSCs according to the following criteria: 1 – MSCs must be purified from the BM stromal population based on plastic adherence under standard culture conditions. 2 – MSCs must be positive for CD105, CD90, and CD73, express low levels of MHC-I, and be negative for MHC-II, CD11b, CD14, CD34, CD45, and CD31. 3 – MSCs must differentiate in vitro into osteocytes, chondrocytes, and adipocytes [3].
The cell-of-origin concept refers to the normal cell type that acquires the first cancer-promoting mutation and initiates tumor formation [4]. During the last years there has been overwhelming evidence that MSC is the cell-of-origin for many types of sarcomas [5, 6]. As these cells either transform directly into cancer stem cells or settle in the tumor stroma to facilitate the process of malignant transformation and increase the ability of cancer cells for aggressiveness. MSCs have been shown to induce Ewing sarcoma [7], osteosarcoma [8], chondrosarcoma [9], synovial sarcoma [10] and gastric cancer [11]. The transforming properties of MSCs and their possible role as sarcoma progenitors make these cells useful as a model for studying sarcomagenesis and progression.
Spontaneous transformation of both murine and human MSCs in long-term cultures, have been reported in many studies [12, 13, 14]. Houghton et al. showed that bone marrow derived MSCs home to the gastric mucosa during Helicobacter infection and differentiate abnormally as dysplastic cells, can initiate, contribute to and become gastric adenocarcinoma. The time can be dramatically decreased by introducing MSCs which have been repeatedly passaged in culture and allowed to spontaneously transform [11].
Although MSC involvement in the carcinogenesis has been investigated and verified that these cells are the origin of cancer stem cells, we have not yet discovered an effective treatment that selectively target cancer stem cells. This is because of our failure to determine the mechanism of MSC transformation and the lack of knowledge of the factors that control this mechanism.
Epigenetic changes are heritable changes in gene expression without alteration of the coding sequence of the gene. Epigenetic mechanisms include, DNA methylation, histone modification and noncoding RNA-mediated processes. In addition to the common hallmarks of cancer which includes sustaining proliferative signaling, evading growth suppressors, resisting cell death, enabling replicative immortality, inducing angiogenesis, and activating invasion and metastasis, these epigenetic changes have recently emerged as common hallmarks of many cancers [15].
Epigenetic regulators are a group of elements that regulate gene expression along the genome. They act antagonistically to either promote (Trithorax group TrxG) or repress (Polycomb group PcG) transcription through regulation of specific amino acid modifications in histones [16]. Dysregulation of these elements may lead to genome wide changes in gene expression which in turn, lead to cancer development [17]. Recently, research has focused on the relationship between the abnormalities in these regulators and many types of cancers. According to the cancer stem cell model, alterations in epigenetic mechanisms of stem cells lead to malignant transformation of these cells [17]. These early epigenetic events in stem cells are also the earliest events in cancer initiation. Epigenetic disruption in stem cells may give rise to a high-risk aberrant progenitor cell population that can undergo transformation upon gain of subsequent genetic mutations [17]. Until now, we have not been able to identify the factors that lead to these epigenetic alterations.
We followed up the expression profile of 84 epigenetic regulators and P53 mutations in early and late passages of bone marrow derived MSC model from C57BL/6J mice, in an attempt to decipher the mechanism of in vitro transformation of MSCs.
Materials and methods
Expansion of cryopreserved mouse MSC passages
We used already established MSCs and progressively passaged for more than a year. The cells were originally obtained from the bone marrow of male C57BL/6J mice. The cells were first established on July, 2004 [11] and have been well characterized on March 2006 [11], where surface marker expression have been characterized by fluorescent activated cell sorting (FACS), and lineage specific marker assessment was determined by immunohistochemistry or reverse transcription polymerase chain reaction (RT-PCR). At each passage, cultures were split, and one plate was held in culture at confluence for detection of foci, one plate was split into the next passage culture, and the remainder of the plates was frozen [18].
Cryopreserved C57BL/6J mouse MSCs from passages 3, 6, 9, 12, 28, A2, B2 and the last passage were removed from the liquid nitrogen. The cells were thawed and 1x Pbs (pH 7.2) was added then centrifuged at 1200 rpm for 10 minutes. The supernatant was removed, and the cells were re-suspended in1x phosphate buffer saline (pH 7.2) and centrifuged again for 10 minutes at 1200 rpm. The supernatant was removed, and the cells were re-suspended in 5 ml Dulbecco’s Modified Eagle Medium (DMEM) containing 10% fetal calf serum (FCS), L-Glutamine and penicillin streptomycin. The cells were left in CO2 incubator at 37
Isolation and culture of control bone marrow MSCs from C57BL/6J mice
Control MSCs used for gene expression and P53 sequence analyses were isolated from the bone marrow of male C57BL/6J mice. The mice used were the same species sex and age from which MSCs used in the study were established. Bone marrow isolation from mice was performed in accordance with relevant guidelines and regulations of the University of Massachusetts Medical School. C57BL/6J mice were purchased from Jackson Laboratories (Bar Harbor, ME, USA) and housed in micro isolator cages in an SPF facility, given free access to chow and water.
At 8 weeks of age, male C57BL/6J mice were euthanized by CO
RNA extraction
Culture media from both control and MSc passages was removed and the cells were washed with 1x phosphate buffer saline (PH7.2). RNA from each passage was isolated using RNeasy plus kit from Qiagen. (Cat no. 74134). The integrity of the RNA samples was determined by 2% denaturing agarose gel electrophoresis. RNA was quantified and subjected to reverse transcription to cDNA.
cDNA synthesis
1121.3 ng of RNA per sample were used for cDNA synthesis. cDNA was synthesized using the RT2 First Strand Kit Qiagen (Cat no. 330401), 5
Real-time reverse transcription polymerase chain reaction for mouse polycomb and trithorax complexes
RT2 SYBR Green ROX qPCR Mastermix (Cat. no. 330520) was briefly centrifuged. In a clean tube 1350
After ending the PCR program, CT (threshold cycles) values were collected and the fold change for each gene was calculated using 2
P53 amplification
PCR amplification for the DNA binding domain of Trp53 was studied on the RNA samples using one step RT-PCR kits Qiagen (cat. no. 210210). The following primers were used Trp53F(96–115), 5
The fold regulation of the 35 dysregulated epigenetic regulators in passage 3 (group1)
The fold regulation of the 35 dysregulated epigenetic regulators in passage 3 (group1)
Gene symbols and fold regulation of the 35 dysregulated epigenetic regulators in passage 3 (group1). Thirty Four genes up and one downregulated.
The fold regulation of the 34 dysregulated epigenetic regulators in passage 6 (group2)
Gene symbols and fold regulation of the 34 dysregulated epigenetic regulators in passage 6 (group2). Twenty six genes up and 8 downregulated.
The fold regulation of the 38 dysregulated epigenetic regulators in passage 9 (group3)
Gene symbols and fold regulation of the 38 dysregulated epigenetic regulators in passage 9 (group3). Thirty genes up and 8 downregulated.
The fold regulation of the 22 dysregulated epigenetic regulators in passage 28 (group4)
Gene symbols and fold regulation of the 22 dysregulated epigenetic regulators in passage 28 (group 4). Fourteen genes up and 8 downregulated.
The fold regulation of the 42 dysregulated epigenetic regulators in the last passage (group5)
Gene symbols and fold regulation of the 42 dysregulated epigenetic regulators in the last passage (group5). Thirty five genes up and 7 downregulated.
The sequences chromatograms were viewed by FinchTV program. TrP53 sequence results were analyzed using mouse blast search (
Results
Most of polycomb and trithorax complexes showed abnormal expression in early and late MSC passages
We studied the fold change of a total of 84 epigenetic regulators including polycomb and trithorax elements in the control and 5 passages of MSCs, these passages are 3, 6, 9, 28 and the last passage (is a late MSC culture passage which has been in vitro cultured for more than a year, it was the last MSC passage number has been passaged during the time of the study). The control cells were MSCs cultured from the bone marrow of 8 week old male C57BL/6J mice. When the cells reached confluent, they were subjected to RNA extraction and analysis. We considered the genes are abnormally regulated if they have at least 2 fold changes (2 folds up or 2 folds down regulation from the control). Out of 84 genes, 62 were abnormally expressed (supplementary materials Fig. S1). We detected these abnormalities in all passages prior beginning from the 3rd passage. In passage 3, there were 34 genes up, and 1 downregulated (Table 1a, Fig. 1a). In passage 6, there were 26 up, and 8 downregulated genes (Table 1b, Fig. 1b). In passage 9, there were 30 up, and 8 downregulated genes (Table 1c, Fig. 1c). In passage 28, there were 14 up, and 8 downregulated genes (Table 1d, Fig. 1d). However, the last passage showed 35 up, and 7 downregulated genes (Table 1e, Fig. 1e).
Among the PcG and TrX elements, we found 9 regulators are abnormally regulated throughout the early and late passages and show significant dysregulation. These regulators are Lysine (K)-specific demethylase 5D (kdm5d), Retinol binding protein 2, cellular (Rbp2), Ribonuclease L (2
The fold regulation of the 9 abnormally expressed genes in all MSC passages
The fold regulation of the 9 abnormally expressed genes in all MSC passages
The fold regulation of RbP2, Kdm5d, RNASEL, Zbtb16, Cbx5, Cbx7, DNMT3L, Phf19 and smarc1a, in the 5 early and late MSC passages.
Kdm5d was upregulated in passage 3 and downregulated in passages 6, 9, 28 and the last passage, while Rbp2 was upregulated in passages 6, 9 and the last passage. However, Chromobox Cbx7 was downregulated in passages 3, 6, 9, 28 and upregulated in the last passages while, dnmt3l was downregulated in passages 3, 6, 9, 28 and upregulated in the last passage (Table 2, Fig. 2a and b).
Substitution mutations were detected in the early and late while deletion and insertion mutations were appeared in the late MSC culture passages
We sequenced the DNA binding domain of the TrP53 gene in a total of 8 passages representing the early and the late passages. We sequenced passages 3, 6, 9 and 12 which represent the early passages and passages 28, A2, B2 (A2 and B2 are late transformed MSC lines isolated after 4 months from sub cloning of the original cell line) and the last passage which represent the late passages.
We did not find any mutation in the control passage. For passage 3, we detected (C
The scatter plot compares the normalized expression of every gene of the 84 epigenetic regulators between each passage and the control passage by plotting them against one another to visualize large gene expression changes. The central line indicates unchanged gene expression. The dotted lines indicate the selected fold regulation threshold. Data points beyond the dotted lines in the upper left and lower right sections meet the selected fold regulation threshold. (a) Scatter plot shows 35 dysregulated epigenetic regulators in passage 3 (group1), 34 abnormally up and 1 abnormally downregulated genes. (b) Scatter plot shows 34 dysregulated epigenetic regulators in passage 6 (group2), 26 abnormally up and 8 abnormally downregulated genes. (c) Scatter plot shows 38 dysregulated epigenetic regulators in passage 9 (group3), 30 abnormally up and 8 abnormally downregulated genes. (d) Scatter plot shows 22 dysregulated epigenetic regulators in passage 28 (group4), 14 abnormally up and 8 abnormally downregulated genes. (e) Scatter plot shows 42 dysregulated epigenetic regulators in the last passage (group 5), 35 abnormally up and 7 abnormally downregulated genes.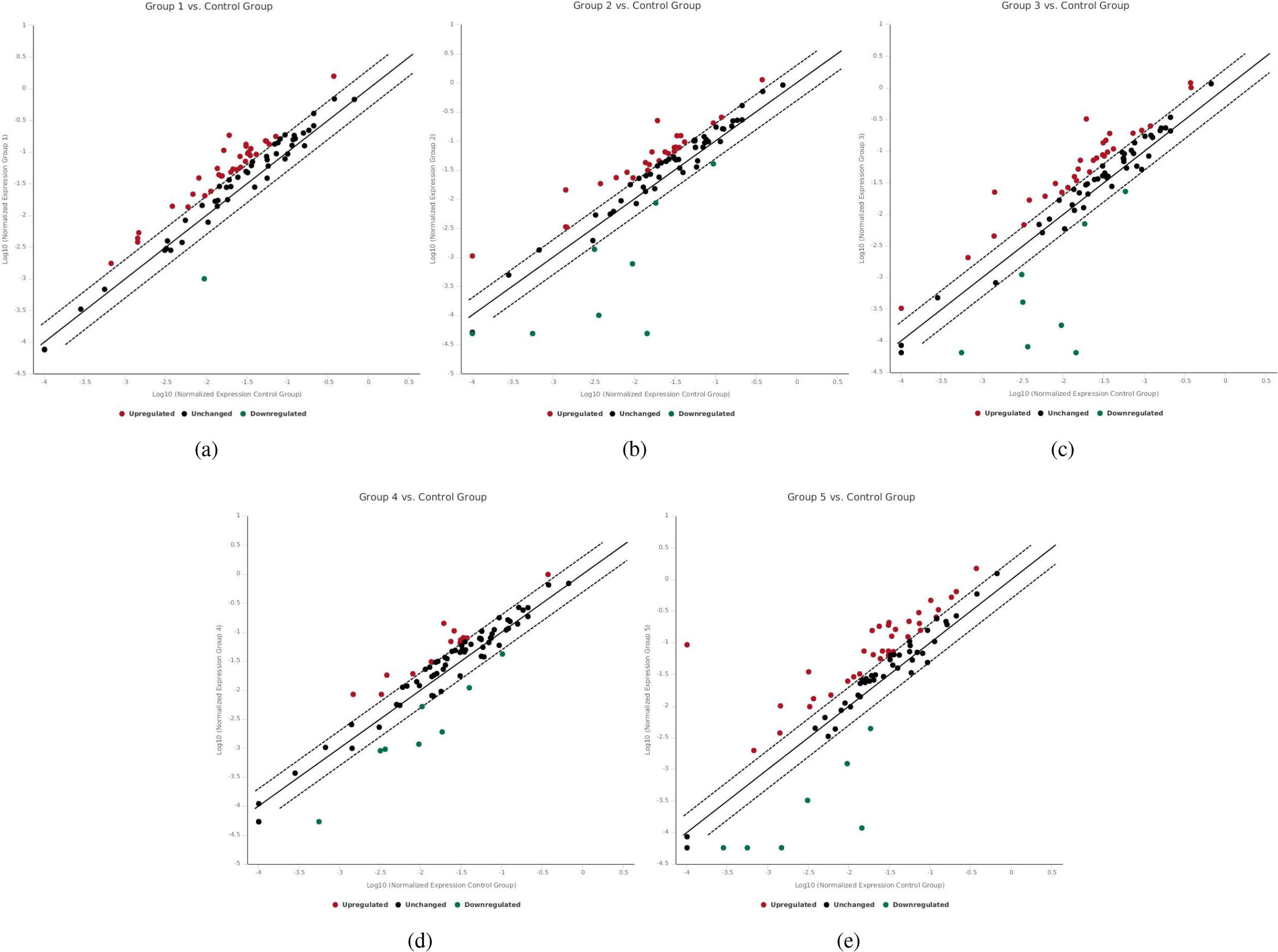
Represents the fold changes of the 9 dysregulated genes in all the 5 early and late passages of the BM derived MSC culture model. (a) Following up the fold change of Kdm5d, RNASEL, Zbtb16, Cbx5, Cbx7, DNMT3L, Phf19 and smarc1a in the 5 early and late MSC passages. (b) Following up the fold change of Rbp2 gene in the 5 early and late MSC passages.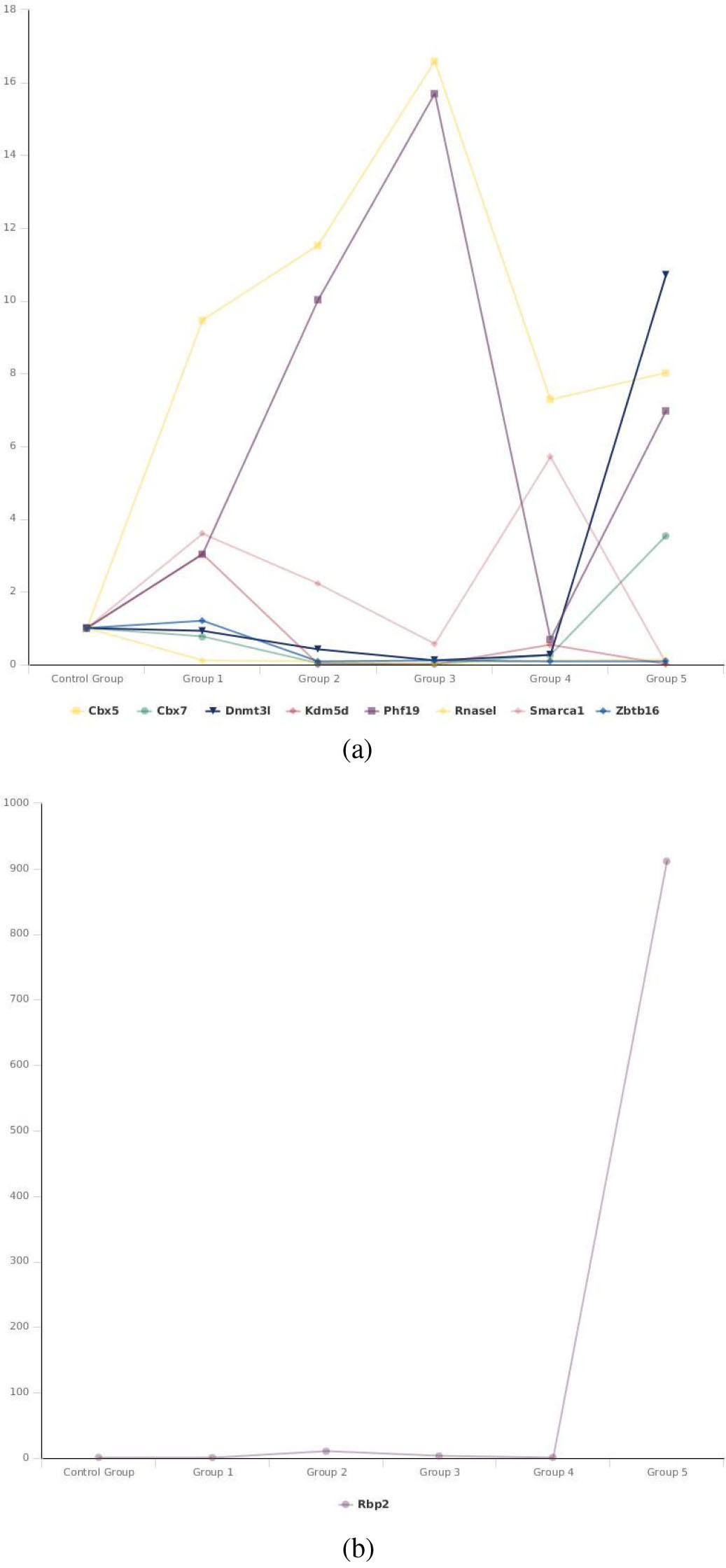
Electropherogram of part of the TrP53 gene sequence for Passages 3, 6, 9, 12 and the last passage of the BM derived MSC culture model from C57BL/6 mice. (a) A is showing normal sequence. B is showing heterozygous c.456C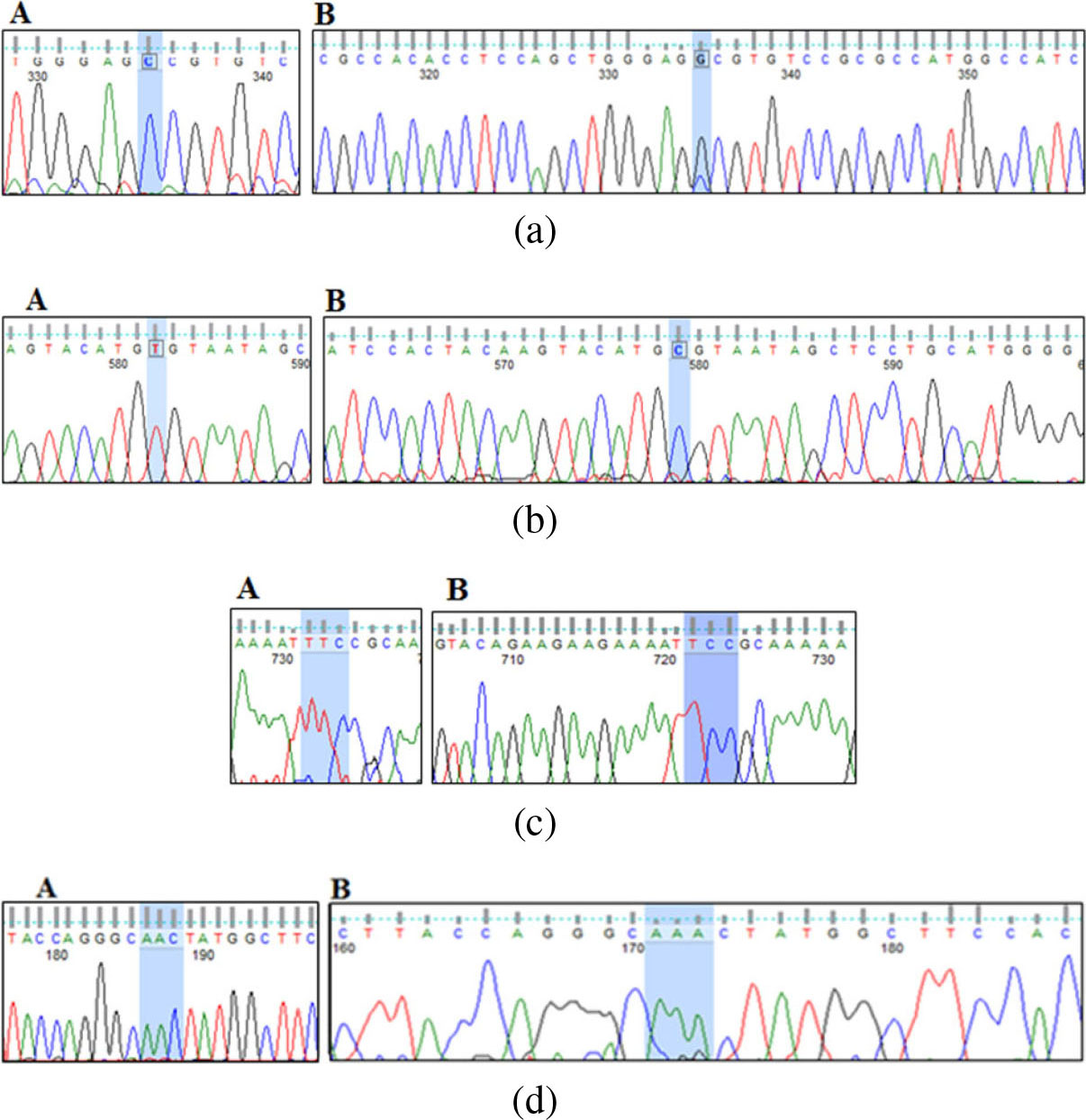
For passages 6, 9, 12 and the last passage, we detected T
The last passage showed the presence of T deletion (TrP53):c.857delT) (Fig. 3c) which results in frame shift mutation changing the protein phenylalanine at position 286 into Serine (p.F286fs*56) (supplementary Fig. S2c) which corresponds in human to the frame shift mutation (TP53):c.866delT (p.L289fs*56).
The last passage also showed the presence of A insertion (TrP53: c.308-309insA) or (c.308dupA) (Fig. 3d) resulting in frame shift mutation changing the protein Asparagine at position 103 into Lysine (p.N103fs*27) (supplementary Fig. S2d) corresponding in human to the frameshift mutation (TP53):c.315insA (p.S106f s*43).
Passage 28 had G and T insertion, passage A2 had G insertion, Passage B2 had G insertion and A deletion (Data not shown).
Several studies have demonstrated the involvement of MSCs in the carcinogenesis process. These studies have made several assumptions about the process of MSC transformation. Despite these assumptions, so far we could not decipher the mysteries of this process. The results of our study can contribute to the interpretation of how malignant transformation of MSCs. The present study has proven that there is an abnormality in the gene expression of most epigenetic regulators in MSC cultures, with different mutations in the P53 gene confirming the transformation of these cells. Recent studies have demonstrated a relationship between the disruption of epigenetic processes and carcinogenesis [17].
Although we have demonstrated an abnormality in the gene expression of 62 epigenetic regulators, 9 are dysregulated in all MSC passages. For the remaining 53 regulators, each was dysregulated in at least one passage. These 9 regulators were found to regulate the expression of many genes that are cornerstones in cancer pathways including Wnt receptor signaling pathway [19], EMT [20], IFN [21], P53 pathway [22], PTEN signaling pathway [23] and AKT-NF-
The presence of abnormalities in the gene expression of these 9 regulators in all passages indicates its important role in MSC oncogenesis and their possible role as drivers in this transformation. Other studies are clearly needed to study the functional characteristics and the therapeutic role of these abnormally expressed epigenetic regulators. This occurs by detecting the impact of targeting these factors on tumor formation in animal models for cancer.
We noticed that the demethylases were active and the methylases were inactive in the early passages, these findings may indicate the importance of demethylation in the early stages of MSC transformation. Many studies documented that demethylation is an early event in carcinogenesis [25, 26]. Also one of the functions of the DNA methyltransferase Dnmt3L is facilitating the methylation of cytosine residues, this is in addition to its ability to stimulate the catalytic activity of DNMT3 A and DNMT3B as well as its ability to bind to chromatin leading to anchoring the DNA methylation machinery onto chromatin DNA templates [27]. This is an important issue in the DNA proofreading mechanism where DNA polymerase can accurately differentiate between the old and the newly synthesized DNA strand through cytosine methylation of the old DNA strand. The downregulation of DNMT3L in the early passages of MSCs may induce a defect in cytosine methylation of these cells, so the DNA polymerase cannot accurately differentiate between the original and the newly synthesized DNA strand. This may lead to defect in the DNA proofreading mechanism and persistence of mutation. This may explain presence of spontaneous P53 mutation in the early and late MSC cultures. Moreover, presence of mutation due to a defect in the DNA proofreading mechanism shows that the mutation likely occurs randomly along the DNA, not only in the P53 gene.
This scenario seems logical, especially because the majority of the 9 dysregulated epigenetic regulators are responsible for methylation and demethylation. One of these 9 regulators is zbtb16 (PLZF), an important regulator induces methylation through activating DNMT1 [28]. This epigenetic regulator is down regulated in all MSC passages. Studies indicated that dysregulated zbtb16 leading to hypo-methylation resulting into activation of zbtb16 target genes including c-myc, c-kit and Crabp1 [28]. Interestingly, both zbtb16 and DNMT3L have a common target; it is the retrotransposon L1 where DNA methylation activity of these 2 regulators induces full silencing of L1 gene. It was found that cell stress inhibit the methylation activity of zbtb16 through translocation from the nucleus to the cytoplasm [29]. So, cell stress resulting from the in vitro expansion of MSCs may be responsible for inactivation of zbtb16 in our MSC culture model.
Presence of P53 mutations in the early and late passages of our MSCs culture model confirms the transformation of these cells. The detected mutations were found in different human tumors [30, 31]. Li et al. studied the P53 mutation in the DNA binding domain of these cells and found 5 point mutations in the late-passages, 3 of which were pathogenic [18]. In the present study, we followed up the progression of P53 mutation in the early and late passages and we found 4 types of mutations. In passage 3, the mutation was non-pathogenic in human and did not appear in the other passages. The other 3 types of mutations were pathogenic in human and one of which was frequent in all tested passages starting from passage 6. Previous studies found that more than half of human cancers carry loss of function mutations of P53 [32] and 95% of these mutations were detectable within the genomic region encoding the DNA-binding domain [33].
Interestingly, many studies found a correlation between P53 mutation and abnormal expression of cbx7 [34], DNMT3L [35], RNASEL [36], Zbtb16 [37] and smarca1 [38]. These elements are abnormally expressed in different types of malignancies and are associated with P53 inhibition.
MSCs used in our study were examined for tumor formation by injection in a C57BL/6J mice model for gastric cancer where tumor was formed after 18 months and by examining the resulting tumor, the injected MSCs were detected (11). Moreover, the authors found that they can decrease the time for tumor formation by introducing MSCs which have been repeatedly passaged in culture for more than a year. The results of the in vivo experiments clearly support our findings assuming the in vitro transformation of MSCs.
The present findings may suggest a model for in vitro transformation of MSCs where the stress on MSCs due to repeated replication in culture, may induce the epigenetic regulators for abnormal expression. The upregulation of demethylases and downregulation of methylases in the early passages may affect the DNA proofreading mechanism resulting into genomic instability and accordingly frequent P53 mutation. Persistence of P53 mutation in parallel to the abnormal expression of epigenetic regulators may guarantee the transformation of MSCs. By time, MSCs accumulate mutations enough to induce a malignant transformation.
Although we did this study on mice MSCs and this may explain the large number of mutations, but we must be aware that we are studying a group of factors that regulates the expression of genes. These epigenetic regulators have a common function in both human and mice. Abnormality in any of these regulators may lead to genomic instability resulting in malignant transformation. Regardless the number of mutations, abnormality in any of these regulators will have an adverse effect on gene expression and accordingly disturbance in many processes and pathways in both mice and human cells. Regarding the P53 mutations in our study, we found the same types of mutations in different human cancers. Therefore, we must take the utmost care and ensure the safety of MSCs before applying these cells in the treatment, especially if the cells used were passaged in culture.
Conclusions
From the present study, we may conclude that BM derived MSCs undergo transformation in culture toward malignancy. The cells are characterized by abnormal expression of 62 epigenetic regulators. Nine epigenetic regulators are dysregulated in all passages. Upregulation of demethylases and downregulation of methylases is characteristic in the early passages. P53 mutation is prominent in all passages. A single base substitution mutation is frequent in the early and late passages, while deletion and insertion mutations are detected in the late passages. Alterations of the epigenetic regulators in the early and late passages of BM derived mouse MSCs result in genomic instability and frequent P53 mutation and, accordingly, malignant transformation. Targeting the abnormally expressed epigenetic regulators in MSC cultures and studying the impact of targeting on in vivo tumor formation is clearly needed to examine the therapeutic role of these regulators.
Funding
Dr. Ayman Metwally received a 6-month post-doctoral fellowship from the Science and Technology Development Fund (STDF) Egypt, under the US-Egypt Junior Scientist Development Visit Grants to Dr. Houghton’s Laboratory at the University of Massachusetts Medical School, USA. The funding agency supported the travel and salary for Dr. Metwally during the fellowship period.
Author contributions
Conception: Ayman Metwally and JeanMarie Houghton
Interpretation or analysis of data: Ayman Metwally
Preparation of the manuscript: Ayman Metwally
Revision for important intellectual content: JeanMarie Houghton and Hanchen Li
Supervision: JeanMarie Houghton
Methodology: Ayman Metwally and Hanchen Li
Validation: Ayman Metwally and JeanMarie Houghton
Supplementary data
The supplementary files are available to download from
sj-docx-1-cbm-10.3233_CBM-203121.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-203121.docx
Footnotes
Acknowledgments
We thank Dr. Eman Bayomi, Department of Medical Molecular Genetic, National Research Center, Egypt for revising and editing the p53 sequence results.
