Abstract
OBJECTIVE:
To explore the effects of the programmed death-1/programmed death-ligand 1 (PD-1/PD-L1) signaling pathway on the intestinal flora in patients with colorectal cancer (CRC).
METHODS:
A total of 30 CRC patients treated with PD-1 monoclonal antibody therapy in the Oncology Department of our hospital from January 2018 to January 2019, and another 30 patients treated with routine non-immune therapy were enrolled. The feces specimens were collected for sequencing, the CRC model was established, and the 16S rRNA gene sequences in intestinal flora in feces specimens of mice were analyzed.
RESULTS:
The 3-month progression-free survival could not be predicted through the gene count or abundance of metagenomic species (MGS) in intestinal microflora of patients. The gene count or MGS abundance was related to the clinical progression-free response. There were abundant unclassified Escherichia coli, s_lactobacillus and s_unclassified parasutterella in patients treated with PD-1. The reflection curve of microbiota had an obvious difference in richness (Chao1), but had no apparent difference in diversity (Shannon).
CONCLUSION:
The PD-1/PD-L1 signaling pathway can regulate the metabolic activity of intestinal flora, thereby promoting immune surveillance of tumors.
Introduction
As a complex ecosystem, the human body is affected by a large number of microorganisms, including bacteria, yeasts, fungi, archaea and viruses, and they jointly constitute the symbiotic flora. It is implicated to the changes in the nutrient transformation, immune response and heterologous biological metabolism, and is considered as an important component of host physiology [1]. These functions are performed through a group of complex and multi-layered controls from the microbiota and host factors. Therefore, such a complex partnership between microorganisms and hosts is considered as a basic element of health. For example, the patients with inflammatory bowel disease (IBD) and colorectal cancer (CRC) have different intestinal bacteria, fungi and viruses compared with healthy subjects [2, 3, 4]. Currently, the effects of microorganisms on the occurrence, development and treatment of diseases have been widely studied. Microorganisms are particularly attractive in the field of disease treatment, partially because great success has been achieved in the treatment of recurrent clostridium difficile infection with fecal microbiota transplantation (FMT) [5]. At present, whether FMT or derived synthetic cocktails can be applied to specific gastrointestinal diseases remains unclear, and such a problem is being solved currently through many clinical trials for treatment of metabolic syndrome, diabetes and IBD.
The microbiota can synthesize or transform a variety of metabolites, including hormones, essential vitamins and other biologically active compounds, which cannot be obtained by the host [6]. These metabolites can regulate various biological functions, especially the immune system and nervous system [7]. It is reported that the changes in normal microbiota can cause many diseases [8]. In the case of cancer, some specific bacteria have been proved to be involved in the carcinogenesis. In addition, it has also been found that the microbiota regulates the efficacy and toxicity of cancer treatment, including chemotherapy and immunotherapy [9].
In the field of cancer, the interaction between bacteria and therapeutic drugs has drawn extensive attention. Bacteria on the other hand present impact on therapeutic drugs, including regulating the efficacy and toxicity of chemotherapeutic and immunotherapeutic agents through the metabolic and immune-mediated mechanisms [10]. In particular, it was recently discovered that the intestinal flora can profoundly affect the response of cancer patients to immune checkpoint-blocking therapy, attracting much attention. In this study, the effects of the programmed death-1/programmed death-ligand 1 (PD-1/PD-L1) signaling pathway on the intestinal flora in patients with CRC were explored using the clinical and animal specimens.
Materials and methods
Clinical specimens
A total of 30 CRC patients treated with PD-1 monoclonal antibody (pembrolizumab) therapy in the Oncology Department of Dongguan Eastern Central Hospital from January 2018 to January 2019, and another 30 patients treated with routine non-immune therapy (chemotherapy) were enrolled, from whom the feces specimens were collected. No probiotics/antibiotics were adopted in both groups. All patients were informed of the study, and the study regarding both patients and animals was approved by the Ethics Committee of Dongguan Eastern Central Hospital.
Animal grouping
The procedures for animal use had been reviewed and approved by the Laboratory Animal Ethics Committee of Dongguan Eastern Central Hospital. In this study, a total of 96 male BALB/c mice weighing about 20 g and aged 5 weeks old were purchased, and they were all adaptively fed for 1 week under pathogen-free conditions and had free access to food and water.
The mouse CRC CT26 cell lines were cultured in the RPMI-1640 medium (Gibco, Gaithersburg, MD, USA) supplemented with 10% heat-inactivated fetal bovine serum in an incubator with 5% CO
16S ribosomal ribonucleic acid (rRNA) gene sequence analysis of intestinal flora in feces specimens
Sequencing was performed by Aksomics (Shanghai, China). The total deoxyribonucleic acid (DNA) was isolated from the feces specimens, and the bacterial 16S rRNA gene V3-4 region was amplified via polymerase chain reaction (PCR) using the forward primer (5’-AYTGGGYDTAAAGNG-3’) and reverse primer (5’-TACNVGGGTATCTAATCC-3’) according to the instructions. Then the PCR products were separated via gel electrophoresis and purified using AP-GX-500 DNA gel extraction kit (Axygen, Corning, USA). The library was built using the resulting products, followed by sequencing on the MiSeq sequencing platform (Illumina, USA) in the way described by Zhao et al.
Sequencing is performed for feces specimens of all cancer patients at the time of diagnosis according to the clinical results. The gene count is expressed as mean 
The trimmed and assembled sequences of each sample were compared with Greengene 16S rRNA database set 10 using the best matching classification option, so as to classify the taxonomic abundance in Quantitative Insights into Microbial Ecology (QIIME). The bacterial operational taxonomic unit (OTU) was generated using the UCLUST function in QIIME. The following statistical data were obtained using the R software: ACE, Chao, Simpson and Shannon indexes were calculated for
Sequencing of feces specimens at the time of diagnosis. The relative abundance of each MGS in responder (R) (partial response or stable disease) relative to that in non-responder (NR) (progressive disease) according to the RECIST 1.1.
Metagenomics assessment and Cochran-Armitage test: detection frequency of patients with Aspergillus mucus in feces according to the clinical status partial response (PR), stable disease (SD) or progressive disease (PD).
SPSS 17.0 (SPSS Inc., Chicago, IL, USA) and GraphPad Prism 5.0 (GraphPad, La Jolla, CA, USA) were used for statistical analysis. First, the normal distribution test was performed for the data set in each experiment. The data were analyzed by one-way analysis of variance (ANOVA) in the case of normal distribution, followed by pairwise comparison according to the homogeneity test of variance and different parameter tests. Otherwise, the data were compared using Kruskal-Wallis H test. In ANOVA, the homogeneous difference was analyzed via post-hoc LSD test, and Dunnett’s test was used if it was not variance.
Results
Patient characteristics and clinical outcomes
From January 2018 to January 2019, 30 CRC patients as treatment group received PD-1 monoclonal antibody (mAb) therapy while another 30 patients as control group underwent routine non-immune therapy. The mean age from two groups was 66.5
Response of cancer patients after 3 months of PD-1 mAb therapy predicted via genomic analysis of feces specimens
Sequencing was performed for feces specimens of all cancer patients at the time of diagnosis according to the clinical results. It was found that the 3-month progression-free survival (PFS) could not be predicted through the gene count or abundance of metagenomic species (MGS) (Fig. 1).
Shotgun sequencing of feces specimens at the time of diagnosis
To explore the composition of the intestinal flora, over 20 million short-read DNA sequences in each specimen were sequenced via quantitative metagenomics, and then 9.9 million genes were analyzed in references. The results showed that the gene count or MGS abundance of the evaluated specimen was related to the clinical progression-free response (Figs 2 and 3).
PD-1/PD-L1 regulated intestinal microflora
The microflora specimens in the intestine (feces) were prospectively collected from tumor-bearing mice. Before the anti-mouse PD-1 mAb therapy, the composition of the intestinal microflora was first evaluated, and the relative diversity of the two communities was clarified (Fig. 4A). The results of principal coordinate analysis (PCA) revealed that the microbiota structure was not clearly separated between the two groups (Fig. 4B). Then whether there are differences in the intestinal microflora between the two groups was determined. Interestingly, in the comparison of composition of common bacterial groups using Wilcoxon rank sum test, at the species level there were abundant unclassified Escherichia coli, s_lactobacillus and s_unclassified parasutterella in patients treated with PD-1. To further explore these findings, high-dimensional comparison was performed via LEfSe, confirming the above results again (Fig. 5).
A) PD-1/PD-L1 regulates composition of intestinal microflora. Stacked bar of composition of common bacterial groups in feces specimens at the start of immunotherapy in animal models. B) PCA of OTU.
LDA score of abundant taxonomic units in the fecal microbiota in animal models. 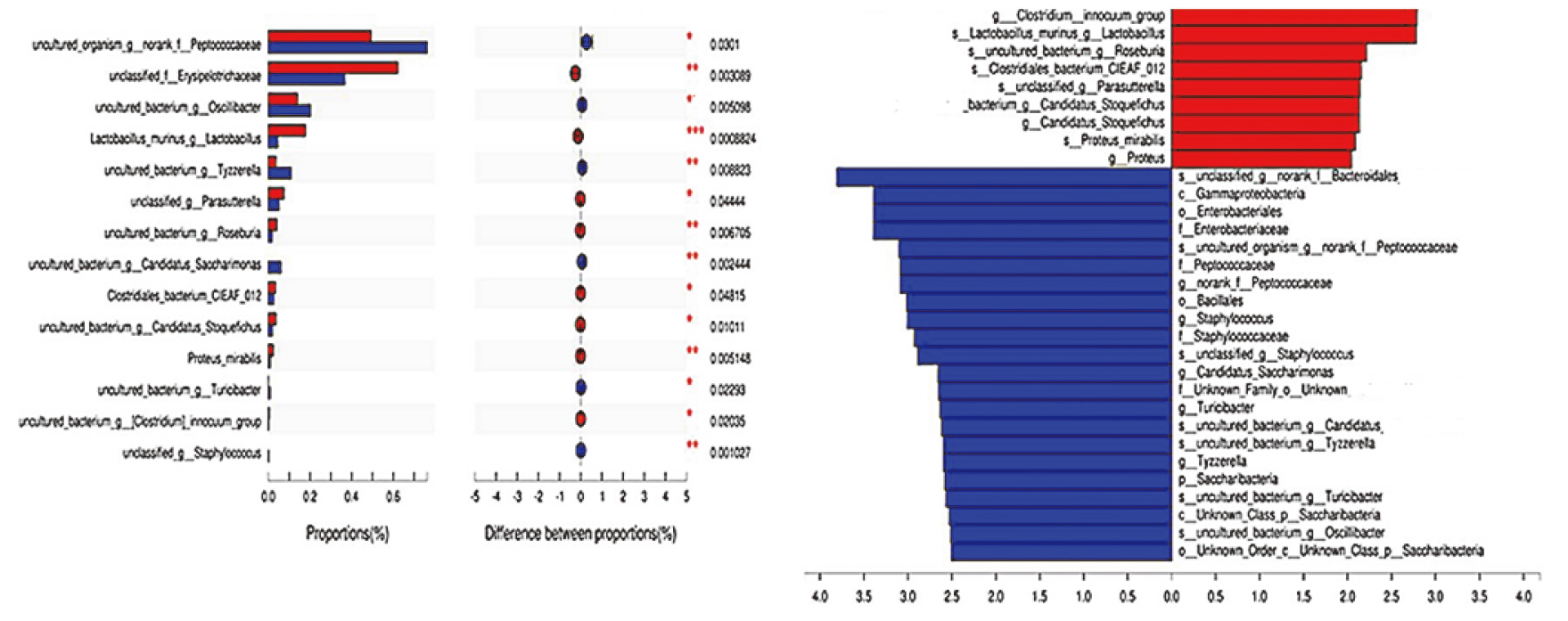
The diversity of microbiota was first analyzed via
Reflection curve of 
Biological function predicted based on the KEGG database. 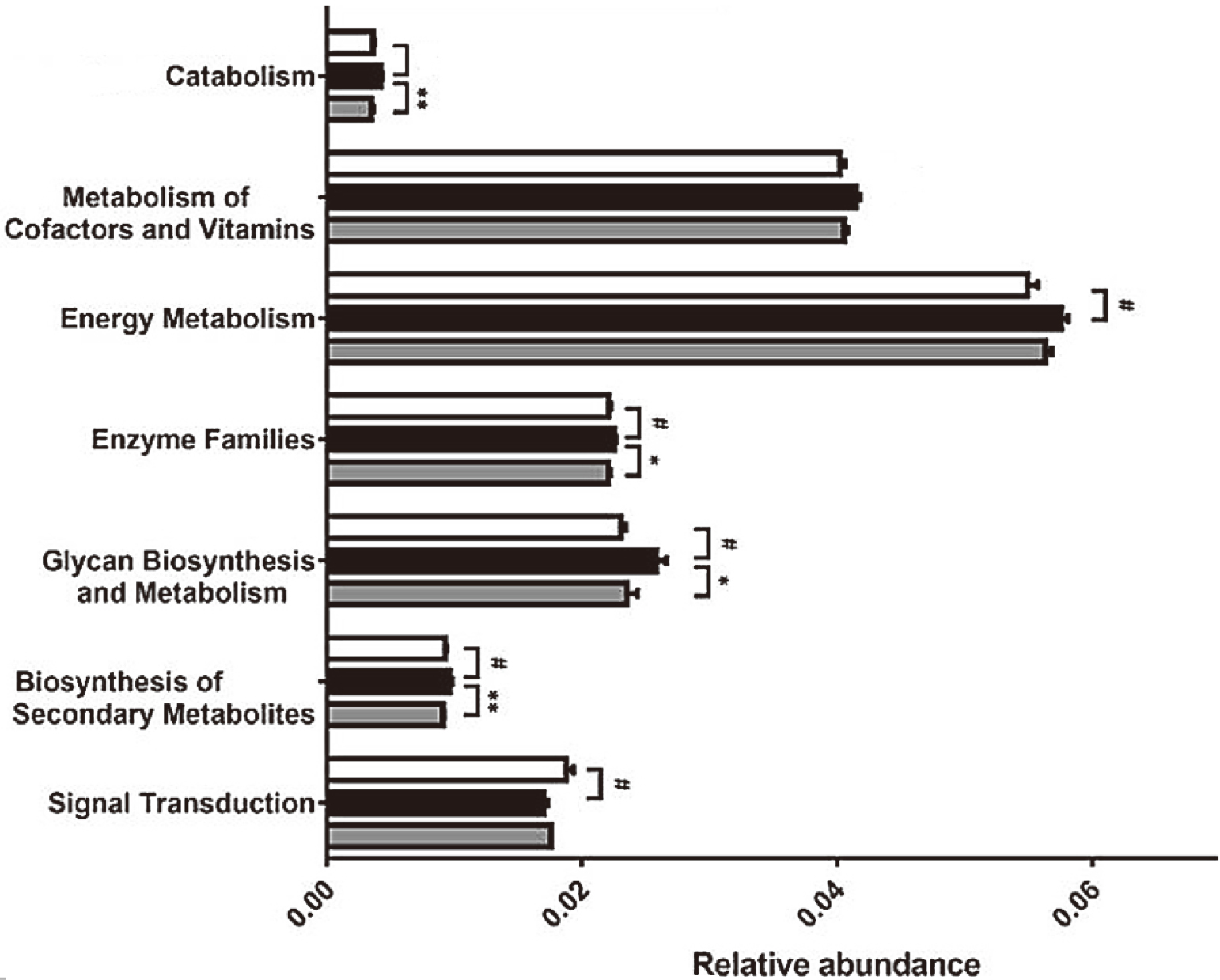
The regulatory effect of PD-1 treatment on microbial function was analyzed via comparison of sequencing data with those collected in the KEGG database (Fig. 7). It was found that the abundance of several genes related to five metabolic pathways was obviously up-regulated in tumors treated with PD-1, including those involved in “biosynthesis of secondary metabolites”, “energy metabolism”, “enzyme family”, “glycan biosynthesis and metabolism”, “cofactor and vitamin metabolism” and “transport and catabolism” (a cellular process pathway), but the abundance of genes involved in “cell mobility” and “signal transduction” pathways was down-regulated. After ESGH treatment, the changes in genes involved in most of these pathways (such as “transport and catabolism”, “enzyme family”, “glycan biosynthesis and metabolism” and “biosynthesis of secondary metabolites”) were remarkably reversed, indicating that PD-1 can regulate the metabolic activity of intestinal flora, thereby promoting the immune surveillance of tumors.
Discussion
The constant development of new clinical trials for various cancers may help better understand the role of the microbiota in mediating the efficacy of immune checkpoint inhibitors [11]. Before the microbiota is sampled and included in these trials, larger cohort studies may also be needed to identify potential microbial markers that define response.
In addition to T-cell targeted therapies and immune checkpoint inhibitors, the relation between microbiota and cancer management remains to be studied. For example, understanding the role of the microbiota in other immunotherapies, such as cancer vaccines, oncolytic viruses and cell-mediated therapies (chimeric antigen receptor T-cell therapy), may help determine the specific treatment-related microbiota components or overlapping signals appropriate for the extensive immunotherapy. Another important viewpoint about the relation between microbiota and immune checkpoint responses is the source of signal. Great efforts have been made to link drug efficacy to bacteria, but other signals may be hidden due to the complexity of microbiota. As mentioned above, fungi and viruses are intrinsic parts of the microbiota, which have been proved to affect the immune response [12, 13, 14]. Therefore, it will be important to study the role of these microorganisms in defining the interaction between microbiota and cancer treatment. To sum up, searching for microbial signals that determine the response to cancer treatment is an ongoing task, which is expected to provide exciting new examples in the research field of cancer in next few years.
CRC is a multifactorial disease associated with a variety of lifestyles. Preclinical models suggest that the different compositions of the intestinal microflora may affect the response to anti-PD-1 therapy at the level of tumor microenvironment [15, 16, 17]. Therefore, the regulatory effect of anti-PD-1 therapy on intestinal flora was explored.
In this study, the results revealed that the intestinal microflora significantly affected the PD-1 blockade in mice and patients, which validated the recent study on cancer immunotherapies that the composition of the intestinal microflora has a major impact on patient prognosis [18]. However, several problems remain unresolved. For example, the mechanism of the immunomodulatory effect of Aspergillus mucus, one of the most abundant bacteria in the ileal microbiota, is unknown. Intestinal barrier dysfunction and systemic endotoxemia may be caused in cancer patients faced with stress. By enhancing the integrity of intestinal barrier and reducing systemic inflammation, Aspergillus mucus and Clostridium may form the “steady-state‘’ entity to prevent colonic and systemic immunosuppression [19, 20, 21]. At the same time, T cells released by PD-1 can reduce the tolerance of small intestine, thereby inducing local and systemic Th1 immune responses to Aspergillus mucus, which may improve the immune surveillance of cancer. Besides, the comprehensive analysis of fecal components may further reveal the functional connection between the microbial ecosystem and the anti-cancer immune surveillance due to the long-term migration of the microbiota caused by anti-cancer microorganisms [22, 23, 24]. The results in this study demonstrate that microorganisms control the immune checkpoint in cancer individuals, and regulate the intestinal ecosystem, which was in line with previous finding concerning the role of PD-1 in intestinal mucosa that the activation of PD-1 signaling pathway promoted host protection at the intestinal mucosa during acute infection with a bacterial gut pathogen [25]. However, the limitation in our study still exists that the predicted genes and their functions according to the KEGG database requires further validation and investigation.
In conclusion, our preliminary data demonstrate that programmed death-1/programmed death-ligand 1 (PD-1/PD-L1) signaling pathway plays a fundamental role on the intestinal flora in patients with colorectal cancer (CRC), which provides new insight for the strategies of immune surveillance towards tumors.
Footnotes
Conflict of interest
None.
