Abstract
BACKGROUND:
Coiled-coil domain containing protein-124 (Ccdc124) is a putative mRNA-binding factor associated with cell division, and ribosome biology. Previous reports mentioned an up-regulation of CCDC124 gene in cancer, and listed its mRNA in a molecular prognostic signature in breast cancer.
OBJECTIVES:
Establishing RNA-binding characteristics of Ccdc124 for a better molecular functional characterization, and carrying-out retrospective studies in order to evaluate its aberrant expression in human cancer samples from various tissue origins.
METHODS:
Bioinformatics calculations followed by RIP and RNA-seq experiments were performed to investigate mRNA targets of Ccdc124. Quantitative studies on arrays of cDNAs from different cancers and IHC assays on tissue arrays were used to assess CCDC124 expression levels in cancers.
RESULTS:
Ccdc124 was characterized as an RNA-binding protein (RBP) interacting with various mRNAs. CCDC124 mRNA levels were high in tumors, with a particular up-regulation in cancers from esophagus, adrenal gland, endometrium, liver, ovary, thyroid, and urinary bladder. IHC assays indicated strong Ccdc124 positivity in endometrial (95.4%), urinary bladder (68.4%), and ovarian cancers (86.8%).
CONCLUSION:
Ccdc124 is a cytokinesis related RBP interacting with various mRNAs. CCDC124 mRNA over-expression and an accompanied increase in Ccdc124 protein accumulation was reported in cancers, indicating this RBP as a novel cancer cell marker.
Keywords
Introduction
Cancer is a complex multifactorial disease typically characterized by uncontrolled cellular proliferation and metastatic spreading throughout the body. Worldwide, endometrial and ovarian cancers account for
Identification of cancer-conducive alterations in cellular molecular interactions has long been a subject of intensive study; and a number of such interactions have been shown to involve RNA-binding proteins (RBP) [2, 3]. RBPs assemble ribonucleoprotein (RNP) complexes with RNA, and they mediate virtually every step of the RNA metabolism [4]. Causal mutations and mis-expression of RBPs affect most stages of RNA lifecycle, including RNA splicing (e.g. SRSF2) [5], 3’ end processing (e.g. CPEB1) [6, 7], editing (e.g. ADAR1) [8], stability (e.g. ZFP36) [9], translation (e.g. eIF4E) [10], and biogenesis of small RNAs such as miRNAs (e.g. AGO2, LIN28) [11]. Alterations of RNA metabolism due to RBP dysfunction can cause global changes in the transcriptome and proteome of the cell, and can affect carcinogenesis related molecular mechanisms involved in cell growth, proliferation, invasion and apoptosis [12, 13, 14, 15, 16, 17, 18].
Coiled-coil-domain-containing protein 124 (Ccdc124) was first discovered in our past studies as a new centrosome and midbody localized protein that regulate cytokinetic abscission during progression of mitosis in cultured human cell models [19]. Our previous RNA-interference experiments have indicated that knocking-down Ccdc124 expression in HeLa cells leads to aberrant cytokinesis, as characterized by time-lapse imaging monitored formation of multinucleated cells [19]. Importantly, Ccdc124 is encoded by a gene that is broadly conserved in eukaryotes, and predictions based on in silico analysis of functional domains in Ccdc124 by the web-based NCBI/UniProt software platform indicated it as a putative RBP (
A comprehensive data mining based study by Kechavarzi and Janga analyzing comparative expression patterns of RBPs in healthy human tissues and in nine human cancers (brain, breast, colon, kidney, liver, lung-adeno, lung-squamous, prostate, and thyroid cancers) by using RNA-seq data available from the Human BodyMap (HBM) and the Cancer Genome Atlas (TCGA) [
In the present study, we initially carried out in silico analysis on Ccdc124 to predict localization of its putative RBDs, and for initial characterization of possible RNA-interaction parameters for this protein. Then, we transfected HeLa cells with a vector expressing eGFP tagged version of CCDC124 under CMV-promoter, and subsequently we used RNA-immunoprecipitation (RIP) followed by RNA-sequencing methods for experimental validations of interactions between Ccdc124 and mRNAs. Furthermore, we investigated the versatility of Ccdc124 as a cancer cell marker, initially by examining CCDC124 expression change in human cancer tissues and normal tissues using a TissueScan array (OriGene, USA) spotted with cDNAs derived from 18 different types of cancer/normal tissue origins (
Materials and methods
Tissue cDNA arrays
TissueScan Cancer Survey cDNA Array (OriGene Technologies Inc., Rockville, MD, USA, Catalog no. CSRT-303) was used as described in manufacturers protocols in order to screen expression level of Ccdc124 in samples of indicated human cancer types. This cDNA array contained human samples from 381 patients, and included both cancerous tissues and normal samples from healthy tissues.
To investigate the mRNA expression level of Ccdc124 in ovarian cancers, a distinct cDNA array containing 48 samples (OriGene Technologies Inc., Rockville, USA, cat. no. HORT 303) covering 6 normal cDNA samples, and cancer cDNAs from 9 different stages (IA, 2-IB, 3-IC, 3-IIB, 1-IIC, 1-II, 3-IIIA, 6-IIIB, 12-IIIC, 1-III,1-IV) was used, as previously mentioned.
Ccdc124 expression analysis by real-time quantitative PCR (qPCR)
Real-time qPCRs were performed with Syber Green using Stratagene Mx3000P. Each reaction was performed at least in triplicate in 25
Fluorescence microscopy
Fluorescence microscopy pictures were taken at 10x magnification with FITC filter using Nikon Eclipse T
Bioinformatics calculations by catRAPID algorithm predicts Ccdc124 as an RNA-binding protein. A) Prediction of interaction propensity of Ccdc124 protein by catRAPID analysis. The graphic indicates classical (0.57), putative (0.35) and non-classical (0.28) RBP classes, and the overall prediction score of Ccdc124 as 0.65. B) RNA-binding propensity profile of Ccdc124 protein showing putative RNA-binding domains and their RNA-interaction propensity. C) The algorithm identifies peaks corresponding to regions between residues 74-124, 54-67, and 152-181 of Ccdc124.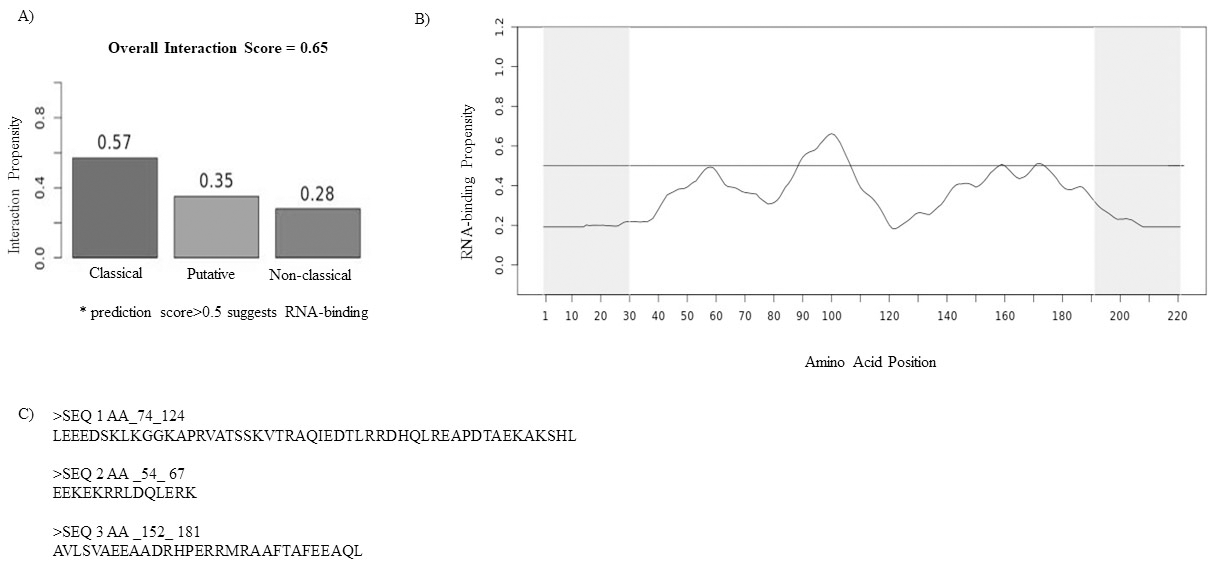
The expression level of CCDC124 in ovarian, endometrium and urinary bladder cancer tissue samples was analyzed by using human cancer tissue microarrays (US Biomax Inc., Rockville, MD; cat. no. OV485, BL 1002a and UT501a). To prevent tissue detachment during antigen retrieval, slides were baked for 1 h at 60
Statistical analysis
Statistical analysis was performed using GraphPad Prism software version 6.1 (GraphPad Software, San Diego, California, USA). The results were presented as data obtained in three different independent experiments. Cancer samples data was analyzed with Mann-Whitney test (
Results
In silico analysis of Ccdc124 to predict its possible RNA binding potential
NCBI database search indicated a possible RNA binding function for Ccdc124 (
Characterization of Ccdc124 as an mRNA binding protein
A reliable technique for identifying targets of a RBP is RNA-immunoprecipitation (RIP) followed by the identification of protein bound RNAs by RNA-sequencing (RIP-seq) [29, 30]. We used a human cervix cancer cell model, HeLa, and transfected the cells with vectors containing either GFP-linked version of CCDC124 under control of cytomegalovirus (CMV) promoter (CMV-CCDC124-GFP), or related empty CMV-GFP vector as control (Fig. S1). Then, we used either anti-Ccdc124 antibody or anti-GFP antibody coated sephadex beads in order to precipitate Ccdc124-RNA complexes from lysates prepared by using untransfected HeLa cells (controls), or cells transfected either with pEGFP-N2 or pCMV-CCDC124-GFP vectors (Fig. S1). As a confirmatory approach, RIPs were performed by using both anti-Ccdc124 antibody and anti-GFP antibody covered beads in two different sets of experiments on lysates obtained from CMV-CCDC124-GFP transfected HeLa cells. Same immunopurification protocols were also followed with lysates obtained from untransfected cells, or cells transfected with CMV-GFP vector, as controls. Following RIP experiments, immunoprecipitated Ccdc124-bound RNA samples were analyzed by RNA-sequencing methods (Suppl. Materials and Methods and
Up-regulation of Ccdc124 mRNA in cancers of different tissue origins
Besides playing important roles in cellular homeostasis by controlling gene expression at the post-transcriptional level, RBPs are also involved in various pathways related with RNA-splicing, translation, cellular stress response, and innate intracellular immunity [31, 32, 33]. Mutations mapping various RBPs and alterations in expression levels of corresponding transcripts were mostly associated with degenerative diseases, but also with cancer [34, 35, 36]. Previously, two different in silico data-mining studies focusing on levels of RBP encoding transcripts in cancers, and on MYB transcription factor dependent gene signatures in breast cancers have indicated that CCDC124 mRNA expression could be significantly upregulated in different types of cancers [22, 23]. Therefore, after proving RNA binding capacity of Ccdc124, we aimed to study in detail the expression level of CCDC124 gene in cancer tissue samples. For this purpose, we took advantage of a ready-made human cancer and normal tissue cDNA arrays (TissueScan, Origene, USA) containing materials from 381 tumor/normal paired human samples coming from 18 different types of cancers in an array ready for qPCR analysis (Supplement Table S4). Thus, we examined CCDC124 mRNA levels in normal/cancer tissue pairs coming from adrenal gland, breast, esophagus, kidney, liver, lung, lymphoid tissue, ovary, pancreas, prostate, stomach, testis, thyroid gland, urinary bladder, uterus, and thyroid gland tissues, as described in Materials and Methods. Our data indicated that as compared to healthy tissue pairs, CCDC124 mRNA expression level was significantly high (from
Expression of CCDC124 mRNA in human normal or cancer tissue samples. Transcript profiling was performed by using a ready-made cancer tissue qPCR array, as described in Methods. cDNA concentrations were normalized to 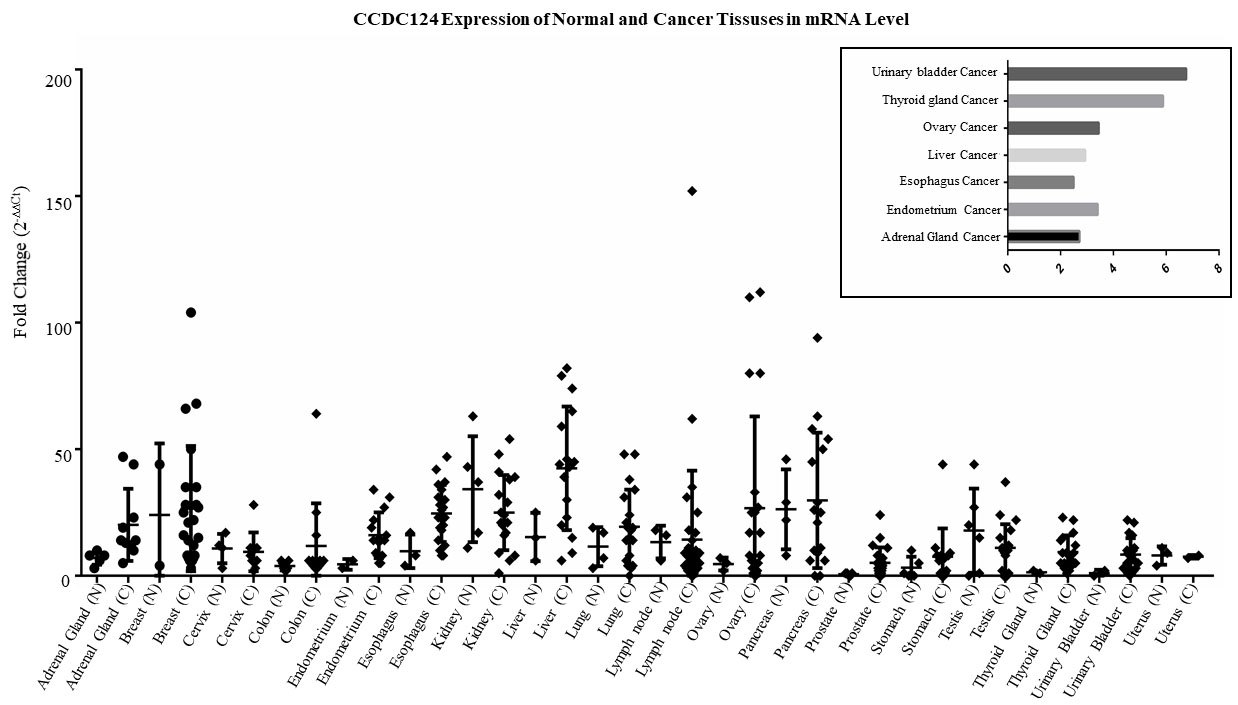
Comparisons of Ccdc124 mRNA expression levels in a selected set of cancer types. A) Comparative analysis of expression of Ccdc124 in normal and cancer tissues originating from endometrium, esophagus, urinary bladder, thyroid and liver. Asterisks indicate statistically significant differences (Mann Whitney test *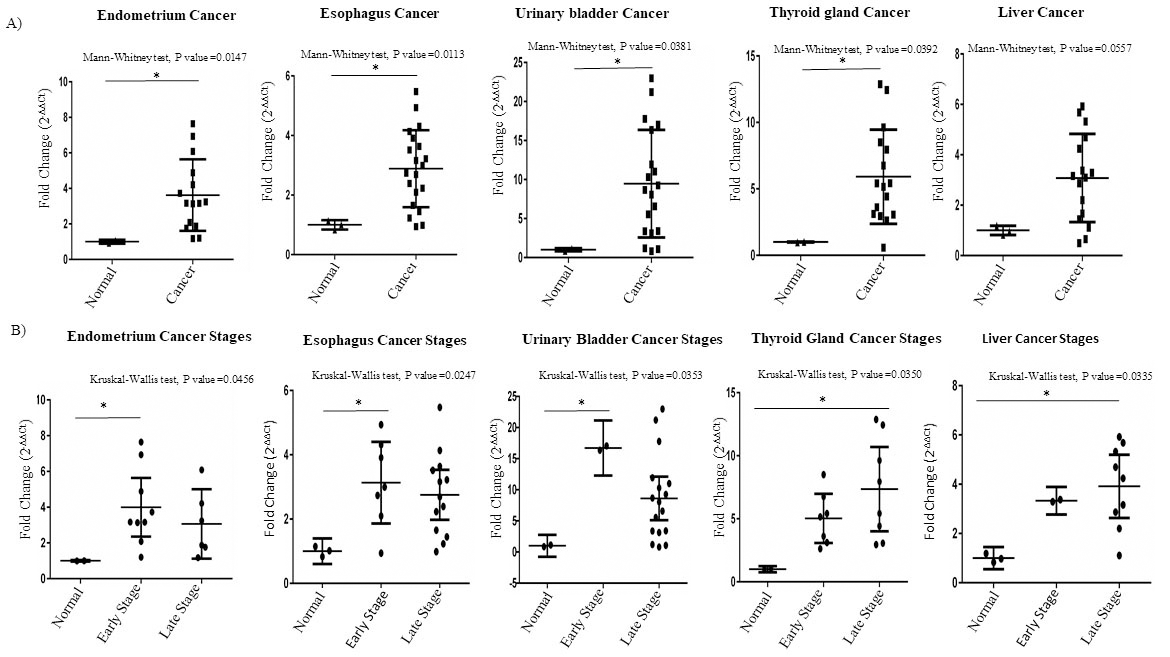
Analysis of Ccdc124 mRNA expression in adrenal gland and ovarian cancers. A) Graphical expressions of fold changes in Ccdc124 mRNA levels in adrenal gland and ovarian cancers. Asterisks indicated statistically significant differences (Mann Whitney test *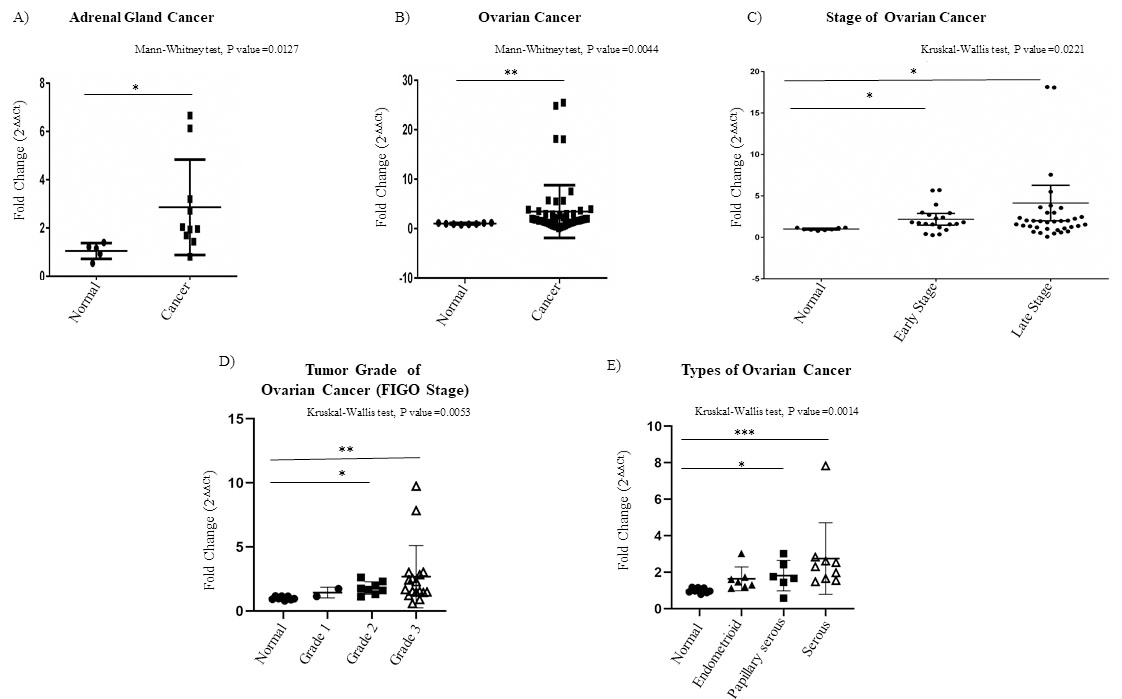
Interestingly, we observed that the CCDC124 mRNA was particularly high at the early stages of endometrium, esophagus and urinary bladder cancers, while it was high at relatively later stages of thyroid and liver cancers (Fig. 3B). Significantly, we detected high CCDC124 mRNA level in cancers originated from adrenal gland and ovary tissue, independent of tumor stage (Fig. 4A–C). Yet, when ovarian cancer samples were compared based on FIGO tumor grading system [37], CCDC124 mRNA expression was more pronounced in Grades 2 and 3 samples which were histologically classified as endometrioid, papillary serous and serous, as compared to normal tissue pairs (Fig. 4D and E).
Correlations between Ccdc124 protein immunohistochemistry staining and clinicopathological data associated with endometrium cancer samples in TMA
Immunohistochemical analysis of Ccdc124 protein using specific antibodies on paraffin embedded endometrial carcinoma and normal tissue sections in ready-made tissue arrays. A) IHC stained sections from normal tissue (magnifications, 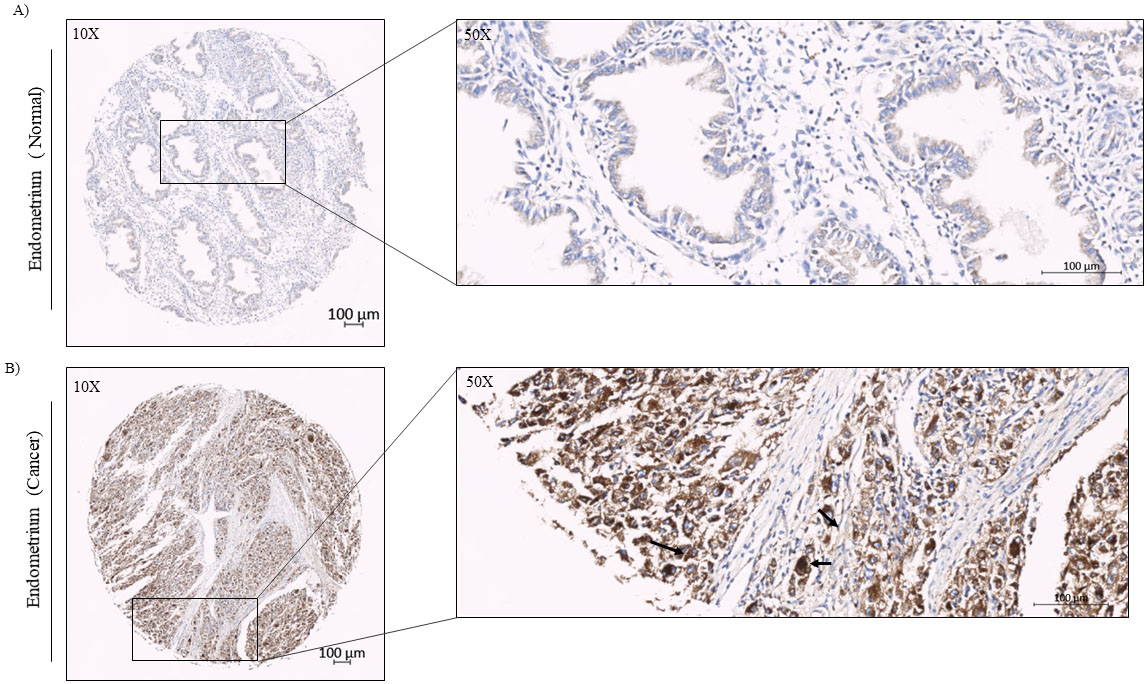
Immunohistochemical analysis of Ccdc124 protein using specific antibodies on paraffin embedded endometrial cancer and normal tissue ready-made protein array samples. Immunohistochemical analysis of Ccdc124 protein in normal endometrium and endometrial cancer microarray tissue samples. A) IHC staining of human endometrial cancer tissues at 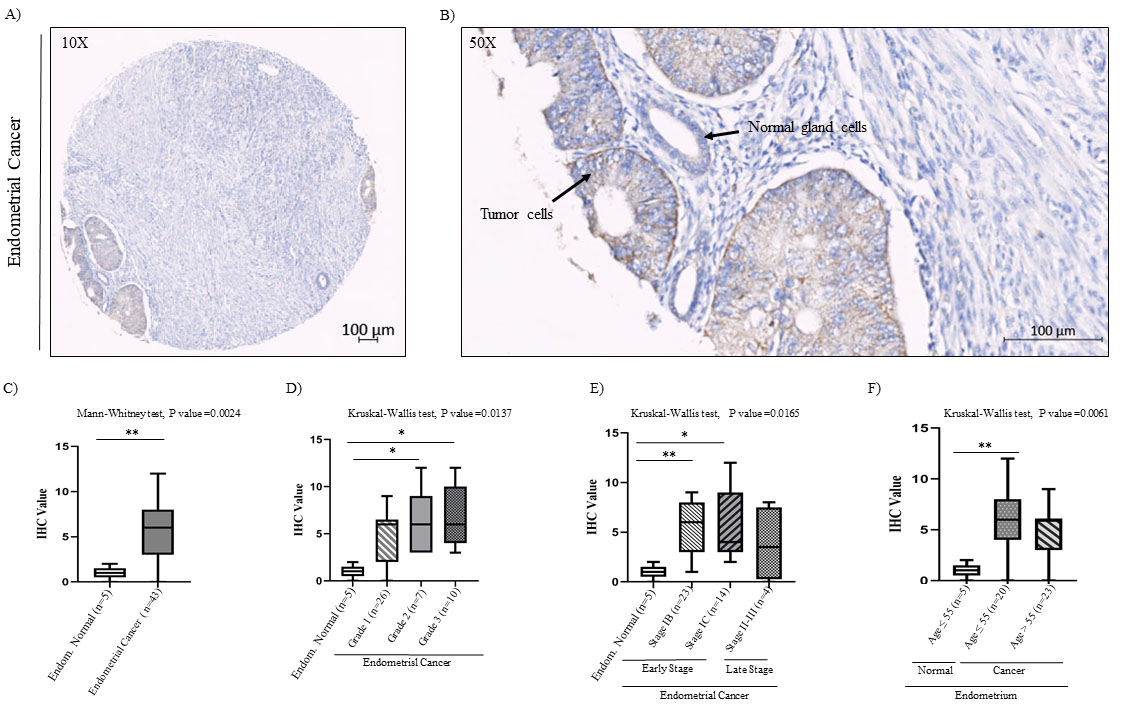
After identifying increased CCDC124 mRNA levels in cancers originated from urinary bladder, thyroid gland, ovary, endometrium, liver, adrenal gland, and esophagus tissues, we decided to examine by immunohistochemistry (IHC) methods the possible increase of Ccdc124 protein in this subgroup of cancers. For this purpose, we used ready-made tissue microarrays (TMA) obtained from commercial sources (US Biomax, USA). For IHC analysis; human endometrial, ovarian, and urinary bladder cancer TMAs were stained with anti-Ccdc124 antibodies (;ioss, USA) suitable for use on paraffin embedded tissue sections. Then, we evaluated tissue-specific Ccdc124 positivity, as it is done routinely by the presence of dark-colored areas in histopathological examination (Supplement Fig. S3). The results showed that regardless of the origin of cancer, Ccdc124 had broadly cytoplasmic reactivity and major epithelial distribution (for example, Figs 5 and 7). In normal endometrial TMA samples, we detected low staining pattern of Ccdc124 antibody in endometrial gland cells and no reactivity in endometrial stromal cells (Fig. 5A). However, in endometrial cancer TMA samples (
Regarding Ccdc124 expression in normal urinary bladder TMA samples, we detected a relatively low staining pattern in transitional epithelium and glomerulus, but not in neighboring smooth muscle tissue (Fig. 7A). However, IHC analysis of Ccdc124 presence in urinary bladder cancer samples indicated 44.7% low, 15.8% moderate, 7.9% strong staining patterns (
Correlations between Ccdc124 protein immunohistochemistry staining and clinicopathological data associated with urinary bladder cancer samples in TMA
Correlations between Ccdc124 protein immunohistochemistry staining and clinicopathological data associated with urinary bladder cancer samples in TMA
Immunohistochemical analysis of Ccdc124 protein levels in normal and urinary bladder cancer tissue microarray samples. A) IHC staining of human normal urinary bladder tissue at 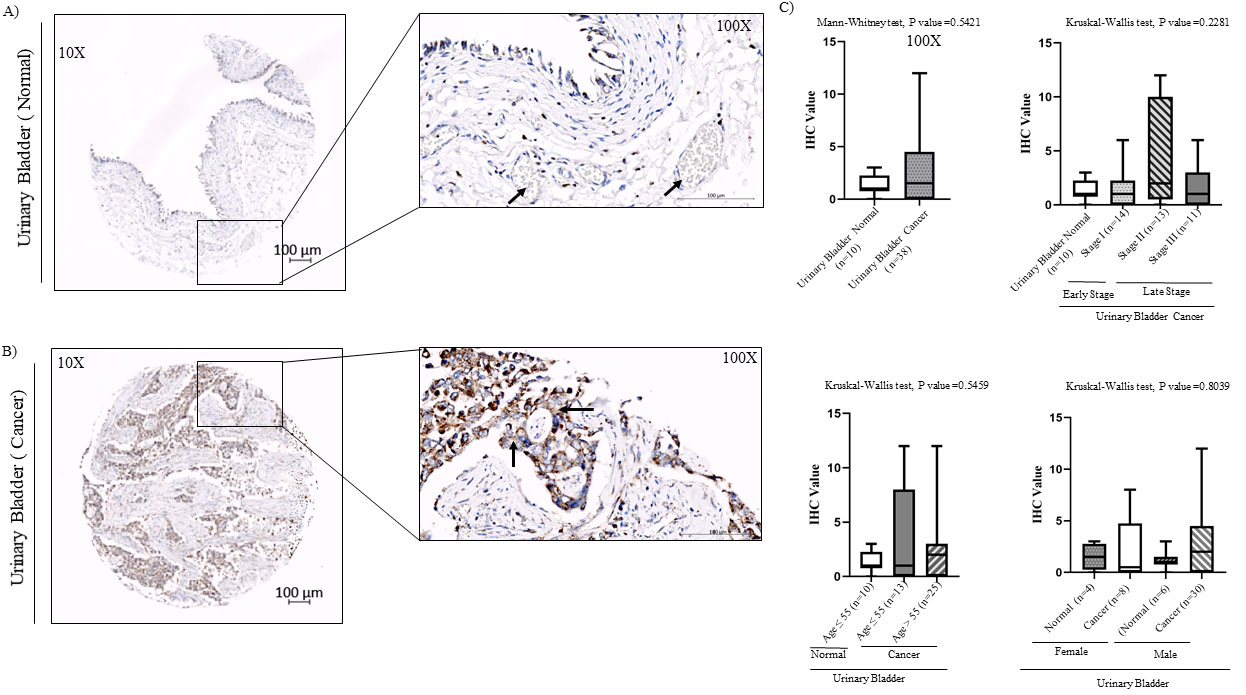
Subsequently, we carried-out similar analysis in ovary cancer TMA samples. In this analysis, we first noticed that Ccdc124 protein was predominantly detectable in the cytoplasm of oocytes in normal ovary tissue sections, but not in the stromal cells of ovaries (Fig. 8A). Relatively low IHC reactivity was detected in ovarian cancer samples (
Correlations between Ccdc124 protein immunohistochemistry staining and clinicopathological data associated with ovarian cancer samples in TMA
Immunohistochemical analysis of cell types expressing Ccdc124 protein in normal and ovarian carcinoma tissues. A) anti-Ccdc124 Ab stained sections in normal ovary tissue (magnification, 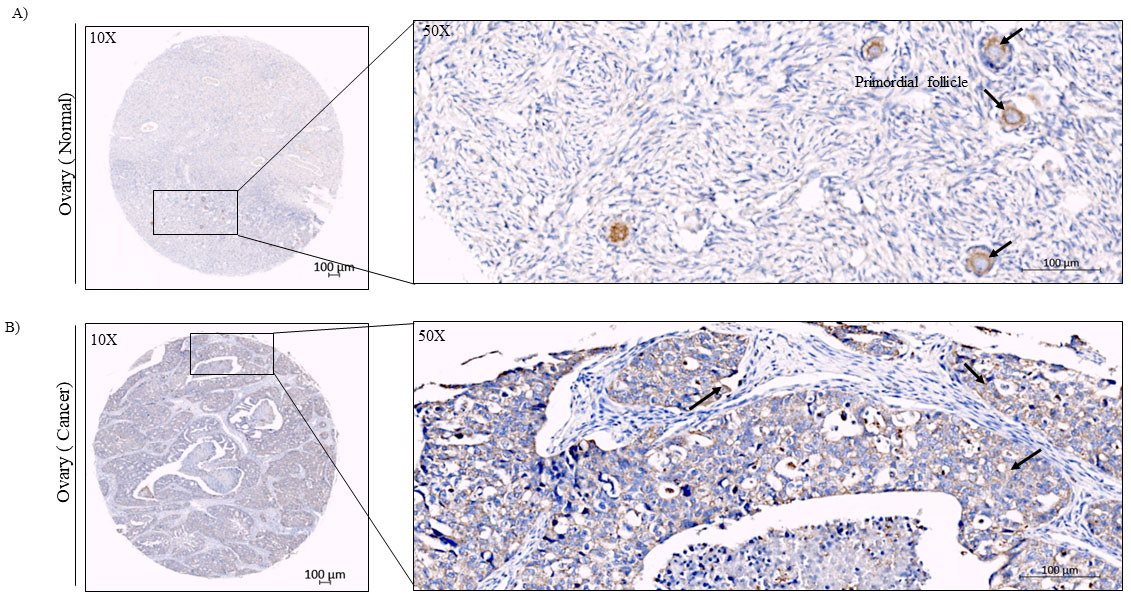
Immunohistochemical analysis of Ccdc124 protein levels in normal and ovarian cancer tissue microarray samples. A) IHC staining of human ovarian mucinous papillary adenocarcinoma tissue at 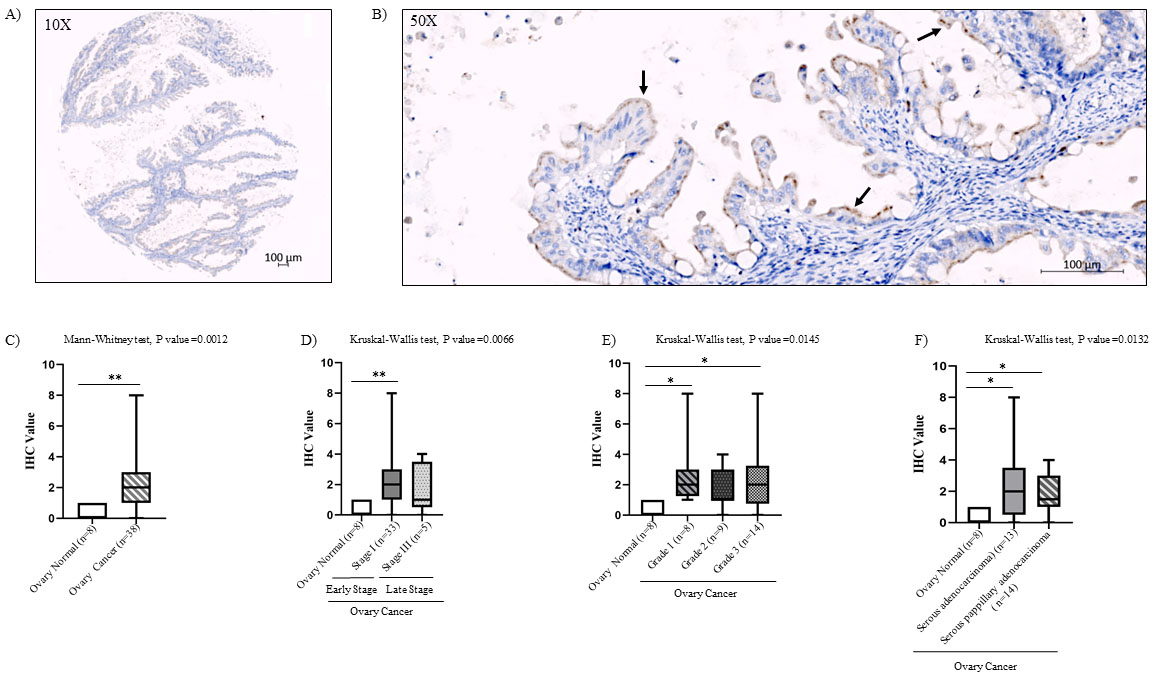
In the present study, firstly we have addressed RNA-binding propensity of Ccdc124 by in silico analysis using CatRapid software platform. This analysis indicated possible intrinsic disorder regions in Ccdc124, together with one major and two minor predicted RNA binding domains (Fig. 1). Subsequent experimental approaches testing RNA binding capacity of Ccdc124 by using RIP, followed by RNA-sequencing methods have not only proven RNA binding ability of Ccdc124, but in the meantime allowed us to identify its potential mRNA targets. By these RNA-sequencing based experimental approaches, we found 27 potential mRNA targets which were physically interacted with Ccdc124 (Supplement Table S3). Interestingly, preliminary predictions on cellular functions of these 27 factors based on STRING interaction network database analysis [38] indicated that at least five of those RBPs might take part in positive regulation of Ras superfamily of small GTP-binding proteins known to be involved in formation and progression of cancers (Supplement Table S3).
Subsequently, by using qPCR methods, we tested in cervix cancer originated HeLa cells the validity of two of the highest propensity value mRNA interactions with Ccdc124, and we obtained results confirming interactions between those candidate mRNA molecules and our protein of interest (Fig. S2). These results have proven unequivocally that Ccdc124 is an RNA binding protein targeting mRNA in addition to rRNA, as the latter was previously shown to interact with the yeast orthologue of Ccdc124 [21, 39].
Recent studies indicated clear roles of RNA-binding proteins in oogenesis by regulating translations of select groups of mRNA [40, 41, 42]. Interestingly, in this study, we observed a pronounced Ccdc124 staining in oocyte follicles in normal tissues, which could indicate a functional role of Ccdc124 in these follicles. As reported previously by Telkoparan et al., Ccdc124 was diffusely distributed in cytoplasm, as expected in cells at interphase [19]. This may also indicate a similar distribution pattern of its target mRNAs. Moreover, our data showed that Ccdc124 protein had a rather epithelial staining pattern, as compared to stromal cells, indicating a primary function for this protein in epithelial cells.
One of our major results in the present study is up-regulation of Ccdc124 expression both at mRNA and protein levels in ovarian and endometrial cancers. Previously, in two independent data mining RNA-seq data, and by high-throughput studies, up-regulation of Ccdc124 was mentioned in cancers of different tissue origins (brain, breast, colon, kidney, liver, lung-adeno, lung-squamous, prostate, and thyroid cancers), but a detailed analysis on this expression was absent [22, 23]. We have now extended these studies, and addressed expression levels of CCDC124 mRNA and the encoded protein in cDNA and tissue arrays corresponding to a large number of human cancer samples with grade and stage specific classifications. In this analysis, we confirmed previously obtained results, and we better characterized cancer tissue specific over-expression of CCDC124 (Fig. 2). Significantly, at the mRNA level, we detected approximately from 2.4 to 7-fold increase in the expression of Ccdc124 in urinary bladder, thyroid gland, ovary, endometrium, liver, adrenal gland, and esophagus cancers (Figs 2–4). This result indicated the promising potential of Ccdc124 as a novel cancer biomarker.
We also observed a very significant increase in CCDC124 mRNA expression in urinary bladder cancer samples by qPCR method (Figs 2 and 3A). Yet, in contrast to results obtained in ovarian and endometrium cancers, in bladder cancers Ccdc124 protein expression levels were relatively mild (compare Fig. 2 inset with Fig. 7). This discrepancy could be related with post-transcriptional regulation mechanisms operating differentially in tumors of different tissue origins [41, 42].
In endometrial cancer samples, we noticed a
Our data indicated nearly 4-fold higher expression of CCDC124 mRNA in ovarian cancers, particularly in papillary serous and serous ovarian carcinoma as compared to normal tissue controls (Figs 2 and 4). This result is also confirmed by IHC staining of tissue-arrays by anti-Ccdc124 antibodies (Fig. 9). Furthermore, we detected a grade dependent increase in Ccdc124 expression at relatively later stages of tumor progression, both by qPCR and by IHC methods (grade-2 and grade-3; Figs 4 and 9).
In this study, we have not assessed CCDC124 mRNA or protein expression levels in tumors in comparison to widely used proliferation markers and/or prognostic factors, such as for instance Ki-67 [43]. Yet, interestingly, similar to Ccdc124, the proliferation marker/prognostic factor Ki-67 is also upregulated in early stage ovarian cancers [44], in urinary bladder cancers [45], and in endometrial cancers [46]. Further studies are required to carry-out a comparative mRNA, protein expression, and immunohistochemical analysis of these two seemingly independent markers, in order to establish whether there exist possible relationships between their expression levels and clinicopathological features, such as degree of malignancy, recurrence, or survival.
Ccdc124 was previously shown to be a factor with a sub-cellular localization at centrosomes and midbody, and it was associated with the separation of dividing daughter cells at the end of mitosis [19]. Aberrant expression of proteins localized at centrosomes and cytokinetic midbody were known to be associated with centrosome disfunctions, aberrations in chromosome segregation, and aneuploidy [47]. Therefore, it is plausible to propose a similar cellular outcome for Ccdc124 over-expression. Indeed, this might also indicate a possible early contribution of aberrant Ccdc124 expression to carcinogenesis, as imbalanced chromosome distribution was previously shown to contribute not only to heterogeneity and drug resistance of cancer cells, but also to DNA damage and to the activation of DNA damage response involving p53 pathway [48, 49].
We have carried-out Ccdc124 mRNA expression and immunohistochemistry-driven evaluation of protein expression analysis in tissue sample sets coming from different but clinically comparable cohorts of patients at similar age intervals. Thus, mRNA and protein expression analysis data were obtained form comparable sample sets, but not from the same patients individually. This was a necessary compromise in order to access to relatively large numbers of samples (381 tumor/normal paired human cDNA samples from 18 different cancer types, and tissue arrays from 119 tumor/normal paired samples) obtained from different sources, and to carry-out a comprehensive study in a reasonable time-period. As clinically followed patient groups were not included to our study, one limitation of this work was the absence of series of tumor samples of different stages coming from patients for an individual grade-per-grade analysis of Ccdc124 expression changes in tumors of different tissue origins. Future studies will elaborate individual stage-specific changes in Ccdc124 expression in detail, and they will define its contribution to tumor type and stage specific molecular profiles.
Conclusion
In conclusion, our experimental assays clearly indicated that as predicted by computational methods, Ccdc124 is an RBP that interacts with select mRNA species in HeLa cells, a widely used cervix cancer cell model (Fig. 1 and Figs S1 and S2). Besides, we demonstrated that CCDC124 mRNA was over-expressed between approximately three to seven folds in human endometrial, ovarian, urinary bladder, thyroid gland, liver, and esophagus, adrenal gland cancer tissues (Fig. 2). In subsequent tests on human cancer tissue arrays by IHC methods, we have shown that this up-regulation of CCDC124 mRNA was also accompanied by a pronounced Ccdc124 protein accumulation, particularly in endometrial and ovarian cancers, and to a lesser degree in urinary bladder cancer samples. Because of significantly high levels of expression, CCDC124 mRNA, and the corresponding protein (Ccdc124) could easily be detected by IHC methods, qualifying it as a strong biomarker protein at least for ovarian, endometrial, and urinary bladder cancers. Future molecular studies on cellular functions of Ccdc124 could lead to a better understanding of roles of RBPs in cancer cell biology and provide novel detection and treatment strategies in fight against this disease.
Author contributions
Conception: Ö.A., P.T.A., U.H.T.
Interpretation and analysis of data: Ö.A., N.K.S., P.T.A., U.H.T.
Preparation of the manuscript: Ö.A., N.K. S., U.H.T.
Revision for important intellectual content: Ö.A., N.K.S., P.T.A., U.H.T.
Supervision: U.H.T.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-200802.
sj-docx-1-cbm-10.3233_CBM-200802.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-200802.docx
sj-xlsx-1-cbm-10.3233_CBM-200802.xlsx - Supplemental material
Supplemental material, sj-xlsx-1-cbm-10.3233_CBM-200802.xlsx
sj-xlsx-2-cbm-10.3233_CBM-200802.xlsx - Supplemental material
Supplemental material, sj-xlsx-2-cbm-10.3233_CBM-200802.xlsx
sj-xlsx-3-cbm-10.3233_CBM-200802.xlsx - Supplemental material
Supplemental material, sj-xlsx-3-cbm-10.3233_CBM-200802.xlsx
sj-xlsx-4-cbm-10.3233_CBM-200802.xlsx - Supplemental material
Supplemental material, sj-xlsx-4-cbm-10.3233_CBM-200802.xlsx
Footnotes
Acknowledgments
The present study was funded by grants from Turkish Scientific and Technological Research Council (TUBITAK) KBAG 114-Z-349 to U.H.T., and by Ministry of Development 2014-K-120430 to Gebze Technical University Centre Research Laboratory (GTU-MAR).
Conflict of interest
Authors declare no conflict of interest.
