Abstract
BACKGROUND:
The advancement of cancer genomics has allowed for multiplex gene assays using next-generation sequencing (NGS) to be practically implemented, however, a clinical practice system remains to be established.
OBJECTIVE:
We evaluated the feasibility of clinical sequencing using NGS-based multiplex gene assays between cooperating medical institutions in patients with advanced cancers.
METHODS:
In this observational study, DNA and RNA samples prepared from existing tumor tissues were subjected to comprehensive genomic profiling using targeted sequencing.
RESULTS:
From January 2017 to March 2019, 36 samples from 33 patients were assessed. Of all patients, 27 (82%) had lung cancer, with the median age of 50 years (range 38–83). Multiplex gene panel tests were successfully carried out on 35/36 (97%) samples. Potentially actionable gene alterations were identified in 10/30 (33%) samples (3 HER2, 2 KRAS, 2 ALK, 1 PIK3CA, 1 RET, and 1 CDKN2A). In the 6 samples examined for resistant mechanisms, ALK I1171N mutation and MET copy number gain were detected in 2 patients with ALK rearrangement-positive lung cancer.
CONCLUSIONS:
Clinical sequencing using NGS-based multiplex gene assays between collaborating domestic medical institutions was feasible, with a success rate of
Keywords
Background
Cancer is thought to arise through the accumulation of acquired gene alterations [1, 2]. Progress in the field of molecular biology has facilitated the identification of multiple gene abnormalities that are associated with malignant tumors. The identification of therapeutic targetable driver mutations, such as the epidermal growth factor receptor (EGFR), anaplastic lymphoma kinase (ALK), c-ros oncogene 1 (ROS1), and v-raf murine sarcoma viral oncogene homolog B1 (BRAF) has been successfully translated into clinical practice for cancer patients [3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]. The clinical application of driver mutations and their inhibitors are expanding.
Tumor genotyping to identify actionable oncogenic driver mutations and mechanisms of resistance to targeted drugs has become increasingly important in the management of cancer patients [20, 21, 22, 23]. Advances in next-generation sequencing (NGS) technologies allow for high-throughput multiplexing of therapeutic targetable gene alterations [24, 25, 26, 27, 28]. While cancer genomic profiling using NGS-based assays has been rapidly adopted in clinical practice, there is an urgent need for the implementation of an established system in a clinical setting [29, 30]. As the expanding number of potentially actionable mutations in cancer patients necessitates tumor genotyping, tissue biopsies are often assessed by NGS-based assays [21, 24]. Thus, we evaluated the feasibility of clinical sequencing using NGS-based multiplex gene assays between cooperating medical institutions for genotype-directed therapy in clinical practice [31].
Methods
Before the NGS-based panel tests, FoundationOne CDx Cancer Genomic Profile (Chugai Pharmaceutical Co., LTD.) and OncoGuide NCC Oncopanel System (Sysmex), received insurance coverage for cancer genome profiling in Japan in February 2019, solid cancer patients, considered eligible by the attending physician, were enrolled in this study and underwent comprehensive genomic profiling for detecting possible molecular targets. No restrictions on tumor, disease stage, subsequent or previous treatment, performance status, or other factors were imposed. All patients provided written informed consent to participate in this study.
After obtaining the consent, 10 sliced, undyed specimen slides were prepared from existing formalin fixed paraffin embedded (FFPE) samples, which were originally fixed for routine pathological diagnosis of patients at Kitasato University Hospital; samples were fixed in 10% neutral buffered formalin within 24–48 hours of being obtained. Tumor slide specimens were sent from Kitasato University Hospital, Kanagawa, to Kindai University, Osaka, for the multiplex NGS-based panel tests.
At the clinical laboratory of Kindai University, DNA and RNA were extracted after confirming if
Mutation detection was performed on 10 ng of DNA, through multiplex PCR amplification using CLv2 or CHPv2. The Ion Xpress Barcode Adapters (Thermo Fisher Scientific) were used for sample identification. Fusion detection with LF was performed on 10 ng of RNA, using reverse transcription followed by library preparation. The Ion Xpress Barcode Adapters were used for sample identification. Fusion detection with Archer CTL was performed on 50 ng of RNA, using reverse transcription followed by library preparation. The MBC Adapters (ArcherDx) were used for sample identification. Purified libraries were pooled and sequenced on an Ion Torrent Proton system (Thermo Fisher Scientific). The panels used for each sample are shown in Supplemental Table 1.
DNA sequencing data were accessed through the latest version of Torrent Suite (5.0. to 5.10) (Thermo Fisher Scientific). Reads were aligned against the hg19 human reference genome, and variants were called using the latest version of variant caller (5.0. to 5.10) (Thermo Fisher Scientific). Raw variant calls were filtered using a quality score of
This study was approved by the Kitasato University Medical Ethics Organization (study number: B16-55) and the ethics committee of Kindai University Faculty of Medicine (study number: 27-169).
Results
Patient characteristics
Between July 2017 and December 2018, written informed consent was obtained from 33 advanced-cancer patients at Kitasato University Hospital. Thirty-six tumor samples from these patients were submitted to Kindai University for genomic testing. The patient characteristics are summarized in Table 1. The median age was 50 years (range, 38–83), and 24 (72%) patients were under 60 years. Fifteen (45%) were female and 10 (30%) were never smokers. Most patients (
Characteristics of the patients subjected to molecular analysis (
33)
Characteristics of the patients subjected to molecular analysis (
EGFR, epidermal growth factor receptor; ALK, anaplastic lymphoma kinase.
Distribution of actionable gene alterations based on the NGS-based assay for the 36 tumor specimens. Fig. 1-1. Genotyping to identify actionable gene alterations. Fig. 1-2. Genotyping to search for resistant mechanisms of molecular targeted therapies. The pie chart shows the number and frequency of patients with potentially actionable gene alterations (dark gray), without specific gene mutations (gray), and those not subjected to NGS analysis (light gray).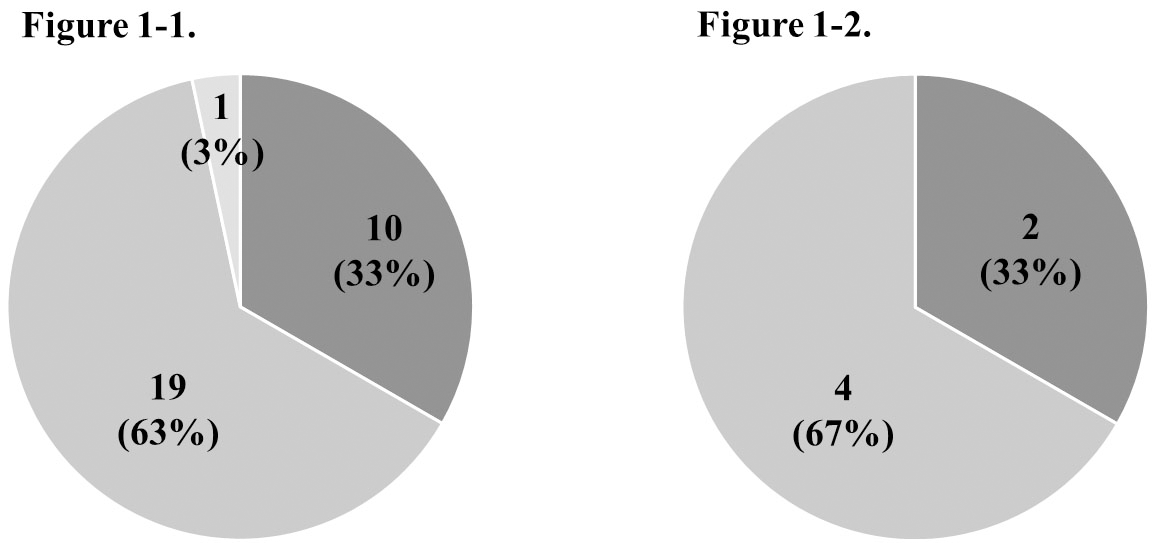
Genotyping results for the 36 tumor specimens
TBLB, transbronchial lung biopsy; TBB, transbronchial biopsy; LN, lymph node; amp. amplification, CN, copy number.
Treatment progress and timing of the multiplex NGS-based assay in the 10 patients with potentially actionable alterations. Each bar represents the time course from the first-line chemotherapy for individual patients with potentially actionable alterations. All patients with lung cancer except those with pancreatic cancer patient indicated as number 30. The color of each bar indicates the line of chemotherapy (dark gray: first, gray: second, light gray: third, gray vertical stripes: fourth, gray horizontal stripes: fifth, white: sixth line chemotherapy). Black circles indicate when the consent was obtained for the multiplex gene assay in this study.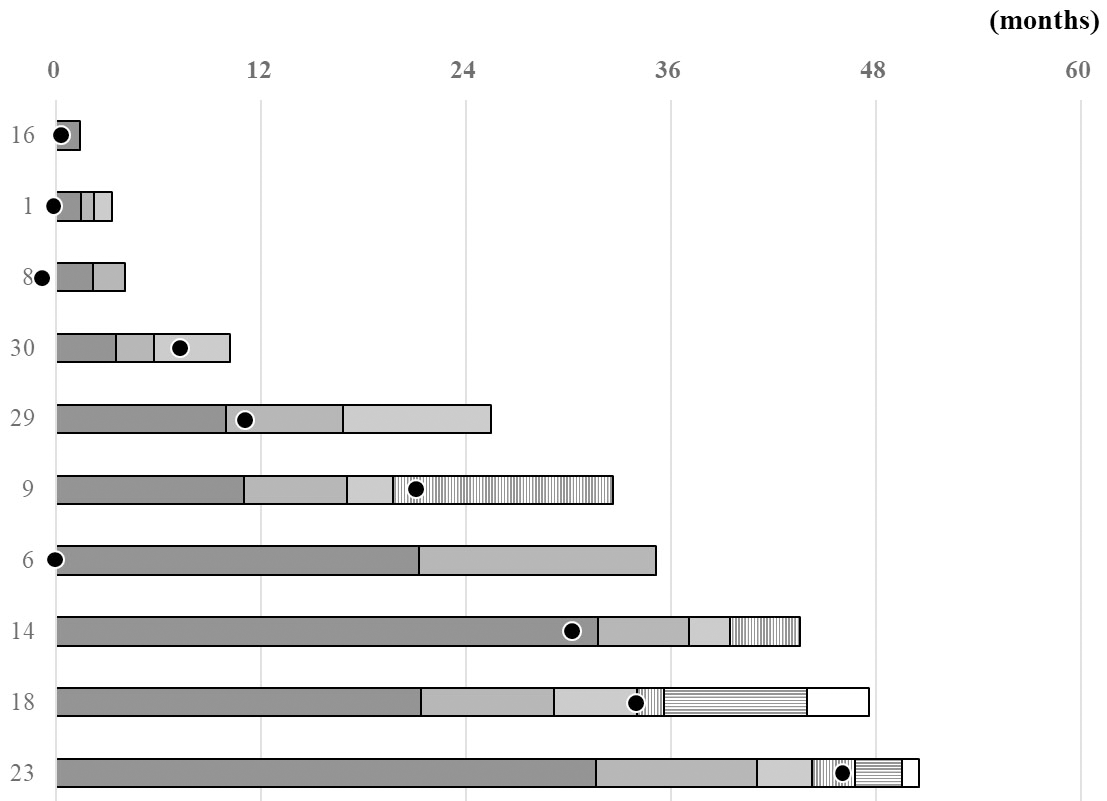
Archived FFPE tumor tissues were available, and DNA and RNA extracted from these tissues were used for genomic profiling using an NGS-based assay. Sequence reads were trimmed based on the quality score (Phred score
The success rate of the NGS assay was 97% (
Prevalence of genomic alterations
Of the 33 patients tested for molecular alterations in this study, 29 patients were investigated for actionable molecular therapeutic targets. Potentially actionable genomic alterations, including HER2 in 3 (11%), KRAS in 2 (7%), ALK in 2 (7%), and PIK3CA, CDKN2A, and RET in 1 each (3%), were identified in 10 (33%) of them (Fig. 1-1, Table 2-1). One of the two patients in whom ALK rearrangement was detected was treated with an ALK inhibitor under their health insurance. However, ALK inhibitor was not administered to the other patient, because the companion diagnostic, ALK immunohistochemistry (IHC), was negative; therefore, the treatment was not approved by the insurance company. The patients for whom the target genes were other than the ALK rearrangements could not be enrolled in the ongoing clinical trials for the experimental drugs based on the genomic information. Deterioration of the general condition of the patient during the course of treatment was the main reason for non-participation in clinical trials, even if the NGS-based panel test was performed early in the course of treatment (Fig. 2).
When the 4 patients were assessed for resistant mechanisms against EGFR tyrosine kinase inhibitor or ALK inhibitors, ALK mutation (p.lle1171Asn) and MET amplification were detected as reported resistant mechanisms of ALK inhibitors (Fig. 1-2, Table 2-2). Based on these results, the patient with the ALK p.lle1171Asn resistant mutation could receive the ALK inhibitor ceritinib for three months as a 7th line chemotherapy.
There was no incidental finding for germline variants to be disclosed, pointed out by experts in this study.
Discussion
Clinical sequencing using multiplex NGS-based gene assays was successful between collaborative domestic medical institutions, using archived FFPE samples from patients with advanced solid cancers. Potential actionable genomic alterations were detected in the 10 of 30 (33%) samples from 29 patients, who could not be enrolled in ongoing clinical trials except for a patient with ALK rearrangement treated with an ALK inhibitor. In this study, EGFR and ALK alternations had been single-gene tested in clinical practice, and twenty-four (72%) of patients under the age of 60 were enrolled, to actively perform genomic profiling for younger patients with advanced cancer. Although previous studies conducted on lung cancer patients at Kindai University identified 22% of actionable gene alterations other than EGFR mutations and ALK fusion, the frequency with which the therapeutic targeted genes can be identified may increase if the patients are selected by the attending physician based on their background [21]. However, cancer genomic profiling in clinical tissue specimens affects detection frequency due to various factors, such as variable mutational burden in different tumors, intra-tumor heterogeneity, and number of tumor cells. In addition, treatment selection after identification of actionable gene alterations poses major challenges, as ongoing clinical trials are only open in limited institutions and selected patients.
In one lung cancer patient, where ALK rearrangement was identified by the multiplex gene panel test, ALK IHC, performed as a companion diagnostic for insurance-approval of the drug use, was negative. The discrepancy between the panel test based on NGS and the IHC results could be due to the amount of sample and the number of tumor cells, intratumor heterogeneity, and test sensitivities. In clinical practice, there are problems with the quantity and quality of the tissue biopsy samples obtained for gene panel tests for cancer diagnosis. Sample management is one of the issues in cancer genomic profiling for targeted inhibitor use in clinical settings.
In order to identify resistance against EGFR or ALK inhibitors, re-biopsy samples after disease progression of the targeted therapy were submitted, and the mechanism for ALK resistance was found in 2 cases [32, 33, 34, 35, 36]. In 1 of the 2 patients, sequential treatment with an ALK inhibitor was administered, based on the genomic profiling results. Almost all patients with metastatic cancers who initially benefit from targeted therapies eventually develop resistance. It is important to elucidate the mechanisms of resistance, and it is hoped that the mechanism-based therapeutic strategy will be widespread in the future with the routine implementation of NGS-based multiplex gene assays.
At present, it is possible to perform multiplex gene panel tests that show the comprehensive genomic characteristics of cancer, and gene profiling tests are being performed in cooperation between facilities that perform cancer medical care. Based on the results of cancer genomic profiling, an expert panel comprising multidisciplinary cancer specialists is needed to ensure meaningful interpretation and accurate reporting of results to each institution to address the matched therapeutic opportunity. The system of precision medicine directed by cancer genomic profiling is currently being developed by medical institutions, from sample operation and gene analysis to the disclosure of results. This study assessed an NGS-based approach between related medical institutions, performed before the insurance approval of multiplex gene panel tests, and demonstrated that this strategy is feasible. In the future, it will be necessary to expand the gene analysis that leads to the determination of appropriate treatment options to provide more clinical benefits for the patients.
This study had certain limitations, including that this was an observational study conducted at two cooperating institutions with a small sample size; hence, the results cannot be considered definitive. Second, in this study, several biases, such as patient selection, should be considered because the target patients for genomic profiling are not limited and patients in various situations have been enrolled. Moreover, it remains unclear whether the genotype-directed therapy of patients positive for actionable gene alterations other than EGFR mutations or ALK rearrangement were affected by tyrosine kinase inhibitors. Although no definitive conclusions can be drawn, genomic profiling using NGS-based assays may be feasible in clinical practice, and could be considered an early test of treatment eligibility based on identified potentially actionable gene alterations. In order to select subsequent treatments based on clinical sequencing results, it is necessary to understand genomic abnormalities, including information on target inhibitors, before the disease progresses.
Conclusions
Clinical sequencing using NGS-based assays is feasible between collaborative medical institutions that cooperate with cancer genomics, and is useful for identifying targeted gene alterations and elucidating resistant mechanisms. While an improvement of the system in clinical practice is urgently warranted, there are several factors that must be considered, including appropriate patient selection for comprehensive genomic profiling, examination time, decision-making for treatment, sequencing costs, and the development of therapeutic drugs. The challenge is to carry out multiplex gene panel tests early and administer effective drugs to patients before their general condition deteriorates.
Author contributions
Conception: T. Fukui, J. Sasaki, K. Nishio
Interpretation or analysis of data: T. Fukui, K. Sakai, J. Sasaki, M. Takeda, T. Takahama, K. Nakagawa, K. Nishio
Preparation of the manuscript: T. Fukui, J. Sasaki, MI Kakegawa, S Igawa, H Mitsufuji, K. Naoki
Revision for important intellectual content: T. Fukui, K. Sakai, J. Sasaki, K. Nishio, K. Naoki
Supervision: J. Sasaki, K. Nakagawa, K. Naoki
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-200781.
sj-docx-1-cbm-10.3233_CBM-200781.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-200781.docx
Footnotes
Acknowledgments
The authors would like to thank Editage (www.editage.jp) for English language editing.
Conflict of interest
The authors have no conflicts of interest.
