Abstract
BACKGROUND:
Smac/DIABLO is a proapoptotic protein deregulated in breast cancer, with a controversial role as a tumor marker, possibly due to a lack of correlative mRNA and protein analyses.
OBJECTIVE:
To investigate the association of Smac/DIABLO gene and protein levels with clinical variables in breast cancer patients.
METHODS:
Smac/DIABLO mRNA expression was analyzed by qPCR in 57 frozen tissues, whereas protein levels were assessed by immunohistochemistry in 82 paraffin-embedded tissues. Survivin mRNA levels were also measured. In vitro assays were performed to investigate possible regulators of Smac/DIABLO.
RESULTS:
Higher levels of Smac/DIABLO mRNA and protein were found in estrogen receptor (ER)-positive samples (
CONCLUSIONS:
mRNA and protein levels of Smac/DIABLO were increased in ER-positive breast tumors in comparison with ER-negative samples, although the mechanism of this regulation is still unknown. Public databases showed a possible clinical relevance for this association.
Introduction
Breast cancer is the leading cause of death by cancer in the female population worldwide [1]. In Mexico, most patients have locally advanced or metastatic disease when first seen at the oncology clinic [2]. Several factors such as limited access to prevention and early detection of breast cancer have been proposed as the cause of this late diagnosis [3]. In addition, Mexican women with breast cancer have a mean age at diagnosis of 52 years [2], which is at least a decade younger than women in the U.S. [4]. In Mexico, as in other countries, most patients have ER-positive breast tumors (60%), whereas 23% are human epidermal growth factor receptor 2 (HER2)-positive tumors [2]. Breast cancer heterogeneity makes hard to give all patients an optimal treatment and predict the individual risk of metastasis. Currently at the time of diagnosis, the potential for tumor recurrence and general prognosis are estimated according to clinicopathological features such as tumor size, grade, involvement of lymphatic vessels, expression of ER, progesterone receptor (PR), and HER2 [5, 6]. However, these criteria have been insufficient to help in the early detection and prognosis of this disease [7].
Apoptosis resistance is an important mechanism for tumor development and progression [8]. Smac (second mitochondria-derived activator of caspases) [9], also known as DIABLO (Direct IAP binding protein with low pI) [10] is a pro-apoptotic protein released into the cytosol in response to noxious stimuli, where it binds and inhibits members of the Inhibitors of Apoptosis Proteins (IAP) family, promoting caspase activation and cell death [9]. The mitochondrial release of Smac/DIABLO during apoptosis can be regulated by survivin, a IAP family member [11]. Altered Smac/DIABLO expression has been observed in several tumors. Low levels of Smac/DIABLO are prognostic for poor prognosis in bladder cancer [12], renal cell carcinoma [13] and lung cancer [14]. However, in cervical cancer [15], gastric adenocarcinomas [16] and leukemia [17] Smac/DIABLO levels are increased as compared to healthy tissue; and in the latter case, high levels of Smac/DIABLO are associated with poor prognosis.
Conflicting results have been found when the expression of Smac/DIABLO was analyzed in breast tumors. Pluta and collaborators found a diminished expression by flow cytometry of Smac/DIABLO in tumor samples in comparison with benign breast tumors. In addition, Smac/DIABLO expression inversely correlated with tumor stage [18]. Moreover, in 102 samples of invasive ductal carcinoma, Zhang and colleagues observed cytoplasmic staining of the protein which was more prevalent in the HER2-positive group. No other association between Smac/DIABLO expression and clinical features was found [19]. Independently, Mansour and coworkers analyzed Smac/DIABLO and survivin mRNA expression by qualitative RT-PCR in 37 benign breast tumors and 48 malignant tumors. This group found a reduced Smac/DIABLO expression in the breast cancer group and an inverse relation with survivin expression. Smac/DIABLO downregulation showed a positive correlation with ER, but no additional correlation with other clinicopathological feature or risk factor [20].
In the present work, we analyzed the Smac/DIABLO gene and protein expression in two different sample groups of Mexican breast cancer patients. We searched for a correlation between Smac/DIABLO expression, survivin expression and clinicopathological features. We found a negligible correlation between Smac/DIABLO and survivin expression levels. Interestingly, we observed a positive correlation between Smac/DIABLO and ER expression at both mRNA and protein levels. Our finding was supported by data from public databases, where a positive correlation between the proteins Smac/DIABLO and ER was found in breast tumors samples. Additionally, we analyzed public microarray gene expression data and found that Smac/DIABLO expression has a prognostic value for breast tumor patients, particularly for patients with ER-positive tumors. Additionally, we performed in vitro assays and found that Smac/DIABLO is neither a target gene of ER nor participates in ER expression regulation.
Materials and methods
Patients
The first group of samples was collected during the 2008–2011 period at the Fundación contra el Cáncer de Mama (FUCAM) after informed consent. Tumor samples were collected during surgery, a total of 57 samples were analyzed. The protocol was reviewed and approved by the Ethics and Research committees of the National Institute of Genomic Medicine (INMEGEN) and FUCAM in Mexico City (CE2009/11). After macroscopic inspection by the pathologist, sections of tumor were frozen in liquid nitrogen and stored at
Tissue microarray construction
For Smac/DIABLO protein analysis, paraffin- embedded tissues were obtained from the Oncology Hospital in the XXI Century National Medical Center; since these tissues constituted residual material, their use does not represent a risk for patient diagnosis. Tumor tissues from 82 patients were embedded in paraffin, and 4 mm sections were stained with hematoxylin and eosin to select morphologically representative areas. Tissues selected from each specimen were punched with a 1 mm needle and transferred to a recipient paraffin block using a Tissue Microarray ATA 100 Chemicon (Temecula, CA, USA). Then, 4 mm sections of these tissue arrays were sectioned and placed onto positive charged slides (VWR Superfrost Plus).
Immunohistochemistry of breast cancer tissues
For Smac/DIABLO protein analysis panel, whole tissue sections were stained with Estrogen Receptor (Ventana CONFIRM anti-ER, Roche, Strasbourg, France), Progesterone Receptor (Ventana CONFIRM anti-PR), HER-2/neu antibody (Ventana, PATHWAY anti-HER-2/neu) and Smac/DIABLO mouse mAb (Cell Signaling, Danvers, MA, USA). Immunohistochemistry was performed with the Ventana Ultraview DAB Detection Kit (Ventana). Diaminobenzidine was used as the chromogen and hematoxylin as the nuclear counterstain. All slides were processed in a Ventana BenchMark XT processor (Ventana, Tuczon, AZ, USA) and mounted with synthetic resin. Slides were scanned in ScanScope CS2 (Aperio) and image analysis was performed in Image J. Two independent observers reviewed the slides and estimated the percentage of tumor cells and staining intensities scores. Smac/DIABLO score expression is the product of the percentage of positive cells multiplied by the immunostaining intensity.
The Cancer Genome Atlas (TCGA) samples
We used Regulome Explorer (
Cell lines
All cell lines were obtained from American Type Culture Collection (Manassas, VA, USA) with the exception of the SKBR-3 cell line, which was kindly donated by PhD Fernando Enriquez Rincon. SKBR-3 cell line was authenticated by short tandem repeat (STR)-profiling using AmpFLSTR
Protein extraction
Total protein was obtained from panel of breast cancer cell lines with RIPA buffer. Briefly, cells were grown up to 80% of confluency and then were washed with PBS 1X, cells were scrapped in PBS 1X and centrifuged to obtain cell pellet. Cellular pellet was resuspended in RIPA buffer, homogenized and incubated in ice for 10 min. After incubation samples were centrifuged at 10,000 rpm for 30 min. The supernatant was recovered as total protein extract. Proteins from long-term estrogen-deprived (LTED) MCF-7 cells and Smac/DIABLO-overexpressing cells were obtained with M-PER Mammalian Protein Extraction Reagent (Thermo Scientific, Waltham, MA, USA) following manufacturer’s instructions, with minor modifications. Briefly, cells were washed with PBS 1X and then were scrapped with PBS 1X and centrifuged at 1,000 rpm for 5 min. The cell pellet was resuspended in 100–200
Western blot
Whole-cell lysates were separated by SDS gel and transferred to 0.22
Chemicals
17
Plasmids
For Smac/DIABLO promoter assays, 1.5 kb from the genomic 5’ untranslated region (UTR) of Smac/ DIABLO were amplified and cloned in the pSEAP2 vector (Clontech, Mountain View, CA, USA) as previously reported [21]. The construct, named pSEAP2-1.5kbsmacpromoter, was verified by capillary sequencing. An absent region of 34 bp was founded in our sequence, and this deletion was also found in the genomic 5’ UTR of Smac/DIABLO from HEK293 cells when the sequences were compared with the Homo sapiens 12 BAC RP11-512M8 sequence (GenBank: AC048338.29). The vector pSEAP2-Basic (Clontech, Mountain View, CA, USA) was used as control. The ORF of Smac/DIABLO was amplified by PCR with the following primers, forward 5’-GCGCGGATCCATGGCGGCTCTGAAGAGTTGGCT-3’; reverse 5’-AGCTCT CTAGACTCAGGCCCTCAATCCTCACGC-3’. The amplified product was cloned into pCR2.1-TOPO vector (Invitrogen, Carlsbad, CA, USA). The construction with the transgene was amplified and the ORF of Smac/DIABLO was obtained with restriction enzymes, the fragment was cloned into the multiple cloning site of pQCXIP (Clontech, Mountain View, CA, USA), in the region flanked by NotI and BamHI sites. The correct insertion of the ORF was verified by enzymatic digestion. pQCXIP empty vector was used as control.
RT and qPCR
1
E2 treatment
MCF-7 and T47D cells cultured in normal conditions were seeded in twelve-well plates, the next day, cells were washed twice with PBS 1X and once with DMEM phenol red-free (Life Technologies, Carlsbad, CA, USA) supplemented with 5% charcoal-stripped bovine serum (Life Technologies, Carlsbad, CA, USA). After 5 days of culture in phenol red-free medium, cells were treated with 10 nM E2 and ethanol, as vehicle, for 24 h. RNA was extracted at this time.
Gene reporter assays
MCF-7 cells were transiently transfected with pSEAP2-Basic vector (Clontech, Mountain View, CA, USA) and pSEAP2-1.5kbsmacpromoter plasmid in a ratio of 1:200 with pMetLuc2-Control vector (Clontech, Mountain View, CA, USA) as an internal control. 24 h postransfection cells were seeded in a 24-well plate and the next day washed twice with PBS1X and once with DMEM w/o phenol red (Life Technologies, Carlsbad, CA, USA) supplemented with 5% of charcoal-stripped fetal bovine serum (Life Technologies, Carlsbad, CA, USA). Cells were maintained in this medium for 3 days before hormone treatment. Cells were stimulated with E2 10 nM for 24 h. Medium was collected for further analysis. Non-transfected cells were stimulated under the same conditions for pS2 mRNA expression analysis by qPCR. SEAP activity was analyzed with GreatEscape SEAP Chemiluminescence Kit 2.0 (Clontech, Mountain View, CA, USA) and luciferase activity with Ready-To-Glow Secreted Luciferase Reporter Assay (Clontech, Mountain View, CA, USA) according to manufacturer’s instructions.
Long-term estrogen-deprived (LTED) MCF-7 cells establishment and experiments
To establish LTED MCF-7 cells, the cells were cultured in DMEM phenol red-free (Life Technologies, Carlsbad, CA, USA) supplemented with 5% charcoal-stripped bovine serum (Life Technologies, Carlsbad, CA, USA) for five to seven months. At this time, cells were exposed to E2 10 nM for 24 h or with Fulvestrant (1
Cell transfection
For transient transfection of MCF-7 cells, 5.8
Statistical analysis
Statistical analysis was performed with the GraphPad Prism 5.0 (GraphPad Software Inc.) software. Intergroup differences were analyzed by using Mann-Whitney
Results
Smac/DIABLO expression is higher in ER-positive breast tumors and is associated with a worse prognosis in breast cancer patients
First, we evaluated Smac/DIABLO mRNA expression by qPCR in a panel of 57 breast cancer samples. Table 1 shows the clinicopathological characteristics of the samples. Infiltrating ductal carcinoma was the most frequent diagnosis (82%), and the mean age of the patients was 55 years. Approximately 68% of the samples were ER-positive, 56% were PR positive and 18% were HER2 positive. All samples expressed Smac/DIABLO mRNA, although at different levels. We found higher levels of Smac/DIABLO in the 50 years old or older group (
Clinicopathological features of mRNA analysis breast cancer group (
57)
Clinicopathological features of mRNA analysis breast cancer group (
Correlation between Smac/DIABLO mRNA expression and breast cancer patients clinicopathological features (
Correlation between survivin mRNA expression and breast cancer patients clinicopathological features (
Correlation between Smac/survivin mRNA expression and breast cancer patients clinicopathological features (
Relationship between Smac/DIABLO score expression (
Smac/DIABLO sensitizes cells to apoptosis by binding and inhibiting IAP function [9]. Survivin is an important member of IAP family and it is overexpressed in several tumor types. Due to the role of these two proteins in cell death regulation, their expression have been associated with poor prognosis in several malignancies [22, 23, 24, 25]. Therefore, we analyzed survivin mRNA expression in our samples. Smac/DIABLO and survivin expression showed a negligible positive correlation (Spearman correlation
We also evaluated the expression of the Smac/ DIABLO protein by immunohistochemistry in a tissue microarray of 82 independent breast tumor samples. Table 5 shows the clinicopathological characteristics of these patients and the correlation with the Smac/DIABLO staining score. The score was calculated multiplying the staining intensity, which was evaluated as negative (0), weak (1), moderate (2), or strong (3) (Fig. 1C) and by the percentage of positive cells, which ranged from 80 to 100%. Smac/DIABLO protein was mostly cytosolic. Interestingly, Smac/DIABLO staining score was higher in ER-positive tumors (
To independently validate our findings, we searched in data from the Cancer Genomic Atlas initiative (TCGA). We found that Smac/DIABLO protein and ER mRNA showed a negligible but significative positive correlation (
Smac/DIABLO mRNA and protein levels are higher in ER positive breast cancer samples. (A) Smac/DIABLO mRNA levels were analyzed by qPCR in breast tumors samples grouped according with ER status. The graph shows the median with interquartile range (black line). (B) Smac/DIABLO protein expression was analyzed by immunohistochemistry in breast cancer samples considering the ER status. The median value of IHC score and interquartile range (black line) are shown in the graph. (C) Images of Smac/DIABLO immunostaining levels (200X). The scale of immunostaining intensity is shown inside the images. 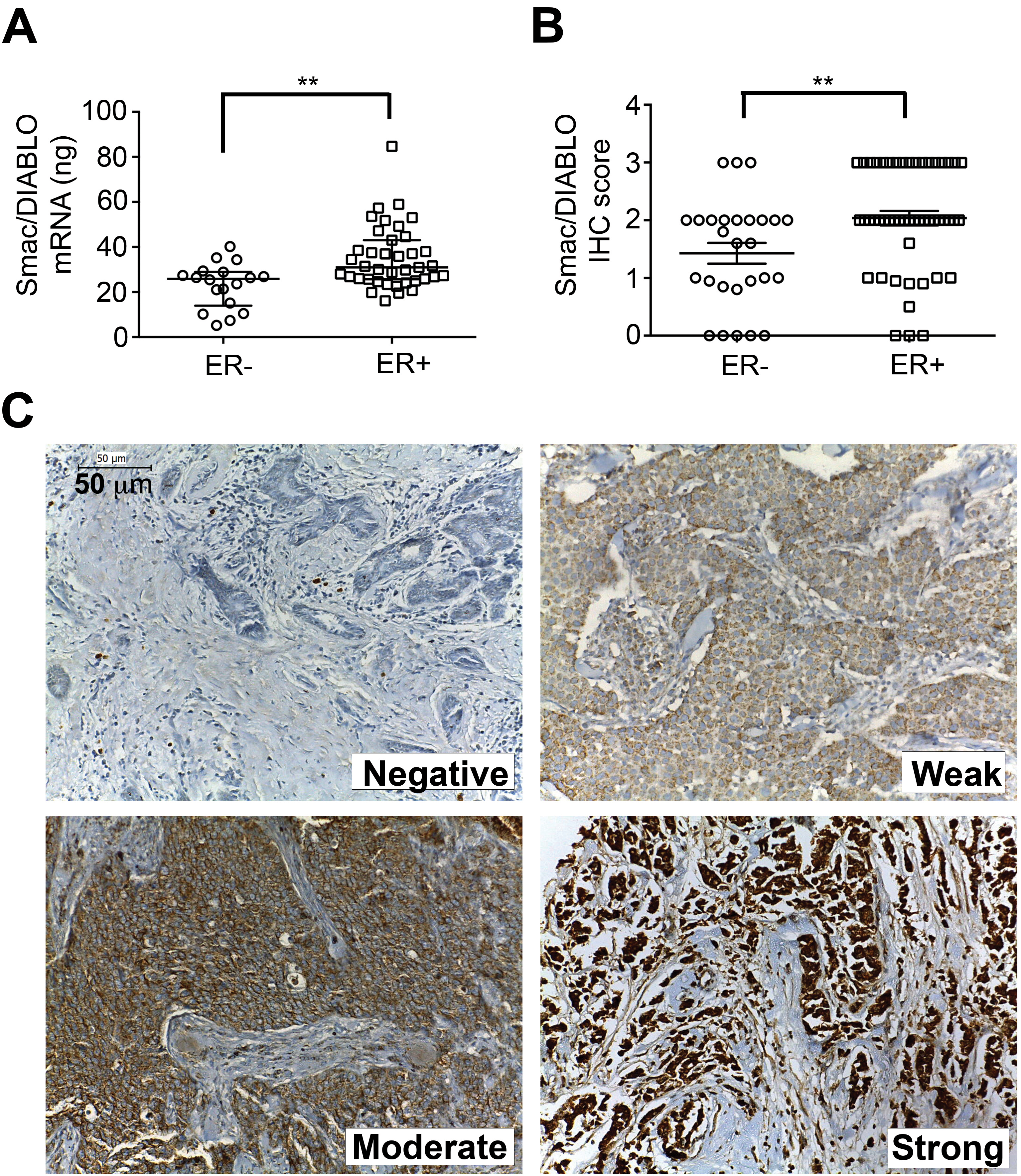
Smac/DIABLO protein correlations with ER in TCGA breast cancer samples. Expression data from the Cancer Genomic Atlas consortium (TCGA) were analyzed with Regulome Explorer tool (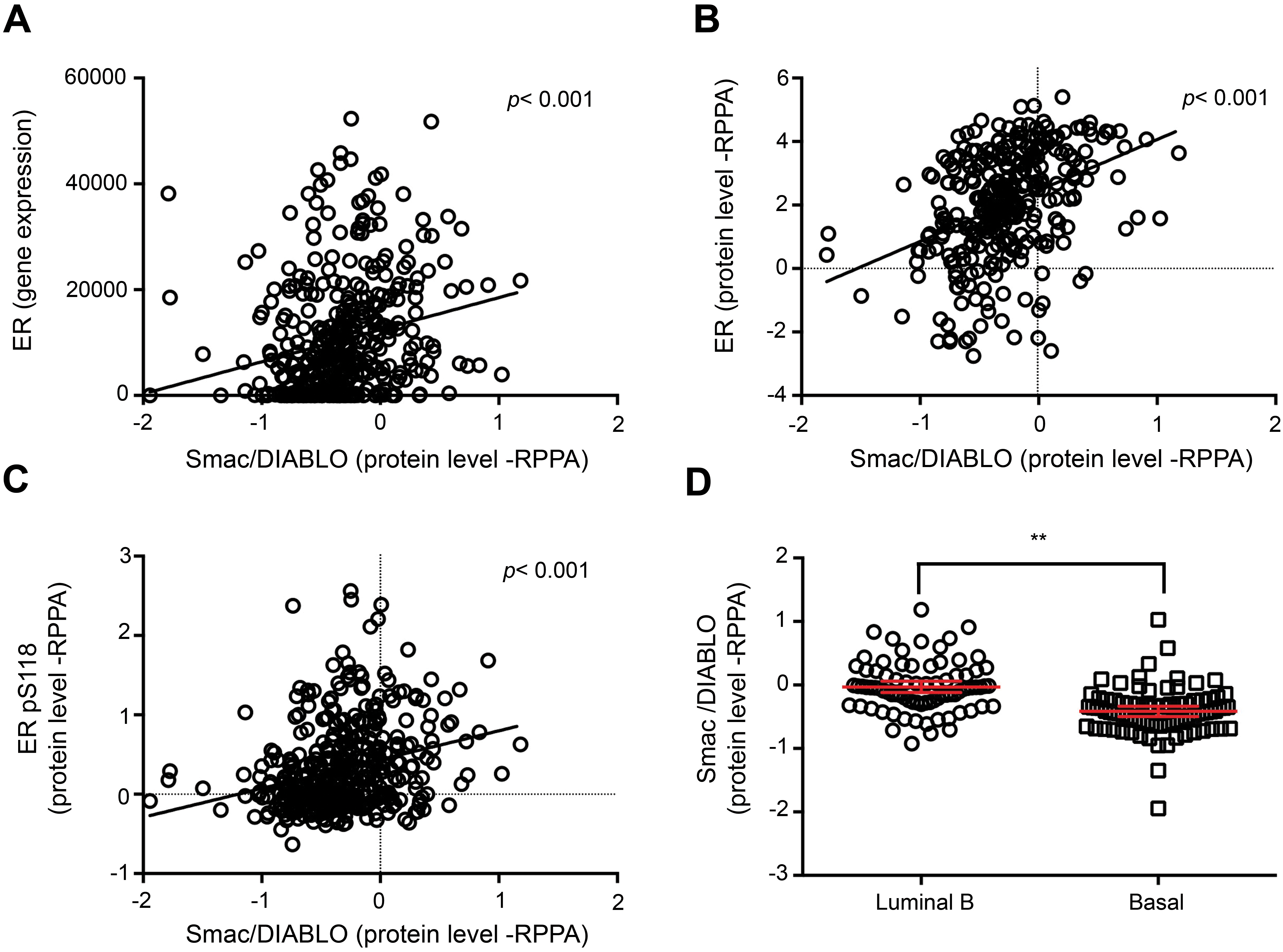
Higher expression of Smac/DIABLO in patients with ER-positive breast cancer is associated to a worse prognosis. Kaplan Meier plots of overall survival (A, C, E) and distant metastases free survival (B, D, F) of all breast cancer patients (A and B), patients with ER positive tumors (C and D), and patients with ER negative tumors (E and F). Samples were splitted according to Smac/DIABLO gene expression in high (red line) and low (black line) expression groups. The data presented in this figure were obtained from the public database KM-plotter (www.kmplot.com). The expression of Smac/DIABLO mRNA (Affymetrix ID 219350_s_at) and its relationship with survival of breast cancer patients was analyzed with this tool.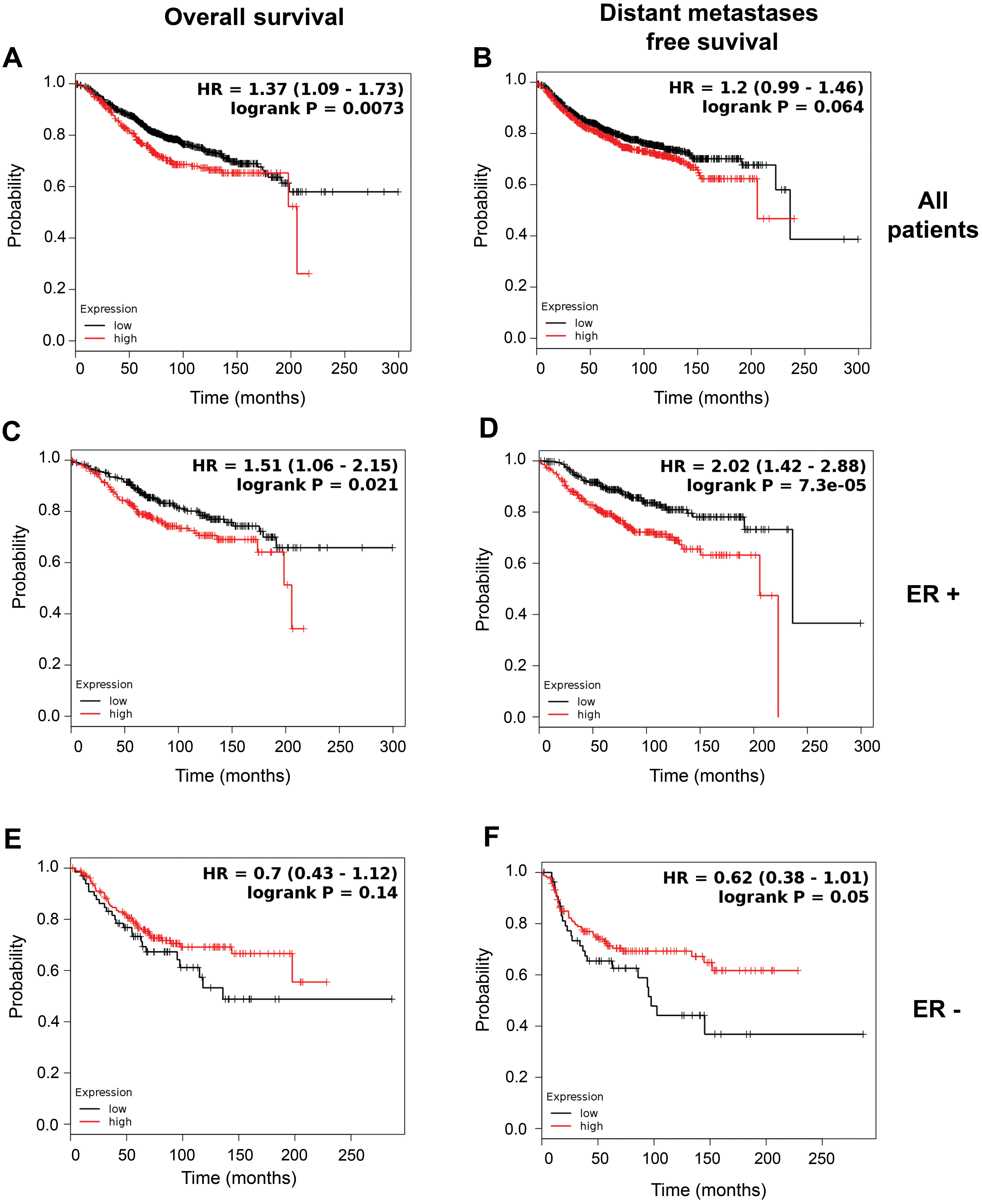
Smac/DIABLO protein levels are higher in ER-positive breast cancer cell lines. (A) Quantitative data of Smac/DIABLO levels were analyzed by immunoblot in breast cancer cell lines with different status of ER. The reported status of ER is indicated in the graph. The experiment was done by duplicate and the data were normalized with GAPDH expression. The graph shows the mean value (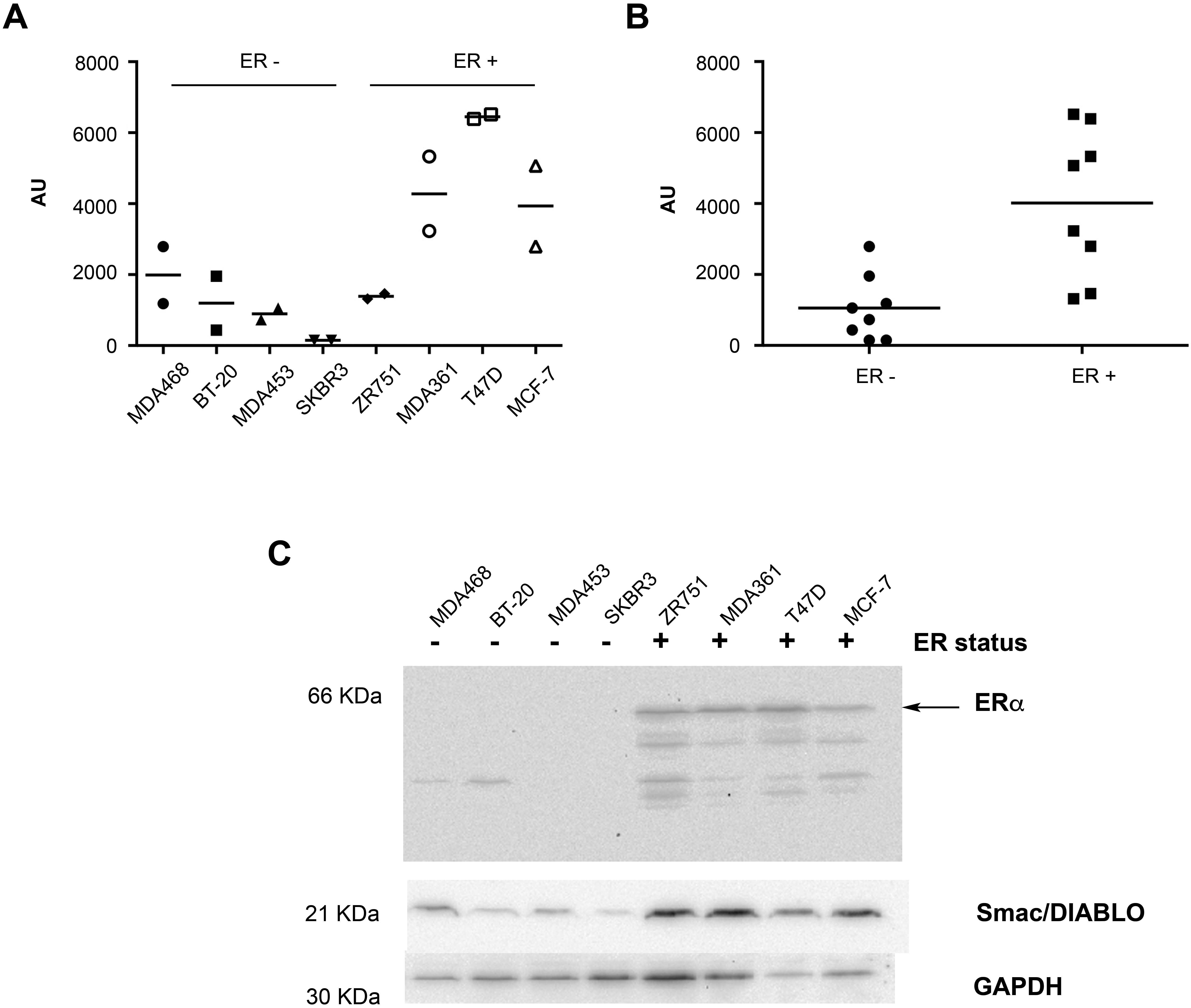
Smac/DIABLO expression is not modulated by estradiol in ER-positive breast cancer cell lines. T47D (A) and MCF-7 cells (B) were exposed to E2 (10 nM) for 24 h and Smac/DIABLO and pS2 expression were analyzed by qPCR after incubation. Graphs show fold change mean 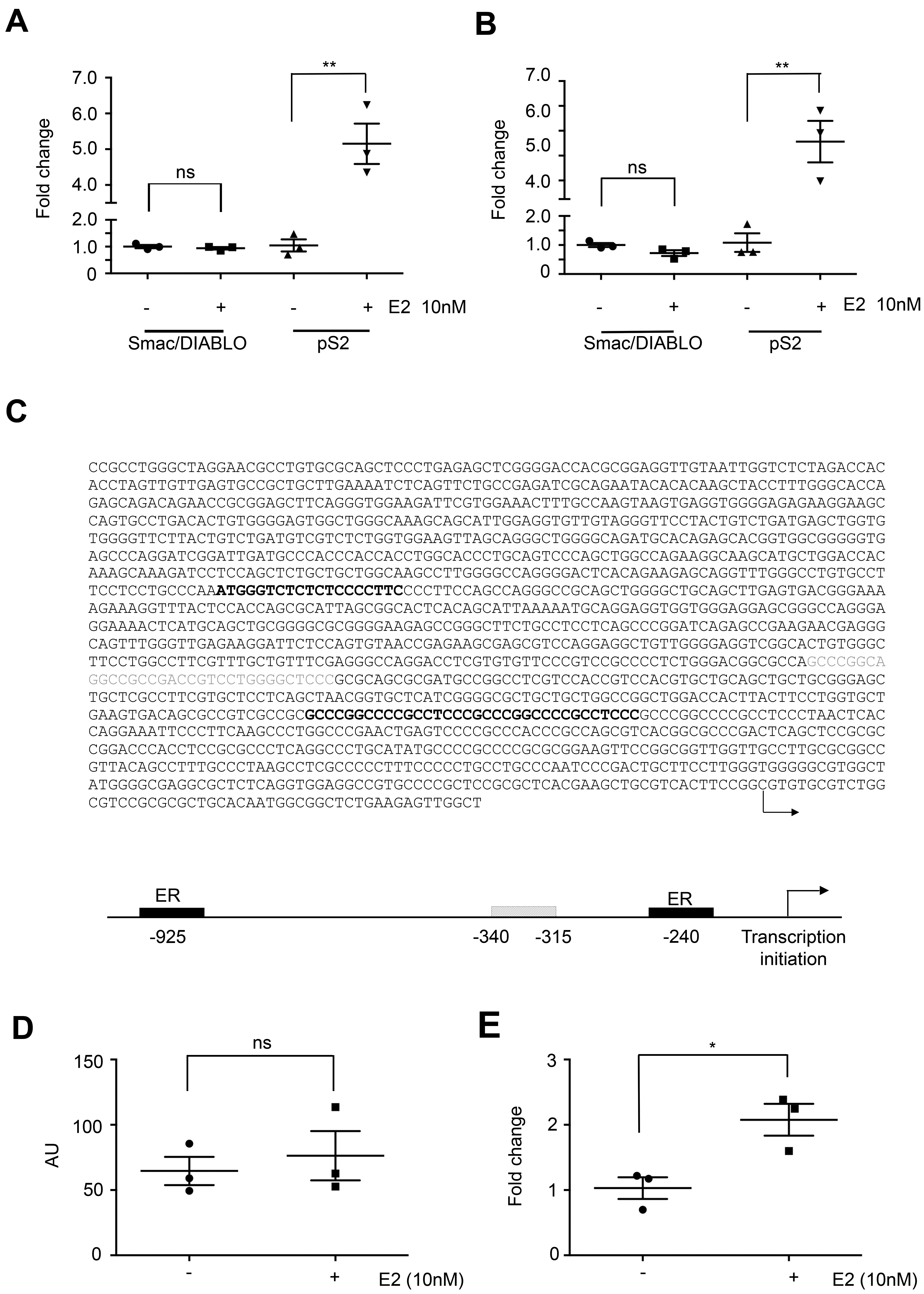
ER does not regulate the expression of Smac/DIABLO in LTED MCF-7 cells. (A) ER levels in MCF-7 cells cultured in absence of estrogen by 5 months were analyzed by immunoblot. 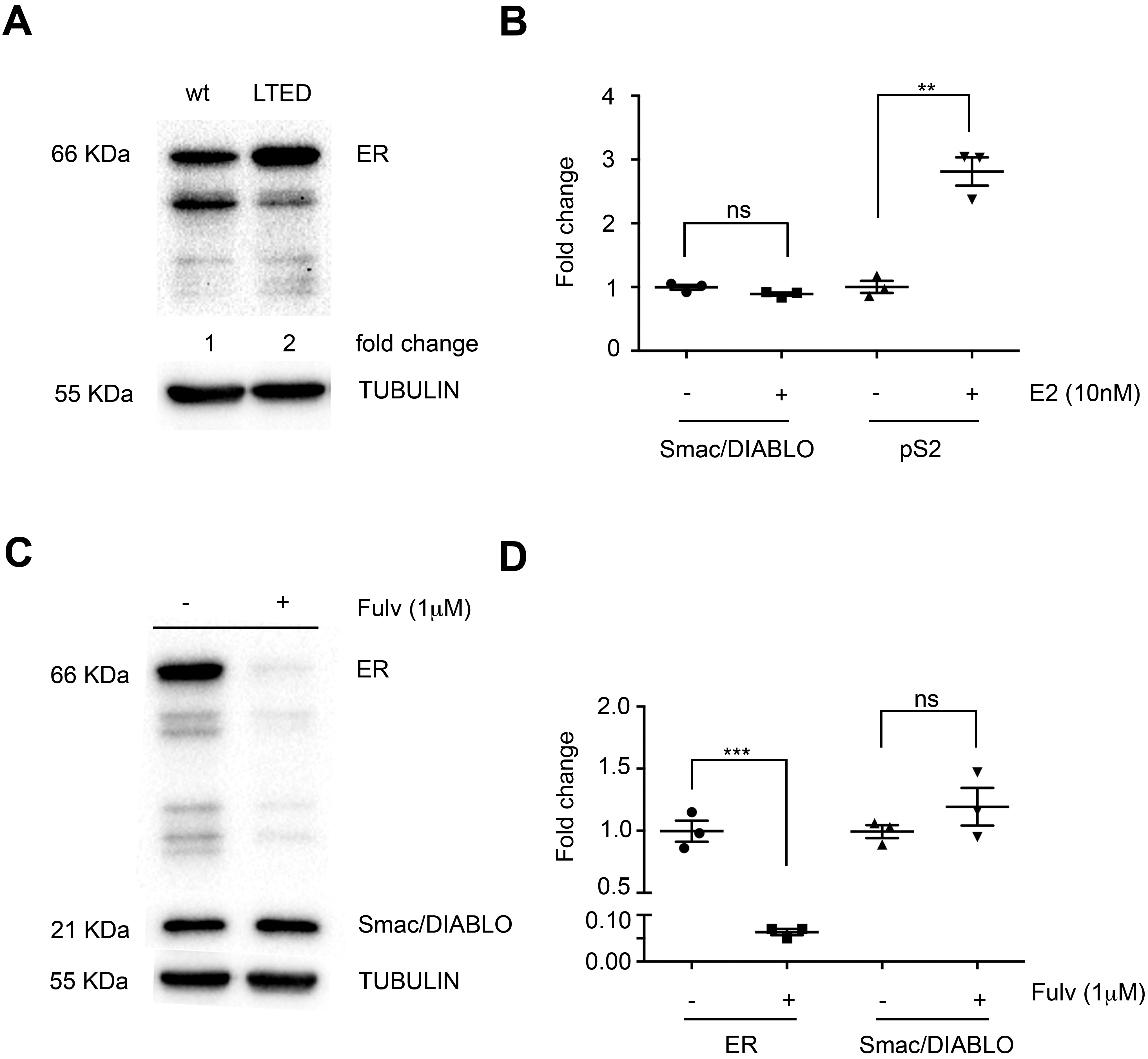
Smac/DIABLO does not regulate ER expression in MCF-7 cells. MCF-7 cells were transiently transfected with the open reading frame of Smac/DIABLO and Smac/DIABLO overexpression and its effect on modulation of ER expression were evaluated after 48 h. (A) Image of representatives immunoblots. (B) Densitometric analysis of immunoblots of Smac/DIABLO and ER. The expression of 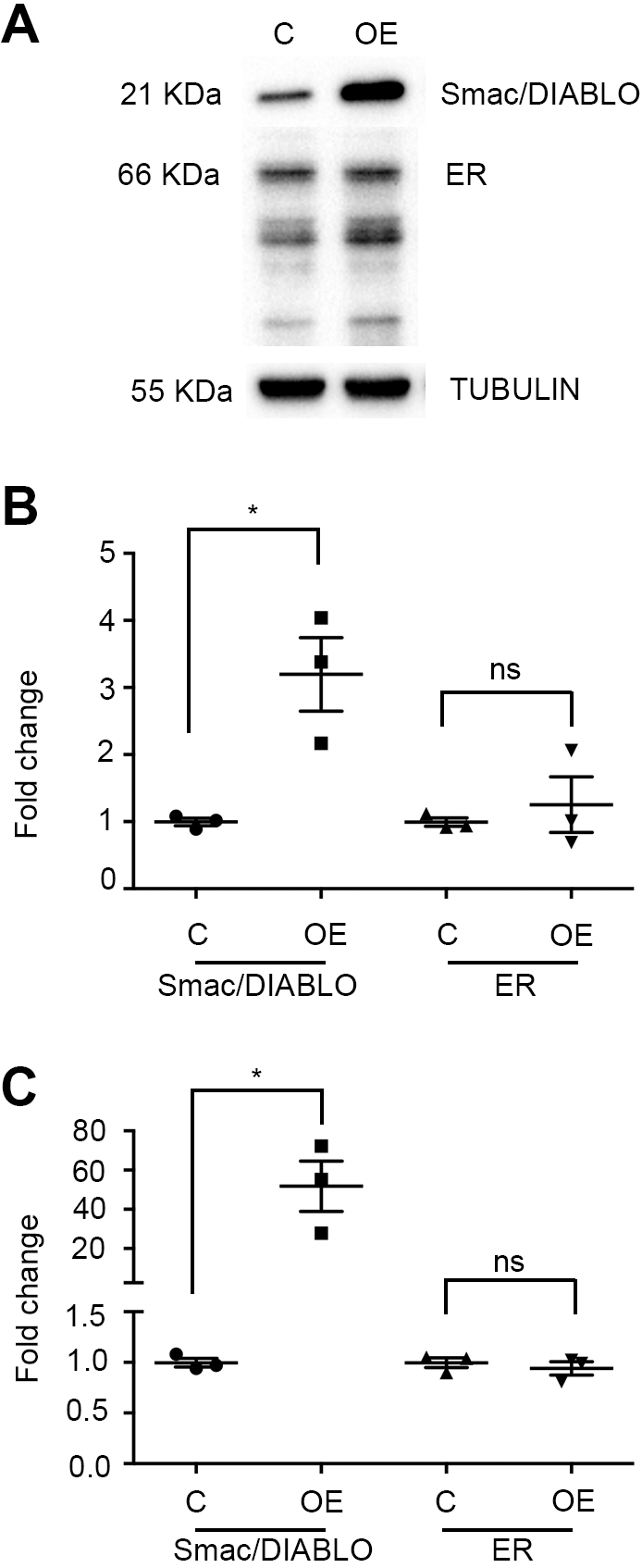
Unfortunately, information regarding to patient’s survival was unavailable; therefore, to assess the effect of Smac/DIABLO expression on breast cancer patients’ survival, we searched an online database (www. kmplot.com) [26]. Data of Smac/DIABLO mRNA expression, assessed by microarrays, were analyzed. First, we investigated whether the levels of Smac/DIABLO were associated with survival of breast cancer patients, irrespective of their ER status. We found that breast cancer patients with high levels of Smac/DIABLO expression had a lower overall survival (OS) (1402 samples,
In conclusion, these data suggested that in breast cancer patients with ER-positive disease, there is an increase in the expression of both Smac/DIABLO mRNA and protein. In addition, high Smac/DIABLO expression is associated with poor prognosis, which depended on ER expression.
In order to gain more insight into the association between Smac/DIABLO and ER in breast cancer, we analyzed the expression of Smac/DIABLO by immunoblots in a panel of breast cancer cell lines with different ER status. We found higher levels of Smac/DIABLO in ER-positive breast cancer cell lines in comparison with ER-negative cells, as revealed by densitometric analyses (Fig. 4A–C). These results suggested for the first time a direct association between Smac/DIABLO and ER in breast cancer.
Estrogen receptor does not regulate Smac/DIABLO expression
Since ER is a known transcriptional factor, we sought to analyze whether Smac/DIABLO could be one of its targets. We exposed the ER-positive breast cancer cell lines MCF-7 and T47D to E2 (10 nM) for 24 h and analyzed Smac/DIABLO mRNA expression by qPCR. As a control, we also evaluated the expression of the pS2 gene, a known ER target gene. The expression of Smac/DIABLO was not regulated by the hormone exposure, while pS2 mRNA was upregulated in the presence of E2 in both cell lines, as shown in Fig. 5A and B. We analyzed whether the promoter of Smac/DIABLO could have estrogen-response elements (EREs). To this purpose, we used the TFBIND tool [27] (
Since it has been reported that overexpression of ER in breast cancer patients correlates with poorer prognosis in breast cancer patients [28, 29], we analyzed whether ER could regulate the expression of Smac/DIABLO in a model of LTED MCF-7 cells, where both an upregulation of ER and a hypersensitivity of its receptor have been observed [30]. To this purpose we cultured MCF-7 cells in estrogen-deprived conditions by five to seven months and as shown in Fig. 6A, we observed a two-fold increase in ER protein expression. When LTED MCF-7 cells were stimulated with E2 for 24 h we did not observe changes in Smac/DIABLO mRNA expression, whereas the known ER target pS2 was effectively upregulated (Fig. 6B). Next, we used the opposite strategy; we inhibited the ER expression in LTED MCF-7 cells with Fulvestrant, a known ER antagonist that induces downregulation of ER. Fulvestrant treatment efficiently inhibited ER protein expression, while Smac/DIABLO expression was not modulated (Fig. 6C and D). Thus, we conclude that Smac/DIABLO is not regulated by ER, neither in normal nor in estrogen-deprived conditions. Of note, that this is the first report where the regulation of Smac/DIABLO expression by ER in these conditions is studied.
Smac/DIABLO does not participate in the regulation of ER expression
Since it has been reported that Smac/DIABLO is able to modulate the expression of a number of proteins [31], we evaluated the participation of this protein on ER expression. For this, we transiently overexpressed Smac/DIABLO in MCF-7 cells and analyzed the ER protein by immunoblot. As shown in Fig. 7A and B, we successfully overexpressed Smac/DIABLO protein in these cells. Nevertheless, no change in ER protein was observed. Additionally, we analyzed ER mRNA levels in these cells by qPCR. As shown in Fig. 7C, mRNA levels of Smac/DIABLO were higher in overexpressing cells, while ER mRNA levels remained unchanged. We thus conclude that Smac/DIABLO does not participate in ER expression regulation in MCF-7 cells. Consequently, Smac/DIABLO role in breast cancer seems not to be directly mediated by ER regulation. These results suggest that another molecule(s) may modulate the function of Smac/DIABLO in breast cancer.
Discussion
Breast cancer remains the most common cancer in the female population worldwide. Clinical and molecular heterogeneity presents a challenge to estimate prognosis as well as the patients’ response to treatment. For these reasons there is a big research effort focused on finding new biomarkers to improve patient management. A widely used biomarker to estimate response to hormone therapy is ER expression, which is expressed in 50–80% of the breast tumors [32]. However, de novo or acquired resistance to hormone therapy remains a major clinical complication [33].
Smac/DIABLO is a proapoptotic protein that exerts its function through IAP inhibition. Altered expression of this protein has been observed in several tumors and, according to its proapoptotic function, reduced levels of Smac/DIABLO have been associated to poor prognosis in bladder, renal, and lung cancers [12, 13, 14]. However, in cervical cancer and leukemias, Smac/DIABLO expression is elevated in comparison with healthy tissue, and in the latter case, this increased expression was associated with poor prognosis [15, 34]. In breast cancer, several groups have analyzed Smac/DIABLO expression, with conflicting results. Pluta et al. [18] reported lower Smac/DIABLO protein levels in breast cancer patients and an inverse correlation with tumor stage. Mansour et al. [20]also found a diminished expression of Smac/DIABLO mRNA in tumor samples and an inverse correlation with survivin and ER [20]. In contrast, Zhang et al. [19] only observed that Smac/DIABLO protein expression was more prevalent in the HER2 positive group. These contradictory results could be attributed to the variability between the different populations analyzed, a reduced number of samples analyzed, the type of molecule (mRNA or protein) studied, and/or the diversity of employed techniques.
In the present study, we assessed the expression of Smac/DIABLO mRNA and protein in two independent groups of Mexican breast cancer patients. We analyzed the Smac/DIABLO mRNA expression by qPCR, while protein levels were assayed with immunohistochemistry. We found a higher expression of Smac/DIABLO mRNA in patients older than 50 years than in younger patients. This contrast with previous works showing a higher expression of Smac/DIABLO in patients younger than 50 years old [18, 35], and a lack of association with age [19, 20, 36]. We also found an inverse correlation between Smac/DIABLO mRNA expression and tumor grade in our samples. This result was in agreement with the recently reported study by Pluta et al., which showed an inverse correlation between Smac/DIABLO and grade [35]; although in previous works the authors did not found this association [18, 19, 20, 36]. It is important to mention that reduced Smac/DIABLO expression has been observed in more advanced tumors of the lung [14], in renal cell carcinoma [13], and germ cell tumors [37]. Even when Smac/DIABLO expression was observed in all samples, we found an increase in its mRNA in ER-positive tumors (
Due to the antagonistic and closely related functions of Smac/DIABLO and survivin, an IAP family member [38, 39], the expression of both proteins in breast cancer has also been previously analyzed. Mansour et al., analyzed the expression of Smac/DIABLO and survivin by semiquantitative PCR and found a significant inverse correlation between these two genes [20]. Accordingly, in a recent study from Zhao and coworkers, the expression of both proteins was analyzed by immunohistochemistry, and a negative association between both proteins was reported (
Next we analyzed Smac/DIABLO levels by immunohistochemistry in a different panel of tumor samples. In agreement with previous reports in breast cancer and other tumors [19, 34, 41], we observed that Smac/DIABLO staining showed cytosolic distribution. Practically all tumor cells (80–100%) presented this protein signal pattern. In contrast to previous reports where an inverse correlation between Smac/DIABLO and ER [20] or, no correlation was observed [18, 19, 35, 36], we found an increase in Smac/DIABLO expression in ER-positive tumors (Table 5; Fig. 1B), as we had previously observed at the mRNA level. These discordant results could be explained, as mentioned before, by the different techniques employed to assess Smac/DIABLO levels and the reduced number of samples analyzed. Interestingly, this is the first report where Smac/DIABLO mRNA and protein levels were positively associated with ER in breast cancer samples. These results were supported by data derived from public databases, where a high number of samples were available, and where a positive correlation between Smac/DIABLO and ER expression was also observed.
In line with the association between Smac/DIABLO and ER, we found that Smac/DIABLO may have an ER-dependent role in the prognosis of breast cancer patients. Our search in an online database revealed that high expression of Smac/DIABLO in ER-positive patients is associated with poor prognosis, whereas in ER-negative patients correlates with a good prognosis. It has been observed that high levels of Smac/DIABLO and lack of ER are independent predictors of pathological complete response (pCR) in patients with locally advanced breast cancer treated with neoadjuvant chemotherapy (anthracyclines) [36]. Additionally, high levels of Smac/DIABLO were associated with a complete remission (pCR) and a better disease-free survival, also in locally advanced breast cancer [35]. As expected, due to its proapoptotic function, high levels of Smac/DIABLO could be associated with a good prognosis since chemotherapy kill cancer cells by inducing apoptosis. However, in accordance with our findings, it has been proposed that apoptosis is important to progression and metastasis of cancer [42]. Even in breast cancer, a high apoptotic index (AI) has been associated with more aggressive tumors [43, 44, 45] and poor prognosis [43, 45]. In support of this role of apoptosis in breast cancer progression, it has been reported that high expression of BCL-2, an antiapoptotic protein, was associated to a good prognosis [46], while high levels of BIK (Bcl-2 interacting killer), a proapoptotic member of the BCL-2 family, were associated to a poor prognosis in this malignancy [47]. Additionally, high expression of caspase-3 protein has been associated to poor prognosis in breast cancer [48]. In addition, in melanoma, low expression of proapoptotic proteins Bax, Bak and Smac/DIABLO was associated with a better prognosis [49]. These counterintuitive results can be explained by several reports that demonstrated that apoptosis is able to induce proliferation and expansion of more aggressive clones to induce tumor growth and invasion [50, 51]. Accordingly, we found in our analysis of TCGA data that Smac/DIABLO protein levels were higher in Luminal B subtype samples in comparison with Basal subtype ones (Fig. 2D). Although Luminal B and Luminal A subtypes of breast cancer are ER-positive, the former shows a higher proliferative index and a higher risk of relapse than Luminal A subtype [52]. In line with these observations, our group reported that overexpression of a new alternative splice variant of Smac/DIABLO increased mammospheres formation derived from the MCF-7 breast cancer cell line. Whole-genome expression analysis from these mammospheres revealed the activation of cell survival and proliferation pathways. ER signaling network was one of them since transcriptional activation of estrogen-responsive genes (PGR, GREB1, PDZK1, and RAB31) was induced by the new Smac/DIABLO variant [53]. Of relevance, this new isoform of Smac lacks of a mitochondrial targeting sequence and of the domain for IAP binding [53], possibly affecting its proapoptotic function. Finally, as mentioned above, BIK can induce a more aggressive phenotype of breast cancer, even though induces a limited cell death [47]. In this regard, even when the pro-apoptotic function of Smac/DIABLO is thought to be IAP-dependent [9, 10, 54] additional mechanisms have been proposed [55]. Moreover, an apoptosis-independent role of Smac/DIABLO in tumor growth inhibition has been described [31] and non-apoptotic functions of Smac/DIABLO related to cancer initiation and progression involving synthesis of lipids, transport of ions, production of exosomes and inflammation have been also proposed [56]. Therefore, Smac/DIABLO functions in cancer have not been fully elucidated.
To validate our initial findings, we assessed the expression of Smac/DIABLO in several breast cancer cell lines with different ER status and found that higher levels of Smac/DIABLO were present in ER-positive breast cancer cell lines. This result agrees with our results derived from tumor samples. Unfortunately, we did not find evidence for an ER-dependent Smac/DIABLO regulation using several approaches. Although it is known that ER regulates genes related to apoptosis, such as Bcl-2, Bcl-xL, survivin, Caspase-9, Puma, Noxa and TRAILR3 [57, 58, 59] to promote cell proliferation, there are no reports about Smac/DIABLO regulation by this transcriptional factor. In agreement with our results, previous data from microarrays performed in breast cancer cell lines aimed to identify target genes for ER did not reveal regulation of Smac/DIABLO [60, 61]. However, the identification of ER target genes is very complex due to the participation of several mechanisms of regulation in its signaling pathway. For this reason, several strategies to identify its target genes have been employed, and there is no consensus between all results [62, 63]. Additionally, a common observation in breast cancer is the overexpression of ER, and higher levels of the receptor have been associated with a poor prognosis in comparison with those patients with low levels of ER. Interestingly, it has been reported that elevated levels of ER can induce aberrant activation of the receptor, even in the absence of the ligand [64]. Clinically, the growth of a breast tumor in an E2-independent way is a mechanism of resistance to hormonal therapy [30]. To study this in vitro, a model for E2 independence has been established by long-term estrogen-deprivation culture of MCF-7 cells, where an increase of twofold in ER levels has been observed [65]. We took advantage of this model to analyze if Smac/DIABLO expression could be regulated by ER. Following our previous experiments, we did not find evidence of ER-mediated regulation of Smac/DIABLO. We conclude that Smac/DIABLO is not a target gene of ER, even in estrogen-deprived conditions, so we do not support this mechanism as an explanation for the poor prognosis of breast cancer patients with ER-positive disease and high levels of Smac/DIABLO. The mechanism of Smac/DIABLO and ER co-expression, and its association with poor prognosis seems be different from that of the proapoptotic protein BIK, which shows a poor prognostic value in ER-positive breast tumors and its expression is regulated by ER [47]. Finally, we explored the possibility that Smac/DIABLO could regulate ER expression. We did not find neither ER mRNA nor protein modulation in response to Smac/DIABLO overexpression. Taking together these results, we conclude that the co-expression of Smac/DIABLO and ER is not due to a direct regulation between them, at least in our experimental conditions. This opens the possibility that another protein or molecule could regulate the expression of both proteins.
It is important to mention that our study has some limitations. We analyzed Smac/DIABLO mRNA and protein expression in two small groups of breast cancer patients; the first one with 57 samples and the second one with 82, so this could be the reason to the difference we observed with regard to previous works. In addition, although we found higher levels of Smac/DIABLO mRNA and protein in ER-positive breast cancer samples, these results were derived from two independent sets of samples. It would be very interesting to perform the analyses in the same samples to find a direct correlation between mRNA and protein levels. In addition, in our study we did not analyze the expression of Smac/DIABLO in adjacent normal tissue or healthy samples, which would add important information about the regulation of Smac/DIABLO in our samples. Finally, an important limitation of our work, was the lack of OS and DMFS information from our patients. This information is essential in order to evaluate the effects of the coexpression of Smac/DIABLO and ER on the survival of the patients involved in the present study. For all the reasons mentioned above, we analyzed public datasets and perform additional assays to support our findings but recognize that further investigation is needed.
Altogether, our data suggest that Smac/DIABLO and ER are co-expressed in breast tumor samples. However, we could not find any direct regulatory mechanism between these two proteins that could explain their association. Interestingly, a prognostic ER-dependent role for Smac/DIABLO was supported by online databases, and further studies are required to delve into this.
Footnotes
Acknowledgments
We thank PhD Oscar Vladimir Orduña Trujillo for his help with statistical analysis. This work was supported by Consejo Nacional de Ciencia y Tecnología (CONACyT) grants 87688 and 79838, and the National Institute of Genomic Medicine grant 013/2008.
Conflict of interest
The authors declare that they have no conflict of interest.
Authors’ contributions
ME, RG, KV, PP, AH, JT, VM INTERPRETATION OR ANALYSIS OF DATA, PREPARATION OF THE MANUSCRIPT.
FL PREPARATION OF THE MANUSCRIPT.
JM REVISION FOR IMPORTANT INTELECTUAL CONTENT.
GC CONCEPTION, INTERPRETATIONS OR ANALYSIS OF DATA, PREPARATION OF THE MANUSC- RIPT.
