Abstract
BACKGROUND:
The CKLF-like MARVEL transmembrane domain-containing family (CMTM) is the protein product of at least one splice variant of each gene contained a Marvel (MAL and related proteins for vesicle trafficking and membrane link) domain, involved in a variety of cellular processes and the pathogenesis of diseases, including tumorigenesis. However, the diverse expression patterns and prognostic values of eight CMTMs have yet to be elucidated.
OBJECTIVE:
We analyzed the expressions and impacts on survival of different CMTM factors in BC patients to determine their potential diagnosis and prognosis values in BC.
METHODS:
In the current study, we examined the transcriptional and survival data of CMTMs in patients with breast carcinoma (BC) from ONCOMINE, GEPIA, Kaplan-Meier Plotter, and cBioPortal databases.
RESULTS:
It was found that CMTM5/7 were down-regulated, whereas CMTM1/6 were up-regulated in BC patients compared with the normal tissues. In survival analyses through the Kaplan-Meier plotter database, increased mRNA expressions of CMTM5/6/7 and decreased mRNA expression of CMTM4 were associated with better relapse-free survival (RFS) of BC patients.
CONCLUSIONS:
These data provided CMTM5/7 as new biomarker and prognostic factors in BC.
Introduction
The CKLF-like MARVEL transmembrane domain-containing family (CMTM) consists of CMTM1, CMTM2, CMTM3, CMTM4, CMTM5, CMTM6, CMTM7, CMTM8 [1]. The gene encoding product of CMTM family is named as MARVEL domain, which regulates transportation of transmembrane proteins and secretory proteins [2]. Previous studies have demonstrated that CMTM family play the important roles in self-stabilization of the immune system and tumorigenesis [3, 4, 5]. Furthermore, CMTMs are involved in cancer cell proliferation, development, metastasis [6, 7, 8]. For example, the results showed that antiproliferative functions were demonstrated for CMTM4 and CMTM8 In HeLa cells [7, 9]. Early studies have found that CMTM5 suppressed tumour growth in human hepatocellular carcinoma and CMTM7 was lowly expressed in digestive system cancer tissue [10, 11].
Breast carcinoma (BC) is the most common malignant tumor among women, which remains the major cause of death from cancer in women worldwide [12]. Because of the frequent recurrence and the unsatisfactory long-term survival of patients with BC [13, 14], it is necessary to further investigate biomarkers and prognostic factors, enhancing diagnosis, prognosis and individualized treatments.
At present, the expression levels and values of CMTM gene family in BC remain unclear. On the basis of the large databases published online and bioinformatics analysis method, we analyzed the expressions and impacts on survival of different CMTM factors in BC patients to determine their potential diagnosis and prognosis values in BC.
Materials and methods
Ethics statement
This study was approved by the Academic Committee of Jiangsu Provincial Center for Disease Control and Prevention, and conducted according to the principles expressed in the Declaration of Helsinki. All data were obtained from published online databases.
ONCOMINE
ONCOMINE (
GEPIA
GEPIA is a RNA sequencing expression data collection and analysis web server, including 9,736 tumors and 8,587 normal samples from the TCGA and the GTEx projects. GEPIA was used to further strengthen differential expression levels of CMTMs in BC/normal [16].
Breast cancer Gene-Expression Miner v4.2
Breast cancer Gene-Expression Miner v4.2 is a data mining tool of published annotated transcriptomic data, consisting 36 genomic datasets [17, 18]. Expressions of candidate genes can be compared according to different clinical criteria, such as ER status, PR status, HER2 status, and so on.
The kaplan-meier plotter
The prognostic value of CMTMs mRNA expression was evaluated using Kaplan-Meier Plotter (
CCLE
The mRNA levels of CMTM7 in BC cells were analyzed by CCLE database (
Results
The transcription levels of CMTMs in patients with BC
Using ONCOMINE databases, the transcriptional levels of CMTMs in cancers with those in normal samples were compared (Fig. 1). The mRNA expression levels of CMTM7 were significantly down-regulated in patients with BC in 14 datasets. In Ma’s dataset [21], the transcription level of CMTM7 in BC was lower than that in the normal samples: invasive Ductal BC with a fold change of
The significant changes of CMTMs expression in transcription level between different types of breast carcinoma (ONCOMINE database)
The significant changes of CMTMs expression in transcription level between different types of breast carcinoma (ONCOMINE database)
The transcription levels of CMTM factors in different types of cancers (ONCOMINE).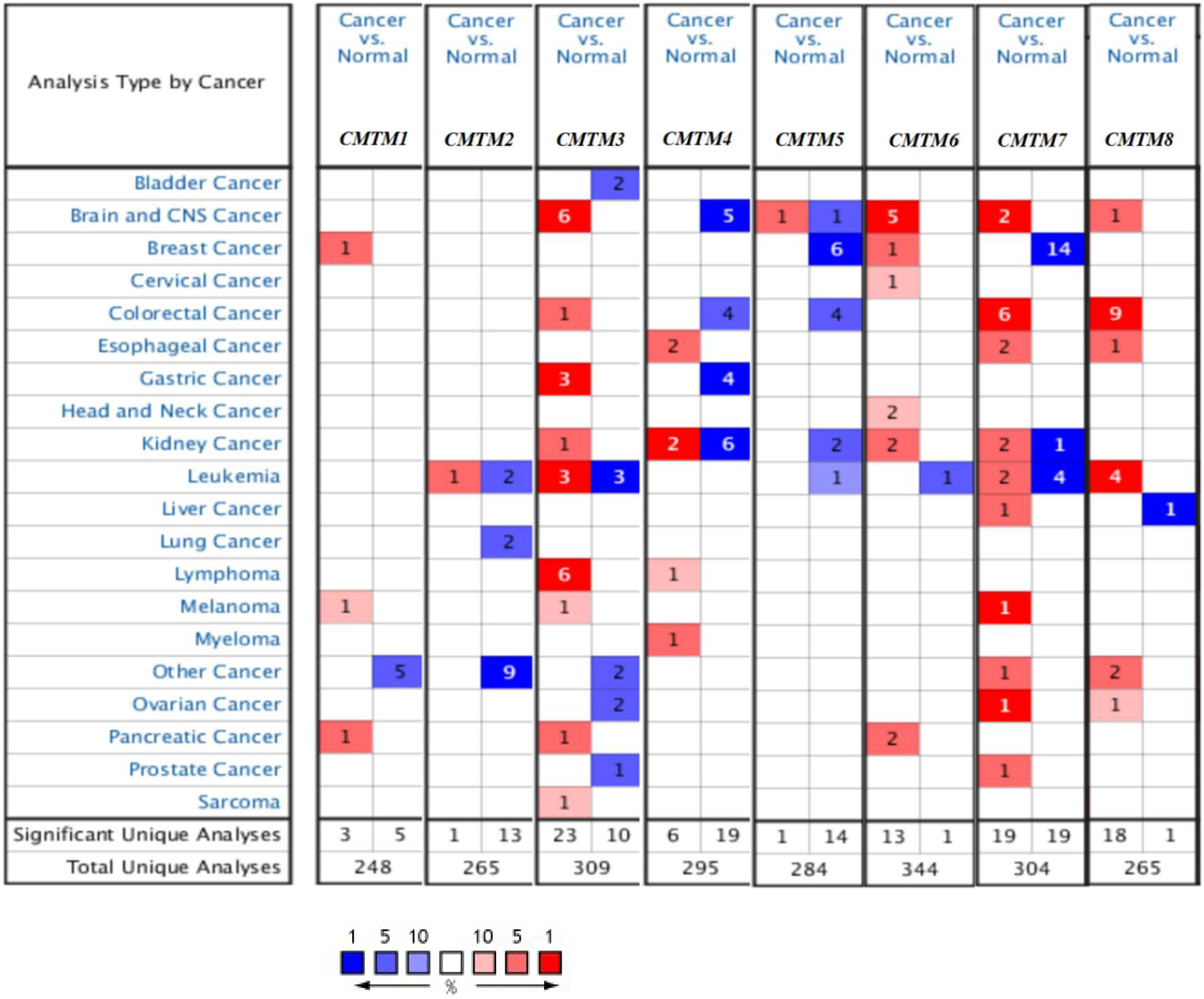
The expression of CMTMs in BC (GEPIA).
In addition, we further compared the mRNA expression levels of CMTMs between breast invasive carcinoma (BRCA) and normal breast tissues by using GEPIA dataset. The result indicated that the expression level of CMTM7 was distinctively lower in BRCA than that in breast tissues (
Breast Cancer Gene-Expression Miner v4.2 was performed to compare the mRNA expression of CMTM factors between BC patients, according to different clinicopathological parameters (Table 2). There was no significant difference between
Increased mRNA expressions of CMTM5/6/7 and decreased mRNA expression of CMTM4 were associated with better RFS of BC patients
We further explored the critical efficiency of CMTMs in the survival of patients with BC. The Kaplan-Meier
The relationship between mRNA expression of CMTMs and clinicopathological parameters of breast carcinoma
The relationship between mRNA expression of CMTMs and clinicopathological parameters of breast carcinoma
curve and log-rank test analyses revealed that the increased CMTM5/6/7 and decreased CMTM4 mRNA levels were significantly associated with RFS (
The prognostic value of mRNA level of CMTM factors in BC patients (RFS in Kaplan-Meier plotter). (A) CMTM1 (1555738_at). (B) CMTM2 (229967_at). (C) CMTM3 (224733_at). (D) CMTM4 (224998_at). (E) CMTM5 (230942_at). (F) CMTM6 (223047_at). (G) CMTM7 (226017_a).
The mRNA levels of CMTM7 in BC cells were analyzed by CCLE database. The results indicated there were 809 CMTM7-related genes, and CMTM7-related genes in the top 20 of positive and negative correlation coefficients were showed (Fig. 4), including SIDT1, FOXA1, SPDEF, IGF2BP2, ETS1, S100A2 and so on.
CMTM7-related genes in the top 20 of positive and negative correlation coefficients.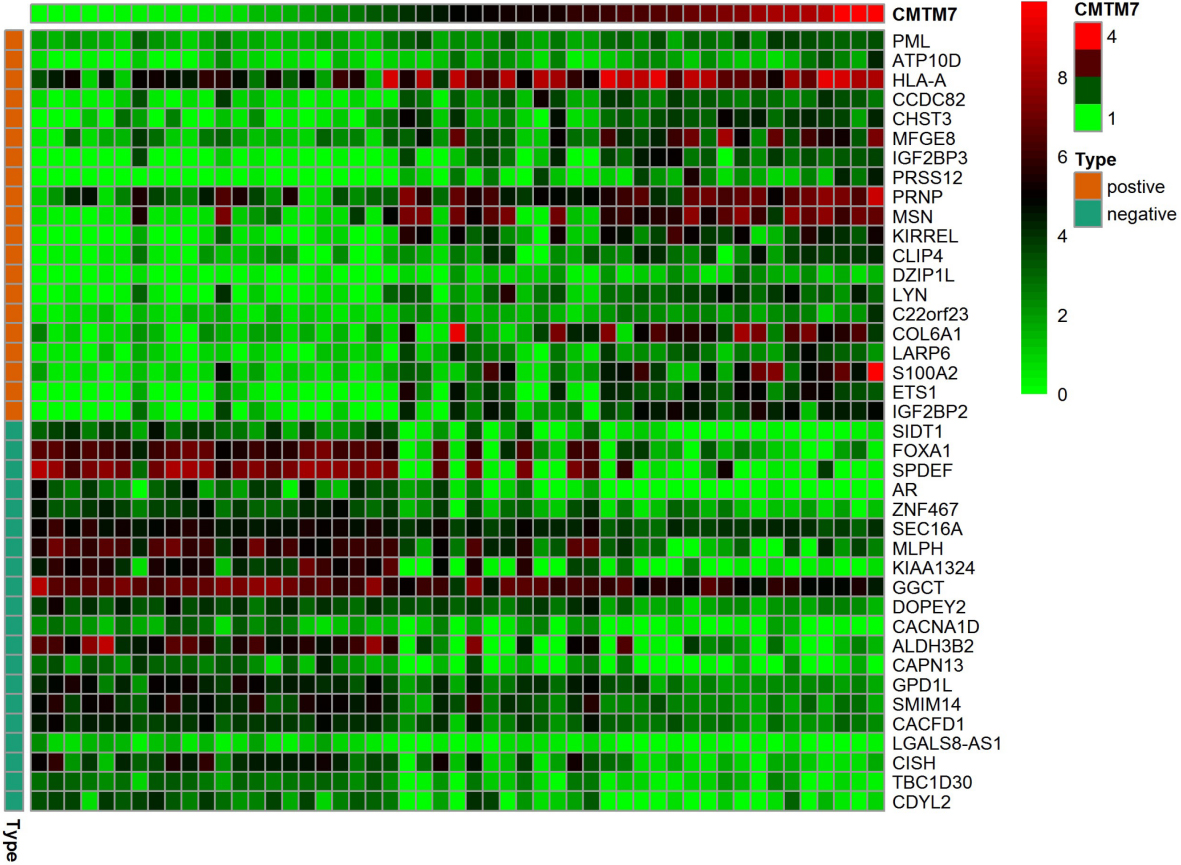
The functions of CMTM7 and CMTM7-related genes were predicted by analyzing Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) in the Database for Annotation, Visualization, and Integrated Discovery (DAVID). GO enrichment analysis predicted the functional roles of target host genes on the basis of three aspects, including biological processes (BP), cellular components (CC), and molecular functions (MF). We found that GO: 0022612 (gland morphogenesis), GO: 0048732 (gland development), GO: 0030879 (mammary gland development), GO: 0010631 (epithelial cell migration), and GO: 0090132 (epithelium migration) were significantly regulated by CMTM7-related genes in BC. GO: 0055037 (recycling endosome), GO: 003101 (extracellular matrix), GO: 0019838 (growth factor binding) were also significantly controlled by CMTM7-related genes, which were associated with BC (Fig. 5A).
The functions of CMTM7 and genes significant associated with CMTM7 alterations were predicted by analysis of GO (A) and KEGG (B) by DAVID tools (
KEGG analysis can analyze the pathways related to the functions of CMTM7-related genes. Five pathways related to the functions of CMTM7-related genes in BC were found (Fig. 5B). All of them, including hsa04510 (Focal adhesion), hsa04151 (PI3K-AKT signaling pathway), hsa04512 (ECM-receptor interaction), hsa01524 (Platinum drug resistance) and hsa04152 (AMPK signaling pathway), were involved in the tumorigenesis and pathogenesis of BC (Fig. 6).
PI3K-AKT signaling pathway regulated by the CMTM7 alteration in BC.
CMTMs is a newly discovered family of proteins linked to chemokines and transmembrane-4 superfamily, and the role of CMTM5/7 in the tumorigenesis of several cancers has been partially confirmed [26, 27]. However, the distinct roles of CMTMs in both mammary gland and BC development are not clear. The present study is the first time to explore the mRNA expressions and prognostic values (RFS) of different CMTM factors in BC by further bioinformatics analysis.
CMTM1 consists of at least 23 alternatively spliced isoforms designated CMTM1_v1-v23, among which, CMTM1_v17 is the most studied in BC. CMTM1_v17 was highly expressed in testis and many tumor tissues including breast tumor [28]. The overexpression of CMTM1_v17 in the BC cell line MDA-MB-231 promoted cell proliferation and resistance to TNF-
It was reported that CMTM5 may exert tumor-suppressive activities on cervical carcinoma [31], pancreatic cancer [32], ovarian cancer [27], leukemia [33] and prostate cancer [34]. CMTM5 expression was significantly downregulated in hepatocellular carcinoma (HCC) tissues as well as cell lines. The expression of CMTM5 was absent in 77.6% of HCC tissues compared with 3.9% in normal liver tissues. Low CMTM5 expression was significantly correlated with poor overall survival in patients with HCC [10]. Not surprisingly, we also found that CMTM5 was suppressed in BC patients. Importantly, the BC patients with higher CMTM5 expression showed better RFS, providing a potential prognostic biomarker for BC. Interestingly, considering hormone receptor criterions, CMTM5 showed positive correlation with TNBC, a more aggressive type with poor prognosis, which was consistent with the early study that patients with TNBC showed p53 overexpression, a known tumor suppressor [35]. In the previous studies on the mechanism of CMTM5 involvement in the tumor suppression, Guo et al. reported that restoration of CMTM5 induced pancreatic cancer cells apoptosis with activation of caspase 3, 8 and 9 [32]. In addition, CMTM5 was found to reduce cell growth and metastasis ability in HCC through regulating PI3K-AKT signaling and its downstream Bcl2, cyclinD1, cyclinE, MMP2 and MMP9 expressions, and an upregulation of p21, Bax, Bad, cleaved caspase3 expressions [10].
CMTM6, a previously uncharacterized protein, was identified as a critical regulator of PD-L1, which was reported as an immune checkpoint inhibitor, to modulate the T cell activities both in vitro and in vivo of some tumors [36, 37]. Guan et al. found that CMTM6 expression in gliomas was dramatically higher than in normal brain tissues, and high CMTM6 expression was associated with reduced survival time and may serve as a strong indicator of poor prognosis in gliomas [38]. By contrary, CMTM6 was downregulated in HCC tissues, supporting a tumor suppressor role of CMTM6 in HCC [39]. In our study, CMTM6 was up-regulated in BC patients. But to our surprise, a high CMTM6 expression was significantly correlated with good RFS in the patients with BC, which seemed inconsistent with the role of CMTM6 as an oncogene. As few evidences focused on CMTM6, the underlining function of CMTM6 needs more researches.
CMTM7 is broadly expressed in normal tissues and cell lines, but frequently silenced or downregulated in some cancer lines and primary tumors [11]. We demonstrated that CMTM7 was down-regulated in BC patients and higher CMTM7 expression predicted better RFS. Furthermore, GEPIA analysis was consistent with that of ONCOMINE, demonstrating that CMTM7 were down-regulated in invasive carcinoma. It was reported that CMTM7 down-regulated the phosphorylation of Akt, inhibited the ERK activation, thereby inhibiting EGFR activation in the P13K/AKT downstream target in KYSE180 cells [40]. Huang et al. found enforced expression of CMTM7 inhibited the cell growth and migration of liver cancer cells, then further analysis revealed that CMTM7 suppressed AKT signaling and induced cell cycle arrest at the G0/G1 phase in the liver cancer cells [41]. Our result of KEGG analysis also highly enriched in PI3K-AKT signaling pathway. Besides, 11 genes enriched in hsa01524 (Platinum drug resistance), including NOXA, FAS, GSTP1, ERK and so on. Chemotherapy resistance, whether inherent or acquired, is a major problem in the management of BC. CMTM1 high expression, another member of CMTM family genes, was also correlated to chemoresistance in non-small cell lung cancer patients treated with neoadjuvant chemotherapy [42].
There was no significant difference between the mRNA levels of CMTM2/3/4/8 in BC and normal tissues, which were consistent with GEPIA analysis. Although there was no significant upregulation of CMTM4 expression level in BC, BC patients with lower mRNA levels of CMTM4 were found to have significantly better RFS. We thus supposed that CMTM4 was a promising target for BC therapy. The new research showed that the downregulation of CMTM4 was found in renal clear cell carcinoma and in the 786-O and A498 cell lines. Also, up-regulated CMTM4 suppressed renal clear cell carcinoma xengraft growth in nude mice [43]. In view of the above ambiguous findings, further research needs to be performed to investigate the role of CMTM4 in tumorigenesis and prognosis.
Conclusions
In this study, we performed bioinformatics analysis on the expressions and prognostic values of the eight CMTMs in BC for the first time. Our results indicated that the decreased expression of CMTM5/7 and increased CMTM6 in BC tissues might play important roles in BC oncogenesis. Our findings supported that CMTM4 was the potential treated target for BC therapy, whereas CMTM5/7 were potential prognostic markers for better survival of BC, providing more accurate prognosis.
Footnotes
Conflict of interest
The authors declare that they have no competing interests.
