Abstract
BACKGROUND:
Breast cancer is the most common malignant tumor and usually occurs in women. Studies have shown that lncRNA nuclear enriched abundant transcript 1 (NEAT1) contributes to breast cancer progression. This study intends to further investigate the molecular mechanism of NEAT1 in breast cancer.
METHODS:
The expression levels of NEAT1, miR-410-3p and Cyclin D1 (CCND1) were detected by quantitative real-time PCR (qRT-PCR) in breast cancer tissues and cells. Kaplan-Meier analysis and the log-rank test were performed to determine the relationship between NEAT1 and overall survival. Cell Counting Kit-8 (CCK-8) assay analyzed cell proliferation. Transwell assay was performed to examine cell migration and invasion. The protein levels of CCND1 and epithelial-mesenchymal transition (EMT)-related proteins (E-cadherin, N-cadherin and Vimentin) were measured by western blot. The target relationship was predicted by bioinformatics analysis, and confirmed by luciferase reporter assay and RNA Immunoprecipitation (RIP) assay. Xenograft analysis was used to evaluate the tumor growth in vivo.
RESULTS:
NEAT1 and CCND1 were upregulated, while miR-410-3p was down-regulated in breast cancer tissues and cells. Higher NEAT1 expression level was associated with lower survival rate of breast cancer patients. Knockdown of miR-410-3p restored silenced NEAT1-mediated the inhibition of on proliferation, migration, invasion and EMT of breast cancer cells. In addition, NEAT1 regulated CCND1 expression by sponging miR-410-3p in breast cancer cells. NEAT1 knockdown blocked the tumor growth in vivo.
CONCLUSION:
NEAT1 induced breast cancer progression by regulating the miR-410-3p/CCND1 axis, indicating that NEAT1 may be a potential therapeutic target in breast cancer.
Introduction
Breast cancer (BC) is the most common malignant tumor worldwide and the leading cause of cancer death in women [1, 2]. Breast cancer caused 535,000 deaths among women and 10,000 deaths among men, making it the fifth-largest cancer deaths in the world in 2016 [3]. Moreover, breast cancer is divided into four subtypes, namely luminal A, luminal B, HER2-positive and basal-like, of which the majority of basal-like subtypes are triple-negative breast cancer (TNBC) [4]. Furthermore, non-coding RNAs (ncRNAs) have been reported to contribute to breast cancer progression [5].
Long non-coding RNAs (lncRNAs) are more than 200 nucleotides in length and have no protein-coding ability. LncRNAs can serve as indicators for the diagnosis and prognosis of breast cancer [6]. Mechanically, lncRNAs act as competing endogenous RNAs (ceRNAs) by modulating target gene expression [7]. Recently, lncRNA nuclear enriched abundant transcript 1 (NEAT1) plays a crucial regulatory role in various cancers [8], including acting as an oncogene, regulating the Wnt and AKT pathways or sponging microRNAs [9]. Several studies have unveiled that NEAT1 participates in the pathogenesis of human malignancies and may be a therapeutic target for many cancers [10, 11, 12]. In previous research, NEAT1 played a tumor-promoting role in triple-negative breast cancer by regulating chemoresistance and cancer stemness [13]. Also, NEAT1 triggered growth, migration, and invasion of breast cancer cells by upregulating ZEB1 and sponging miR-448 [14]. However, the mechanism of lncRNA NEAT1 is poorly investigated in breast cancer.
MicroRNAs (miRNAs) consist of 18 to 25 nucleotides and are highly conserved. In terms of mechanism, miRNAs are capable of regulating the expression of target genes at the post-transcriptional level [15]. Previous studies have revealed that miR-410-3p functioned as a tumor promoter or suppressor in cancer. For example, miR-410-3p was upregulated in prostate cancer and down-regulated in breast cancer [17, 17]. Moreover, miR-410 targeted endoplasmic reticulum lipid raft-associated 2 (ERLIN2) to impede the progression of ER-positive breast cancer [18]. Among them, ERLIN2 regulates cell cycle progression associated with breast cancer by interacting with mitotic promoting factors [19]. Many studies have elucidated the correlation between lncRNAs and miRNAs in breast cancer. Nevertheless, the relationship between NEAT1 and miR-410-3p in the progression of breast cancer has not been reported.
Cyclin D1 (CCND1) is associated with cell cycle dysregulation in cancers, including Ewing sarcoma [16], oral carcinoma [21] and breast cancer [22]. Previous studies have indicated that cell cycle regulator CCND1 was a target of miR-142, and miR-142 inhibited proliferation of endometrial cancer cells by targeting CCND1 [23]. In breast cancer, LINC00473 induced cell proliferation by activating CCND1 expression [24].
In this study, we examined the expression levels of NEAT1, miR-410-3p and CCND1 in breast cancer tissues and cells. Further, the function and molecular mechanism of NEAT1 and miR-410-3p were investigated.
Materials and methods
Tissue samples
All 37 pairs of breast cancer tissues and corresponding adjacent normal tissues were obtained from patients who received surgery at the First Affiliated Hospital of Nanchang University. All patients did not receive chemotherapy or radiotherapy prior to surgery. The research was approved by the Ethics Committee of the First Affiliated Hospital of Nanchang University. Written informed consent was signed by all participants. All tissues were immediately frozen in liquid nitrogen and then stored at
Cell culture
Human non-tumorigenic epithelial cell line (MCF-10A) and two breast cancer cell lines (SKBR3 and MCF7) were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). The cells were seeded in Dulbecco’s Modified Eagle Medium (DMEM; Gibco, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS; Gibco) at 37
Plasmids and cell transfection
Small interfering RNA (siRNA) of NEAT1 (si-NEAT1) and the control siRNA (si-NC), NEAT1 overexpression plasmid (NEAT1) and the negative control pcDNA, shRNA of NEAT1 (sh-NEAT1) and the negative control (sh-NC), miR-410-3p mimic (miR-410-3p) and the control mimic (miR-NC), miR-410-3p inhibitor (in-miR-410-3p) and the control inhibitor (in-miR-NC), pcDNA-CCND1 (CCND1) and the control pcDNA (pcDNA) were purchased from RiboBio (Guangzhou, China). Cells were transfected by Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, USA) following the protocols of manufacturer.
Quantitative real-time PCR (qRT-PCR) assay
Total RNA was isolated from tissues and cells using TRIzol reagent (Invitrogen) following the manufacturer’s protocol. The first-strand cDNA was synthesized using the High-Capacity cDNA Reverse Transcription Kits (Thermo Fisher Scientific, Waltham, MA, USA). qRT-PCR was performed using SYBR Green PCR Master Mix (Thermo Fisher Scientific). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and U6 were identified as endogenous controls. Primers as follows: NEAT1 (forward, 5’-CTTCCTCCCTTTAACTTATCC ATTCAC-3’; reverse, 5’-CTCTTCCTCCACCATTACC AACAATAC-3’), CCND1 (forward, 5’-GCTGCGAAG TGGAAACCATC-3’; reverse, 5’-CCTCCTTCTGCAC ACATTTGAA-3’), GAPDH (forward, 5’-TGCACCAC CAACTGCTTAGC-3’; reverse, 5’-GGCATGGACTGT GGTCATGAG-3’), U6 (forward, 5’-CTCGCTTCGGC AGCACA-3’; reverse, 5’-AACGCTTCACGAATTTG CGT-3’), 18S (forward, 5’-AAACGGCTACCACATCC A-3’; reverse, 5’-CACCAGACTTGCCCCTCCA-3’). The primers for miR-410-3p were purchased from Sangon Biotech (Shanghai, China).
Luciferase reporter assay
The wild-type (WT) or mutant (MUT) NEAT1 containing miR-410-3p binding sites was cloned into pGL3 vector (Promega, Madison, WI, USA), respectively. Then, the control mimic (miR-NC) or miR-410-3p mimic (miR-410-3p) was cotransfected with pGL3-NEAT1-WT or pGL3-NEAT1-MUT into SKBR3 and MCF7 cells. In addition, to explore whether miR-410-3p may bind to CCND1 3’-UTR, pGL3 plasmids harboring the predicted miR-410-3p binding sites (CCND1-WT) or the mutant (CCND1-MUT) were constructed and cotransfected with the control mimic (miR-NC) or miR-410-3p mimic (miR-410-3p) into SKBR3 and MCF7 cells using Lipofectamine 2000 (Invitrogen). Luciferase activity was analyzed using the Dual-Luciferase Reporter Assay System (Promega) at 48 h post-transfection according to the manufacturer’s protocol.
RNA Immunoprecipitation (RIP) assay
Magna RNA-Binding Protein Immunoprecipitation Kit (Millipore, Billerica, MA, USA) was used to perform the RIP assay following the manufacturer’s instructions. Briefly, SKBR3 and MCF7 cells were harvested after transfection with the control mimic (miR-NC) or miR-410-3p mimic (miR-410-3p). Next, the cells were lysed with RIP lysis buffer containing protease inhibitor and RNase inhibitor. The cell lysates were collected and incubated with magnetic beads coated with Ago2 antibody (Millipore) for 4 h at 4
Cell Counting Kit-8 (CCK-8) assay
The transfected cells (3.0
Transwell assay
Interestingly, transwell chambers for cell invasion were coated with Matrigel (BD Biosciences, Franklin Lakes, NJ, USA), while the transwell chambers for cell migration were uncoated. Firstly, cells were suspended in serum-free medium and then added into the upper chamber containing 8
Western blot assay
Total protein was extracted using RIPA buffer (Thermo Fisher Scientific) containing protease inhibitor cocktail (Roche Molecular Biochemicals, Indianapolis, IN, USA). Protein concentration was determined using the Easy II Protein Quantitative Kit (TransGen Biotech, Beijing, China). Subsequently, the protein samples were separated by SDS-PAGE and transferred to PVDF membranes (Millipore). The membranes were blocked by 5% non-fat milk for 2 h at room temperature and then incubated at 4
LncRNA NEAT1 was upregulated, while miR-410-3p was down-regulated in breast cancer tissues and cells. (A) and (D) qRT-PCR was carried out to detect the expression levels of NEAT1 and miR-410-3p in breast cancer tissues and adjacent normal tissues. (B) and (E) The expression levels of NEAT1 and miR-410-3p were analyzed in non-tumorigenic epithelial cells (MCF-10A) and breast cancer cell lines (SKBR3 and MCF7) using qRT-PCR. (C) and (F) The correlation between NEAT1 expression or miR-410-3p expression and overall survival were determined by Kaplan-Meier analysis. GAPDH or U6 was used as a loading control. 
BALB/c nude female mice (6 weeks old) were maintained in pathogen-free (SPF) conditions. SKBR3 cells were transfected with sh-NC or sh-NEAT1, and then subcutaneously injected into nude mice to establish a xenograft model. Tumor volume was measured every 7 days, and tumor weight was detected after mice were sacrificed. Partial breast tissues were snap-frozen for RNA and protein extraction. The xenograft analysis was approved by the Animal Ethics Committee of the First Affiliated Hospital of Nanchang University.
Statistical analysis
All experiments were performed at least three times independently. All data were expressed as mean
NEAT1 directly targeted miR-410-3p. (A) The predicted binding sites of NEAT1 and miR-410-3p. (B) and (C) SKBR3 and MCF7 cells were cotransfected with NEAT1-WT or NEAT1-MUT luciferase reporter vectors and miR-NC or miR-410-3p mimic (miR-410-3p), and luciferase activity was examined at 48 h after transfection. (D) RIP assay was performed to determine NEAT1 enrichment in Ago2 immunoprecipitation complex. (E) The correlation between NEAT1 and miR-410-3p in breast cancer tissues. (F) and (G) SKBR3 and MCF7 cells were transfected with pcDNA, pcDNA-NEAT1 (NEAT1), si-NC and si-NEAT1, respectively. The expression levels of NEAT1 and miR-410-3p were detected by qRT-PCR. 
LncRNA NEAT1 was upregulated while miR-410-3p was down-regulated in breast cancer tissues and cells
Firstly, the levels of NEAT1 and miR-410-3p in breast cancer tissues and cells were detected using qRT-PCR. The results of qRT-PCR exhibited that NEAT1 expression was dramatically increased in breast cancer tissues and cell lines (SKBR3 and MCF7) compared to adjacent normal tissues and non-tumorigenic epithelial cells (MCF-10A) (Fig. 1A and B). However, miR-410-3p expression was remarkably decreased in breast cancer tissues and cells (Fig. 1D and E). We also detected the expression levels of NEAT1 and miR-410-3p in non-tumorigenic epithelial cells (MCF-10A), triple-negative (BT549, MDA-MB-231), HER2-positive (ZR-75-30, SKBR3) and estrogen receptor-positive (MCF7) breast cancer cells. The results of qRT-PCR exhibited that NEAT1 expression was strikingly increased, and miR-410-3p expression was remarkably reduced in five breast cancer cell lines compared to MCF-10A cells, with the most significant changes in SKBR3 and MCF7 cells (Fig. S3). In addition, to analyze the correlation between NEAT1 expression and breast cancer patient prognosis, the Kaplan-Meier analysis was performed and the results showed that higher NEAT1 or lower miR-410-3p expression significantly associated with a poor prognosis in the enrolled breast cancer patients (Fig. 1C and F). These data revealed that NEAT1 might play an oncogenic role in breast cancer. Besides, we predicted miRNAs bound to NEAT1 and screened five miRNAs (miR-495-3p, miR-507, miR-329-3p, miR-129-5p and miR-410-3p) that were down-regulated in BC. Then, the expression levels of five miRNAs in BC tissues and adjacent normal tissues were verified by qRT-PCR. The results suggested that the down-regulation of miR-410-3p in BC tissue was the most significant among the five differentially expressed miRNAs, so miR-410-3p was selected for further research (Fig. S1A).
NEAT1 directly targeted miR-410-3p
Firstly, to explore the subcellular localization of NEAT1 in BC cells, nuclear and cytoplasmic fraction assay was performed. The results exhibited that NEAT1 was mainly distributed in the cytoplasm of BC cells (Fig. S2). To investigate the interaction between NEAT1 and miR-410-3p, LncBase Predicted v.2 was utilized. Bioinformatics analysis exhibited that miR-410-3p might be a target of NEAT1 (Fig. 2A). Then luciferase reporter assay was used to validate that NEAT1 targeted miR-410-3p. The results demonstrated that miR-410-3p mimic strikingly reduced the luciferase activity of SKBR3 and MCF7 cells transfected with NEAT1-WT, whereas did not modulate the luciferase activity when the binding sites were mutated (Fig. 2B and C). To further validate the relationship between NEAT1 and miR-410-3p, RIP assay was performed. RIP assay indicated that NEAT1 was significantly enriched in miR-410-3p group compared with the control group (Fig. 2D). Moreover, the correlation between NEAT1 and miR-410-3p was negatively correlated in breast cancer tissues (Fig. 2E). Next, the expression levels of NEAT1 and miR-410-3p in SKBR3 and MCF7 cells transfected with pcDNA, pcDNA-NEAT1 (NEAT1), si-NC or si-NEAT1 were detected, respectively. The results suggested that overexpression of NEAT1 markedly elevated NEAT1 expression and decreased miR-410-3p expression, while NEAT1 was
Knockdown of miR-410-3p reversed silenced NEAT1-mediated the inhibition of proliferation, migration, invasion and EMT of breast cancer cells. (A)–(F) SKBR3 and MCF7 cells were transfected with si-NC, si-NEAT1, si-NEAT1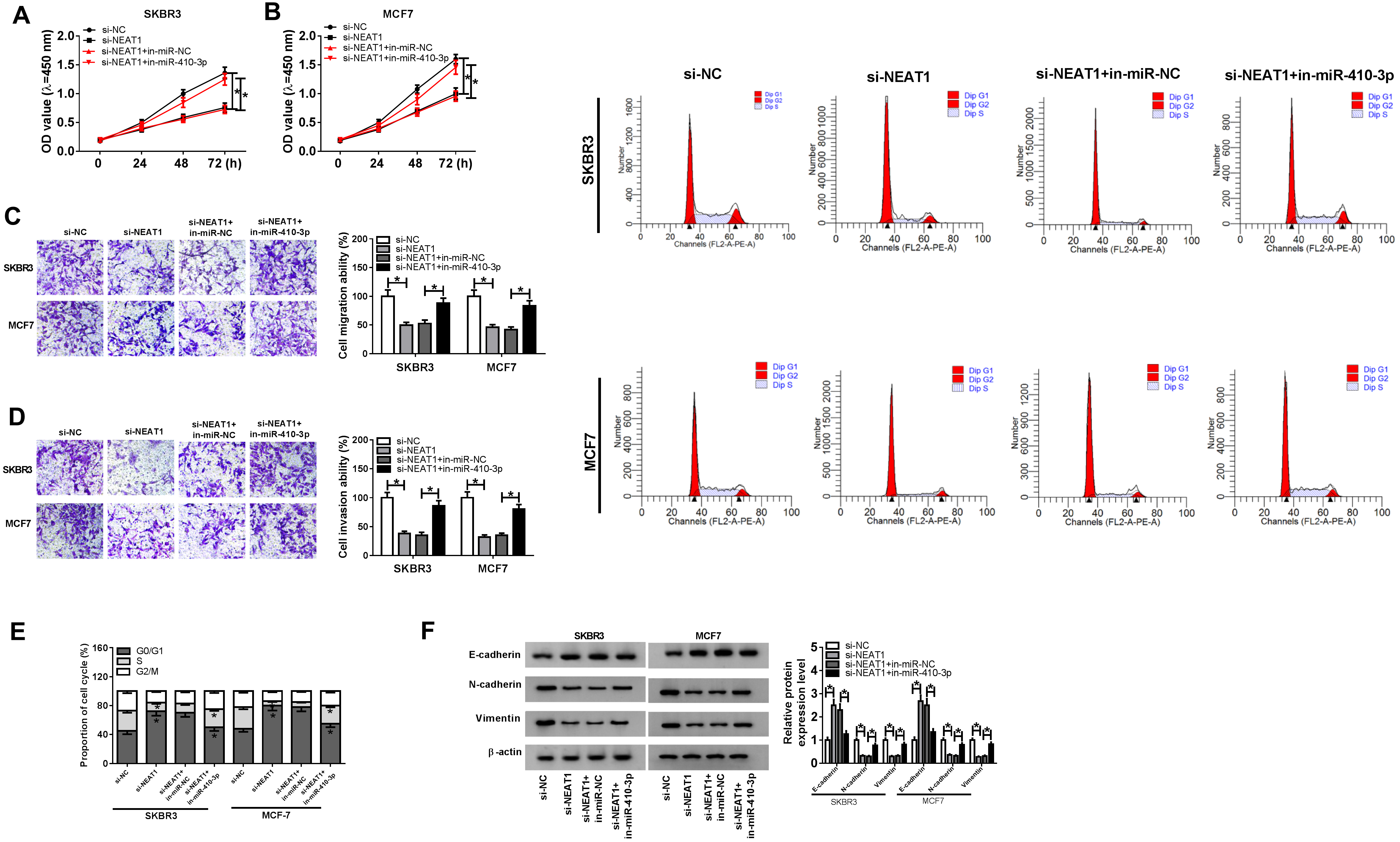

CCND1 was a direct target of miR-410-3p in breast cancer. (A) The putative binding sites for miR-410-3p and CCND1 3’-UTR, and mutations in the binding sites. (B) and (C) SKBR3 and MCF7 were cotransfected with CCND1 3’-UTR-WT or CCND1 3’-UTR-MUT luciferase reporter plasmids and miR-NC or miR-410-3p mimic (miR-410-3p), and luciferase activity was examined at 48 h post transfection. (D) CCND1 expression was detected in breast cancer tissues and adjacent normal tissues using qRT-PCR. (E) The expression of CCND1 in BC tissues and adjacent normal tissues was analyzed using IHC. (F) The expression of CCND1 was examined in non-tumorigenic epithelial cells (MCF-10A) and breast cancer cell lines (SKBR3 and MCF7) by qRT-PCR. (G) The correlation between miR-410-3p and CCND1. (H) SKBR3 and MCF7 cells were transfected with miR-NC or miR-410-3p mimic (miR-410-3p), and CCND1 expression was detected by western blot.
remarkably down-regulated and miR-410-3p was strikingly upregulated in SKBR3 and MCF7 cells transfected with si-NEAT1 (Fig. 2F and G). Thus, these results demonstrated that NEAT1 targeted and negatively regulated miR-410-3p.
To explore the functional effects of NEAT1 and miR-410-3p on breast cancer progression, SKBR3 and MCF7 cells were transfected with si-NC, si-NEAT1, si-NEAT1
CCND1 reversed the inhibition of miR-410-3p overexpression on proliferation, migration, invasion and EMT of breast cancer cells. (A)–(F) SKBR3 and MCF7 cells were transfected with miR-NC, miR-410-3p, miR-410-3p
6 NEAT1 regulated CCND1 expression through miR-410-3p in breast cancer cells. (A) and (B) SKBR3 and MCF7 cells were transfected with si-NC, si-NEAT1, si-NEAT1
Knockdown of NEAT1 blocked the tumor growth in vivo. (A)–(E) SKBR3 cells were transfected with sh-NC or sh-NEAT1 and subcutaneously injected into nude mice. (A) Tumor volume was examined every 7 days. (B) Tumor weight was detected after mice were sacrificed. (C) and (D) The levels of NEAT1 and miR-410-3p were examined by qRT-PCR. (E) CCND1 expression was detected using IHC. (F) The protein level of CCND1 was measured by western blot. 
To investigate the molecular mechanism of miR-410-3p in breast cancer, we first predicted the target of miR-410-3p using microT-CDS. CCND1 might be a target of miR-410-3p (Fig. 3A). In addition, we predicted the target genes of miR-410-3p and screened five target genes (HMGB1, STAT3, IRS1, SOX2 and CCND1) that were upregulated in BC. Next, qRT-PCR was used to confirm the expression levels of five target genes in BC tissue and adjacent normal tissues. The results showed that among the five differentially expressed target genes, CCND1 was most remarkably upregulated in BC tissues, so CCND1 was selected for subsequent research (Fig. S1B). To verify whether miR-410-3p directly targeted CCND1, luciferase reporter assay was performed. The results showed that miR-410-3p mimic significantly decreased the luciferase activity of the CCND1 3’-UTR-WT group, but not of the CCND1 3’-UTR-MUT group (Fig. 3B and C). Furthermore, the results of qRT-PCR and immunohistochemistry (IHC) exhibited that CCND1 expression was remarkably higher in breast cancer tissues than adjacent normal tissues (Fig. 3D and E). Also, CCND1 expression was markedly increased in SKBR3 and MCF7 cells compared to MCF-10A cells (Fig. 3F). The expression levels of miR-410-3p and CCND1 were negatively correlated in breast cancer tissues (Fig. 3G). To further confirm the correlation between miR-410-3p and CCND1, CCND1 expression was detected in SKBR3 and MCF7 cells transfected with miR-NC or miR-410-3p mimic. The results showed that miR-410-3p mimic dramatically decreased CCND1 expression (Fig. 3H). All of these results indicated that CCND1 was a direct target of miR-410-3p in breast cancer.
CCND1 reversed the inhibition of miR-410-3p overexpression on proliferation, migration, invasion and EMT of breast cancer cells
To investigate the effects of miR-410-3p and CCND1 on breast cancer progression, SKBR3 and MCF7 cells were transfected with miR-NC, miR-410-3p, miR-410-3p
NEAT1 regulated CCND1 expression in breast cancer cells through miR-410-3p
In order to study the effects of NEAT1 and miR-410-3p on CCND1 expression, SKBR3 and MCF7 cells were transfected with si-NC, si-NEAT1, si-NEAT1
Knockdown of NEAT1 blocked the tumor growth in vivo
To investigate the effect of NEAT1 on the tumorigenesis of breast cancer in vivo, xenograft nude mouse model was constructed. SKBR3 cells transfected with sh-NC or sh-NEAT1 were subcutaneously injected into nude mice. After injection, tumor volume and weight were greatly decreased in the sh-NEAT1 group compared to the sh-NC group (Fig. 6A and B). Moreover, we also determined the expression levels of NEAT1, miR-410-3p and CCND1 in tumor tissues by qRT-PCR, IHC or western blot assay after the mice were sacrificed. These results showed that NEAT1 and CCND1 were distinctly down-regulated (Fig. 6C and E), while miR-410-3p was obviously upregulated in the sh-NEAT1 group compared with the sh-NC group (Fig. 6D). Thus, xenograft assay indicated that NEAT1 knockdown blocked the tumor growth in vivo.
Discussion
According to the presence or absence of molecular markers of estrogen or progesterone receptors and HER2, breast cancer is divided into three main subtypes: hormone receptor positive/HER2 negative, HER2 positive, and triple-negative [25]. The patient’s treatment strategy is based on breast cancer subtypes and tumor stages [26].
LncRNAs have previously been shown to be involved in the tumorigenesis and progression of various tumors [27]. Increasing evidence suggested that lncRNA NEAT1 was upregulated and acted as tumor-promoting function in most tumors [9], such as hepatocellular carcinoma [28], non-small cell lung cancer [29] and colorectal cancer [30]. Yuan et al. demonstrated that NEAT1 functioned as a competing endogenous RNA via sponging miR-133a and regulating SOX4 to promote cervical cancer development [31]. Qian et al. revealed that in breast cancer, NEAT1 positively regulated EZH2 through sponging miR-101, thereby inducing the progression of breast cancer [32]. Moreover, NEAT1 expression was positively correlated with poor prognosis, and NEAT1 facilitated cell proliferation and invasion via targeting miR-218 in breast cancer, suggesting that NEAT1 might be a therapeutic target for breast cancer treatment [33]. It has been previously shown that NEAT1 promoted epithelial-mesenchymal transition (EMT) and 5-fluorouracil (5-FU) resistance by the NEAT1/miR-211/HMGA2 axis [34]. In this study, we found that NEAT1 was remarkably upregulated in breast cancer tissues and cells, and NEAT1 directly targeted miR-410-3p to regulate breast cancer progression.
The present study revealed that NEAT1 bound to miR-410-3p. MiR-410-3p has been reported to play vital effects in various cancers, such as prostate cancer [16] and breast cancer [17]. MiR-410-3p was greatly upregulated in prostate cancer tissues and cell lines and contributed to cell proliferation, migration and EMT by downregulating PTEN expression [16]. In breast cancer, miR-410-3p was evidently down-regulated and inhibited cell proliferation and EMT by targeting Snail [17]. The contradictory role of miRNA-410-3p in prostate cancer and breast cancer is related to tumor microenvironment [35]. The present study further confirmed that miR-410-3p was significantly down-regulated in breast cancer tissues and cells. This research showed that NEAT1 was a sponge of miR-410-3p and miR-410-3p knockdown could abolish the inhibition on proliferation, migration, invasion and EMT caused by NEAT1 knockdown of breast cancer cells.
Furthermore, we further investigated the molecular mechanism of miR-410-3p in breast cancer. The targets of miR-410-3p were predicted by microT-CDS, and CCND1 was selected. CCND1, a cell cycle regulator, played a crucial role in tumorigenesis and was upregulated in breast cancer [36]. Li et al. found that miR-520a-3p inhibited the expression of CCND1 and CD44 at post-transcriptional levels, and repressed cell proliferation, migration and invasion, and induce apoptosis through targeting CCND1 and CD44 in breast cancer [37]. The present study showed that CCND1 was a direct target of miR-410-3p in breast cancer, and CCND1 reversed the miR-410-3p overexpression induced the inhibition of proliferation, migration, invasion and EMT of breast cancer cells.
In conclusion, we found that NEAT1 accelerated breast cancer cell proliferation, migration, invasion and EMT through sponging miR-410-3p and regulating CCND1 (Fig. S6), and NEAT1 knockdown blocked the tumor growth in vivo. Therefore, these results indicated that NEAT1 is a potential therapeutic target for breast cancer treatment.
Footnotes
Conflict of interest
There are no conflicts of interest to declare.
Supplementary data
(A) The expression of five miRNAs associated with NEAT1 and down-regulated in BC was detected in BC tissues and adjacent normal tissues using qRT-PCR. (B) The expression of five target genes bound to miR-410-3p and upregulated in BC was measured in BC tissues and adjacent normal tissues using qRT-PCR. 
The expression of NEAT1 in the nucleus and cytoplasm was examined by qRT-PCR. U6 and 18S were used as positive controls for nuclear and cytoplasmic fractions, respectively. 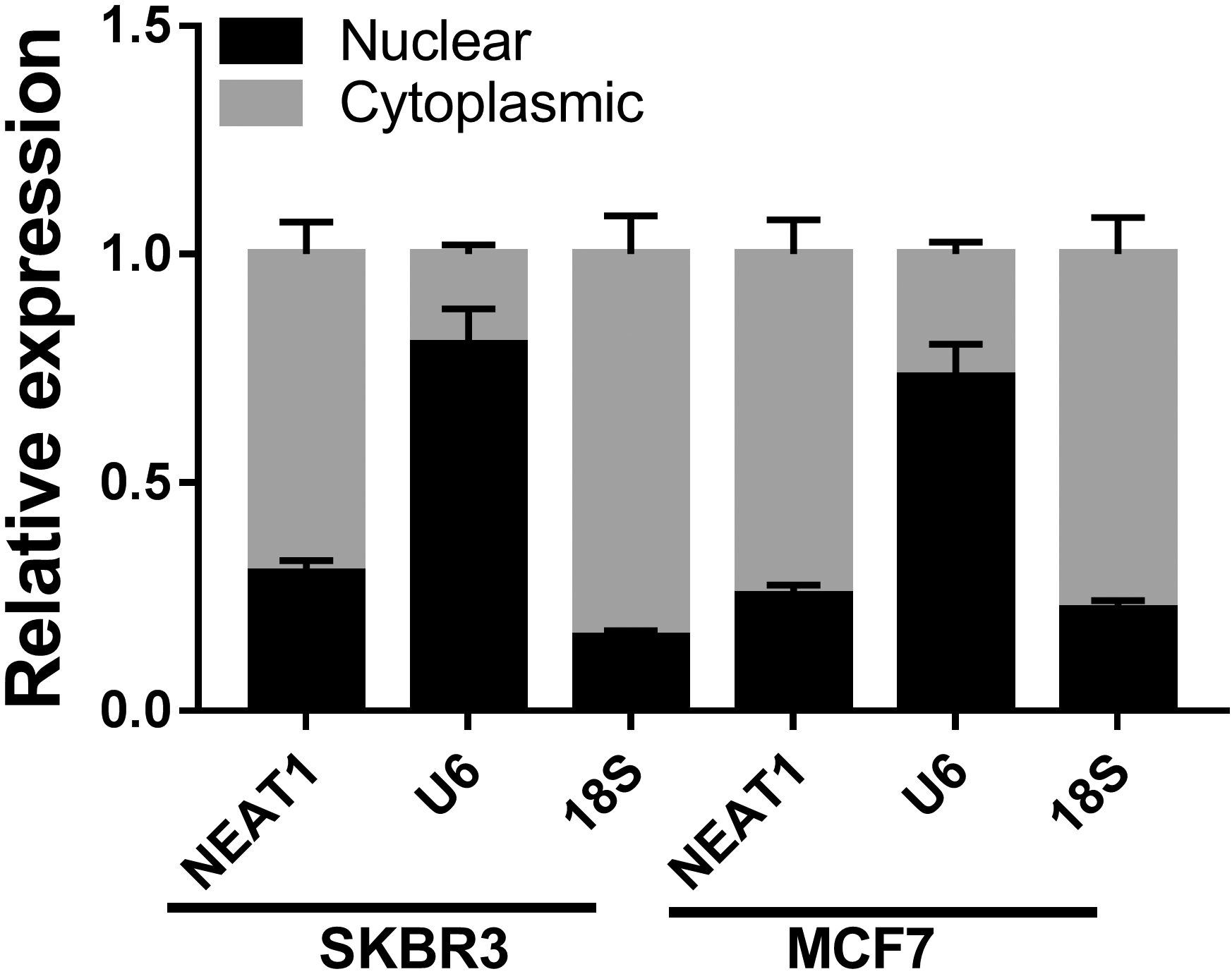
The levels of NEAT1 and miR-410-3p in MCF-10A, ZR-75-30, BT549, MDA-MB-231, SKBR3 and MCF7 cells were measured by qRT-PCR. 
SKBR3 and MCF7 cells were transfected with si-NC, si-NEAT1, si-NEAT1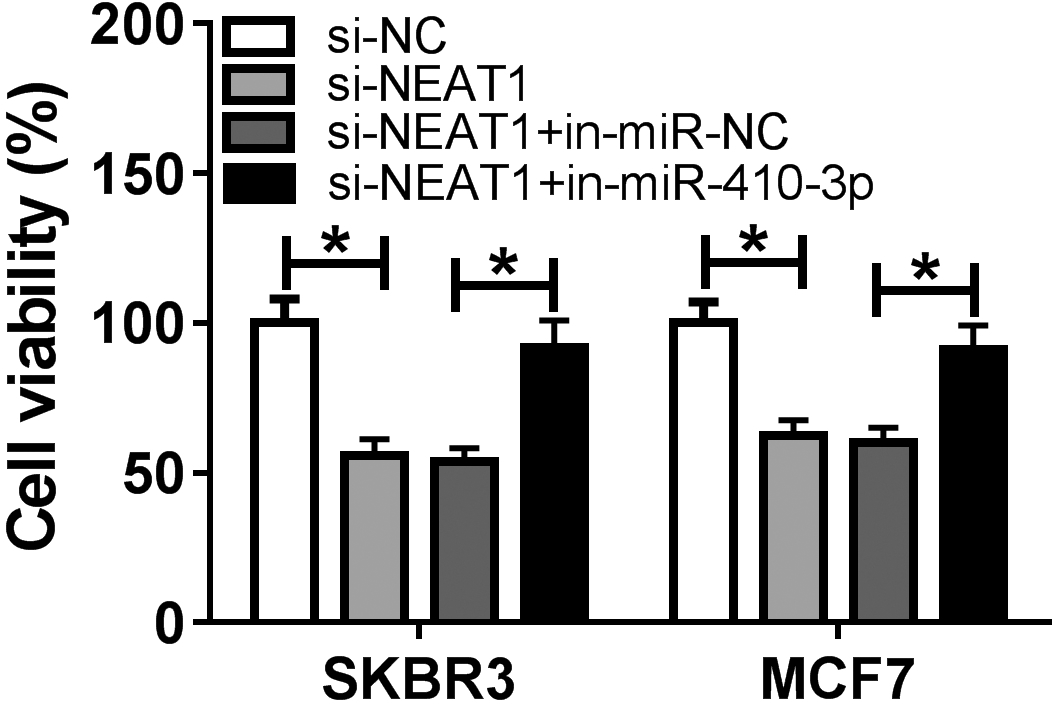
SKBR3 and MCF7 cells were introduced with miR-NC, miR-410-3p, miR-410-3p
Mechanism of NEAT1 in breast cancer development. Red arrows represent an increase and blue arrows represent a decrease.
