Abstract
Osteosarcoma (OS) is one of the most primary bone malignancies, often occurring in adolescents or children. Numerous scientific findings have introduced that long noncoding RNAs (lncRNAs) can be involved in tumor occurrence and development. Although DSCAM-AS1 has been studied in several cancers, its role and mechanism in OS are poorly understood. In this work, high level of DSCAM-AS1 was validated in OS cell lines. Depleting DSCAM-AS1 inhibited cell proliferation, migration and EMT process in OS. Subsequently, we disclosed that DSCAM-AS1 was mainly observed in the cytoplasm of OS cells and could bind with miR-186-5p in OS. Moreover, inhibiting miR-186-5p rescued the impact of silenced DSCAM-AS1 on OS progression. Additionally, GPRC5A was verified as the target downstream of miR-186-5p, and it was negatively modulated by miR-186-5p but positively regulated by DSCAM-AS1. More importantly, DSCAM-AS1 enhanced GPRC5A level in OS by sequestering miR-186-5p. Finally, up-regulating GPRC5A reversed the influences of DSCAM-AS1 repression on the oncogenic behaviors of OS cells. Knockdown of DSCAM-AS1 suppressed NPC tumor growth in vivo. All findings uncovered that DSCAM-AS1 aggravated OS progression through sponging miR-186-5p to up-regulate GPRC5A expression. Thus, we proposed DSCAM-AS1 as a probable target for OS treatment.
Introduction
As one of the most common and aggressive bone cancers, osteosarcoma (OS) usually occurs in adolescents or children [1, 2]. Although great improvements in therapies, the outcomes of most recurrent or metastatic OS patients are still poor [3, 4]. Therefore, molecular regulatory mechanisms involved in OS remain an urgent need to be identified for opening new biomarkers.
Long noncoding RNAs (lncRNAs) are a subgroup of noncoding RNAs, possessing over 200 nt, without the ability to encode proteins [5, 6]. More and more evidence demonstrated that lncRNAs taken part in almost all physiological processes, including tumorigenesis. For example, lncRNA FEZF1-AS1 is prognosis-indicator and promotes tumorigenesis in gastric cancer by Wnt pathway [7]. Also, the participation of lncRNAs in OS has already been discovered. Wang et al. unveiled lncRNA DANCR as a ceRNA of ROCK1 to accelerate cell proliferation and metastasis in OS [8]. Ba et al. elucidated that lncRNA CASC2 regulates miR-181a to exert a tumor-inhibiting function in OS [9]. Shen et al. also suggested lncRNA MEG3 as a negative regulator of OS development by targeting miR-361-5p and FoxM1 [10]. As a new lncRNA, DSCAM-AS1 has also been unveiled to participate in cancer progression. For instance, DSCAM-AS1 modulates BCL11A to play a facilitating part in non-small cell lung cancer [13]. DSCAM-AS1 accelerates tumorigenesis in breast cancer by regulating miR-204-5p and RRM2 [14]. DSCAM-AS1 contributes to malignant phenotypes in ovarian cancer via targeting SOX4 [15]. Nevertheless, the role of DSCAM-AS1 is poorly understood in OS.
The purpose of present work was to study the function of DSCAM-AS1 in OS. To sum up we found that DSCAM-AS1 facilitated cell proliferation and migration in OS through modulating miR-186-5p/GPRC5A signaling. This may provide a promising target for OS treatment.
Materials and methods
Cell culture and transfection
OS cell lines (MG63, U2OS, SAOS2 and HOS) and human normal osteoblasts (hFOB1.19) were acquired from Cell Bank of the Chinese Academy of Sciences (Shanghai, China). Cells were maintained with Dulbecco’s modified eagle’s medium (DMEM; Corning Life Sciences, Tewksbury, MA, USA) containing 10% fetal bovine serum (FBS; Gibco, Gaithersburg, MD, USA) at 37
Quantitative real-time PCR (RT-qPCR)
Total RNA was extracted with TRIzol reagent, which was obtained from Invitrogen. The total RNA was reversely transcribed to cDNA with the Reverse Transcription kit (TaKaRa, Shiga, Japan) based on the manufacturer’s suggestions. RT-qPCR was conducted by utilizing SYBR Select Master Mix (Applied Biosystems, Foster City, CA, USA) with the ABI7300 system (Applied Biosystems). Relative expression level was calculated by 2
CCK-8 assay
MG63 and U2OS cells (1
Colony formation assay
MG63 or U2OS cells were added into 6-well plates. After growing for 2 weeks, each well was rinsed by phosphate buffer saline (PBS; Sigma-Aldrich, St. Louis, MO, USA). Upon that, colonies were fixated by 4% paraformaldehyde (PFA; Sigma-Aldrich) and dyed in 0.5% crystal violet (Sigma-Aldrich). Colonies covering over 50 cells were eventually calculated manually.
Nucleo-cytoplasmic separation
The PARIS
Microarray analysis
Differential expression miRNAs were revealed via Affymetrix Human Genome U133 plus 2.0 Array (Thermo Fisher Scientific). R software was used to distinguish miRNAs with significant differences in expression on microarrays. Differentially expressed miRNAs were identified with discriminating parameters (fold change
Western blot
MG63 or U2OS cells were lysed with RIPA lysis buffer (Thermo Fisher Scientific). The bicinchoninic acid assay (BCA) (Thermo Fisher Scientific) was then adopted to examine protein concentration. Proteins were subjected to separation via sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE; Millipore, Bedford, MA, USA), followed by transferring into the polyvinylidene difluoride (PVDF) membranes (Millipore). Afterwards, membranes were processed with 5% nonfat milk and sequentially co-cultured overnight with primary antibodies (all from Acbam, Cambridge, USA) including anti-E-cadherin (ab76055), anti-N-cadherin (ab76057), anti-GPRC5A (ab155557) and anti-GAPDH (ab8245) at 4
Cell migration assay
Transwell chambers containing the pore size of 8
RNA pull-down assay
DSCAM-AS1 probes were synthesized by Thermo Fisher Scientific. The biotinylated probes were cultivated overnight with cell lysates, after which the streptavidin-coated magnetic beads (Invitrogen) were added. The probes with no biotin labelled were used as the negative control. Finally, RT-qPCR was applied to assess the enrichment of miRNAs in each group.
Luciferase reporter assay
DSCAM-AS1-Wt/Mut or GPRC5A-Wt/Mut was sub-cloned into the pmirGLO dual-luciferase plasmid (Promega, Madison, WI, USA) for forming pmirGLO-DSCAM-AS1-Wt/Mut or pmirGLO-GPRC5A-Wt/Mut. The pmirGLO-DSCAM-AS1-Wt/Mut was co- transfected into MG63 or U2OS cells with miR-186-5p mimics or NC mimics. The pmirGLO-GPRC5A-Wt/Mut was co-transfected into MG63 or U2OS cells with miR-186-5p mimics, miR-186-5p mimics+ pcDNA3.1/DSCAM-AS1 or NC mimics. Upon 48 h of co-transfection, the relative luciferase activity was examined via dual-luciferase reporter assay system (Promega).
RNA immunoprecipitation (RIP) assay
A Magna RIPTM RNA Binding Protein Immunoprecipitation Kit (Millipore) was employed to conduct RIP assay. Briefly, cell lysates were processed with RIP buffer (Invitrogen) containing magnetic beads that were coated with anti-Ago2 antibody (Millipore) or negative control anti-IgG (Millipore). RNAs in precipitated compounds were determined with RT-qPCR.
Nude mouse xenograft model
BALB/c nude mice (six-week-old) were bought from the Laboratory Animal Center of Nanjing Medical University, followed by raising under pathogen-free conditions. Were with MG63 cells with or without DSCAM-AS1 silence were injected subcutaneously into the groin of each mouse. The volume of tumors was recorded every four days. Four weeks later, mice were euthanized and the weight was measured after tumors were excised.
Statistical analysis
Statistical analysis was conducted by use of GraphPad Prism 5 software (GraphPad Software, La Jolla, CA, USA). Data were denoted as mean
DSCAM-AS1 was upregulated in OS cell lines, and knockdown of DSCAM-AS1 suppressed OS tumorigenesis. (A) RT-qPCR assay detected the overexpressed DSCAM-AS1 in OS cells compared to normal hFOB1.19 cells. (B) RT-qPCR validated the efficient knockdown of DSCAM-AS1 by sh-DSCAM-AS1#1/2 in both MG63 and U2OS cells. (C) and (D) CCK-8 and colony formation assays were performed to detect cell proliferation in two OS cells with or without DSCAM-AS1 silence. (E) Transwell assays were performed to detect cell migration in OS cells with DSCAM-AS1 silence or not. (F) Western blot assay measured the expression of EMT related proteins (E-cadherin and N-cadherin) in OS cells under DSCAM-AS1 inhibition. 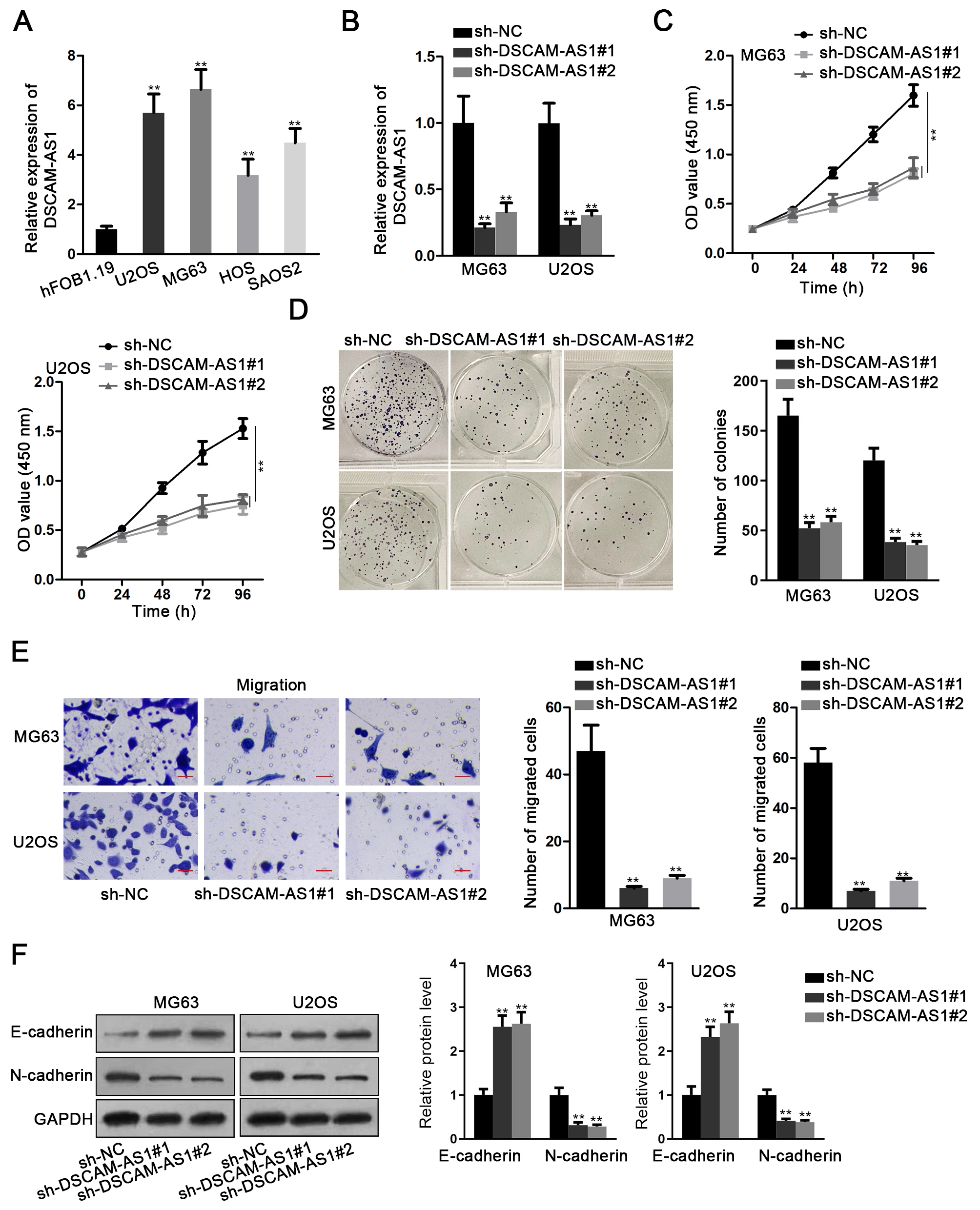
DSCAM-AS1 acted as a sponge for miR-186-5p in OS. (A) Nuclear-cytoplasmic extraction assay was used to confirm the location of DSCAM-AS1 in MG63 and U2OS cells. (B) Microarray analysis was used to search differentially-expressed miRNAs in U2OS cells after DSCAM-AS1 was deleted, and 5 miRNAs (miR-186-5p, miR-150-5p, miR-650, miR-877-5p and miR-3126-5p) presented upregulation with most significance. (C) RNA pull down assay was applied to verify the interaction of DSCAM-AS1 with above 5 miRNAs in MG63 and U2OS cells. (D) RT-qPCR analyzed the expression of miR-186-5p in OS cells relative to normal hFOB1.19 cells. (E) RT-qPCR detected the overexpression efficiency of miR-186-5p in OS cells after transfecting with miR-186-5p mimics. (F) RT-qPCR assessed the change of DSCAM-AS1 and miR-186-5p expression in different transfected OS cells. (G) The binding sites between DSCAM-AS1 and miR-186-5p were predicted by starBase. (H) Luciferase reporter assay was conducted to verify the binding ability between DSCAM-AS1 and miR-186-5p in two OS cells. (I) and (J) CCK-8 and colony formation assays were performed to detect cell proliferation in OS cells with DSCAM-AS1 depletion or together with miR-186-5p inhibition. (K) Transwell assays were performed to detect the migration of cells under above conditions. (L) Western blot measured the expression of E-cadherin and N-cadherin in indicated OS cells. 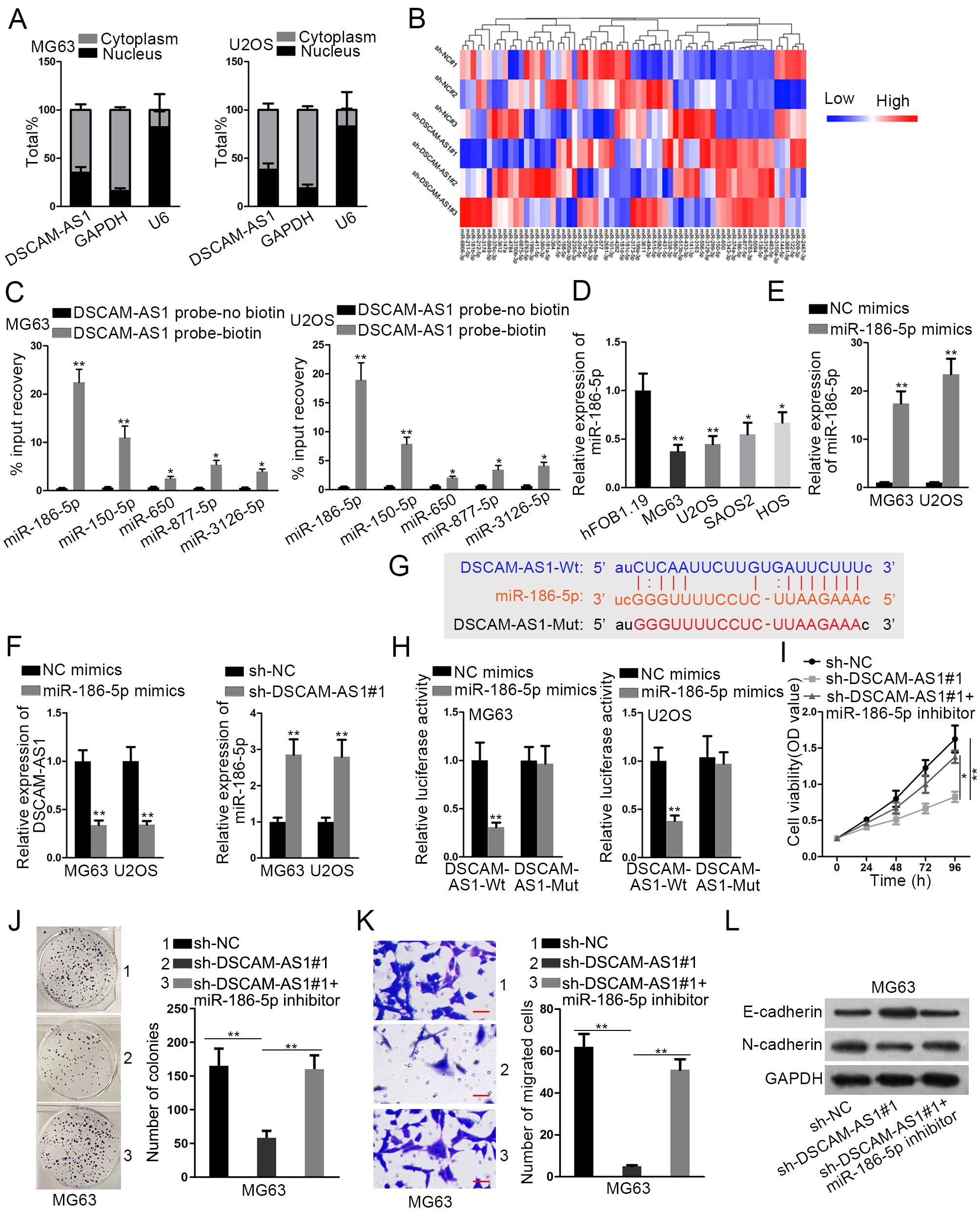
GPRC5A directly interacted with miR-186-5p. (A) The mRNAs bound with miR-186-5p were predicted by three tools (microT, miRanda and RNA22) in starBase and then the results were showed in Venn diagram. (B) RT-qPCR analysis showed the changes on the expression of 5 mRNAs in OS cells after enhancing miR-186-5p level. (C) RT-qPCR and western blot were performed to detect the expression of GPRC5A in OS cells relative to hFOB1.19 cells. (D) RT-qPCR and western blot were performed to detect GPRC5A expression in OS cells with miR-186-5p upregulation or DSCAM-AS1 deficiency. (E) The binding sites between GPRC5A and miR-186-5p were obtained from starBase. (F) Luciferase reporter assay was conducted to verify the relationship among DSCAM-AS1, miR-186-5p and GPRC5A in OS cells. (G) RIP assay verified the co-existence of DSCAM-AS1, miR-186-5p and GPRC5A in RISCs in both MG63 and U2OS cells. 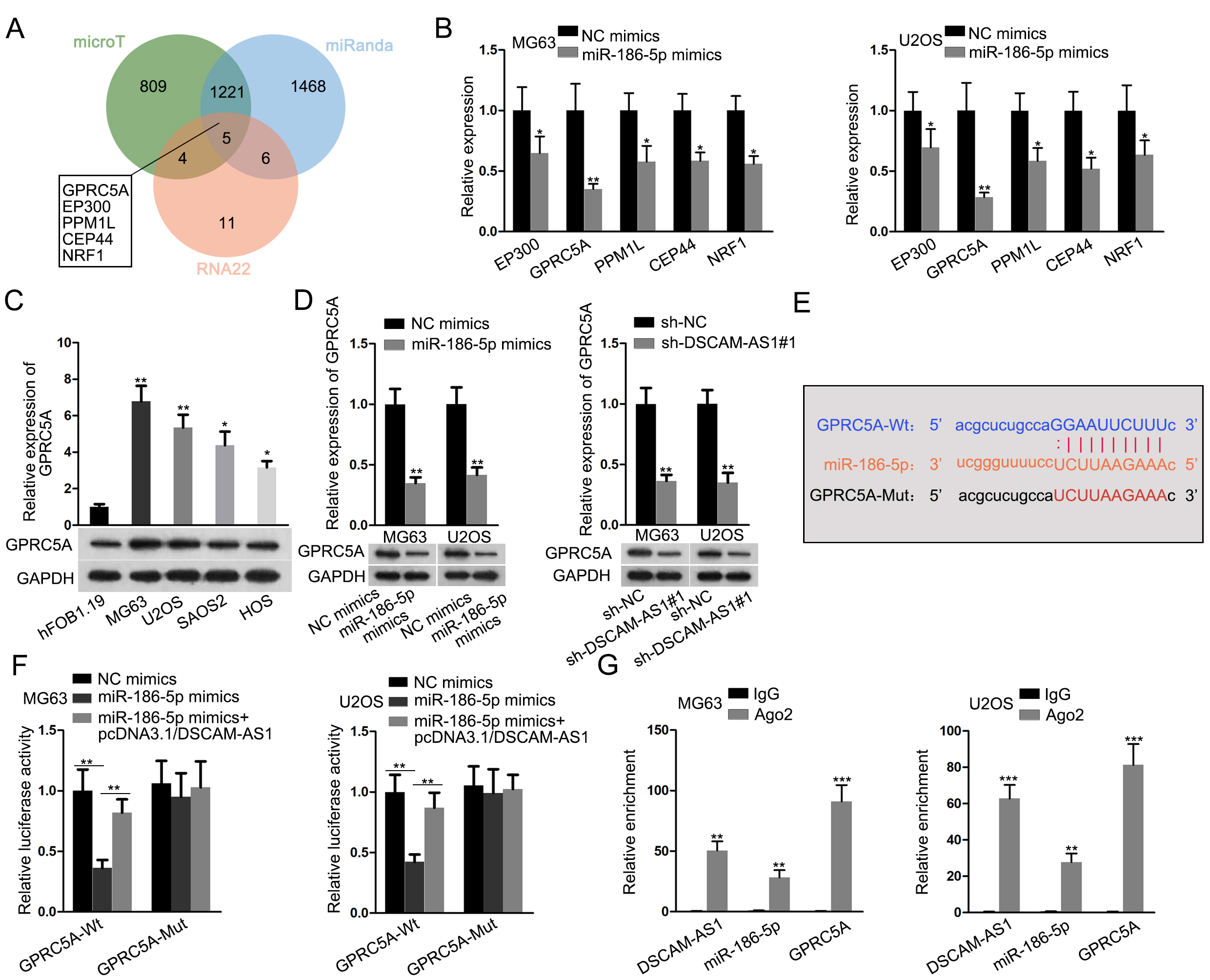
DSCAM-AS1 promoted OS progression by the regulation of GPRC5A. (A) RT-qPCR and western blot analyses were used to detect the overexpression efficiency of GPRC5A in MG63 cells after transfecting with pcDNA3.1/GPRC5A. (B) and (C) CCK-8 and colony formation assays were performed to detect the proliferation of MG63 cells under following transfections: sh-NC, sh-DSCAM-AS1#1, and sh-DSCAM-AS1#1+pcDNA3.1/GPRC5A. (D) Transwell assays were performed to detect cell migration under above contexts. (E) Western blot determined the expression of E-cadherin and N-cadherin in above-mentioned MG63 cells. (F)–(H) Representative images, the growth curve and weight of tumors originated from MG63 cells transfected with sh-NC or sh-DSCAM-AS1#1. (I) The expression of DSCAM-AS1 in the tumor xenografts obtained from above two groups was examined by RT-qPCR. (J) Western blot examined the protein levels of E-cadherin, N-cadherin and GPRC5A in tumors from sh-NC or sh-DSCAM-AS1#1 group. 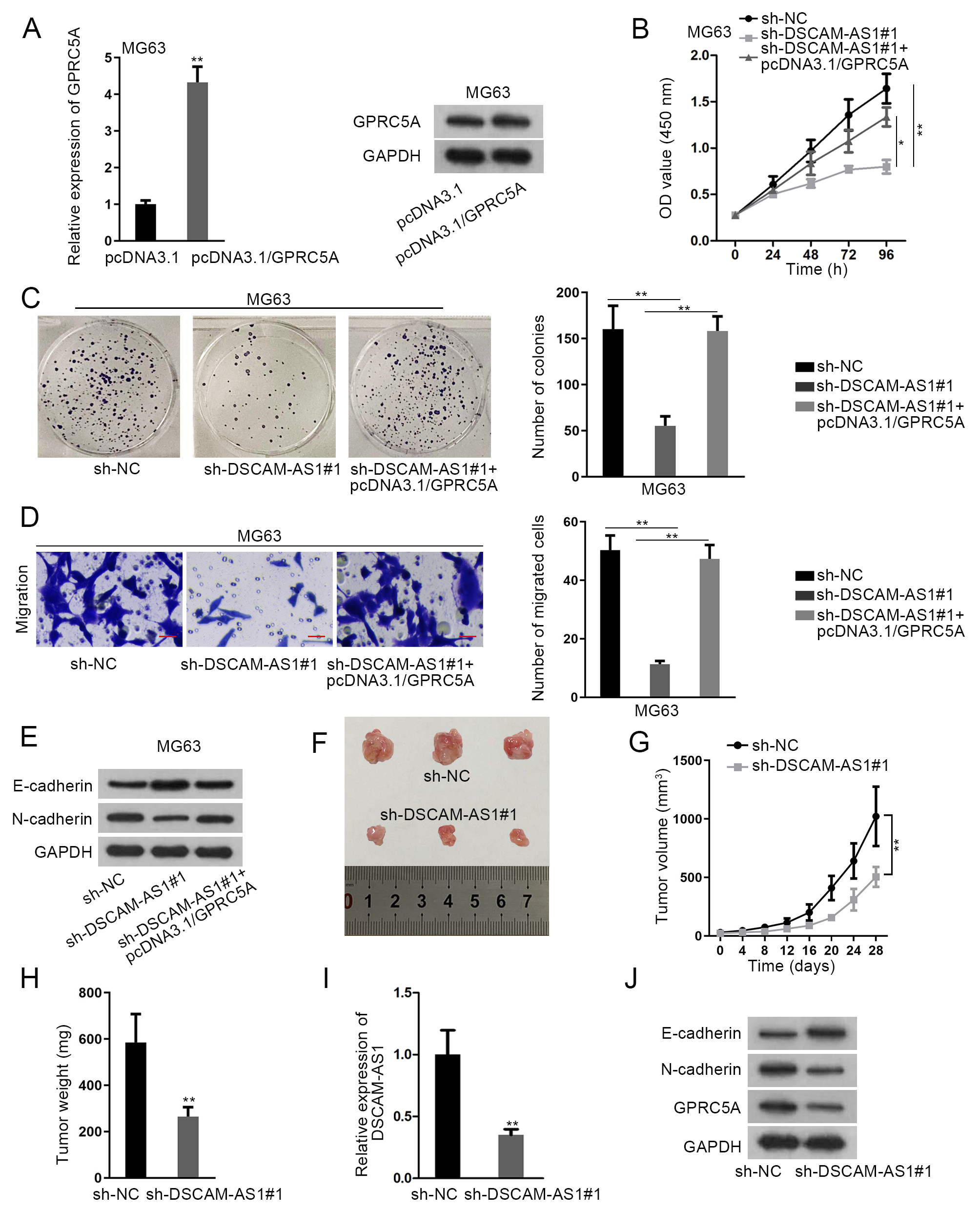
LncRNA DSCAM-AS1 was upregulated in OS cell lines, and silencing DSCAM-AS1 blocked OS progression
To probe the possible function of DSCAM-AS1 in OS carcinogenesis, its expression was firstly measured by RT-qPCR in OS cell lines (MG63, U2OS, SAOS2 and HOS). Results showed that DSCAM-AS1 expression was apparently heightened in OS cells in comparison to that in the normal hFOB1.19 cells (Fig. 1A). As shown in Fig. 1B, it was manifested that in contrast to the sh-NC group DSCAM-AS1 level was dramatically lessened in cells with sh-DSCAM-AS1#1/2. Moreover, downregulating DSCAM-AS1 prominently hampered MG63 and U2OS cell proliferation (Fig. 1C and D). Meanwhile, DSCAM-AS1 knockdown caused a dramatic reduction in OS cell migration ability (Fig. 1E). Western blot analysis demonstrated that DSCAM-AS1 inhibition obviously led to augmented E-cadherin expression and lowered N-cadherin level (Fig. 1F). In brief DSCAM-AS1 up-regulated in OS accelerated cell malignancy.
MiR-186-5p was sponged by DSCAM-AS1 in OS
Nuclear and cytosolic separation assay indicated that DSCAM-AS1 was primarily probed in the cytoplasm of both MG63 and U2OS cells (Fig. 2A). Consistently, FISH assay also presented the main localization of DSCAM-AS1 in cytoplasm (Fig. S1A). Thus, we speculated that FBXL19-AS1 might operate in OS as a ceRNA. Through starBase tool (
GPRC5A directly interacted with miR-186-5p
As shown in Venn diagram, there were five mRNAs possibly targeted by miR-186-5p after searching from microT, miRanda and RNA22 biological prediction websites (Fig. 3A). Further, it manifested that among these five candidates, GPRC5A presented the biggest drop on expression in both two OS cells with elevated miR-186-5p (Fig. 3B). GPRC5A expression was up-regulated in OS cells relative to normal controls (Figs 3C and S1D). Moreover, we proved that miR-186-5p overexpression and DSCAM-AS1 knockdown could remarkably decline the expression of GPRC5A mRNA and protein in both MG63 and U2OS cells (Figs 3D and S1E). The binding sequences between miR-186-5p and GPRC5A were exhibited in Fig. 3E. Further, the luciferase activity of GPRC5A-Wt reporters was unveiled to be impaired in response to miR-186-5p mimics and such impairment was then counteracted by DSCAM-AS1 overexpression (Fig. 3F). Furthermore, DSCAM-AS1, miR-186-5p and GPRC5A were all significantly harvested in anti-Ago2-precipitated complexes (Fig. 3G), indicating their co-existing in RNA-induced silencing complexes (RISCs). Altogether, DSCAM-AS1 was a ceRNA of GPRC5A via its competitive interaction with miR-186-5p.
DSCAM-AS1 facilitated OS development through regulating GPRC5A
Subsequently, we intended to verify whether GPRC5A was implicated in DSCAM-AS1-regulated OS development. Hence, rescue assays were carried out after confirming the efficient upregulation of GPRC5A by pcDNA3.1/GPRC5A in MG63 cells (Figs 4A and S1F). Then, we discovered that the reduced proliferation ability of DSCAM-AS1-inhibited cells was saved by overexpressing GPRC5A (Fig. 4B and C). Also, the decreased cell migration in DSCAM-AS1-depleted MG63 cells was reversed by GPRC5A up-regulation (Fig. 4D). Similarly, the influences of DSCAM-AS1 knockdown on E-cadherin and N-cadherin expression were offset under GPRC5A enhancement (Figs 4E and S1G). Meanwhile, we also characterized the function of DSCAM-AS1 in vivo. Results presented that the tumor size growth rate and weight were all declined in sh-DSCAM-AS1#1 group in comparison to that in sh-NC group (Fig. 4F–H). Additionally, the expression of DSCAM-AS1 was markedly decreased in tumors from sh-DSCAM-AS1#1 group (Fig. 4I). Furthermore, it was indicated that tumors in sh-DSCAM-AS1#1 group expressed higher E-cadherin along with lower N-cadherin and GPRC5A levels than those in control group (Figs 4J and S1H). In sum, DSCAM-AS1 promoted OS progression by targeting GPRC5A.
Discussion
LncRNAs have been considered to play vital parts in cancers, including colorectal cancer [16] cervical cancer [17], liver cancer [18] breast cancer [19], and so on. There are also a lot of researches linking lncRNAs with OS progression. LncRNA SNHG16 contributes to OS cell motility via absorbing miRNA-340 [20]. LncRNA AFAP1-AS1 facilitates the malignancy in OS by targeting miR-4695-5p/TCF4-
Multiple literatures revealed that lncRNAs can absorb miRNAs to release mRNAs, resulting in accelerated cancer progression [16, 25, 26]. MiR-186-5p has been regarded as an inhibitor in diverse cancer types MiR-186-5p upregulation inhibits malignant behaviors of colorectal cancer cells through modulating ZEB1 [27]. CircRNA circPDSS1 accelerates development in gastric cancer via modulating miR-186-5p andNEK2 [28]. HOXD-AS1 accelerates EMT in ovarian cancer through regulation of miR-186-5p and PIK3R3 [29]. MiR-186-5p has also been found to participate in OS development through its repression on FOXK1 [30]. Subsequently, we unveiled that DSCAM-AS1 sponged miR-186-5p in OS. Further, suppressing miR-186-5p countervailed the function of DSCAM-AS1 silence in OS cells G Protein-coupled receptor class C group 5 member A (GPRC5A) has been characterized as a contributor in many malignancies. GPRC5A facilitates cell growth and relates to bone metastasis in prostate cancer [31]. CircGprc5a contributes to the progression of bladder cancer via Gprc5a-targeting peptide [32]. Elevated GPRC5A in colorectal cancer aggravates tumor progression through modulating VNN-1 induced oxidative stress [33]. Nevertheless, GPRC5A has not been studied in OS. In present study, GPRC5A was discovered to be targeted by miR-186-5p in OS Besides, DSCAM-AS1 could positively regulate GPRC5A expression through absorbing miR-186-5p. Finally, we proved that up-regulating GPRC5A reversed the function of DSCAM-AS1 repression in OS cellular processes. More importantly, in vivo experiments also indicated that tumors with silenced DSCAM-AS1 possessed decreased GPRC5A levels and slowed growth rates.
In conclusion, DSCAM-AS1 accelerated OS development through sequestering miR-186-5p to enhance GPRC5A expression. This may indicate DSCAM-AS1 as a promising target for OS treatment.
Footnotes
Acknowledgments
We appreciate all contributors involved in this study.
Conflict of interest
None.
Supplementary data
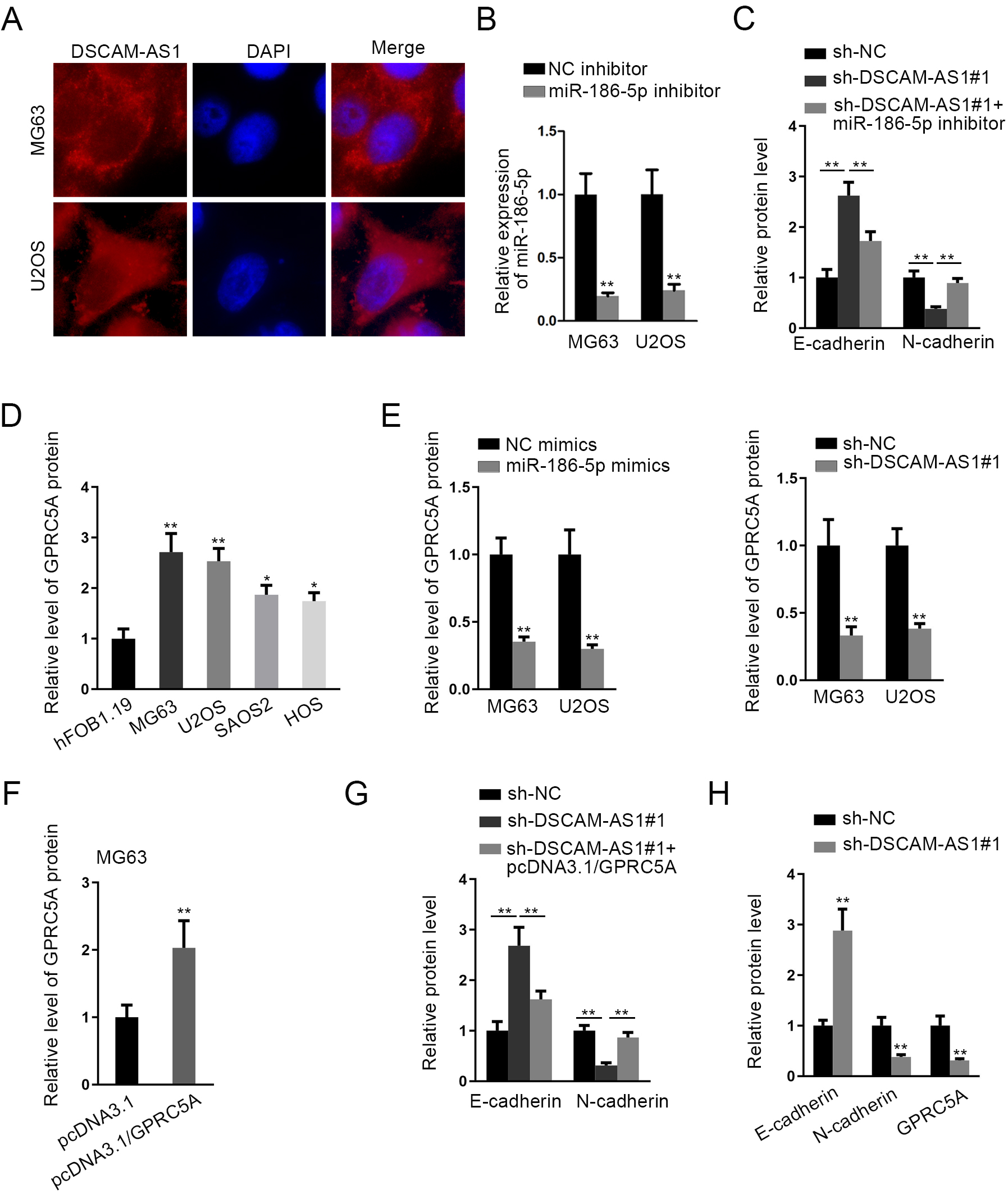
(A) FISH assay determined the cellular distribution of DSCAM-AS1 in MG63 and U2OS cells. (B) RT-qPCR evaluated the expression of miR-186-5p in MG63 and U2OS cells transfected with NC inhibitor or miR-186-5p inhibitor. (C) Quantitative histogram of E-cadherin and N-cadherin protein bands assessed by western blot in Fig. 2L. (D) and (E) The quantification of indicated bands determined by western blot in Fig. 3C and D. (F)–(H) The quantification of indicated bands determined by western blot in Fig. 4A, E and J.