Abstract
BACKGROUND:
Head and neck squamous cell carcinoma (HNSCC) is the seventh most common type of cancer around the world. The aim of this study was to seek the long non-coding RNAs (lncRNAs) acting as diagnostic and prognostic biomarker of HNSCC.
METHODS:
Base on TCGA dataset, the differentially expressed mRNAs (DEmRNAs) and lncRNAs (DElncRNAs) were identified between HNSCC and normal tissue. The machine learning and survival analysis were performed to estimate the potential diagnostic and prognostic value of lncRNAs for HNSCC. We also build the co-expression network and functional annotation. The expression of selected candidate mRNAs and lncRNAs were validated by Quantitative real time polymerase chain reaction (qRT-PCR).
RESULTS:
A total of 3363 DEmRNAs (1822 down-regulated and 1541 up-regulated mRNAs) and 32 DElncRNAs (13 down-regulated and 19 up-regulated lncRNAs) between HNSCC and normal tissue were obtained. A total of 13 lncRNAs (IL12A.AS1, RP11.159F24.6, RP11.863P13.3, LINC00941, FOXCUT, RNF144A.AS1, RP11.218E20.3, HCG22, HAGLROS, LINC01615, RP11.351J23.1, AC024592.9 and MIR9.3HG) were defined as optimal diagnostic lncRNAs biomarkers for HNSCC. The area under curve (AUC) of the support vector machine (SVM) model, decision tree model and random forests model and were 0.983, 0.842 and 0.983, and the specificity and sensitivity of the three model were 95.5% and 96.2%, 77.3% and 97.6% and 93.2% and 97.8%, respectively. Among them, AC024592.9, LINC00941, LINC01615 and MIR9-3HG was not only an optimal diagnostic lncRNAs biomarkers, but also related to survival time. The focal adhesion, ECM-receptor interaction, pathways in cancer and cytokine-cytokine receptor interaction were four significantly enriched pathways in DEmRNAs co-expressed with the identified optimal diagnostic lncRNAs. But for most of the selected DEmRNAs and DElncRNAs, the expression was consistent with our integrated analysis results, including LINC00941, LINC01615, FOXCUT, TGA6 and MMP13.
CONCLUSION:
AC024592.9, LINC00941, LINC01615 and MIR9-3HG was not only an optimal diagnostic lncRNAs biomarkers, but also were a prognostic lncRNAs biomarkers.
Introduction
Head and neck squamous cell carcinoma (HNSCC) is one of the most malignant tumors, ranks among the seventh most common type of cancer worldwide [1]. In spite of considerable advances in surgery, radiotherapy, chemotherapy and multimodality therapy, prognosis of HNSCC remains poor, and the 5-year overall survival is
With advances in gene expression profiles, lncRNAs are reported to be as pivotal regulators involved in various biological processes in cancer, including tumor occurrence, development and metastasis [3, 4]. Recently, many lncRNAs are considered to be novel candidate biomarkers for the diagnosis and prediction of various cancers [5, 6, 7]. However, there are few study on lncRNA diagnostic and prognostic biomarkers for HNSCC is rare. Therefore, the identification of key tumor markers of HNSCC is required.
The aim of this study is to obtain biomarkers of lncRNAs for diagnostic and prognostic HNSCC patients, we used the TCGA datasets to obtain the differentially expressed mRNAs (DEmRNAs) and lncRNAs (DElncRNAs). We performed the machine learning and survival analysis to evaluate the potential diagnostic and prognostic value of lncRNAs for HNSCC. The DElncRNA-DEmRNA co-expression network and the functional annotation of the DEmRNAs co-expressed with the identified optimal diagnostic lncRNAs in HNSCC was performed. The expression levels of candidate genes were verified by qRT-PCR. To our knowledge, this is first time to find diagnostic and prognostic lncRNAs biomarkers in HNSCC by using machine learning.
Materials and methods
TCGA data
The lncRNA and mRNA gene expression profiles and clinical data of HNSCC were download by the Cancer Genome Atlas (TCGA) (
Identification of DEmRNAs and DElncRNAs
The undetectable lncRNAs and mRNAs (with read count value
Identification of the optimal diagnostic lncRNAs biomarkers for HNSCC
To further identify the optimal diagnostic lncRNA biomarkers for HNSCC, we performed feature selection procedures as follows. (1) The importance value of each lncRNA was ranked according to mean decrease in accuracy from large to small by random forest algorithm. (2) The optimum number of features was found by adding a DElncRNA at a time in the top down forward-wrapper packaging method. (3) By using support vector machine (SVM) at each increment and the optimal diagnostic lncRNA biomarkers were identified for HNSCC.
The ‘random Forests’ packet (
Survival analysis of optimal diagnostic lncRNAs biomarkers for HNSCC
To determine the potential association between the identified DElncRNAs and survival in HNSCC patients, survival analysis (
DEmRNAs co-expressed with the identified optimal diagnostic lncRNAs
The correlation between the optimal diagnostic lncRNAs and DEmRNAs were analyzed by the Pearson correlation coefficient. The threshold for DElncRNA-DEmRNA co-expression pairs was
Functional annotation of DEmRNAs co-expressed with the identified optimal diagnostic lncRNAs
Gene Ontology (GO) classification and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis were performed using GeneCoDis (
Primer sequences used for qRT-PCR
Primer sequences used for qRT-PCR
Base on the results of TCGA integration analysis, ITGA6, MMP13 and FOXC1, LINC00941, LINC01615 and FOXCUT were screened as candidate mRNAs and lncRNAs. A total of 6 HNSCC patients were enrolled in this study. Twelve tissues samples of HNSCC patients (
Total RNA was extracted from samples with TRIzol reagent (Tiangen, China) and then subjected to reverse transcription and PCR reactions using Super Real PreMix Plus SYBR Green (Tiangen, China). The 2-
Top 40 DEmRNAs
Top 40 DEmRNAs
All of DElncRNAs
Hierarchical clustering analysis of DElncRNAs and top 100 DEmRNAs between HNSCC and normal tissues. A. DEmRNAs. B. DElncRNAs. Row and column represented DElncRNAs/DEmRNAs and tissue samples, respectively. The color scale represented the expression levels.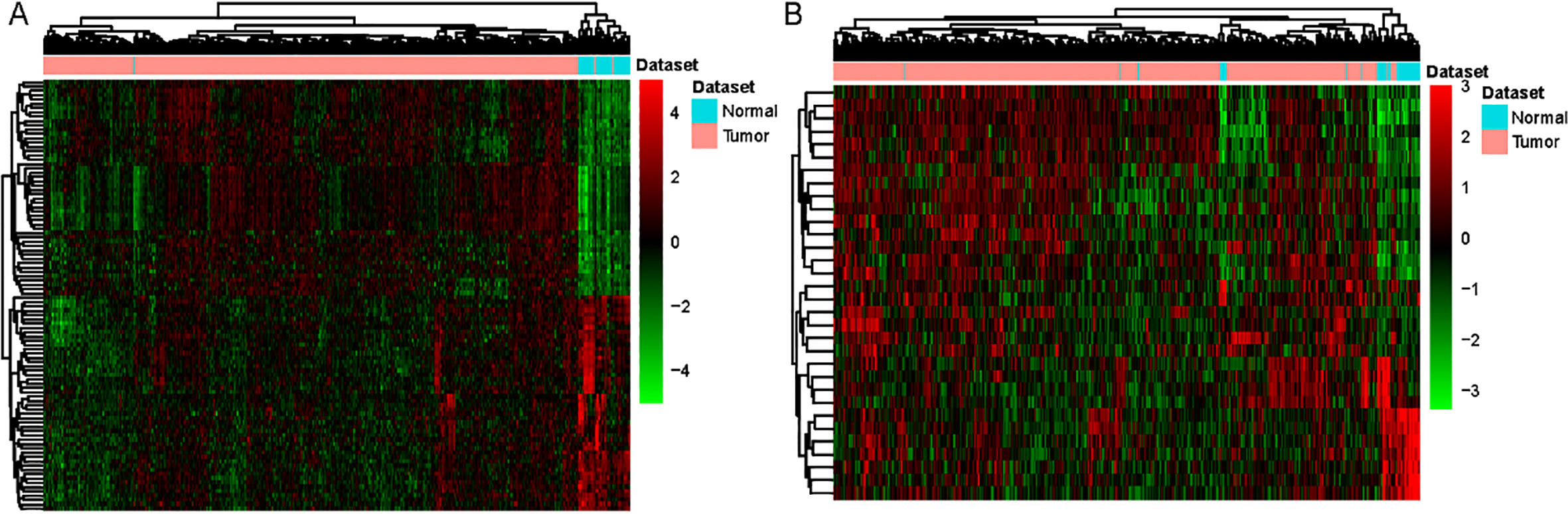
Identification of optimal lncRNA biomarkers for HNSCC. A. The importance value of each DElncRNA ranked according to the mean decrease in accuracy by using the random forest analysis. B. The variance rate of classification performance when increasing numbers of the predictive DElncRNAs. C. Hierarchical clustering analysis of 13 lncRNAs biomarkers.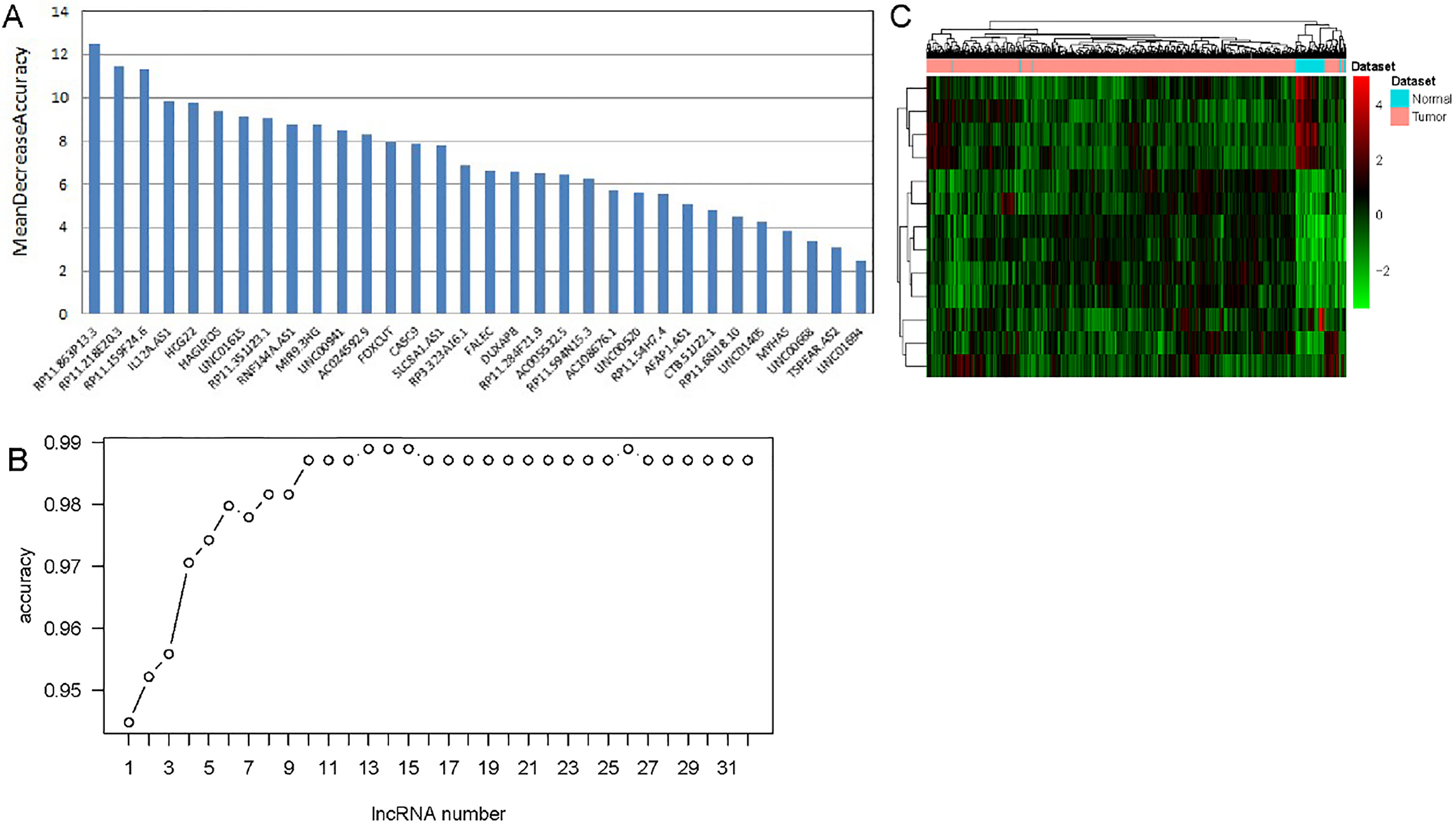
DEmRNAs and DElncRNAs in HNSCC
Base on the DEmRNAs analysis, we found 3363 DEmRNAs in HNSCC patients compared with normal, including 1822 down-regulated mRNAs and 1541 up-regulated mRNAs. Base on the DElncRNAs analysis, we defined 32 DElncRNAs in HNSCC patients compared with normal, including 13 down-regulated lncRNAs and 19 up-regulated lncRNAs. Hierarchical clustering analysis of top 100 DEmRNAs and all of DElncRNAs were illuminated in Fig. 1A and Fig. 1B, respectively. Top 40 DEmRNAs and all of DElncRNAs were listed in Tables 2 and 3, respectively.
Identification of the optimal diagnostic lncRNAs biomarkers for HNSCC
The random forest analysis was used to rank the 32 DElncRNAs according to the mean decrease in accuracy (Fig. 2A). Ten-fold cross-validation result showed that the average accuracy rate of 13 DElncRNAs (IL12A.AS1, RP11.159F24.6, RP11.863P13.3, LINC 00941, FOXCUT, RNF144A.AS1, RP11.218E20.3, HCG22, HAGLROS, LINC01615, RP11.351J23.1, AC024592.9 and MIR9.3HG) reached the higher score for the first time (Fig. 2B). Hierarchical clustering analysis of the 13 DElncRNAs were displayed in Fig. 2C. Thence, these 13 DElncRNAs were identified as the optimal diagnostic lncRNAs biomarkers for HNSCC which were used to build the random forests, decision tree and SVM models. Violin Plot showed the expression levels of these 13 DElncRNAs between HNSCC and normal tissues (Fig. 3). Among which, 5 DElncRNAs (IL12A.AS1, FOXCUT, HCG22, RP11.351J23.1 and AC024592.9) were down-regulated in HNSCC tissues compared with normal, and 8 DElncRNAs (RP11.159F24.6, RP11.863P13.3, LINC00941, RNF144A.AS1, RP11. 218E20.3, HAGLROS, LINC01615 and MIR9.3HG) were up-regulated in HNSCC tissues compared with normal. The AUC of the SVM model was 0.983 and the specificity and sensitivity of this model were 95.5% and 96.2% (Fig. 4A). The AUC of the decision tree model was 0.824 and the specificity and sensitivity of this model were 77.3% and 97.6%, respectively (Fig. 4B). The AUC of the random forests model was 0.983 and the specificity and sensitivity of this model were 93.2% and 97.8%, respectively (Fig. 4C). The ROC of all these 13 lncRNAs (IL12A.AS1, RP11. 159F24.6, RP11.863P13.3, LINC00941, FOXCUT, RNF144A.AS1, RP11.218E20.3, HCG22, HAGLROS, LINC01615, RP11.351J23.1, AC024592.9 and MIR9.3HG) were displayed in Fig. 4D–P. Taken together, the AUC of all these three lncRNAs and their combination were all greater than 0.567 which indicated these 13 DElncRNAs and their combination were related to HNSCC and could predict the development of HNSCC.
Violin Plot displayed the expression levels of 13 lncRNAs biomarkers between HNSCC and normal tissues. The X-axis represented normal and HNSCC groups. The Y-axis represented gene expression levels.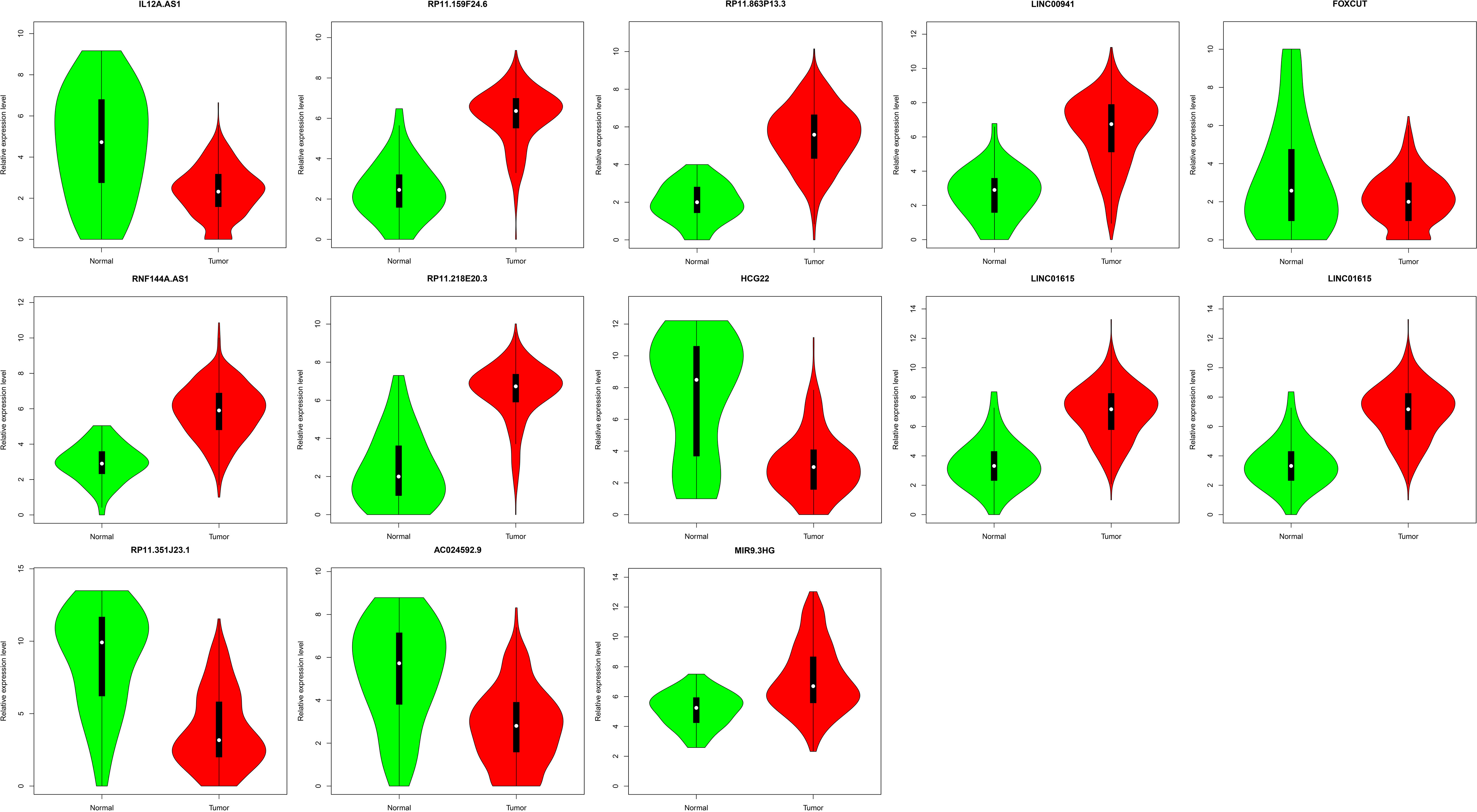
ROC analysis of three HNSCC-specific lncRNAs biomarkers. The x-axis shows 1-specificity and y-axis shows sensitivity.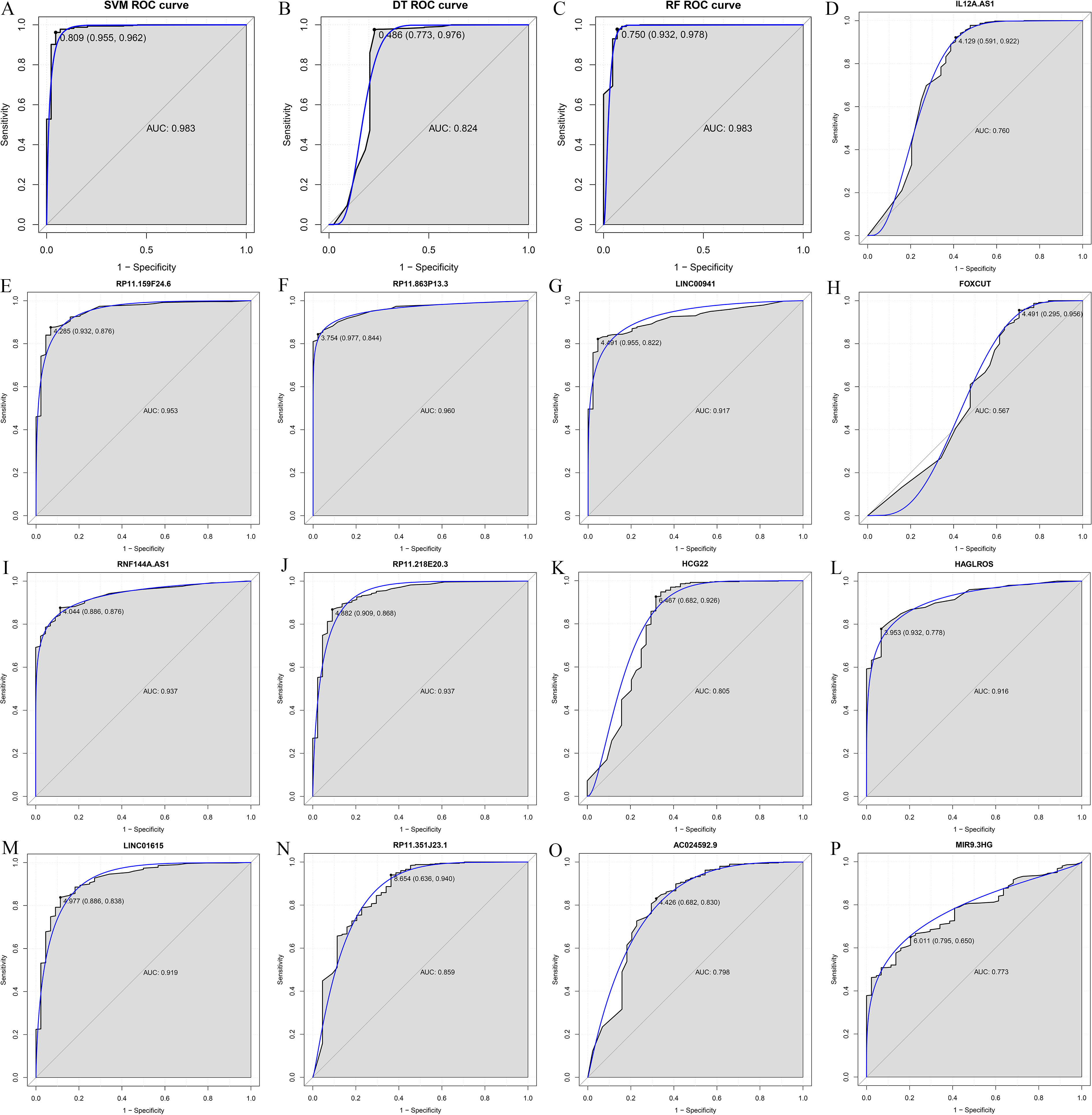
Survival analysis of lncRNAs biomarkers between HNSCC and normal tissues.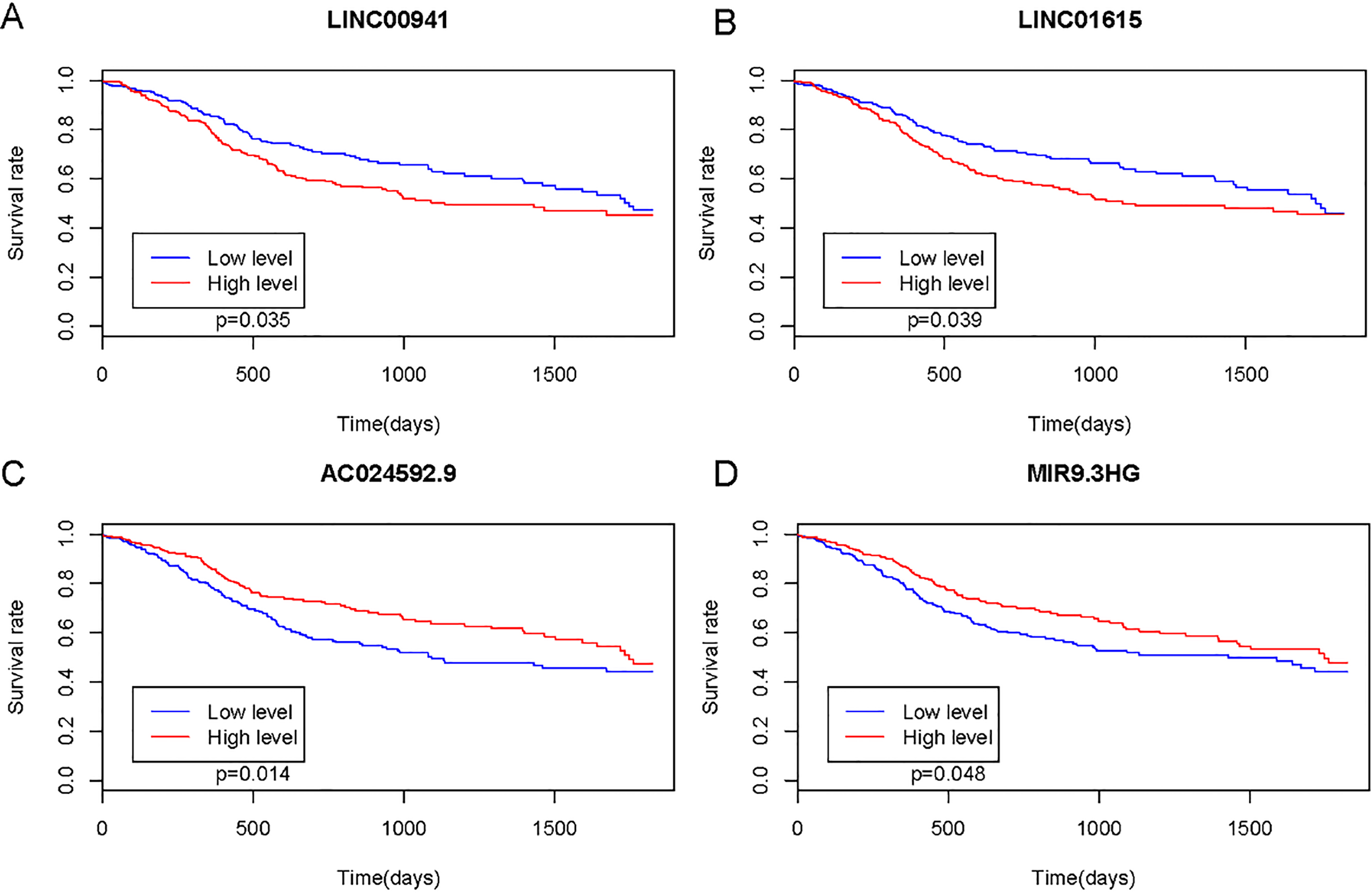
According to the survival analysis, we found AC024592.9, LINC00941, LINC01615 and MIR9-3HG was significantly associated with the prognosis of patients with HNSCC (Fig. 5A–D).
DEmRNAs co-expressed with the identified optimal diagnostic lncRNAs
A total of 13 optimal DElncRNA biomarkers for HNSCC were co-expressed with 610 DEmRNAs, accounting for 1096 DElncRNA-DEmRNA co- expression pairs (Fig. 6). IL12A.AS1, RP11.159F24.6, RP11.863P13.3, LINC00941, FOXCUT, RNF144A. AS1, RP11.218E20.3, HCG22, HAGLROS, LINC 01615, RP11.351J23.1, AC024592.9 and MIR9.3HG were co-expressed with 9, 79, 184, 139, 4, 123, 105, 47, 3, 109, 74, 137 and 83 DEmRNAs, respectively.
lncRNAs-mRNAs co-expression network. The ellipses and rhombuses were represented the mRNAs and lncRNAs, respectively. Green and pink color represented up-regulated and down-regulated, respectively. The black border indicates DEmRNAs and DElncRNAs.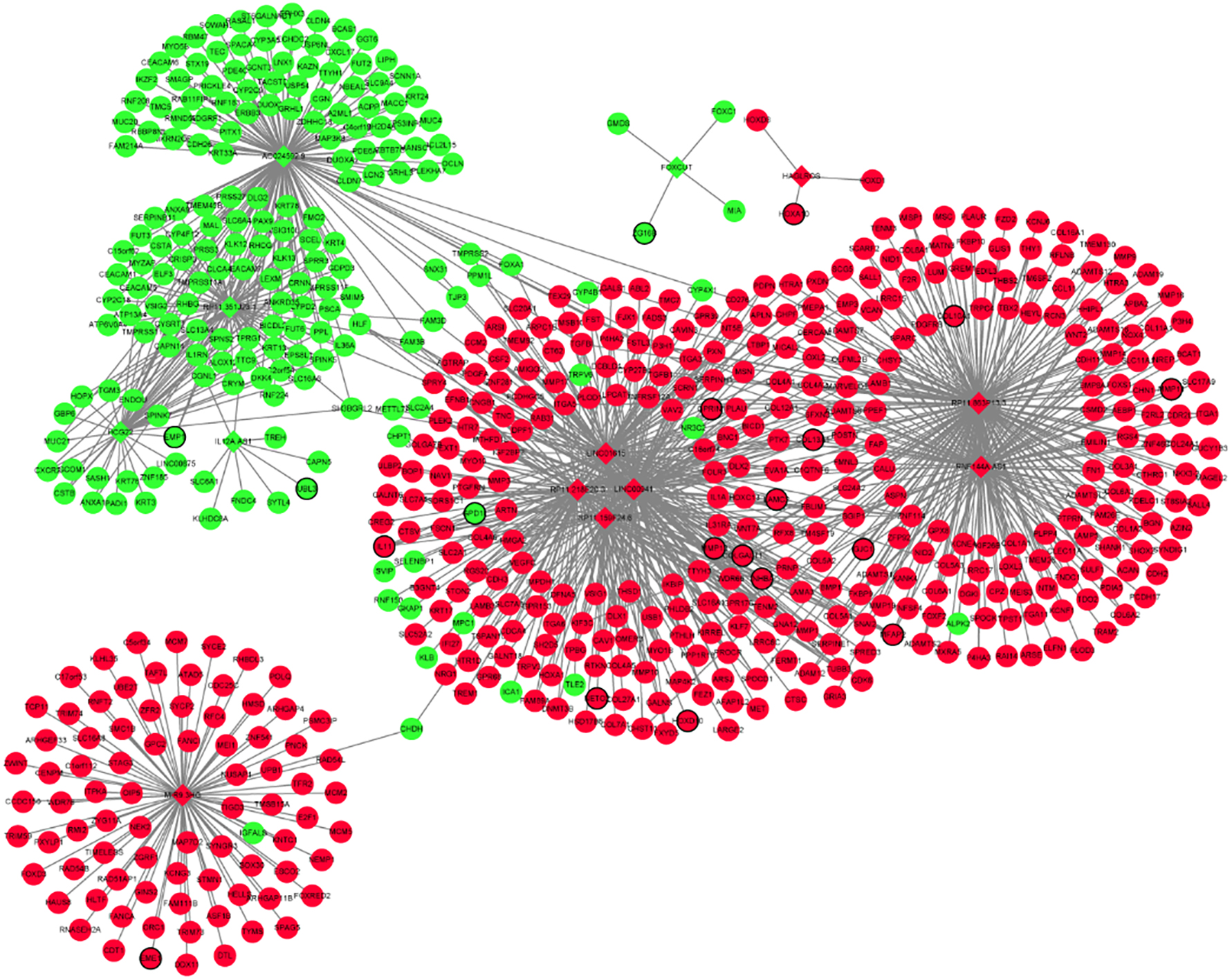
The function annotation of DEmRNAs co-expressed with the identified optimal diagnostic lncRNAs. The x-axis shows -log P and y-axis shows GO terms or KEGG pathways. A. GO terms. B. KEGG pathways.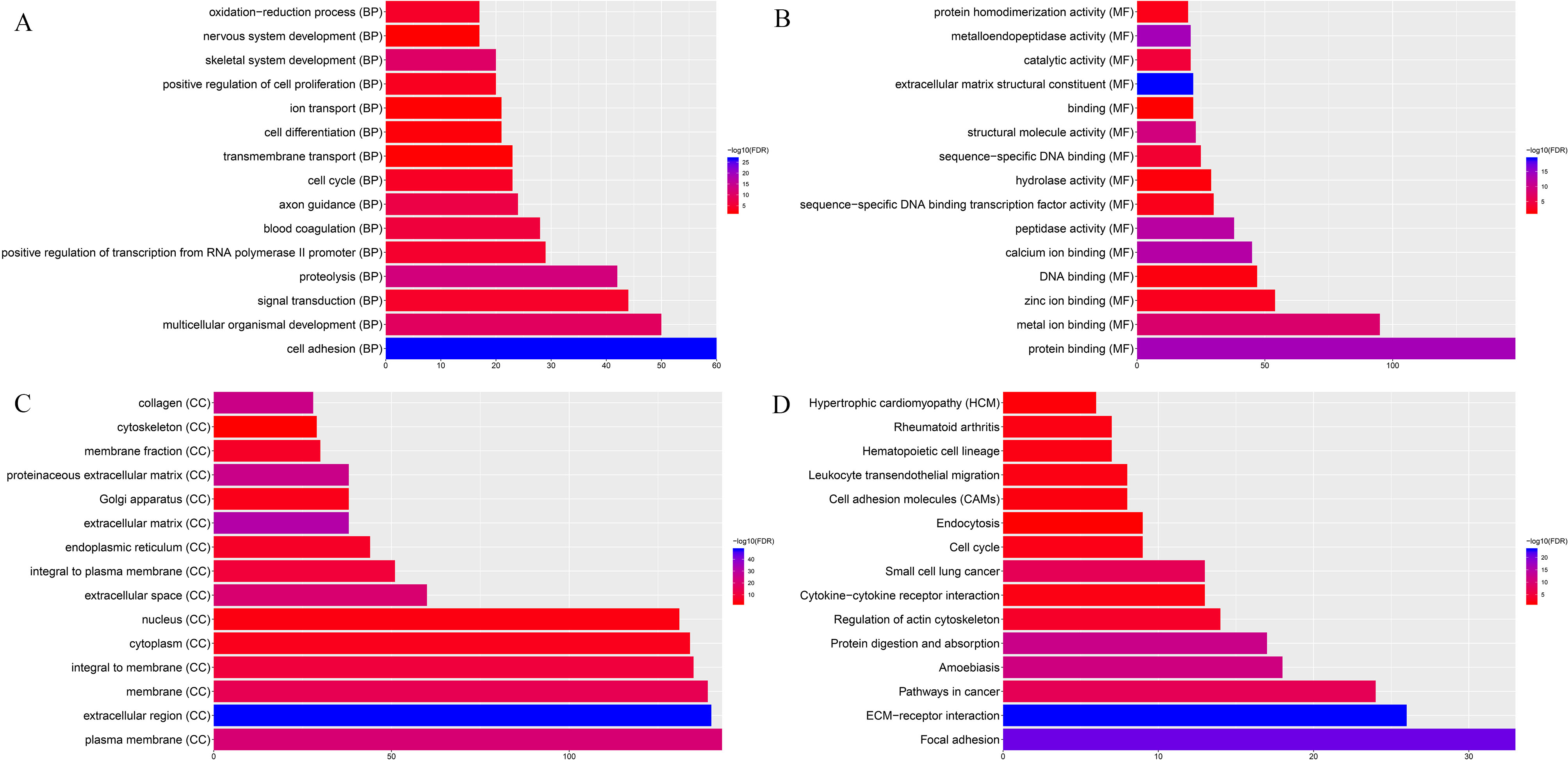
Base on GO enrichment analysis, we found that cell adhesion (FDR
Verification by qRT-PCR
Three DElncRNAs and three DEmRNAs were chose for confirmation. As Fig. 8 displayed, FOXCUT was down-regulated and LINC00941, LINC01615, ITGA6, MMP13 and FOXC1 were up-regulated in HNSCC compared with adjacent tissues. Base on the TCGA results, FOXCUT and FOXC1 were down-regulated while the other lncRNAs and DEmRNAs were up-regulated in HNSCC compared to adjacent tissues. In generally, the confirmation results of qRT-PCR were consistent with our integrated analysis.
Validation optimal lncRNA biomarkers in HNSCC tissue by qRT-PCR. 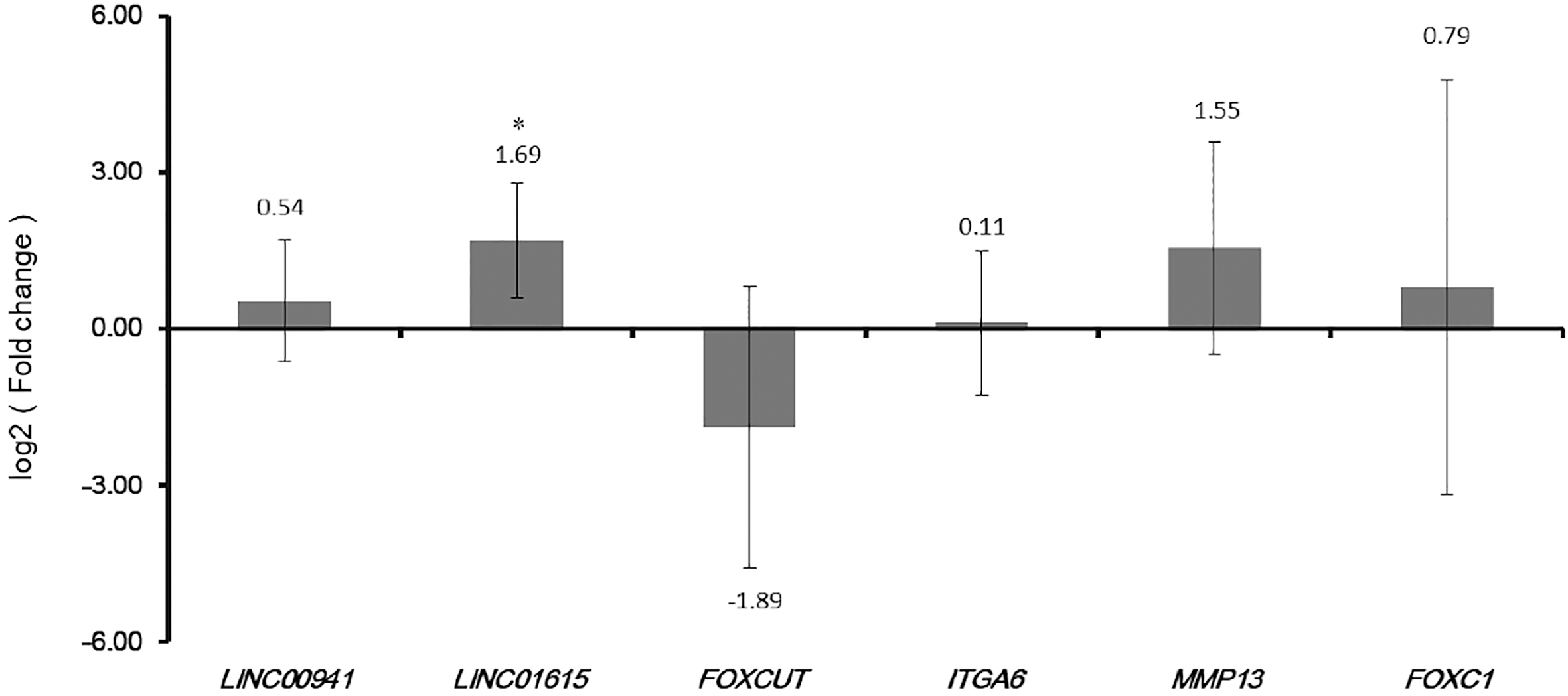
HNSCC, a frequent cancer of the head and neck region, has the high incidence and mortality rate in developing countries nowadays [8]. Although the diagnosis and treatment have been advanced during recent years, HNSCC still ranks among the seventh most common type of cancer worldwide [1]. Thence, exploring for diagnosis and prognosis biomarkers of HNSCC is an urgent problem. Through the present study, we aimed to obtain DEmRNA and DElncRNA in HNSCC patients compared with normal base on TCGA dataset. The machine learning and survival analysis were used to uncover the potential diagnostic and prognostic value of lncRNAs for HNSCC. We obtained 13 optimal diagnostic lncRNA biomarkers including IL12A.AS1, RP11.159F24.6, RP11.863P13.3, LINC00941, FOXCUT, RNF144A.AS1, RP11.218E 20.3, HCG22, HAGLROS, LINC01615, RP11.351J 23.1, AC024592.9 and MIR9.3HG. Among which, AC024592.9, LINC00941, LINC01615 and MIR9-3HG were significantly associated with the prognosis of patients with HNSCC.
To our knowledge, except of IL12A.AS1, RP11.159 F24.6, LINC00941, FOXCUT, HCG22, HAGLROS, LINC01615 and AC024592.9, the present study was the first to identify the RP11.863P13.3, RNF144A.AS1, RP11.218E20.3, RP11.351J23.1 and MIR9.3HG in HNSCC. Li et al. reported that IL12A.AS1 and HCG22 were down-regulated and RP11.159F24.6 was up-regulated between laryngeal squamous-cell carcinoma and adjacent tissues [9]. Zou et al. found that HCG22 and AC024592.9 were down-regulated in HNSCC carcinoma compared with adjacent tissues [10]. HAGLROS was elevated in nasopharyngeal carcinoma, and HAGLROS contributes to nasopharyngeal carcinoma progress via regulating miR-100/ATG14 axis-mediated PI3K/AKT/mTOR signals [11]. Sassenberg et al. displayed that LINC01615 was up-regulated in HNSCC carcinoma compared with adjacent tissues [12]. In this study, IL12A.AS1, HCG22 and AC024592.9 were down-regulated and RP11.159F24.6, LINC01615 and HAGLROS was up-regulated in HNSCC patients compared with normal, which were consistent with reports of other researchers, indicating the our integration analysis data were reliable.
Long intergenic non-protein coding RNA 941 (LINC00941) is an lncRNAs considered as a candidate biomarkers of multiple cancer. LINC00941 is overexpressed in liver cancer and correlated with poor prognosis, and LINC00941 overexpression promoted epithelial to mesenchymal transition of liver cancer [13]. In lung cancer, LINC00941 showed prognostic values and regulated focal adhesion and PI3K-AKT signaling pathway [14]. LINC00941 is overexpressed in gastric cancer tissues and may be a potential novel biomarker for therapeutic of gastric cancer [15]. Liu et al. uncovered that LINC00941 plays a key oncogenic function in gastric cancer and may be used as a potential biomarker for diagnosis and prognosis of gastric cancer [16]. It is reported that LINC00941 was up-regulated in HNSCC carcinoma compared with adjacent tissues [17]. In this study, LINC00941 was up-regulated in both TCGA integration analysis and qRT-PCR validation. The machine learning showed that LINC00941 was a one of optimal diagnostic lncRNAs biomarkers. The survival analysis results displayed that LINC00941 was significantly associated with the prognosis of patients with HNSCC. Therefore, LINC00941 was considered as a potential biomarker for diagnosis and prognosis of HNSCC.
Here, integrin subunit alpha 6 (ITGA6), belongs to the integrin family, was enriched in focal adhesion and ECM-receptor interaction pathway. Silencing of ITGA6 expression was shown to inhibit cell migration and invasion in head and neck cancer cells [18]. Yang et al. reported that ITGA6 in ECM-receptor interaction signaling pathway may play an crucial role in HNSCC [19]. ITGA6 was identified to be significantly correlated with HNSCC [20]. In this study, ITGA6 co-expressed with LINC00941 and RP11.159F24.6 was enriched in focal adhesion and ECM-receptor interaction pathway. Thence, we speculated that LINC00941 and RP11.159F24.6 might be involved in the occurrence of HNSCC by regulating focal adhesion and ECM-receptor interaction pathway.
Jin et al. showed that matrix metallopeptidase 13 (MMP13) was DEGs in HNSCC, suggesting that MMP13 may play a key roles in HNSCC progression [21]. MMP13 was up-regulated in HNSCC, and it has potential value as a marker predictive of the decreased efficacy of anti-PD-1 therapy [22]. In the study, MMP13 was up-regulated in HNSCC and co-expressed with RP11.863P13.3, RP11.218E20.3 and LINC01615. Therefore, we concluded that RP11. 863P13.3, RP11.218E20.3 and LINC01615 might be involved in the progression of HNSCC by regulating MMP13.
Collagen type IV alpha 1 chain (COL4A1), belongs to collagen family, is associated with cell adhesion. Chen et al. found that COL4A1 might be as a biomarker to distinguish oral squamous cell carcinoma and controls [23]. COL4A1 was high expressed in HNSCC which indicates that COL4A1 may participate in HNSCC cell adhesion [24]. In this study, COL4A1 was co-expressed with RP11.159F24.6, RP11.863P13.3, LINC00941 and RNF144A.AS1, and COL4A1 was enriched in pathway of focal adhesion, ECM-receptor interaction and pathways in cancer. These findings demonstrated that COL4A1 might play a pivotal role in the progression of HNSCC by regulating pathway of focal adhesion, ECM-receptor interaction and pathways in cancer.
It has been reported that the fork box C1 (FOXC1) is involved in various cancer cell proliferation, migration and invasion [25, 26, 27]. Gao et al. displayed that FOXC1 is involved in the invasion and metastasis of laryngeal squamous cell carcinoma, suggesting that FOXC1 might be a risk factor for laryngeal squamous cell carcinoma [28]. In this study, FOXC1 co-expressed with FOXCUT in HNSCC. FOXCUT may play a key role in the tumor progression and survival of esophageal squamous s cell carcinoma patients by modulating FOXC1 [29]. Functional of FOXC1 and FOXCUT is correlated with tumor progression of oral squamous cell carcinoma, and FOXC1 and FOXCUT may serve as novel biomarkers in oral squamous cell carcinoma patients [30]. One study found that FOXCUT expression contributed to the proliferation and migration of nasopharyngeal carcinoma by targeting FOXC1 and that FOXCUT may be as a novel nasopharyngeal carcinoma biomarker [31]. We provide further evidence that FOXCUT was a novel diagnosis biomarkers of HNSCC.
In conclusion, 13 DElncRNAs were identified as potential biomarkers of HNSCC. Among which, 4 DElncRNAs were significantly associated with the prognosis of patients with HNSCC. Overall, the confirmation results of qRT-PCR were generally consistent with our integrated analysis. Several limitations still detected in our study, the sample size for qRT-PCR confirmation was small, and large numbers of HNSCC samples are needed for further research. Identifying diagnostic and prognostic biomarkers for HNSCC is a pilot study, and further experiments are needed to understand the biological of key lncRNAs in HNSCC.
Footnotes
Acknowledgments
This study was supported by the Natural science foundation of hunan province (No: 2017JJ2178).
Conflict of interest
The authors declare that they have no conflict of interest.
