Abstract
BACKGROUND:
As the most prevalent primary bone malignancy in children and adolescents, osteosarcoma (OS) has attracted increasing attention. The role of circRNAs in OS has been elucidated in some reports, but many circRNAs remain unexplored. Circ_0000337 has only been revealed as an oncogenic circRNA in esophageal squamous cell carcinoma. Yet whether circ_0000337 exerts any specific function in OS has not been unmasked.
METHODS:
RT-qPCR was used for measurement of circ_0000337, miR-4458 and BACH1 mRNA levels. Western blot was conducted to detect BACH1 protein. CCK-8 assay, Casepase-3 activity assay and transwell assay were utilized to assess changes on cellular processes. Cytoplasmic/nuclear fractionation assay was conducted for circ_0000337 localization in OS cells. Luciferase reporter assay and RIP assay were performed to validate the interaction between miR-4458 and circ_0000337 or BACH1.
RESULTS:
Circ_0000337 expression was upregulated in OS cell lines and it silence hindered OS cell proliferation and migration. MiR-4458 was downregulated in OS cells and miR-4458 upregulation suppressed OS cell growth and migration. Importantly, circ_0000337 sponged miR-4458 to elevate BACH1 expression, thus facilitating OS development.
CONCLUSIONS:
This research for the first time documented that circ_0000337 promoted OS progression via sponging miR-4458 and thus elevating BACH1 expression, offering rational therapeutic target for OS.
Introduction
Osteosarcoma (OS) is known as the most prevalent primary bone malignancy in children and adolescents [1]. OS patients always have a poor outcome due to the aggressiveness of OS [2]. Although OS prognosis has been improved in response to development of surgical resection and chemotherapy, the overall survival rate of OS patients remains unfavorable [3, 4]. Thus it is urgent to identify new targets for OS therapy.
Circular RNAs (CircRNAs) are a group of recently researched RNAs which are omnipresent in organisms and characterized with loop structure formed through back-splicing events [5]. The pivotal role of circRNAs in a wide range of human cancers has been manifested in many reports. For example, hsa_circ_0020397 facilitates the progression of colorectal cancer through upregulatingmiR-138 target genes expression [6]. Hsa_circ_0001982 elicits promoting effect on breast cancer progression through inhibiting miR-143 expression [7]. Hsa_circ_0009910 promotes osteosarcoma development by targeting miR-449a/IL6R axis [8]. Circ_0000337, which is also named as circRNA_100872, is a circRNA derived from PTPRF interacting protein alpha 1 (PPFIA1), which has been reported as amplified and facilitative in chronic myeloid leukemia, head and neck squamous cell carcinoma, invasive ductal carcinoma of the breast etc. [9, 10, 11]. Circ_0000337 has only been proofed as an oncogenic circRNA in esophageal squamous cell carcinoma [12]. The role of circ_0000337 in OS has not been researched yet.
This study is aimed to probe into the specific function and the detailed mechanism of circ_0000337 in OS.
Materials and methods
Cell lines and cell culture
Four human OS cell lines including U2OS, HOS, 143B and MG-63, one human noncancerous osteoplastic cell line NHOst as well as one human osteoblastic cell line hFOB1.19 were all supplied by American Type Culture Collection (ATCC, VA, USA). Cells were kept in Dulbecco’s modified Eagle’s medium (DMEM, Gibco, MA, USA) supplemented with penicillin-streptomycin and 10% FBS. The incubating environment was set with 5% CO
Cell transfection
Interference of circ_0000337 was performed using short hairpin RNAs (shRNAs; 250 ng/ul) against circ_0000337 (sh-circ_0000337-1/2/3) synthesized by GenePharma (Guangzhou, China). For overexpression of BACH1, full sequence of BACH1 was subcloned into pcDNA3.1 plasmids manufactured by Invitrogen (CA, USA), constructing pcDNA3.1/BACH1. MiR-4458, miR-4458 mimics (100 nM) or the negative control (NC-mimics) were commercially obtained from Genepharma. All the constructed products were transduced into U2OS or HOS cells using Lipofectamine 2000 (Invitrogen) in accordance with the guidebook and transduction efficiency was tested by RT-qPCR.
Real-time quantitative PCR (RT-qPCR)
RNA isolated using TRIzol reagent (Invitrogen) was subjected to precipitation using phenol – chloroform plus ethanol and resuspension using non-RNase H
Western blot
Total protein was harvested through cell lysis with RIPA lysis buffer (Beyotime) and quantified in SDS. Protein samples separated on an SDS-PAGE gel were subsequently transferred onto nitrocellulose membranes (GE Healthcare) electrophoretically. Next the membranes were incubated with primary antibodies of BACH1 (1:1000, Santa Cruz, CA, USA) and GAPDH (1:1000, Abcam, Cambridge, UK) at 4
Cell counting kit-8 (CCK-8) assay
Cell Counting Kit-8 (CCK-8) provided by Beyotime (Jiangsu, China) was used to quantify cell viability. In brief, U2OS and HOS cells seeded in 96-well plates with the concentration of 1
Caspase-3 activity assay
A caspase-3 colorimetric assay kit was utilized to assess caspase-3 activity. Cells were preserved for 48 hours after transfection. After being lysed, cells were subjected to centrifugation and the supernatant collected after centrifugation was injected to 96-well plates which were then added with reaction buffer which contained caspase-3 substrate (Ac-DEVD-pNA). After 4 hours of incubation, absorbance at 405 nm was measure by a microplate reader.
Circ_0000337 silence retards cell proliferation and migration in OS. (A) RT-qPCR was carried out for measurement of circ0000337 in four OS cell lines (U2OS, HOS, 143B, MG-63) and a human noncancerous osteoplastic cell line NHOst. (B) RT-qPCR was conducted for validation of sh-circ_0000337-1/2/3 knockdown efficiency. (C) CCK-8 assay was used for detection of proliferative capacity after transduction of sh-circ_0000337-1, sh-circ_0000337-2 or sh-NC. (D) Caspase-3 activity assay was utilized for assessment of cell apoptosis. (E) Transwell assay was carried out for evaluation of cell migration. (F) Subcellular distribution of circ_0000337 was confirmed through cytoplasmic/nuclear fractionation. All the experiments were performed in triplicate. 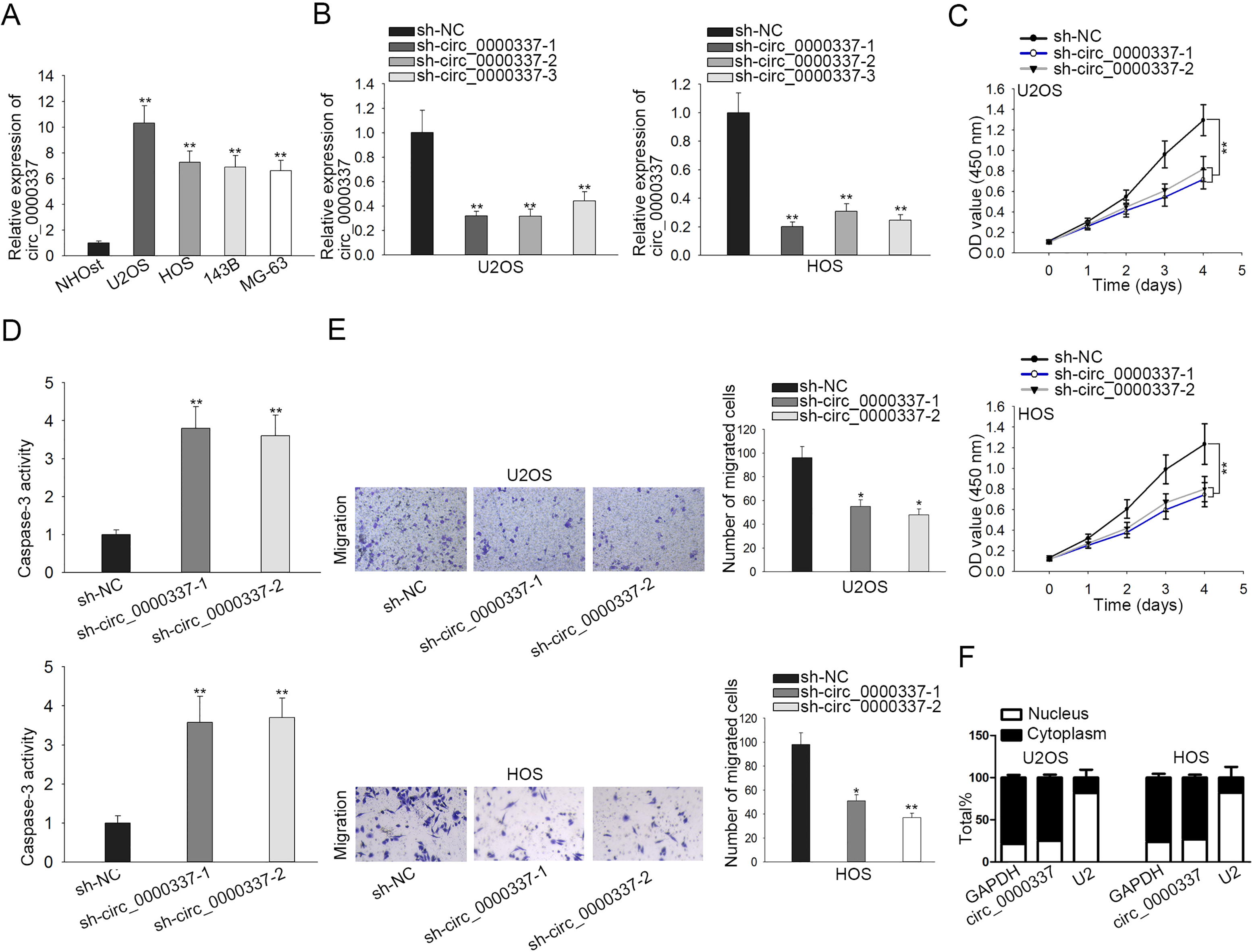
Cell migration was detected using chambers with 8
Nuclear/cytoplasmic fractionation
Subcellular localization of circ_0000337 in U2OS and HOS cells was ascertained using Cytoplasmic and Nuclear RNA Purification Kit (Norgenbiotek Corporation, ON, Canada) based on the manufacturer’s protocol. Cytoplasmic and nuclear RNAs were determined by conducting RT-qPCR with GAPDH or U2 serving as the cytoplasmic or nuclear control.
MiR-4458 is sponged by circ_0000337 and inhibits OS progression. (A) The expression of miR-4458 in OS cell lines and a normal osteoplastic cell line was detected by RT-qPCR. (B) The expression of miR-4458 in U2OS and HOS cells transfected with sh-circ_0000337-1 or sh-NC was assessed by RT-qPCR. (C) The binding sequence was predicted by starBase2. (D) Luciferase reporter assay was used to confirm the interaction between circ_0000337 and miR-4458. (E) RIP assay was conducted and RNAs precipitated were measured by RT-qPCR. (F) CCK-8 assay evaluated cell proliferation after transfection with NC-mimics or miR-4458 mimics. (G) Caspase-3 activity assay measured cell apoptosis. (H) Transwell assay detected cell migration. All the experiments were conducted as least in triplicate. 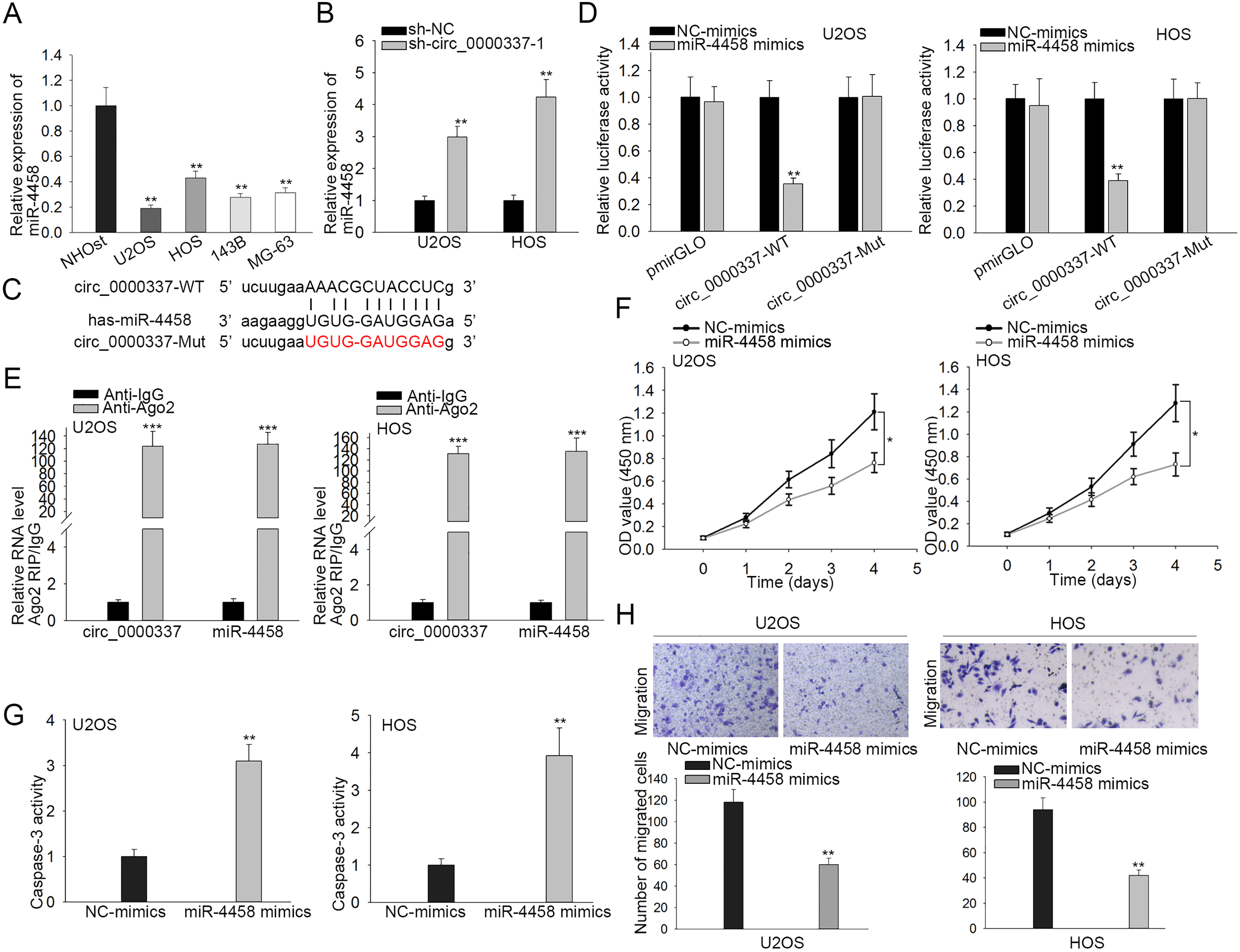
The putative binding sequences of miR-4458 and circ_0000337 or BACH1 were predicted by starBase2 (
RNA immunoprecipitation (RIP) assay
RNA-Binding Protein Immunoprecipitation kit (Millipore, Burlington, MA, USA) was used in RIP assay. Anti-Ago2 and anti-IgG were used for RNA immunoprecipitation and RT-qPCR was used for RNA measurement.
Statistical analysis
Statistical analysis was carried out using SPSS 22.0 (SPSS, IL, USA). Data collected from assays at least in triplicate were expressed as mean
3 BACH1 is negatively regulated by miR-4458. (A) BACH1 mRNA level in OS cell lines and the noncancerous osteoplastic cell line was assessed by RT-qPCR. (B) BACH1 protein level in OS cell lines and normal osteoplastic cell line was measured by western blot. (C) BACH1 mRNA level in U2OS and HOS cells treated with NC-mimics or mir-4458 mimics was assessed by RT-qPCR. (D) BACH protein level in U2OS and HOS cells treated with NC-mimics or miR-4458 mimics was detected by western blot. (E) The binding sequence between miR-4458 and BACH11 was predicted by starBase2. (F) Luciferase reporter assay was used to confirm the interaction between miR-4458 and BACH1. (G) RIP assay was carried out and RNA enrichment was determined by RT-qPCR. All the experiments were conducted at least in triplicate. 
Circ_0000337 facilitates OS progression via elevating BACH1 expression. (A) BACH1 mRNA level in U2OS and HOS cells treated with sh-NC or sh-circ_0000337-1 was detected with RT-qPCR. (B) BACH1 protein level in U2OS and HOS cells treated with sh-NC or sh-circ_0000337-1 was detected with western blot. (C) Upregulation efficiency of pcDNA3.1/BACH1 at mRNA level was validated by RT-qPCR. (D) Upregulation efficiency of pcDNA3.1/BACH1 at protein level was validated by western blot. (E) CCK-8 assay was used for evaluation of cell proliferation. (F) Caspase-3 assay was carried out for evaluation of cell apoptosis. (G) Transwell assay was used for assessment of cell migration. All the experiments were conducted at least in triplicate. 
Circ_0000337 silence retards cell proliferation and migration in OS
The expression profile of circ_0000337 in OS cells was detected. The results of RT-qPCR indicated that circ_0000337 was overexpressed in OS cell lines than that in normal NHOst cell line (Fig. 1A and Supplementary Fig. S1F). To investigate the function of circ_0000337 in OS, we transduced sh-circ_0000337-1, sh-circ_0000337-2 or sh-circ_0000337-3 into U2OS and HOS cells, the knockdown efficacy of which was confirmed by RT-qPCR (Fig. 1B). The follow-up loss-of-function assays were conducted using sh-circ_0000337-1/2 due to their better efficiency. After circ_0000337 depletion, the proliferative capacity of U2OS and HOS cells was strikingly impaired (Fig. 1C). Conversely, Caspase-3 activity assay demonstrated that cell apoptosis depending on activated Caspase-3 was facilitated after circ_0000337 silence (Fig. 1D). Additionally, cell migration was retarded when circ_0000337 was silenced (Fig. 1E). Afterwards, we localized circ_0000337 in U2OS and HOS cells and observed predominant distribution of circ_0000337 (Fig. 1F). To conclude, circ_0000337 silence has inhibitive effect on OS cell proliferation and migration.
MiR-4458 is sponged by circ_0000337 and inhibits OS progression
Considering mounting evidence proving that circRNAs work as miRNA sponges and the cytoplasmic abundance of circ_0000337 in OS cells, we reasoned that circ_0000337 might regulate OS cell proliferation and migration via sponging miRNAs. Thereupon, we utilized starBase2 (
BACH1 is negatively regulated by miR-4458
We further searched for downstream cascade of miR-4458 and found BACH1, whose oncogenic function has been proven by a large amount of studies [16, 17, 18]. Through RT-qPCR and western blot, we found BACH1 was significantly overexpressed in OS cell lines, not only at the mRNA level but also at the protein level (Fig. 3A–B and Supplementary Fig. S1H). After treatment with miR-4458 mimics, BACH1 mRNA and protein levels were both markedly cut down (Fig. 3C–D). The binding site between miR-4458 and BACH1 was predicted via starBase2 (Fig. 3E). As illustrated by Fig. 3F, co-transfection with miR-4458 mimics resulted in an obvious decrease of BACH1-WT luciferase activity, but not the mutant type. Moreover, RIP assay demonstrated that both miR-4458 and BACH1 were significantly enriched in anti-Ago2 group but not in anti-IgG group, further validating the direct binding potential of miR-4458 and BACH1 (Fig. 3G). To sum up, BACH1 is negatively regulated by miR-4458.
Circ_0000337 facilitates OS progression via elevating BACH1 expression
To explore the regulatory effect of circ_0000337 in BACH1 expression, cells transfected with sh-circ_0000337-1 or sh-NC were subjected to RT-qPCR and western blot. The results indicated that the BACH1 mRNA and protein levels were both markedly reduced in circ_0000337 depleted cells (Fig. 4A–B). To examine whether BACH1 was required for circ_0000337 to affect cellular activities in OS, we conducted rescue assays after overexpressing BACH1 in U2OS and HOS (Fig. 4C–D, Supplementary Fig. S1A–B). Co-transfection with pcDNA3.1/BACH1 rescued the attenuating effect of circ_0000337 silence on U2OS and HOS cells proliferation and the promoting effect of circ_0000337 silence on U2OS and HOS cells apoptosis (Fig. 4E–F, Supplementary Fig. S1C–D). In addition, the reduced cell migration ability in response to circ_0000337 knockdown was partly revived by BACH1 upregulation (Fig. 4G, Supplementary Fig. S1E). Taken together, circ_0000337 facilitates OS progression via upregulating BACH1 expression.
Discussion
OS is a highly malignant neoplasm in bone featured with rapid progression and high metastasis, leading to poor prognosis of OS patients [19, 20]. Despite of emerging researches on the pathogenesis of OS, the molecular mechanisms under osteosarcoma have not been fully clarified. Thus investigation of novel biomarkers of OS is badly needed.
Recently the participant role of circRNAs in OS has drawn increasing attention. For example, circFAT1 mediates OS progression by sponging miR-375 and regulating Yes-associated protein 1 expression [21]. Circ_0001721 upregulation facilitates the tumorigenesis of OS cells through sponging miR-569 and miR-599 [22]. Hsa_circ_0081001 serves as a promising biomarker for OS treatment [23]. Although circ_0000337 has been unveiled as an oncogenic circRNA in esophageal squamous cell carcinoma [12], its performance in OS remains elusive. In the present study, we detected the high expression levels of circ_0000337 in OS cells and we also observed that circ_0000337 depletion restrained cell proliferation and migration in OS.
The ceRNA hypothesis elucidated a new way in which coding and non-coding transcripts communicate with each other via competitively binding to miRNAs to post-transcriptionally construct a regulatory network [21, 24, 25]. MiR-4458 has been revealed as tumor-suppressive gene in some types of human cancers [13, 15, 26]. In this study, miR-4458 expression was signally downregulated and circ_0000337 was directly bound to miR-4458. Besides, miR-4458 upregulation impaired the proliferation and migration of OS cells.
BTB domain and CNC homolog 1 (BACH1) is a downstream target of miR-4458 and has been accepted as an oncogene in a wide range of cancers. For example, silenced BACH1 inhibits prostate cancer cell invasion and migration [16]. BACH1 facilitates colorectal cancer progression via BACH1/CXCR4 pathway [17]. BACH1 knockdown represses migration of HT-29 colon cancer cells by inhibiting metastasis-related genes expression [18]. In this research we not only confirmed the direct interaction between miR-4458 and BACH1 but also the negative correlation between circ_0000337 and BACH1. Moreover, rescued-function assays revealed that BACH1 overexpression could offset the repressive effect of silenced circ_0000337 on OS cell proliferation, apoptosis and migration. All the findings indicated that circ_0000337 elevated BACH1 expression to facilitate OS progression by competitively binding with miR-4458.
In summary, this research uncovers a circ_0000337/ miR-4458/BACH1 pathway in regulation of OS progression, which initially documents the oncogenic function and molecular mechanism of circ_0000337 in OS and offers potential therapeutic targets for OS.
Footnotes
Acknowledgments
We appreciate the supports of the Affiliated Hospital of Qingdao University.
Conflict of interest
None to report.
Supplementary data
(A) Upregulation efficiency of pcDNA3.1/BACH1 at mRNA level was validated by RT-qPCR. (B) Upregulation efficiency of pcDNA3.1/BACH1 at protein level was validated by western blot. (C) CCK-8 assay was used for evaluation of cell proliferation. (D) Caspase-3 assay was carried out for evaluation of cell apoptosis. (E) Transwell assay was used for assessment of cell migration. (F–H) The expression of circ_0000337, miR-4458 and BACH1 in OS cell lines and human osteoblastic cell line hFOB1.19 was detected via RT-qPCR. All the experiments were conducted at least in triplicate. 
