Abstract
OBJECTIVE:
Renal cell carcinoma (RCC) is the most common malignancy involving the kidneys and a major cause of cancer mortality. The involvement of microRNA (miRNA) expression in the tumorigenesis and progression of RCC still not been previously highlighted. We aimed to explore the potential role of miR-222-3p in renal cell carcinoma (RCC).
METHOD:
We first found that miR-222-3p was elevated significantly in the RCC tissues as compared to the non-tumor counterparts. We also found that a higher level of miR-222-3p in different RCC cell lines than the HK-2 cells.
RESULTS:
In vitro validation experiment using miR-222-3p mimic molecules significantly induced expression of EMT marker vimentin and downregulated E-cadherin in both 769-P and 786-O RCC cells. In contrary, when miR-222-3p was downregulated by its inhibitor, the reverse observations were made. We then demonstrated a reversal association between the expression level of miR-222-3p and TIMP2/ERK where TIMP2 functions as a tumor suppressor. In a small cohort of 45 clinical samples, we found that miR-222-3p expression level was elevated and was associated with a poorer survival of the patients. Patients with higher miR-222-3p expression showed had a statistically shorter overall survival than those patients of lower miR-222-3p level (HR, 5.789;
CONCLUSION:
Collectively, we showed that miR-222-3p functioned as a tumor progression marker and could be a target for future drug development.
Introduction
Renal cell carcinoma (RCC) represents a highly prevalent kidney malignancy with roughly 65, 000 new diagnoses and may claim as many as 15,000 lives in the USA [1]. RCC has been shown to be extremely resistant towards standard chemo- and radiotherapy; surgery remains the standard and feasible treatment for localized RCC cases [2]. And, the 5-year overall survival is less than 50% [3]. This low survival rate may be contributed by the high frequency of distance metastasis (approximately 25% of the patient at diagnosis). Thus, it is urgent to develop reliable biomarker(s) and/or druggable target for metastatic RCC [4].
MicroRNAs (miRNAs) are RNA molecules which do not code for proteins and play an essential gene-silencing role in many cellular processes including cancer. The miRs suppresses gene expression via their binding to the 3’-UTR of mRNA of their targeted genes and leads to the degradation of the target mRNAs [5]. Accumulating evidence showed that in cancer cells, miRNAs may be dysregulated and resulted in aberrant suppression of tumor suppressors or increased expression of oncogenes [6]. Mounting evidence demonstrate that miRNAs are involved in many aspects of tumorigenesis including increased epithelial-to-mesenchymal transition (EMT, a pre-requisite for cancer cells to metastasize), development of drug resistance, promotion of angiogenesis and anti-apoptosis [7]. Erroneous regulation of miRNAs is virtually found in all cancer types, and RCC is not an exception [8]. Previous evidence has demonstrated that miR-222 played an essential role in the tumorigenesis of gastric [9], brain [10] and lung cancer [11]. However, the role of miR-222 in RCC has not been fully explored.
Based on these premises, in this study, we obtained clinical samples from patients diagnosed with RCC and cell lines to explore the functional role(s) of miR-222 as well as its potential as prognostic marker for RCC patients.
Materials and methods
Clinical sample
RCC tissues and corresponding normal renal tissues of 45 patients were obtained from Department of Urology, Zhejiang Provincial People’s Hospital, People’s Hospital of Hangzhou Medical College (Hangzhou, China). This study was approved by the Ethics Committee of Zhejiang Provincial People’s Hospital, and informed consents were obtained from all patients.
RNA isolation and real-time PCR
Total RNA of clinical samples and cancer cell lines was isolated using Trizol reagent (Invitrogen; Thermo Fisher Scientific, Inc.) and cleaned by RNeasy kit (Qiagen GmbH, Hilden, Germany) based on the protocols provided by the vendor. The isolation of miRNA performed using miRNeasy Kit (Qiagen GmbH, Hilden, Germany) according to the manufacturer’s specifications. Approximately 1
Cell culture and transfection
Human renal cell carcinoma (RCC) cell lines (786-O, ACHN, 769-P and Caki-1) and renal tubular epithelial cell line HK-2 were all purchased from the American Type Culture Collection (ATCC). All cell lines were cultured under the protocols recommended by the ATCC. miR-222-3p mimics, negative control (NC), miR-222-3p inhibitor and inhibitor negative control (inhibitor NC) (Genepharma, Shanghai, China) were transfected into cells at a concentration of 50 nM using Lipofectamine 3000 (Invitrogen; Thermo Fisher Scientific, Inc.).
Migration assay
769-P and 786-O transfected with miR-222-3p mimics, inhibitor and scramble control were seeded into the upper chamber of Boyden chamber (2
Bioinformatics and target prediction analysis
The TCGA kidney renal clear cell carcinoma and papillary carcinoma cohorts were analyzed. The expression range of the probe (for miR-222) is from 6-760 and the cutoff value used was 41 in the analysis. Target prediction was conducted for miR-222-3p with PicTar (
SDS-PAGE and western blotting
Cells were seeded in 6-well plates and harvested. The total protein lysates were then extracted using the RIPA lysis buffer. A standard SDS-PAGE was performed to separate proteins (20
Immunohistochemistry
RCC tumor tissues and adjacent normal non-tumor tissue paraffin blocks were sectioned and mounted onto slides (4-
Flow cytometry for apoptosis measurement
Cells transfected with scramble, inhibitor and mimic molecules of miR-222-3p were collected and suspended with binding buffer. Cells were stained with Annexin V-kFluor488 and PI (Kaiji Biology Corp., China) at room temperature, 30 minutes and analyzed by a flow cytometer (Cyto Flexz, USA). Annexin V+/PI- cells were counted as early apoptotic cells while the annexin V+/PI+ cells counted as late apoptotic cells. The apoptotic ratio was calculated as the ratio of total apoptotic cells/total number of cells. Cells transfected with inhibitor and mimic were normalized against cells transfected with scramble.
Statistics
The results of the experiment were analyzed by SPSS 19.0 (IBM SPSS, Armonk, NY, USA) and presented as the mean
Result
Increased level of miR-222-3p in RCC
We first analyzed The Cancer Genome Atlas (TCGA database) and found that an increased miR-222-3p level was correlated with a shorter survival of the patients (Fig. 1A). To provide in vitro evidence, we compared the expression level of miR-222-3p among different RCC cell lines against normal cell line, HK2. Our results from qPCR analyses were similar to the TCGA data where RCC cells contained a significantly higher level of miR222-3p as compared to HK2 cells (Fig. 1B). Thus, the results indicate that miR-222-3p may serve as an onco-miR in RCC.
Expression of miR-222 is associated to RCC tumorigenesis and prognosis. (A) Kaplan-Meier curve showing the association of miR-222 expression and overall survival analyzed from The Cancer Genome Atlas (TCGA) dataset. Patients with lower levels of miR-222 had significantly higher overall survival compared to those with higher miR-222 expression. Please refer Supplementary Table 2 for clinical information on the patient cohort used in this analysis. (B) Comparative miR-223-3p profiles among different RCC cell lines as normalized to the HK-2 cells. 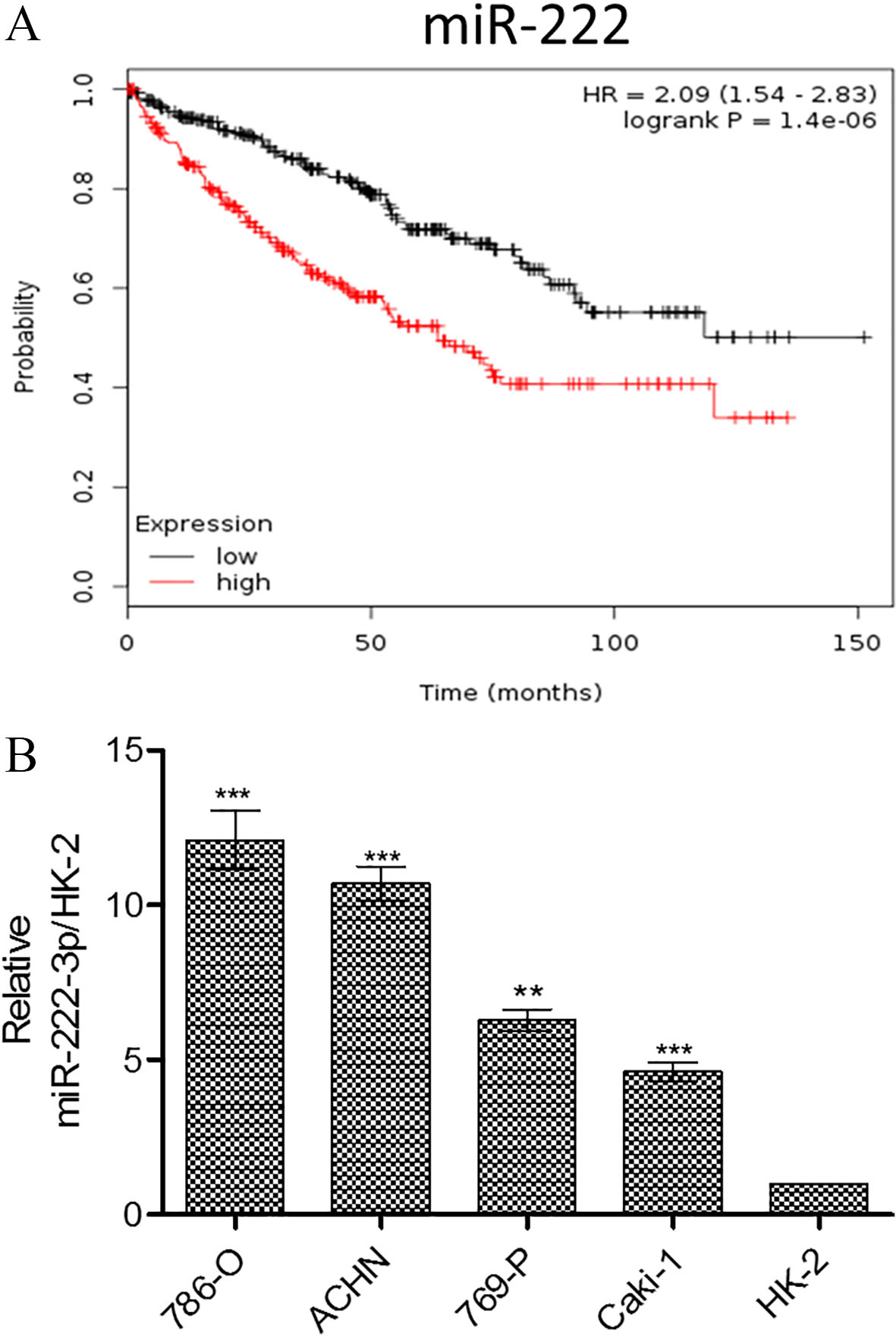
Next, we examined the effect of miR-222-3p on the survival of RCC cell lines using miR-222-3p mimic molecules and inhibitors. We found both 786-O and 769-P cells showed lowered apoptotic ratio when transfected with the mimic molecules (exogenous miR-222-3p) while an increased percentage of apoptotic cells were observed in cells transfected with miR-222-3p inhibitor (Fig. 2A). In support, Western blots of 786-O and 769-P cells transfected with miR-222-3p mimic or inhibitor molecules demonstrated the similar phenomenon. When the level of miR-222-3p was increased by the mimic molecules, there was a lower expression level of the pro-apoptotic marker, PARP in both cells’ lines; the reverse is true in the case of inhibitor (Fig. 2B). These results strongly suggested that miR-222-3p may function to prevent apoptosis in the RCC cells.
Association between miR-222-3p status
and pathological variables in renal clear cell carcinoma
Association between miR-222-3p status
In vitro apoptosis assay. (A) The presence of miR-222-3p reduced the degree of apoptosis in both 786-O and 769-P cells. The data expressed represents results from three independent experiments. ***, 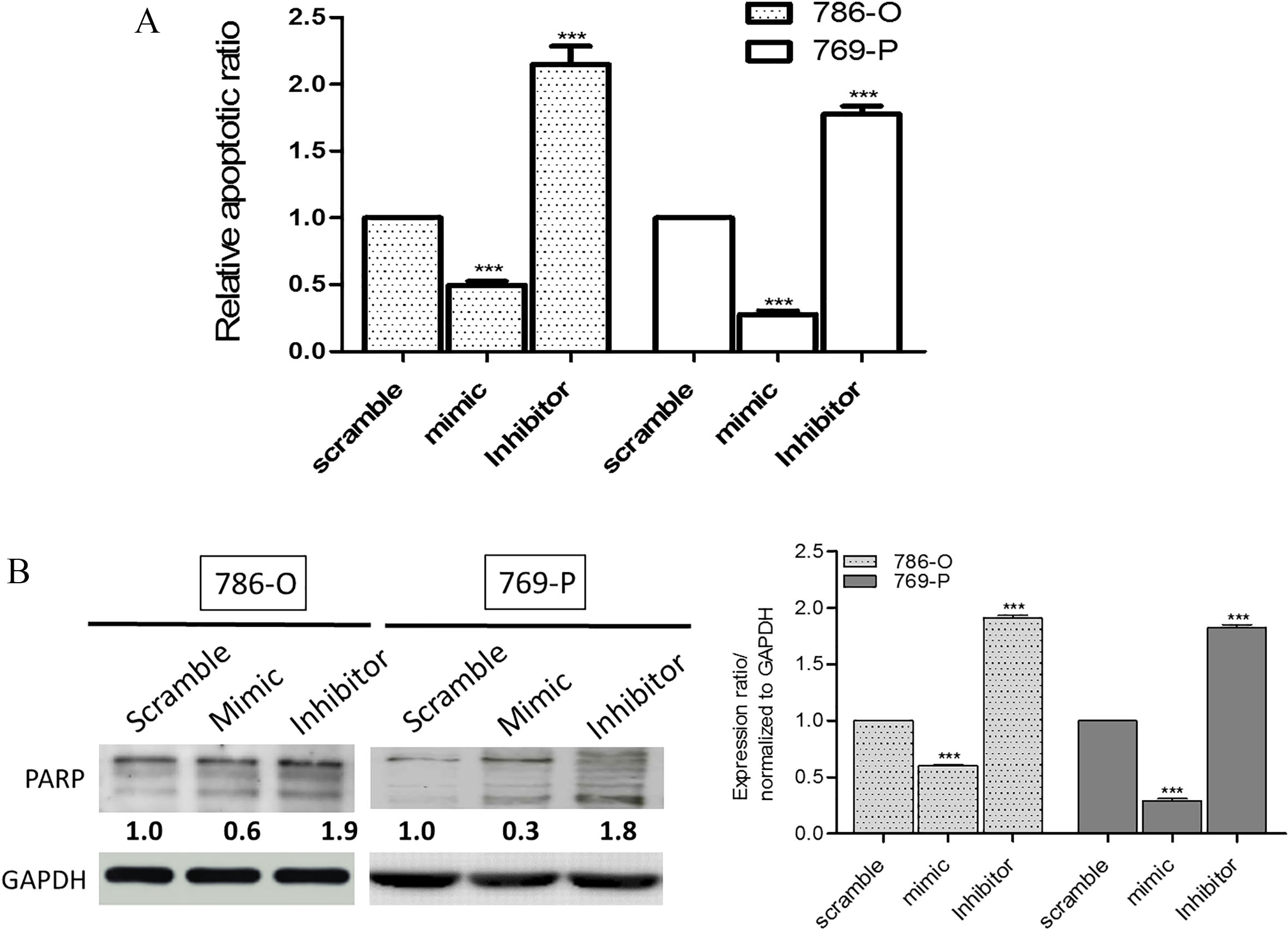
We performed a microRNA database search and found one of the targets of miR-222-3p was TIMP2, a key molecule for cancer metastasis. It was found that miR-222-3p could bind to the 3’UTR of TIMP2 (Fig. 3A). Notably, we showed that miR-222-3p mimic-transfection led to a decreased expression of TIMP2 as well as the phosphorylated ERK1/2 (Fig. 3B) and the reversed observations were made in the case of miR-222-3p inhibitor. In support, we used trans-well assay to evaluate the effect of miR-222-3p on the metastatic ability. Our data demonstrated that silencing of miR-222-3p promoted the migration of both 786-O and 769-P cells (Fig. 3C) while its inhibition led to the reduced the migration (Fig. 3C). In addition, the expression of EMT markers, vimentin (mesenchymal marker) and E-cadherin (epithelial marker) were examined. As expected, the increased miR-222-3p by mimic molecules led to an increased vimentin expression; while the inhibitor of miR-222-3p resulted in the opposite (Fig. 3D). Our findings suggested that miR-222-3p plays an essential role in regulating the migratory ability of RCC via the TIMP2 associated signaling.
The level of miR-222-3p is associated with RCC migratory ability and TIMP2/ERK signaling. (A) TIMP2 is a direct target of miR-222-3p. (Upper panel) The predicted complementary sequence interaction between human miR-222-3p 3’-UTR and TIMP2. (Lower panel) An increased miR-222-3p (mimic) significantly inhibited the luciferase activity of wild-type 3’-UTR of TIMP2 and decreased miR-222-3p (inhibitor) resulted in reverse. (B) Western blot analysis demonstrated the link between miR-223-3p and the expression of TIMP2 and ERK1/2. In both cell lines, the increased of miR-222-3p led to an increased expression of TIMP2 and phosphorylated ERK1/2 and the opposite was observed in the inhibitor group. (C) Wound-scratch experiments showed markedly increased migratory ability in RCC cells (786-O and 769-P cells) transfected with exogenous miR-222-3p mimic molecules as compared to their scramble counterparts. The lower panels represent the comparative migratory ability of RCC cells transfected with miR-222-3p inhibitor. Quantitative measurement of protein expression. The expression level of the protein of interest were compared and normalized against GAPDH (where the level of GAPDH was set at 1). The protein expression level was estimated using ImageJ software. These ratios were derived from three independent western blot experiments. *, 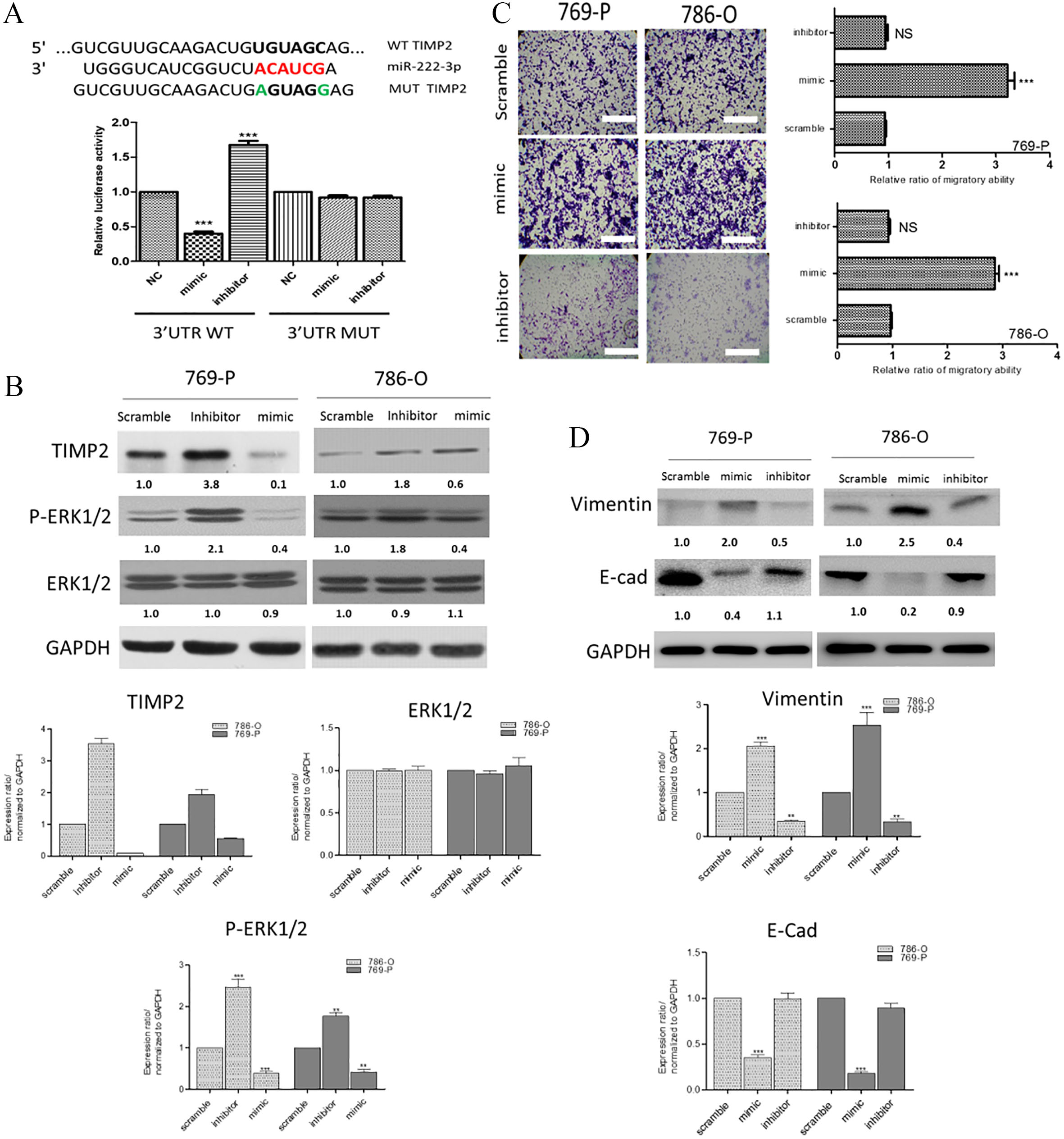
To validate our in vitro analyses, the expression level of miR-222-3p in 45 clinical samples was analyzed using real-time PCR reaction (Table 1). Some significant association was found between miR-222-3p expression level and the following parameters such as sex (
Correlation between miR-222-3p and overall survival
Correlation between miR-222-3p and overall survival
Increased miR-222-3p and decreased TIMP2 is associated with a poor survival in patients with renal clear cell carcinoma (RCC). (A) Immunohistochemical analysis of TIMP2 in RCC samples. Paired tumor (T) and non-tumor (NT) samples were collected from patients with renal clear cell carcinoma. The sections were stained with TIMP2 antibody and scored for the intensity. We found that non-tumor samples (NT) showed a significantly higher TIMP2 staining as compared to a markedly elevated staining of TIMP2 in the tumor (T) samples. (B) Kaplan–Meier curve of metastasis-free survival. According to our qPCR analysis, patients with a higher expression miR-222-3p (above the median) showed a significantly lower metastasis-free survival than their counterparts with a lower miR-222-3p level. 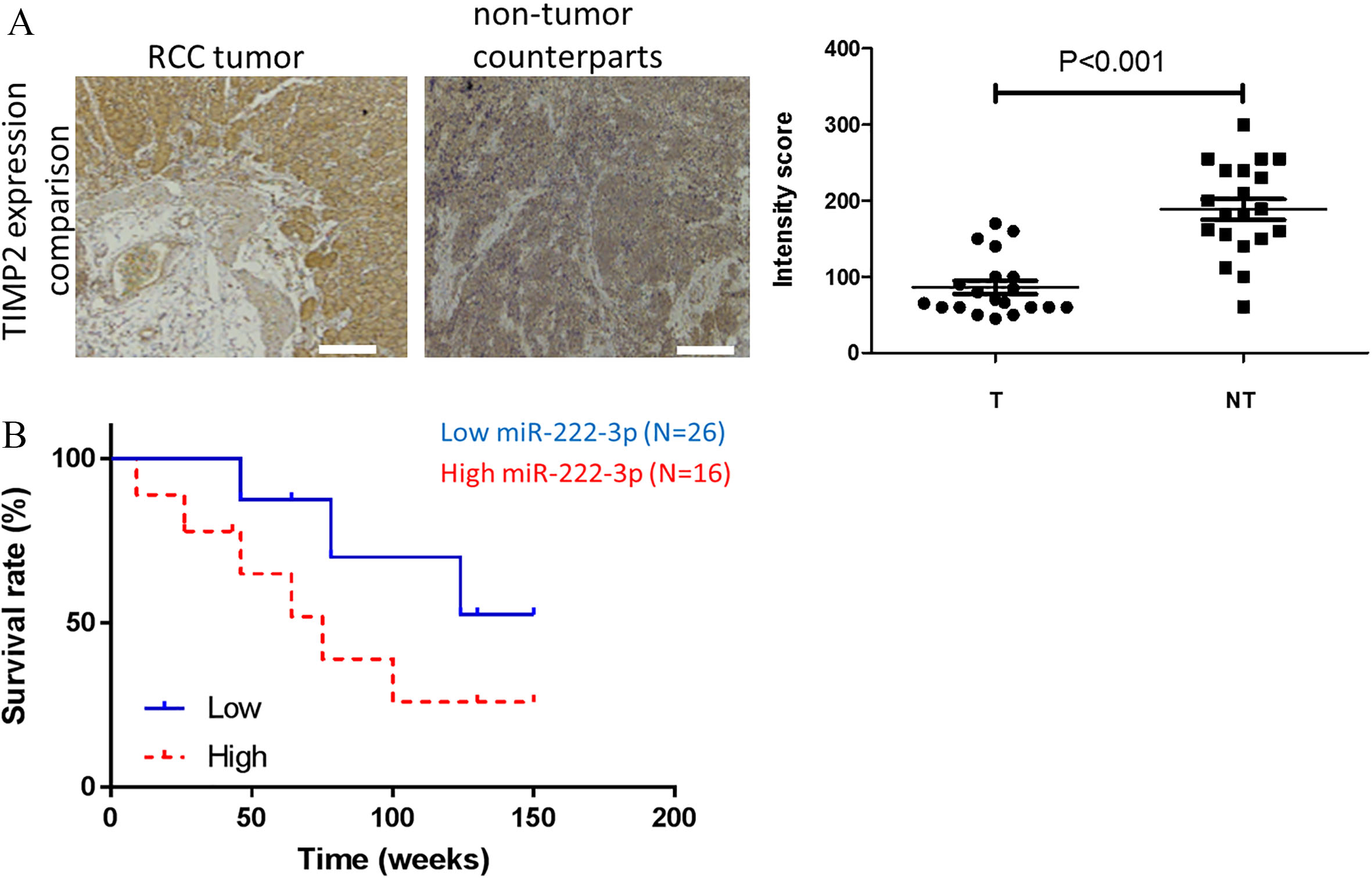
Since we identified TIMP2 as the target of miR-222-3p and its expression was negatively associated with miR-222-3p level in our in vitro experiments, we examined TIMP2 expression using patients’ samples via immunohistochemical analysis. We found that the RCC samples showed a significantly lower intensity for TIMP2 as compared to their non-tumor counterparts (Fig. 4A) and established a clinical correlation between the level of miR-222-3p for a poorer prognosis of the RCC patients (Fig. 4B). The quantitative analysis of the staining intensity was found to be statistically significant. This finding reinforced our previous in vitro study and corroborated with our correlational analysis.
Discussion
Renal cell carcinoma (RCC) represents a prevalent cancer of kidneys. The incidence is growing rapidly, and the outcomes of the patients are often poor despite aggressive treatments [14]. There is an urgency for the development of new approaches for diagnosis and/or therapeutics. In the recent years, microRNAs (or miRs) have gained interests in cancer biologists and physicians due to their roles in tumorigenesis. Accumulating evidence has demonstrated that these non-coding messenger RNA molecules actually play essential roles in genome regulation. MicroRNAs were shown to either promote or suppress major signaling transduction cascades associated with virtually all cellular processes including tumorigenesis [15]. One of the most sought out features for miRs is their role as a surrogate in tumorigenesis.
Our research showed that the level of miR-222-3p was highly expressed in the RCC cell lines but not in the normal tubular epithelial cells, indicating its specificity in the RCC. Our finding is supported by previous studies. For instance, an increase in the level of miR-222 was found to be associated with an increased extracellular signal-regulated kinase 1/2 (ERK1/2) activity resulting in the exacerbation of neointimal hyperplasia in diabetes mellitus [16]. Another report showed that lncRNA Gas5 inhibited glioma cells via miR-222 down-regulation both in vitro and in vivo [17]. More importantly, miR-222-3p was shown to promote the proliferation, invasion and suppress apoptosis by targeting HIPK2 in the case of gastric cancer [18]. It is worth noting that we also identified that the overall level of miR-222, including both miR-222-3p and miR-222-5p (described in this study) predicts a poor overall survival ratio in patients with either clear or papillary cell subtype of renal carcinoma (Supplementary Fig. S1, Tables S2 and S3). Although it was not investigated in this study, a previous report has indicated that the circulating levels of miR-130b-5p, miR-151a-5p, miR-206, and miR-222-3p were significantly higher in sera collected from patients with breast cancer and the level of these circulating miRs was significantly downregulated after patients received surgical interventions [19]. Furthermore, miR-221 and miR-222 were found significantly upregulated in patients with metastatic RCC and associated with a poor progression free survival on sunitinib treatment [20]. Together, these findings and ours strongly suggested that circulating miR-222-3p could also play a prognostic role in the patients with RCC and our group is currently investigating this issue. Collectively, we demonstrated the increased expression of miR-222-3p in RCC clinical samples and cell lines and its positive correlation in promoting the metastatic potential of RCC cells.
Subsequently, we identified and verified TIMP2 as the target of miR-222-3p in both 769-P and 786-O RCC cell lines. Interestingly, TIMP2 has been previously shown to suppress both growth and metastasis in non-small cell lung cancer cells via suppressing the expression of metalloproteases [21]. This is in line with our observation where a decreased TIMP2 expression (by the increased miR-222-3p) was associated with an increased metastatic potential in both RCC cell lines examined. The increased expression of phosphorylated form of ERK1/2 was also correlated to the decreased miR-222-3p, suggesting its role as tumor suppressor. Our finding was also supported by a previous study in the prostate cancer, where phosphorylated ERK suppressed tumorigenesis via down-regulating STAT3 signaling and was associated with a better prognosis in the patients [22].
Our clinical association study demonstrated that miR-222-3p could be used as a prognostic marker for RCC where an increased level correlates a poor overall survival and tumor stage. Accumulating reports on miR-222 also support our view that the level of miR-222 was found to be higher in several cancer types including lung [23], pancreas [24], breast [25], bladder [26], and oral [27]. In light of this, we added another dimension where the EMT status can also be linked. For instance, when miR-222-3p expression was inhibited, vimentin expression was down-regulated, and E-cadherin expression was up-regulated, suggesting miR-222-3p could also be used as a prognostic marker for metastasis. However, the precise underlying mechanism by which miR-222-3p utilized to influence EMT expression requires further investigation. It is also cautioned that since there are several isoforms of miR-222 and selected 3’ isoforms could increase the prognostic capacity of this miRNA in RCC should be investigated in detail in future studies.
In summary, our study demonstrated that miR-222-3p functioned to promote renal cell carcinogenesis. Molecularly, a high miR-222-3p expression was associated with an increased metastatic potential and reduced apoptosis in vitro. We identified that miR-222-3p targets TIMP2 (a tumor suppressor) and ERK1/2 to convey its oncogenic functions. Collectively, our results strongly suggest that miR-222-3p can serve as a biomarker and therapeutic target for RCC.
Footnotes
Acknowledgments
This study was supported by grants from Zhejiang Province Bureau of Health (2015106257, and 2018ZB019) and the National Science Foundation of China (81502541).
Conflict of interest
The authors declare that they have no competing interests regarding this study.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-190264.
