Abstract
BACKGROUND:
The identification of high-sensitivity biomarkers for detection of hepatocellular carcinoma (HCC) from high-risk individuals is essential.
OBJECTIVE:
The present study was undertaken to identify and validate serum microRNAs (miRNAs) as potential biomarkers for hepatitis C virus (HCV)-related HCC.
METHODS:
Illumina sequencing was employed to screen the expression profiles of miRNAs in serum samples of HCV-related HCC patients and liver cirrhosis (LC) patients. RT-qPCR was used to confirm the altered miRNAs between the two groups. Moreover, candidate miRNAs were examined in serum samples of 40 HCC patients, 54 LC patients, 55 patients with chronic HCV hepatitis and 45 healthy controls. Receiver operating characteristic (ROC) curve analysis was used to evaluate the diagnostic performance of the miRNAs for the detection of HCC.
RESULTS:
Four miRNAs (miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p) were significantly increased and two miRNAs (miR-185-5p, miR-23b-3p) were significantly decreased in HCC patients compared to LC patients. ROC curve analysis demonstrated that the six miRNAs could be used as potential biomarkers for HCC detection. Combination of the six miRNAs could efficiently detect HCC in LC patients with the area under the ROC curve (AUC) of 0.995 and combination of the six miRNAs also provided high diagnostic accuracy (AUC
CONCLUSIONS:
The six serum miRNAs can be utilized as a surrogate and non-invasive biomarker for HCV-related HCC diagnosis.
Introduction
Hepatocellular carcinoma (HCC), the most prevalent type of liver cancer, has a very poor prognosis [1]. Hepatitis C virus (HCV) is a major etiologic agent of chronic hepatitis, cirrhosis and HCC in the world [2, 3, 4]. In China, the number of individuals infected with HCV is increasing and the incidence of HCV infection has increased from 0.7 to 15.0 cases per 100 000 persons in the past 10 years [3].
While great progress has been made in the treatment of HCC, the HCC prognosis remains poor and adverse effects of chemotherapy are common [1]. Early diagnosis of HCC and timely treatment can greatly improve life expectancy and reduce mortality. However, only 30% to 40% of patients with HCC are qualify for potential curative intervention at diagnosis because of the lack of clinical presentation and the abscence of potential early identification biomarkers [5]. Thus, new tumor markers with high sensitivity and specificity for HCC are urgently needed.
MicroRNAs (miRNAs) are small, noncoding regulatory RNAs. They inversely regulate the expression of their target genes at the posttranscriptional level by inhibiting translation or causing the degradation of the target messenger RNA (mRNA) [6]. MiRNAs are involved in various biological processes including proliferation, differentiation and apoptosis but also play an important role in the occurrence and development of a variety of diseases including HCC [7, 8, 9, 10]. Evidence has shown differential expression of miRNAs between tumor and normal tissue and miRNAs may act as tumour suppressors, oncogenes, or even possess a dual nature playing both roles depending on the cellular requirements [8, 10]. MiRNAs are tissue-specific and are stable in plasma or serum even after being subjected to severe conditions such as extreme temperature and low pH, which establish the role of miRNAs as non-invasive biomarkers in disease diagnosis [11, 12, 13, 14, 15].
In this study, we performed profiling of miRNAs by high throughput small RNA sequencing in order to identify a set of miRNAs served as efficient biomarkers for detection of HCV-related HCC from high-risk individuals. Candidate miRNAs were selected and validated by using a stem-loop RT-qPCR assay. A combined panel of six miRNAs (miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p, miR-185-5p, miR-23b-3p) was demonstrated could discriminate HCC efficiently from high-risk individuals.
Material and methods
Study design and patients
In the initial stage, pooled serum samples from 5 HCV-related HCC patients and 5 liver cirrhosis (LC) patients served as controls underwent Illumina sequencing to identify miRNAs that showed significant differences between the HCC patients and LC controls. Subsequently, we performed a biomarker confirmation analysis with an RT-qPCR assay. This analysis was carried out in 2 phases: (a) 15 HCV-related HCC patients and 15 LC patients formed the training set. (b) The selected miRNAs were examined by RT-qPCR in the serum samples from 40 HCC patients, 54 LC patients, 60 HCV carriers and 45 healthy controls.
HCV-related HCC patients, HCV-related LC patients and patients with chronic hepatitis C (CH) were recruited from our institution during the study period. All HCC patients were on top of HCV cirrhosis and HCC diagnosis was based on American Association for the Study of Liver Diseases (AASLD) Practice Guidelines [16] and staged by the Barcelona Clinic Liver Cancer (BCLC) system [17]. Patients with HCV-related LC were diagnosed by abdominal ultrasonography and confirmed by histopathology examination. Patients with CH were characterized by the persistent increase of the alanine aminotransferase (ALT) values (more than three times normal) for at least 6 months [18]. Patients with other neoplasm, virus infection or live disease were excluded from the study. 45 healthy volunteers without liver diseases and any tumors were served as healthy controls. The study is approved by the ethic committee of You’an Hospital Affiliated to Capital Medical University and patients signed the informed consent.
Serum collection and RNA extraction
Before curative treatment, blood samples were obtained. After centrifugation at 3000 revolutions/min for 15 min at room temperature, the supernatant was transferred into a new tube. The supernatant was then centrifuged at 15,000 revolutions/min using a high-speed refrigerated centrifuge for 30 min. Then, the supernatant sera were stored at
Demographic and clinical characteristics of the patients for Illumina sequencing in the profiling study
Demographic and clinical characteristics of the patients for Illumina sequencing in the profiling study
Length of distribution of total clean reads.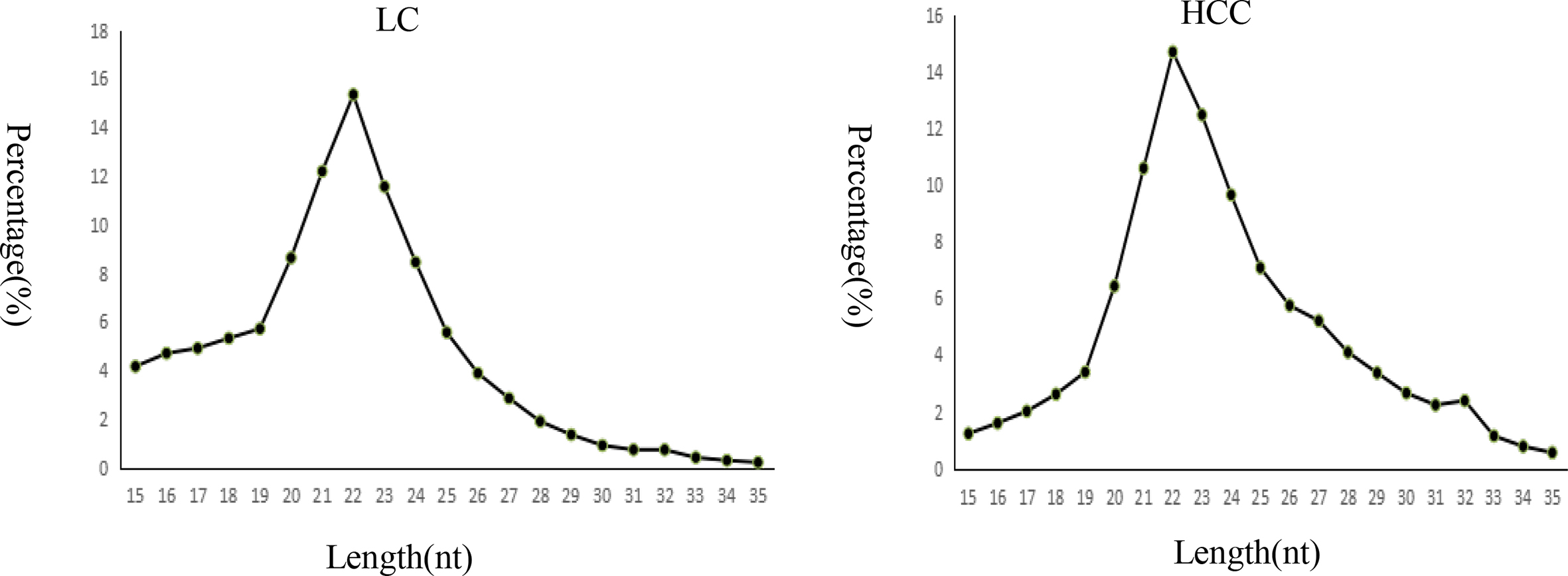
Through PAGE purification, total small RNA mole-cules under 30 bp were isolated. After ligating a pair of adaptors to their 5’ and 3’ ends, the small RNA molecules were amplified for 17 cycles and then fragments about 90 bp were isolated after gel purification. Illumina HiSeq 2500 was used for sequencing analysis according to the manufacturer’s instructions. Then we processed the data by computational analysis.
Quantification of miRNAs by RT-qPCR analysis
To validate the miRNAs discovered by sequencing, RT-qPCR was used. The RNA concentration was quantified with a DU 800 spectrophotometer (Beckman Coulter), Reverse transcription was carried out using a TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems) according to the manufacturer’s protocol: 16
Demographic and clinical characteristics of the patients in the first confirmation analysis set
Demographic and clinical characteristics of the patients in the first confirmation analysis set
MiRNAs values were normalized to cel-miR-39 and are expressed as 2-
Demographic and clinical characteristics of the patients in the second confirmation analysis set
Demographic and clinical characteristics of the patients in the second confirmation analysis set
Patients and sample characteristics
In the profile study, we included 5 HCV-related HCC patients and 5 LC patients as controls. The characteristics of the patients are shown in Table 1. In the first confirmation analysis set, 15 HCV-related HCC patients and 15 LC patients were included. The characteristics of the patients are shown in Table 2. There were no significant differences in the age, gender and transaminase between the two groups. The level of AFP was significantly higher in HCC patients than in LC patients. In the second confirmation analysis set, 40 HCC patients, 54 LC patients, 60 CH patients and 45 healthy controls (HC) were included. The characteristics of the patients are shown in Table 3. As showed by the table, patients differed across groups with regard to age (
Sequencing of serum miRNAs
We obtained 8,892,160 clean reads (60.84%) from 14,616,725 total reads in the LC group and 10,825,202 clean reads (72.49%) from 14,933,961 total reads in the HCC group (Table 4). Length distribution analysis showed that most reads were distributed around 22 nt (Fig. 1).
Sequencing data
Sequencing data
Ten miRNAs were selected from sequencing data
The expression quantities of the miRNAs (2-
Relative expression of miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p, miR-185-5p and miR-23b-3p in different groups (2-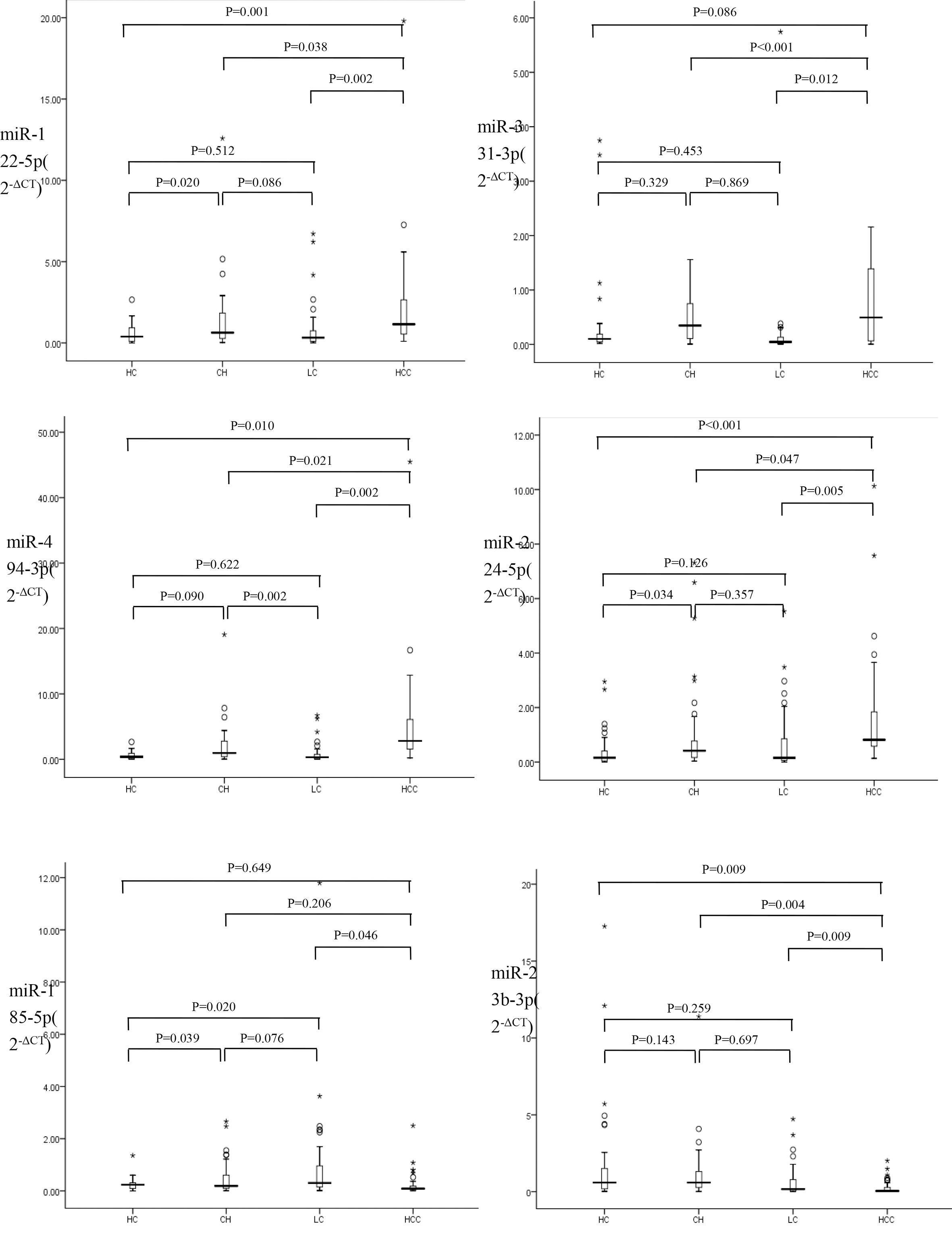
ROC curves for miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p, miR-185-5p and miR-23b-3p for discrimination of HCC from LC.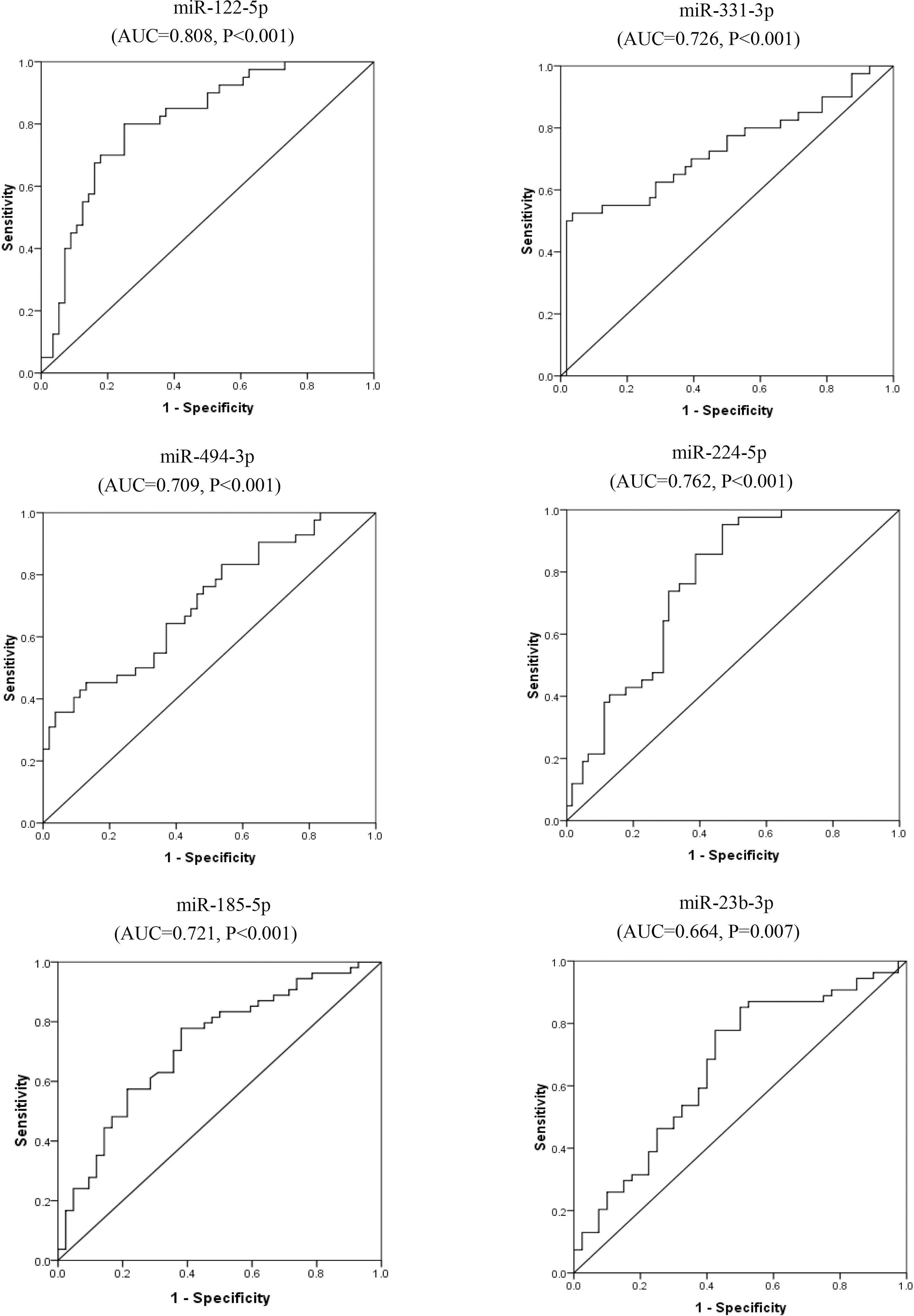
ROC curves for miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p, miR-185-5p and miR-23b-3p for discrimination of HCC from non-malignant controls (HC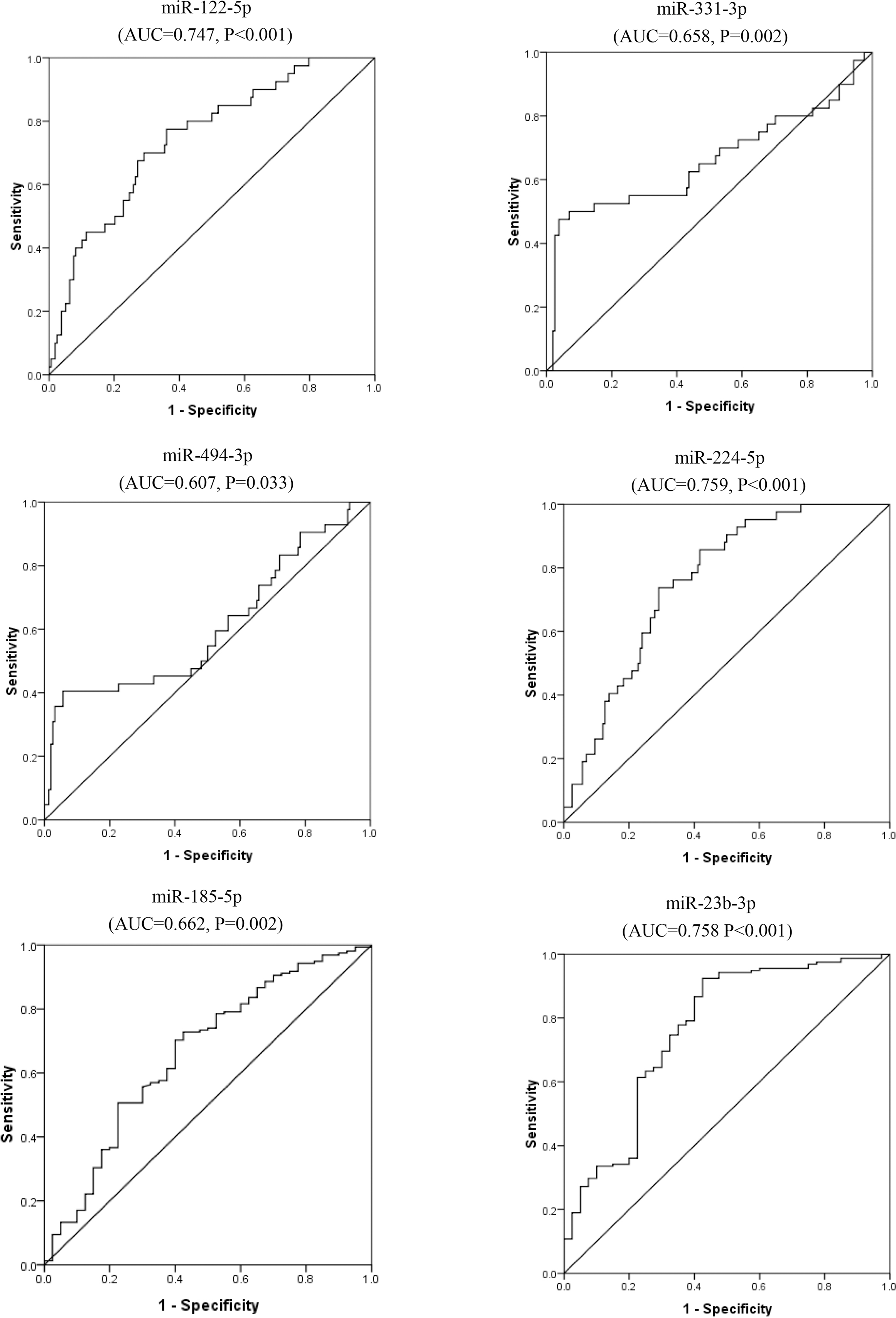
Significant relations among enrolled serum miRNAs in hepatocellular carcinoma patients
ROC curves for combination of miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p, miR-185-5p and miR-23b-3p for discriminate of HCC. A diagnosis of HCC against LC B diagnosis of HCC against non-malignant controls (HC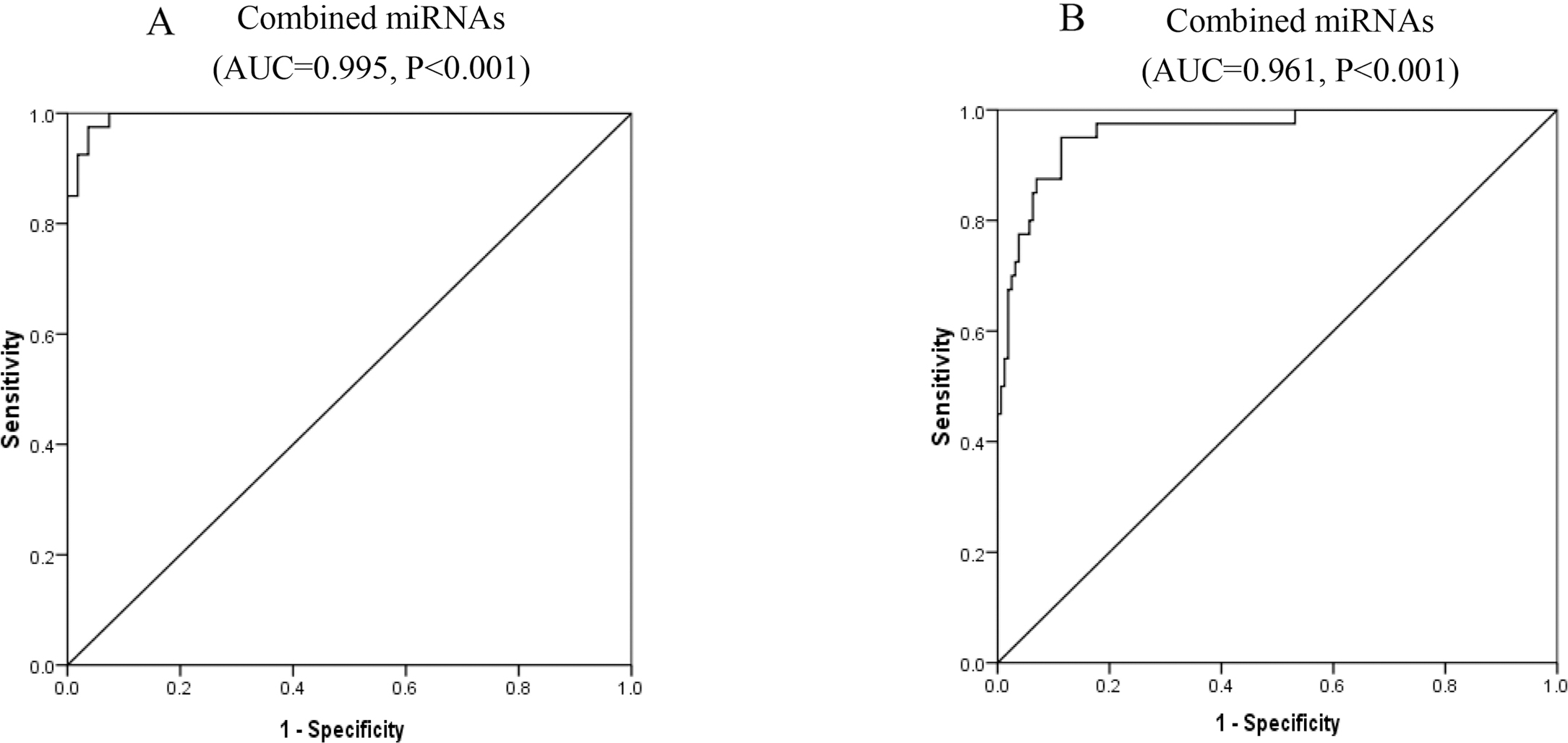
MiRBase database (release 21) was used to identify known miRNAs. 502 miRNAs were found in LC patients and 839 miRNAs were found in HCC patients. A miRNA was considered altered if sequencing detected 500 copies in any patient group and the miRNA showed at least a 1.5 log2 fold change or less than
We used RT-qPCR assay to confirm the expression of candidate miRNAs. In the first confirming set, 15 HCV-related HCC patients and 15 LC patients were used (Table 2).
The expression quantities of the miRNAs (2-
In the second conforming phase, the six selected miRNAs of miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p, miR-185-5p and miR-23b-3p were measured in all groups of HCC, CH, LC and HC. MiR-122-5p, miR-494-3p and miR-224-5p were all upregulated significantly early in CH patients versus health controls. The level of miR-494-3p was then significantly downregulated in LC patients compared to CH patients. The levels of the four miRNAs of miR-122-5p, miR-494-3p, miR-224-5p and miR-331-3p were all significantly higher in HCC patients versus LC patients or CH patients. Except miR-331-3p, levels of the other three miRNAs were also significantly higher in HCC patients versus health controls. The level of miR-185-5p was found significantly downregulated in CH patients and then upregulated in LC patients. In HCC patients the level of miR-185-5p was significantly downregulated compared to LC patients. No significant differences were found in levels of miR-23b-3p between CH, LC and health groups. MiR-23b-3p was significantly downregulated in HCC patients compared to LC patients, CH patients or health controls (Fig. 2).
ROC curve analysis
ROC analysis revealed that studied miRNAs could discriminate between HCC and LC with AUC
For discriminating HCC from non-malignant controls (HC
AUC of combined the six miRNAs for discriminating HCC from LC patients was 0.995 (95% CI 0.987–1.003,
Serum miRNAs’ correlation in the HCC study group
MiR-331-3p was correlated significantly with miR-122-5p (
Discussion
Patients with LC are known to be at high risk of developing HCC. Early diagnosis of HCC among these patients is important to reduce the mortality of the disease. Imaging examinations and biopsy are the major means for diagnosis of HCC at present. But imaging examinations can not distinguish the benign cancer and malignant cancer well. [23] and biopsy has the risk of tumor cells seeding along the needle track [24]. So biomarkers in peripheral blood can efficiently distinguish HCC are urgently needed.
MiRNAs, an integral component of gene regulation networks controlling tumorigenesis are stably maintained in a variety of biological fluids, including human serum/plasma [14, 15]. The quantities of circulating miRNAs reflect the physiological and pathological conditions of the human body and circulating miRNAs have been suggested candidates for a blood-based biomarker for diagnosis and prognosis in many cancers [11].
In the present study, we used small RNA sequencing technology as a discovery step, followed by RT-qPCR-based validation steps to discovery HCV-related HCC specific miRNAs. Our miRNA profiling analysis in sera from LC patients and HCC patients found tent miRNAs with different expressions. The further validation analysis confirmed that six miRNAs of miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p, miR-185-5p, miR-23b-3p were differentially expressed in serum of HCV-related HCC patients versus LC patients, implicating these miRNAs as surrogate biomarkers of HCC.
MiR-122, regulating lipid and cholesterol metabo-lism, comprises 70% of the total miRNAs in the liver. MiR-122 is important for HCV stability in liver cells and is known to regulate HCV stability by binding to an unstructured 5’UTR of HCV-RNA [25]. It has been reported that miR-122 serves as a tumor suppressor in HCC development by targeting multiple target genes such as metalloprotease 17 (ADAM17) [26], Bcl-w (anti-apoptotic gene) [27] and cyclin G1 (CCGN1) [28] and miR-122 expression was downregulated in HCC tumor tissues [28] while its serum levels were demonstrated significantly upregulated in HCC patients in comparison to LC patients or healthy controls [18, 29, 30, 31]. These contrary results may be due to its increased release [31, 32]. In the present study, miR-122-5p was demonstrated to be significantly upregulated in CH patients compared to healthy controls and the serum levels of miR-122-5p were upregulated significantly in HCV related HCC patients compared to LC patients, CH patients or healthy controls. Our findings also confirmed the role of miR-122 as a diagnostic marker for detection HCC in the LC group and control group.
In our study, levels of serum miR-494-3p were found upregulated significantly in HCC patients compared to all controls including LC patients, CH patients and healthy controls. MiR-494-3p was identified as a tumor driver in lung cancer [33], glioma [34], and HCC [35]. In HCC, miR-494-3p promotes cell proliferation, migration and invasion by targeting phosphatase and tensin homolog (PTEN) [35]. MiR-224-5p was also supposed an oncogenic miRNA during hepatocarcinogenesis and was demonstrated higher expressed in HCC tissue than paired adjacent noncancerous liver tissue [36]. Consistent with its tissue expression, in the current study, serum MiR-224-5p was demonstrated to be significantly unregulated in HCC patients compared to all controls. MiR-331-3p, another tumorigenic miRNA, promotes proliferation and metastasis of HCC through suppression of leucine-rich repeat protein phosphatase (PHLPP) mediated dephosphorylation of protein kinase B (AKT) [37]. In the present study, we found serum levels of miR-331-3p were upregulated significantly in HCC patients compared to CH patients and LC patients. Similar results were previously reported by Chen L, et al. [38] who demonstrated higher levels of serum miR-331-3p in HCC patients than patients with benign liver diseases. In the present study, we demonstrated higher levels of serum miR-494-3p, miR-224-5p and miR-331-3p in HCC patients, suggesting the roles of the miRNAs as potential biomarkers for HCC detection.
MiR-185-5p and miR-23b-3p were two downregulated serum miRNAs in HCC patients in our study. It has been reported that miR-185 acts as a tumor suppressor in HCC by targeting multiple genes [39, 40, 41]. Besides, recently Li et al. [42] found miR-185-5p was downregulated by HCV core protein, participating in the occurrence of liver steatosis which is a frequent histologic finding in CH. In the present study, serum miRNA-185-5p was downregulated significantly in HCC patients compared to LC patients, suggesting the diagnostic role of miRNA-185-5p in discriminating HCC from LC patients. MiR-23b-3p is a tumor suppressor which may regulate HCC migration and invasion by targeting Pyk2 via regulation of epithelial-mesenchymal transition (EMT) and may be a therapeutic target for HCC treatment [43]. In tissue, miR-23b-3p was reported aberrantly downregulated in HCC in contrast with the counterpart adjacent non-cancerous HCC [44]. Consistent with its reported tissue expression, levels of serum miRNA were significantly downregulated in HCC patients compared to all controls including LC patients, CH patients and healthy controls in our study. This suggests that serum miRNA may be a novel, non-invasive biomarker for the diagnosis of HCC.
To further analyze the diagnostic capacities of miR-122-5p, miR-331-3p, miR-494-3p, miR-224-5p, miR-185-5p, miR-23b-3p, receiver operating characteristic curves were done. Areas under the curves showed all of the six miRNAs could significantly discriminate HCV-related HCC from LC or from non-malignant controls. For discriminating HCC from LC, the AUC of miR-122-5p was largest with 0.808, second by miR-224-5p with AUC of 0.762. For discriminating HCC from non-malignant controls, the AUC of miR-224-5p was largest with 0.758, second by miR-23b-3p with AUC of 0.762. Combined, the six miRNAs could efficiently discriminate HCC from LC with AUC of 0.995 and AUC of 0.961 to discriminate HCC from non-malignant controls.
In conclusion, six miRNAs (miR-122, miR-331, miR-494, miR-224, miR-185 and miR-23b) were identified that may be useful as biomarkers of HCV-related HCC, especially for high-risk individuals. When used in combination the six miRNAs had a high degree of accuracy in detecting HCC in high risk individuals. Further studies may be indicated to validate these findings.
Footnotes
Acknowledgments
This study was funded by the National Grand Program on Key Infectious Diseases (No. 2015ZX100048 01).
Conflict of interest
The authors declare no conflict of interest.
