Abstract
N-myc downstream regulated gene 2 (NDRG2) is frequently down-regulated in various cancers and functions as a candidate tumor suppressor gene. NDRG2 has been shown to be SUMOylated on the lysine 333 residue, which promoted its ubiquitination and sequentially degradation by the SUMO-targeted ubiquitin E3 ligase RNF4. However, how to regulated NDRG2 deSUMOylation process remains largely unknown. Here, we report that Sentrin/SUMO specific protease (SENP2) was down-regulated in clinic gastric cancer samples and possessed a tumor-suppressive role in gastric cancer. At the molecular level, we found that SENP2 interacts with NDRG2 and mediates the de-SUMOylation process of NDRG2. Overexpression of SENP2 stabilized NDRG2, whereas silencing SENP2 caused rapid NDRG2 SUMOylation and degradation, indicating SENP2 antagonizes NDRG2 ubiquitination and degradation, thereby promoting the stability and function of this protein. Thus, our study reveals that SENP2 acts as a tumor suppressor which is deregulated in gastric cancer and the specific de-SUMOylation activity of SENP2 for NDRG2 is critical for it stabilization as well as gastric cancer cells proliferation.
Introduction
Gastric cancer (GC) is the third leading cause of cancer-related death worldwide and most gastric cancer patients are already diagnosed with late stage disease [1, 2, 3]. Despite research on GC has made some progress in recent years, the 5-year survival of patients with GC still not as satisfactory as expected [4, 5]. Thus, the exact molecular mechanisms still need further study.
SUMO is a small ubiquitin-like protein that can covalently modify a large number of proteins to alter their function, and localization and stabilization [6, 7]. SUMOylation is a highly dynamic and reversible process that is catalyzed by a activating (E1), conjugating (E2) and ligating (E3) enzyme cascade and reversed by a family of six SUMO-specific proteases (SENPs) [8, 9]. SENP2 is a nuclear envelope-associated protease that deconjugates a large number of SUMOylated proteins
N-myc downstream-regulated gene 2 (NDRG2) is involved in multiple biological processes, including cell growth, differentiation, as well as apoptosis [16, 17]. NDRG2 functions as a tumor suppressor and its expression is down-regulated in various cancers, including breast cancer, colon cancer, melanoma, glio- blastoma, thyroid cancer and pancreatic cancer [16]. It has been reported that NDRG2 can be transcriptionally activated by p53 and involved in p53-mediated apoptosis pathway in certain tumor cells [18, 19]. Recently, NDRG2 is found to be a SUMOylated protein and modified by SUMO1 at K333 [20]. SUMOylated NDRG2 was rapidly underwent ubiquitination and sequentially degradation by the ubiquitin E3 ligase RNF4 [20, 21]. However, the specific de-SUMOylation protease that can reverse the SUMOylation of NDRG2-mediated processes is still unidentified.
Here, we show that SENP2 interacts with NDRG2 and mediates the de-SUMOylation process of NDRG2 to stabilize NDRG2. Moreover, SENP2 was down-regulated in gastric cancer samples which in turn caused NDRG2 destabilization in gastric cancer.
Materials and methods
Cell lines and clinic samples Seven human GC cell lines (NCI-N87, MKN74, AGS, NUGC-3, MKN45, MGC803, HGC-27) were obtained from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). All cell lines were maintained in RPMI-1640 (Gibco BRL, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (FBS) except AGS in Ham’s F12 medium (Cellgro, Manassas, VA, USA) and incubated at an atmosphere containing 5% CO
Seven human GC cell lines (NCI-N87, MKN74, AGS, NUGC-3, MKN45, MGC803, HGC-27) were obtained from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). All cell lines were maintained in RPMI-1640 (Gibco BRL, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (FBS) except AGS in Ham’s F12 medium (Cellgro, Manassas, VA, USA) and incubated at an atmosphere containing 5% CO
Five human GC cell lines (AGS, BGC823, MGC803, NCI-N87, and SGC7901) were obtained from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). BGC823, MGC803, NCI-N87, and SGC7901 cells were maintained in RPMI-1640 (Gibco, NY, USA) supplemented with 10% fetal bovine serum (FBS); AGS cells were maintained in F12 medium (Gibco, NY, USA) with 10% FBS; 293T cells were maintained in DMEM with 10% FBS. All cells were incubated at an atmosphere containing 5% CO
Cell proliferation assay
The cell proliferation was detected by 3-(4, 5-dime- thylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) assay. AGS cells were seeded into 96-well plates at 2000 per well and incubated at 37
SENP2 is down-regulated in clinic gastric cancer samples. A. The mRNA expression levels SENP2 in 20 paired GC tissues and adjacent normal tissues were normalised to the expression of beta-actin and calculated by the 2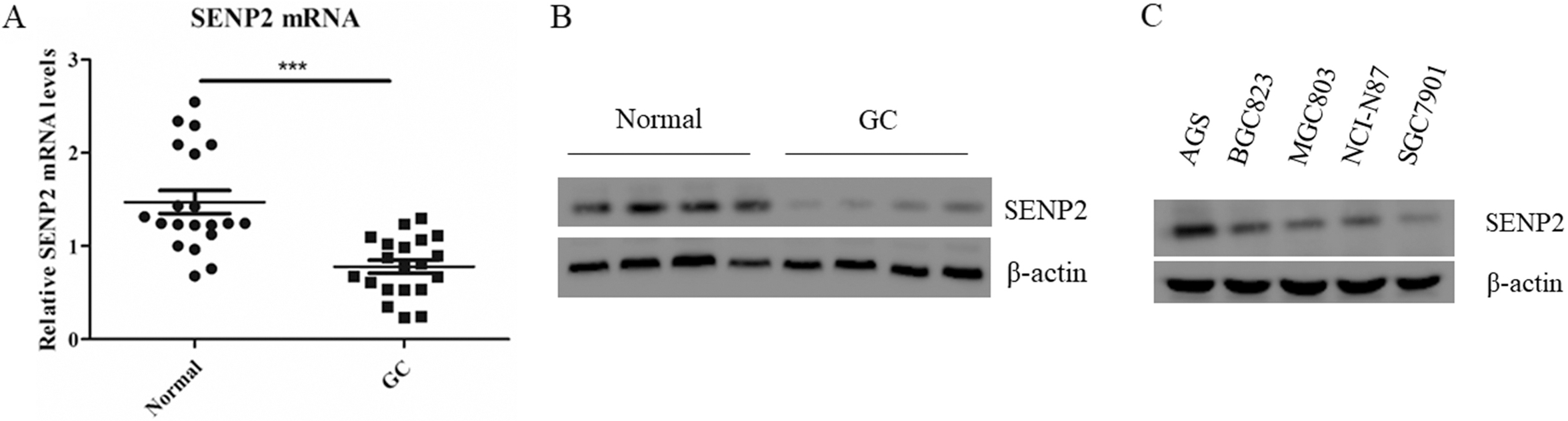
Total RNA was extracted using RNAeasy™ Animal Total RNA Isolation Kit with Spin Column (Beyotime, China) according to the manufacturer’s instructions. Reverse transcription PCR (RT-PCR) and quantitative PCR was carried out by using a SYBR Green Premix Ex Taq (Takara) on Light Cycler480 (Roche, Switzerland). Primers were selected from PrimerBank (
Immunoprecipitation (IP)
Cells were lysed in RIPA buffer for 30 min with gentle rocking at 4
Western blots
Cells were lysed by 2
Colony formation
For colony formation assay, AGS cells were seeded in six-well plates contained a base layer consisting of 0.5 ml culture media (0.6% agar). After appropriate time of growth, the cells were washed with PBS, fixed with 4% paraformaldehyde, and stained with 0.1% crystal violet. Photographs were captured and cell clones were counted.
Statistical analysis
Values are shown as mean
SENP2 inhibits gastric cancer cells proliferation. A. The mRNA levels of SENP2 were determined by Real-time PCR in AGS cells stable expressing the indicated shRNAs. B. AGS cells stable expressing the indicated shRNAs were harvested and subjected to western blot with anti-SENP2 antibody. C. The cell growth curve of AGS cells stable expressing the indicated shRNAs. D. The colony formation ability of AGS cells stable expressing the indicated shRNAs.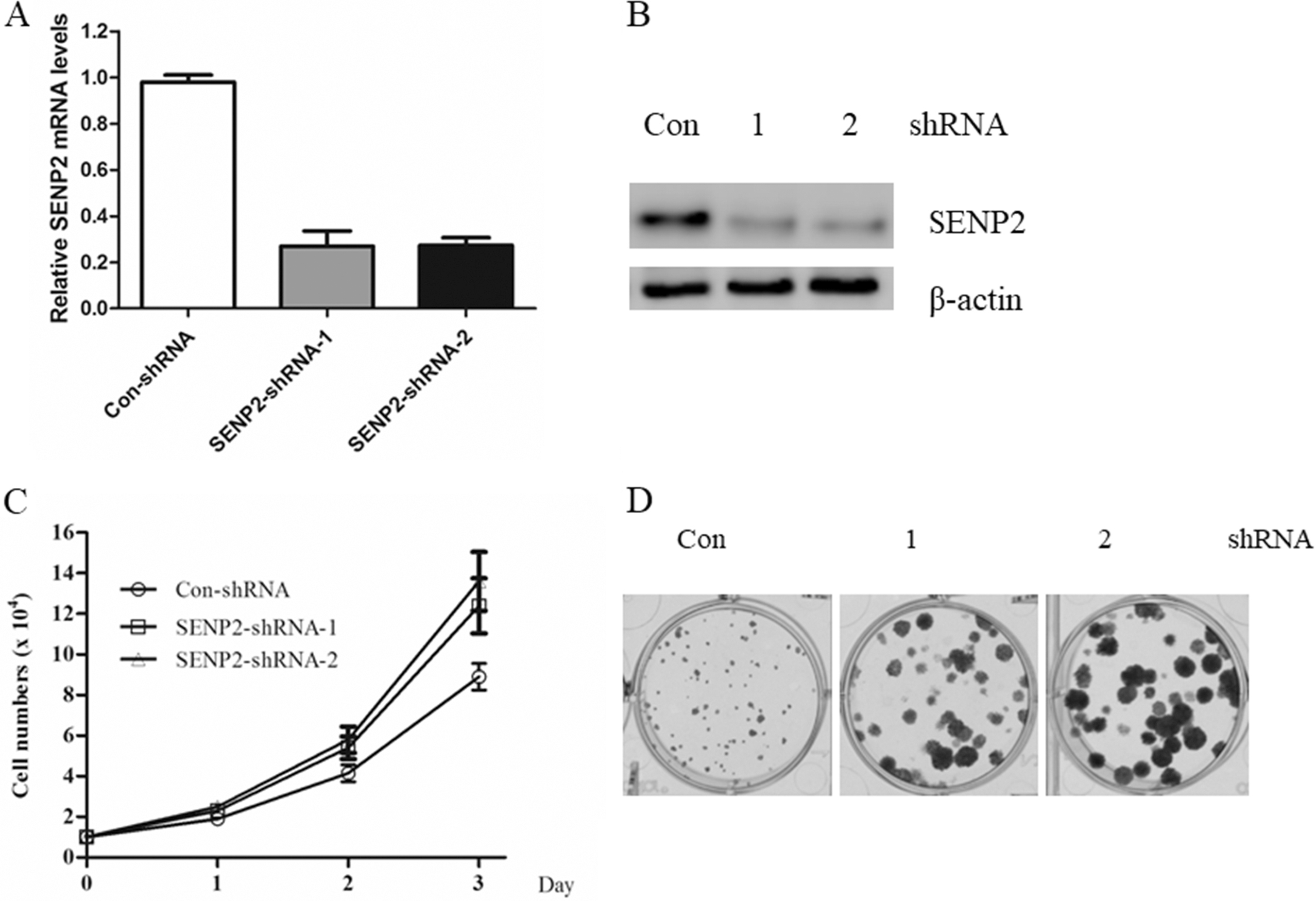
SENP2 interacts with NDRG2 in gastric cancer cells. A. 293T cells were transfected with indicated plasmids for 48 hours. Cell lysate were subjected to Flag M2 beads IP followed by western blot with HA antibody. B. AGS cells lysate was subjected to IgG or NDRG2 IP followed by western blot with anti-SENP2 antibody. C. BL21 bacteria expressing His or His-NDRG2 were incubated with Flag-SENP2 cell lysate and then subjected to western blot with anti-Flag antibody.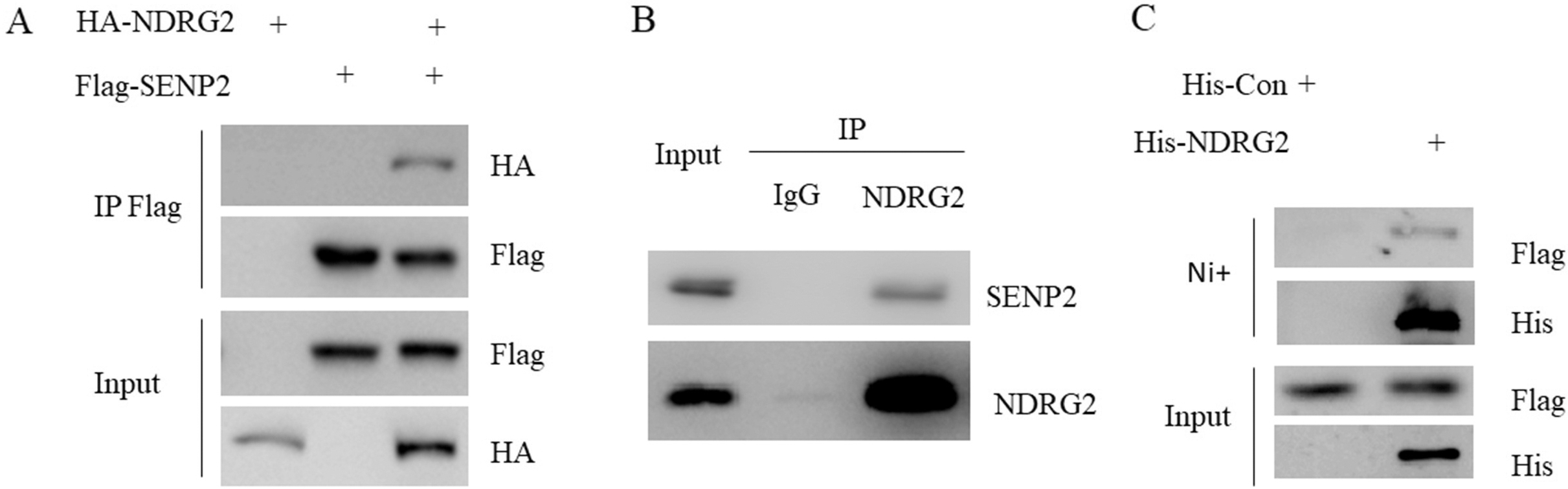
Overexpression of SENP2 increased NDRG2 abundance. A–F: AGS cells were transfected with Flag-SENP1(A), Flag-SENP2(B), Flag-SENP3(C), Flag-SENP5(D), Flag-SENP6(E), Flag-SENP7(F), respectively and subjected to western blot with anti-NDRG2 antibody.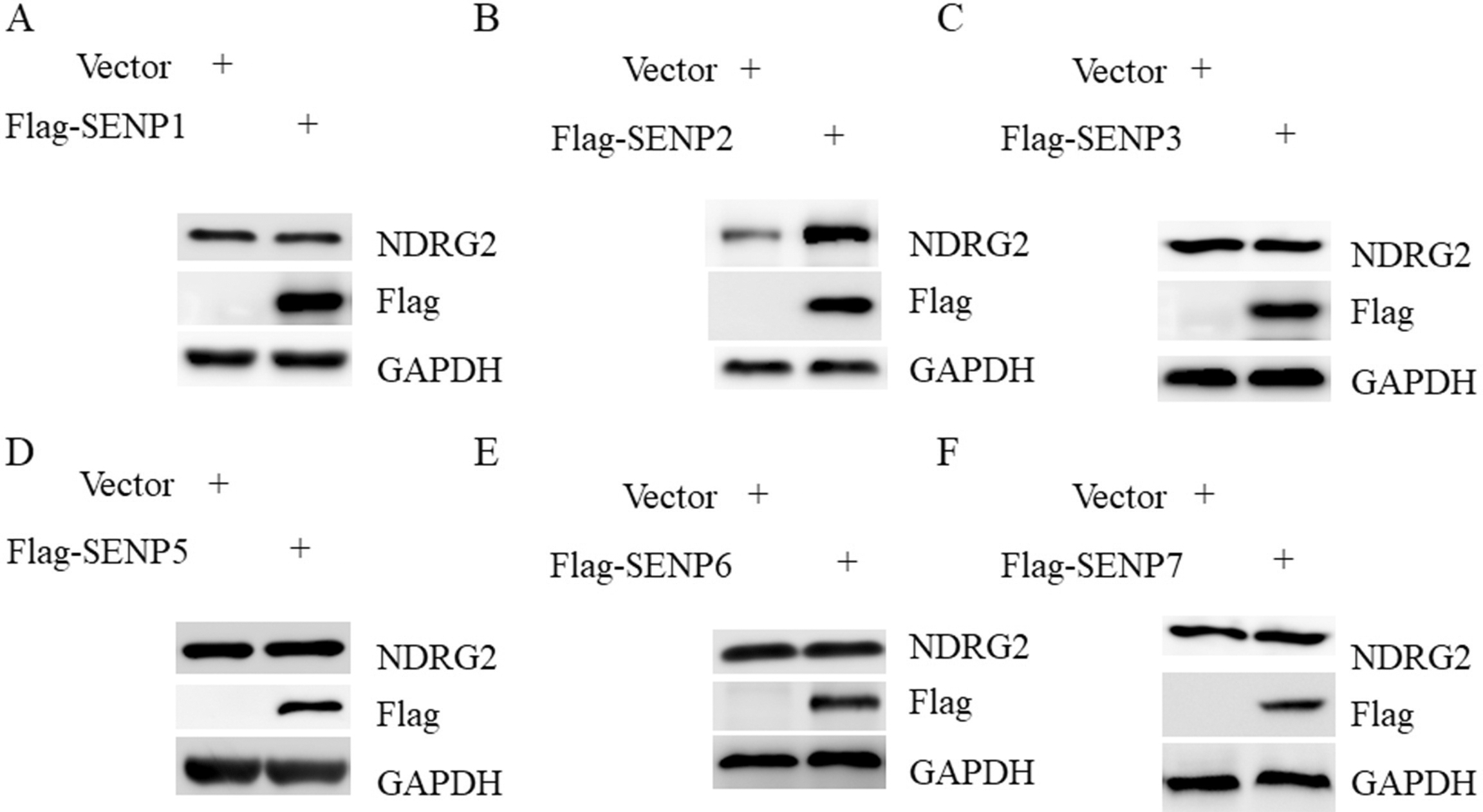
Silencing of SENP2 destabilized NDRG2. A. AGS cells with control or SENP2 knockdown were subjected to western blot with indicated antibodies. B. AGS cells with SENP2 knockdown were treated with proteasome inhibitor MG132 for 4 hours and subjected to western blot with indicated antibodies. C. AGS cells with control or SENP2 knockdown were treated with 50 ug/ml CHX for the indicated time and subjected to western blot with indicated antibodies. D. AGS cells lysate with control or SENP2 knockdown were subjected IP with anti-NDRG2 antibody followed by western blot with anti-ubiquitin antibody.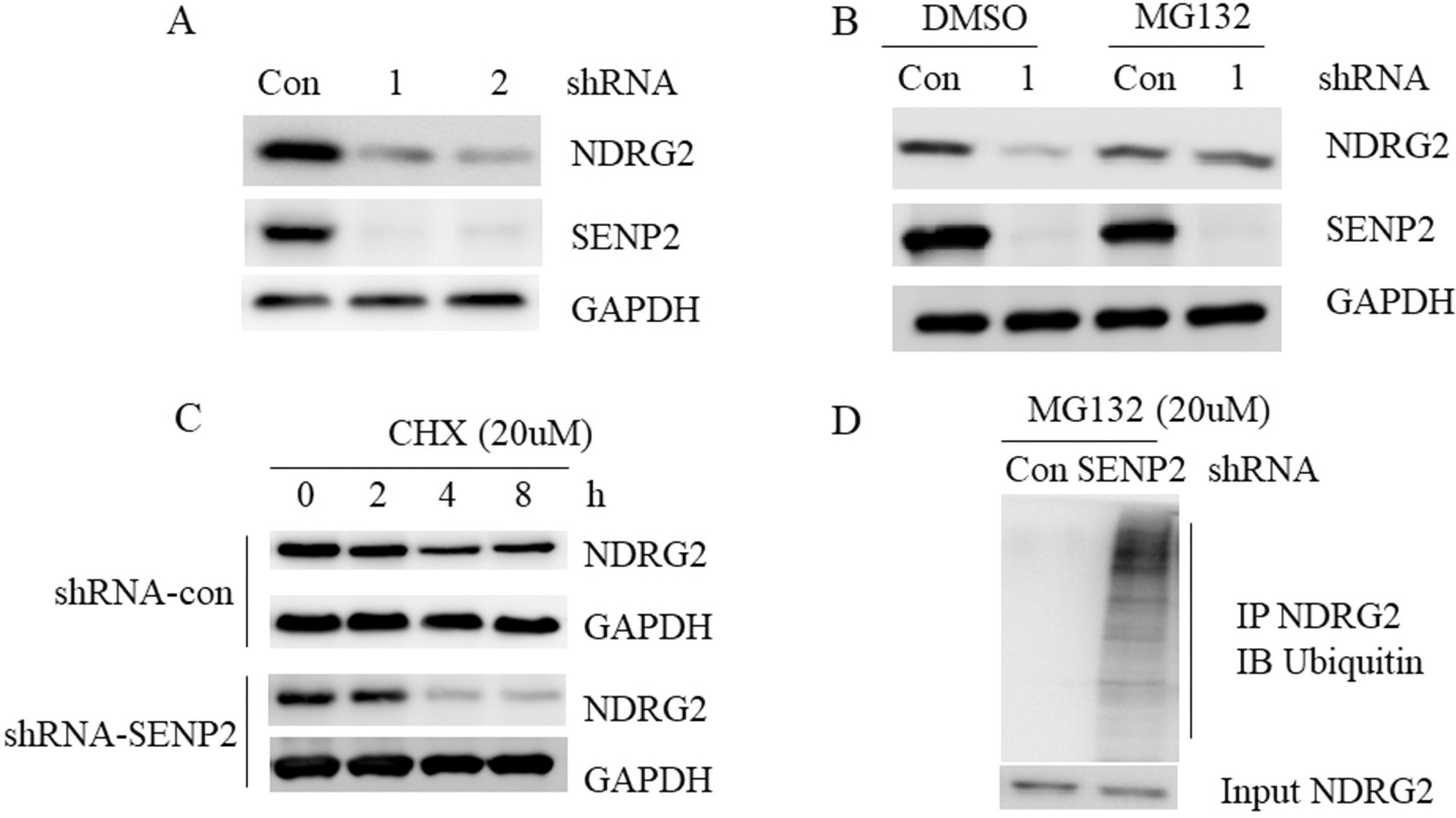
SENP2 deSUMOylates NDRG2. A. 293T cells were transfected with indicated plasmids for 48 hours. Cell lysate were subjected to Flag M2 beads IP followed by western blot with Myc antibody. B. AGS cells lysate with control or SENP2 knockdown were subjected IP with anti-NDRG2 antibody followed by western blot with anti-SUMO1 antibody.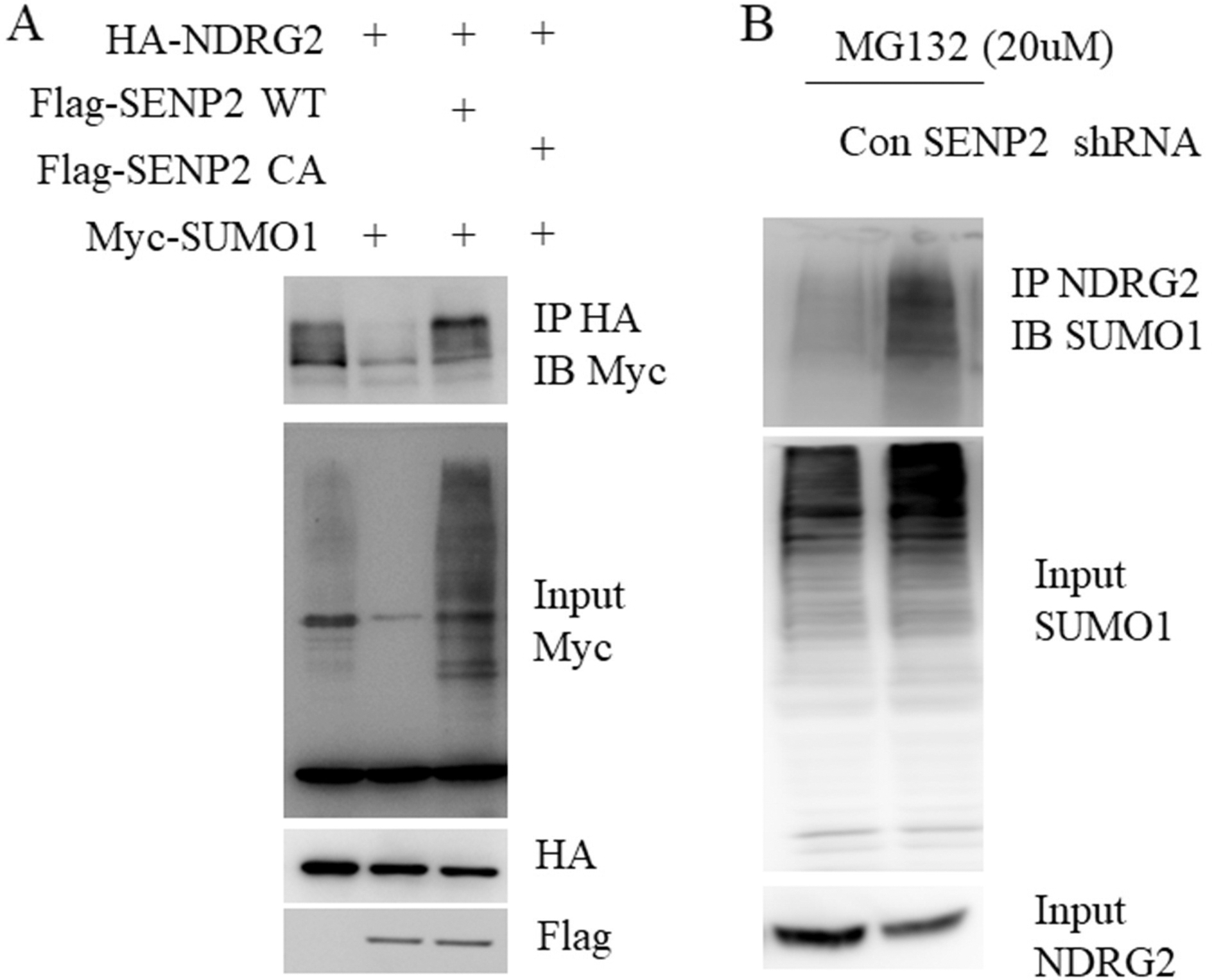
SENP2 is down-regulated in clinic gastric cancer samples
The mRNA expression of SENP2 was determined by real-time PCR in 20 GC and paired adjacent normal tissues. We found that SENP2 was down-regulated in 20 GC samples compared with paired adjacent normal samples (
SENP2 inhibits gastric cancer cells proliferation
To understand the biological role of SENP2 in GC cells, we silenced its expression in GC cell line AGS cells by two different shRNAs against human SENP2. The knockdown efficiency of SENP2 was determined by real-time PCR as well as western blot. We found that both shRNAs decreased the expression of SENP2 efficiently (Fig. 2A and B). Thus, both shRNAs were used to investigate the biological roles of SENP2 in GC cells. As shown in Fig. 2C, silencing of SENP2 significantly promoted cells growth. The colony formation ability of AGS cells were also enhanced by SENP2 knockdown (Fig. 2D). Taken together, these data indicated a critical role of SENP2 in the regulation of GC cells growth.
SENP2 interacts with NDRG2 in gastric cancer cells
By searching several large protein-protein interaction databases, we found that NDRG2 was one of the binding partners of SENP2 [22]. 293T cells were transfected with Flag-SENP2 or HA-NDRG2 or both for 48 hours. Cell lysate were subjected to Flag M2 beads immunoprecipitation followed by western blot with HA antibody. Indeed, we found that HA-NDRG2 was detected in Flag-SENP2 immunoprecipitant (Fig. 3A). Next, by using AGS cell lysate and NDRG2 antibody, we found that endogenous SENP2 was also presented in endogenous NDRG2 immunoprecipitant (Fig. 3B). To test whether NDRG2 interacted directly with SENP2, we further performed in vitro pull down assay. Protein complexes pulled down with nickel beads from BL21 bacteria expressing His or His-NDRG2 incubated with Flag-SENP2 were subjected to western blot. Flag-SENP2 was ready to be detected in His-NDRG2 but not His alone complexes (Fig. 3C). These data indicated that SENP2 interacted with NDRG2 directly.
Overexpression of SENP2 increased NDRG2 abundance
Because both SENP2 and NDRG2 exhibited tumor suppressor function, we then sought to identify the potential mechanisms. NDRG2 has been found to be an unstable protein in tumors, we then asked whether SENP2 could regulate the stability of NDRG2. To this end, we overexpressed all the six SENPs into AGS cells, respectively. We found that SENP2 was the only member that increased the protein amount of NDRG2 in GC cells when overexpressed (Fig. 4A–F). These data suggested a highly specific role of SENP2 in the regulation of NDRG2 in GC cells.
Silencing of SENP2 destabilized NDRG2
Next, we found that NDRG2 protein was hard to detect in AGS cells with SENP2 knockdown (Fig. 5A), but recurrent in the present of proteasome inhibitor MG132 (Fig. 5B), suggesting that SENP2 could prevent NDRG2 from ubiquitin-mediated destruction. Indeed, our cycloheximide (CHX) experiment showed that SENP2 knockdown facilitated NDRG2 protein degradation (Fig. 5C). In agreement, knockdown of SENP2 dramatically enhanced polyubiquitination of endogenous NDRG2 (Fig. 5D). Taken together, these data suggested that SENP2 plays a critical role in the regulation of NDRG2 stabilization.
SENP2 deSUMOylates NDRG2
Recently, NDRG2 was shown to be SUMOylated by RNF4 and we have observed that SENP2 could bind to and stabilize NDRG2. Therefore, we reasoned that SENP2 could be a deconjugation protease for SUMOylated NDRG2. To test this possibility, we performed a de-SUMOylation assay in the transfected 293T cells. As shown in Fig. 6A, the expression of SENP2 wild-type (WT) but not the SENP2 catalytic mutant (CA) significantly disappeared the SUMOylated form of NDRG2. Moreover, we could also observed the accumulation of the SUMOylated NDRG2 in SENP2 knock down (KD) AGS cells in the present of MG132 (Fig. 6B).
Discussion
The NDRG family has three members, including NDRG1, NDRG2, NDRG3 and NDRG4 [23]. This protein family is characterized by an esterase/lipase/ thioesterase active site serine and an
In the present study, we found that SENP2 was associated with tumor suppressor NDRG2 and mediated NDRG2 deSUMOylation in GC cells. As RNF4-mediated NDRG2 is required for its sequential ubiquitination and degradation, our data suggest SENP2 could stabilize NDRG2 by removing its SUMO chain and preventing the attachment of ubiquitin chain. In consistent with it, the half-life of NDRG2 was largely extended and the protein content of NDRG2 was greatly increased in the present of SENP2. In contrast, NDRG2 protein could be barely detected in SENP2 KD cells and its SUMOylation form was easily showed up in the present of a proteosomal inhibitor. We also observed that SENP2 was down-regulated in GC samples and cell lines and SENP2 obviously exhibited a tumor suppressor role in GC cells.
Thus, our study reveals that SENP2 acts as a tumor suppressor and SENP2-mediated NDRG2 deSUMOylation is critical for NDRG2 stabilization as well as GC cells proliferation.
Footnotes
Conflict of interest
None.
