Abstract
Studies have shown that microRNAs (miRNAs) can promote or suppress tumor growth and therefore act as targets for cancer therapy. Hsa-miR-503-5p, a mature miRNA derived from 5’ ends of pre-miR-503, has been proved to regulate cell proliferation, transformation, migration and invasion. However, the biological function of miR-503-3p derived from 3’ ends of pre-miR-503 has never been reported. In current study, we found that miR-503-3p inhibits lung cancer cell viability and induces cell apoptosis. To better understand the molecular mechanism underlying the miR-503-3p participating in this process, PCR array and RNA-sequencing (RNA-seq) were performed and some differential expression genes were discovered between NC and miR-503-3p treated groups. Biological interaction network showed that p21 and CDK4 are the most important proteins involving miR-503-3p signal pathway. Dual-luciferase assay results shown miR-503-3p directly regulates the expression of p21 by targeting 3’-UTR of its mRNA. These results shed light on the potential roles of miR-503-3p, indicating that it may act as an anti-oncogene factor to inhibit lung cancer cell viability.
Introduction
miRNAs belong to the heterogeneous class of ncRNAs, of 22–24 Int RNAs that play important regulatory roles by targeting mRNAs for cleavage or translational repression [1]. Frequent aberrant expression of miRNAs has been reported to commonly occur in several human cancers [2, 3, 4, 5, 6]. The dysfunction of miRNAs in cancer has raised these small cellular components to the ranks of preferred drug targets. Modulating miRNA activities in cancer may provide exciting opportunities for cancer therapy [7, 8, 9].
In humans, the primary transcript of mir-503 is excised and generated two different mature miRNA, hsa-miR-503-5p and hsa-miR-503-3p. These 5p and 3p miRNAs, generated from a single primary transcript, have different target mRNAs and different, even inverse roles in biological process [10, 11]. miRNA-503-5p has been proved to regulate cell proliferation, transformation, migration and invasion by targeting L1CAM, DDHD2, IGF1R and BCL2, PI3K p85 and IKK-
In this study, we found that the over-expression of miR-503-3p significantly inhibited lung cancer cell viability and induced cell apoptosis. To better understand the molecular mechanism underlying the miR-503-3p participating in this process, PCR array and RNA-sequencing (RNA-seq) were preformed to search for differential expression genes between miR-503-3p mimics and negative control (NC) treated cell groups. Our results indicate that p21 and is the target genes of miR-503-3p and may play an important role in the process of miR-503-3p regulating the cell growth.
Materials and methods
Cells and cell culture
Human NSCLC cell lines H292, H358 and H1975 were propagated in RPMI-1640 (Gibco, USA). The medium were supplemented with 10% fetal bovine serum (Hyclone, USA), 100 U/ml penicillin, and 100 U/ml streptomycin (Amresco, USA). Cell culture was incubated at 37
Transient miRNA transfection
According to the sequence of hsa-miR-503-3p (Accession number: MIMAT0022925), a miRNA mimics (sequence: GGG GUA UUG UUU CCG CUG CCA GG) was designed and synthesized by Genepharma (Shanghai, China). Double-stran-ded scrambled RNA (sequence: UUC UCC GAA CGU GUC ACG UTT) was used as the negative control (NC). For transfection, cells were seeded into each well and grown for 24 h until they were 30–50% confluent. Cells were washed and then placed in Opti-MEM serum-free medium and transfected with oligo-nucleotides using Lipofectamine™ 2000 Reagent (Invitrogen, USA) according to the manufacturer’s protocol. After 4–6 h, the medium was changed to normal RPMI-1640 or DMEM medium containing 10% serum, and the cells were continuously cultured at 37
MTT cell proliferation assay
Cells were plated in 96-well plates and transfected with NC and miR-503-3p mimics. After 3–5 days, relative viable cell levels were determined by the MTT assay. The cells were incubated with MTT solution (0.5 mg/mL, 10
Colony formation assay
The clonogenic survival assay was conducted as described previously [16]. In brief, cells were harvested by trypsinization and resuspended in RPMI-1640 medium. An appropriate number of cells were plated into each 60 mm dish to produce colonies. After incubating for 14 days, the formed cell colonies were fixed and stained with 0.5% crystal violet for 20 min. Colonies containing
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was extracted using TRIzol (Invitro-gen, USA) according to the manufacturer’s instructions. The expression of mature miR-503-3p was quantified by All-in-OneTM miRNA qRT-PCR Detection Kit (Genecopoeia, China). Primers for mature miR-503-3p (HmiRQP3217) were purchased from genecopoeia (Guang-zhou, China). U6 was used as an internal control to normalize RNA input, and U6 primer (F: 5’-CTCGCTTCGGCAGCACA-3’, R: 5’-AACGCTTCACGAATTTGCGT-3’) were synthesized by Sheng-gong Biotech (Shanghai, China). The transcript levels of other genes were quantified by RT2 Profiler PCR Arrays Kit (Qiagen, German) according to the manufacturer’s instructions. PCR array analysis was performed in three independent experiments, each using three independent samples. The results of real-time were analyzed by the
Apoptosis assay
Apoptosis analysis was performed in transf-ection and control cells by staining with the Annexin V-FITC Apoptosis Detection kit (Bes-tBio, China). In brief, cells were harvested at a density of 1
Cell-cycle analysis
For cell cycle analysis, transfected cells were harvested by trypsinization 48 h post-trans-fection, wash-ed with PBS twice, fixed in 70% ethanol overnight at 4
RNA-seq
RNA-seq was performed by GUANGZHOU RIB-OBIO CO., LTD (China). Briefly, mRNAs were isolated from total RNA and fragmented to appr-oximately 200 bp. Subsequently, the collected mRNAs were subjected to first strand and second strand cDNA synthesis following by adaptor ligation and enrichment with a low-cycle acc-ording to instructions of TruSeq
Analysis of RNA-seq data
Before read mapping and assembly, poor quality data were filtered out so as to obtain high quality data (Clean Data). For differentially expressed genes, we carried out functional annotation analysis using DAVID [17]. Differentially expressed genes were used as input gene list, and all human genes that were expressed in the 786-O cells were used as the background. We looked for enrichment for genetic association with disease class, KEGG pathways, and biological processes in Gene Ontology (GO). Multiple testing was adjusted using Benjamini approach, and enrichment was declared if Benjamini adjusted
Construction of a protein interaction network
Protein interaction network was constructed accord-ing following steps: firstly, we integrate the famous protein interaction network database [18] and quantify correlations of protein-protein interaction between two protein coding 10954 genes with respect to their relative positions in genomes [19]. Next, proteins interaction prediction was performed in accordance with a computational method to investigate the int-eraction of protein pairs [20] and the interaction network was optimized on the basis of the messenger RNA co-expression database [21]. Combined with the data from Gene Ontology, KEGG, Pfam, and so on, the protein interaction network was constructed [22, 23, 24].
Analysis of miR-503-3p suppressing cell proliferation by MTT assay. A. The effect of miR-503-3p on the proliferation of lung cancer cell lines. Different concentration of miR-503-3p mimics was used for transfection, and NC mimics (100 nM) act as negative control. B. The transcript level of miR-503-3p was detected in H292, H358 and H1975 cells. Each experiment was conducted at least three times independently. (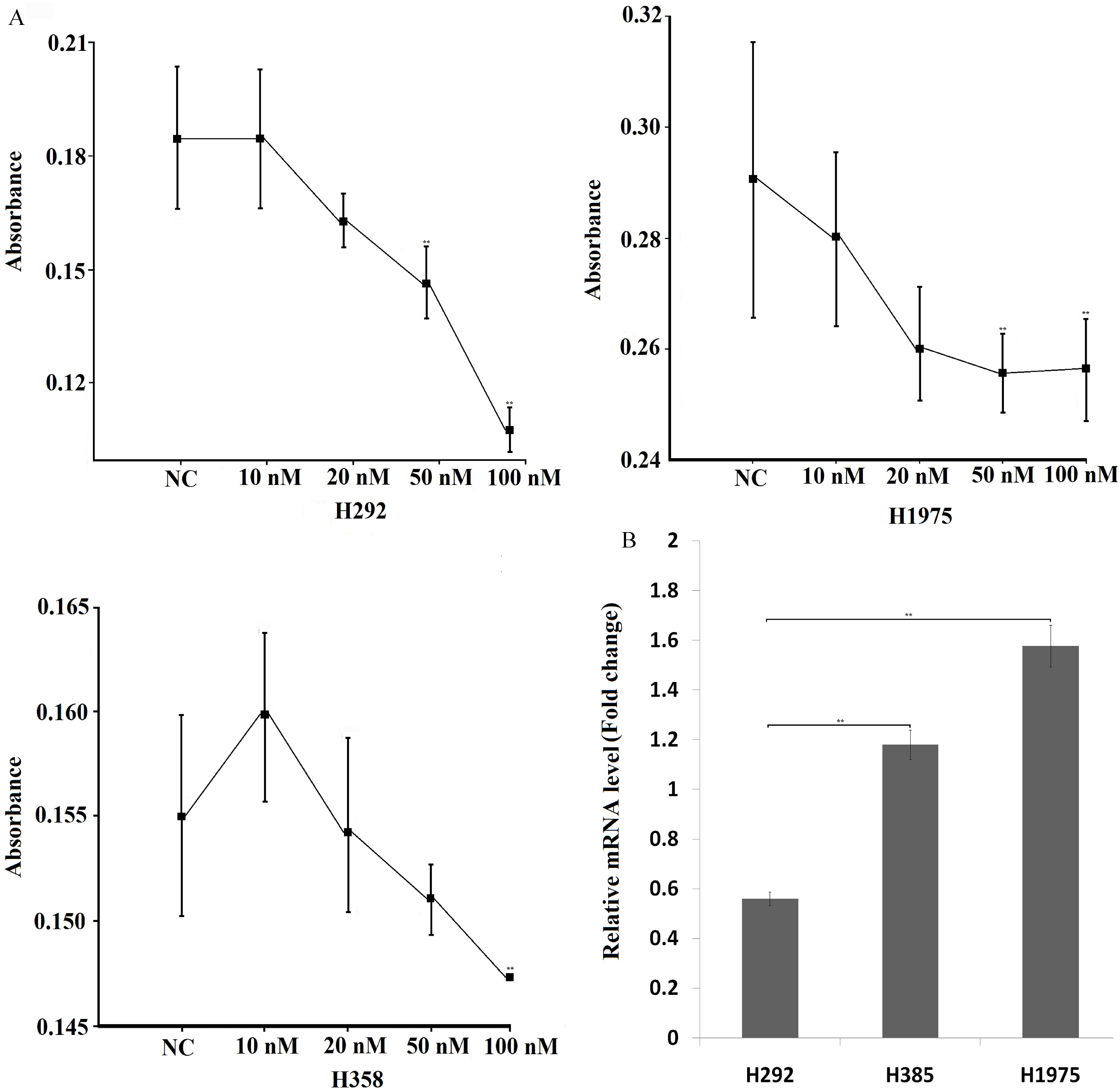
P21-3’-UTR fragment containing predicted target site of miR-503-3p and p21-3’-UTR fragment with mutated target site of miR-503-3p were chemically synthesized from Sangon Bio-tech (Shanghai, China). These fragments were annealed and inserted into the pmirGLO Vector (Promega). 786-O cells were then co-transfected with 200 ng reporter vector and 50 nM miR-503-3p mimics by using Lipofectamine 2000 in a 96-well plate (Corning). The activities of firefly and renilla luciferase in cell lysates were assayed by using Dual-Glo Luciferase Assay System (Promega) at 24 h post-transfection, and the firefly luciferase activity was normalized by the renilla luciferase activity.
Statistical analysis
The statistical significance (
Investigation of miR-503-3p suppressing H292 cell viability. A, B. Analysis of cell proliferation of H292 and H1975 at 24, 48, 72 and 96 h after transfecting with 20 nM miR-503-3p mimics and NC by MTT assay. C. Colony formation of H292 cell after transfected with 10, 20 nM of miR-503-3p, respectively. D. Microscope image (4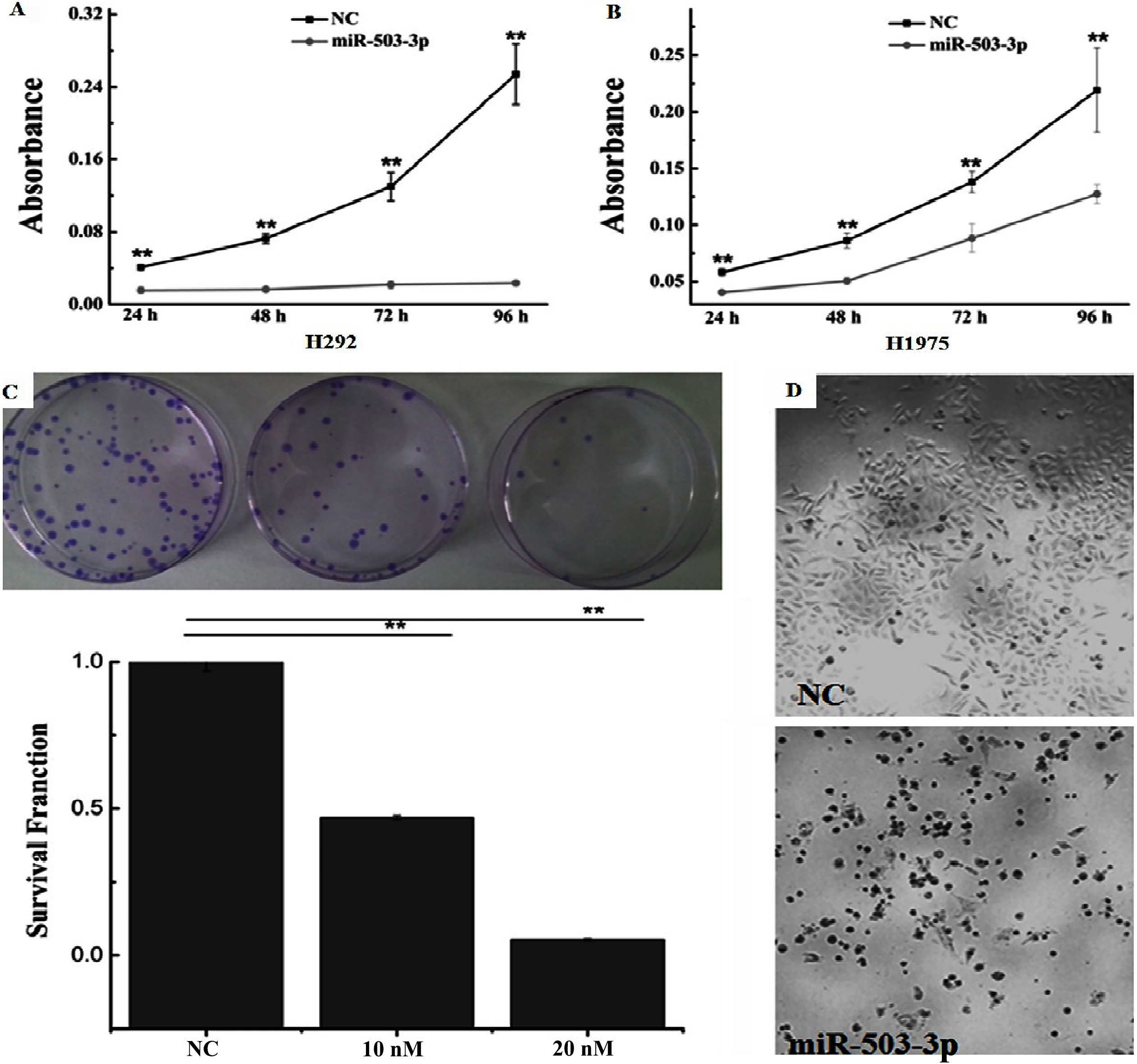
miR-503-3p suppresses cancer cell proliferation
It has been reported that miR-503-5p play an important role in regulating tumor cell angiogenesis, growth, proliferation and apoptosis. So far, the function of miR-503-3p has yet not been elucidated. To investigate whether miR-503-3p is involved in these processes, H292, H358 and H1975 cells were transfected with 10, 20, 50 and 100 nM miR-503-3p mimics, respectively. NC mimics (100 nM) is considered as negative control. At 72 h after transfection, cell viability was measured using MTT assay. Figure 1 shows that the proliferations of all examined cell lines were inhibited after treated with 100 nM of miR-503-3p mimics. Compared to other types of treated cells, the cell proliferation of H292 was extremely suppressed by miR-503-3p mimics (Fig. 1A). Furthermore, the transcript level of miR-503-3p was detected in three cells by RT-PCR, the RT-PCR result showed that the expression of miR-503-3p in H292 was lower than that in H358 and H1975 cells (Fig. 1B).
miR-503-3p influences the cell cycle distribution of H292 and H1975 cells. Cells were transfected with (right)/without (left) miR-503-3p mimics, after 48 h later, cells were collected and the percentage of cell cycle phase was detected by flow cytometry. miR-503-3p slightly decreases the percentage of G1 phase cells and increases the ratio of G2/M phase cells in both H292 and H1975 cells.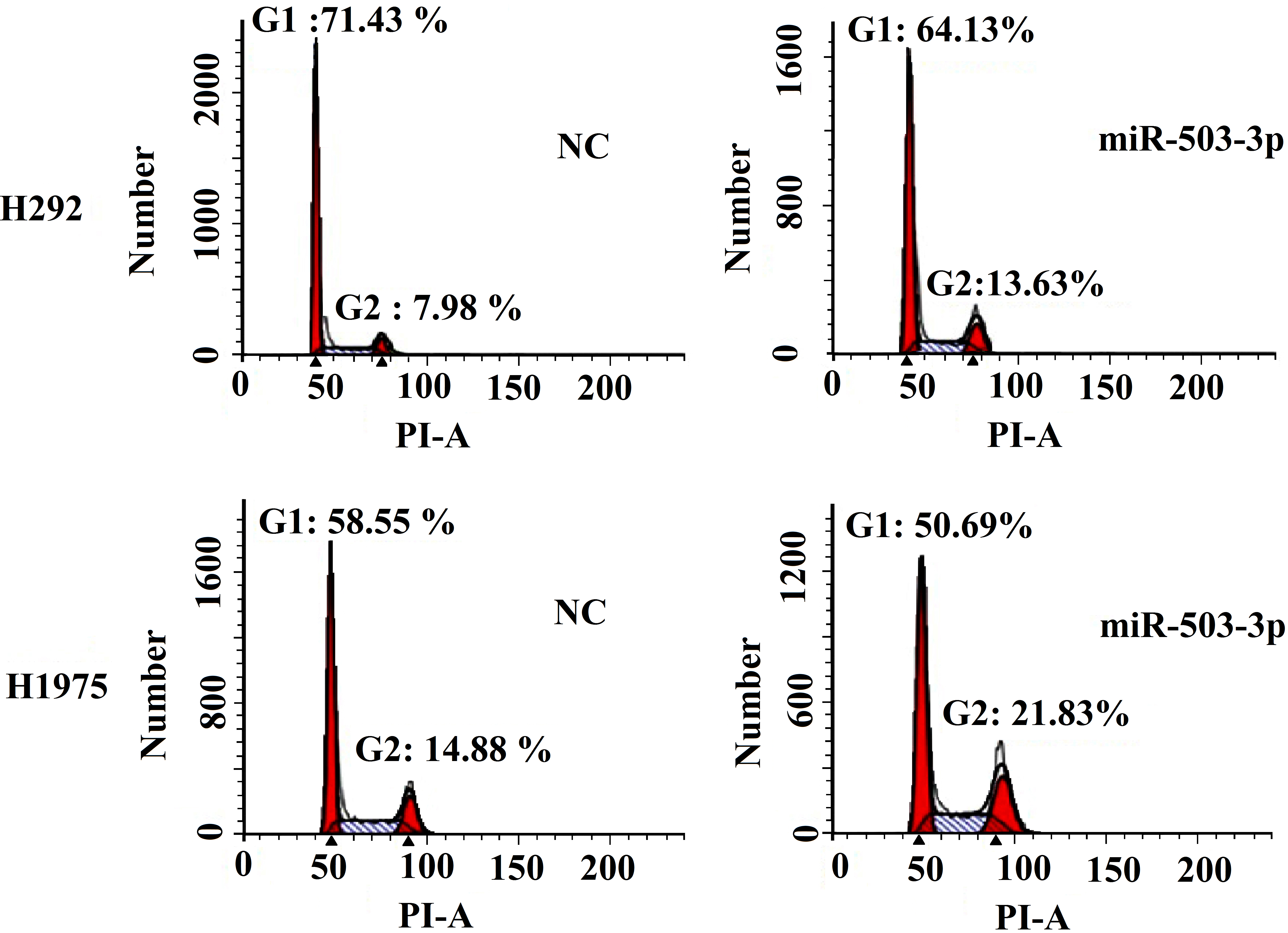
miR-503-3p increases the early apoptotic cell populations in H292 and H1975 cells. H292 and H1975 cells were treated with (right)/without (left) miR-503-3p mimics for 48 h, then collected the PI and Annexin V stained cells for apoptosis analysis by flow cytometry. miR-503-3p dramatically increased the early apoptotic cell populations in H292 cells.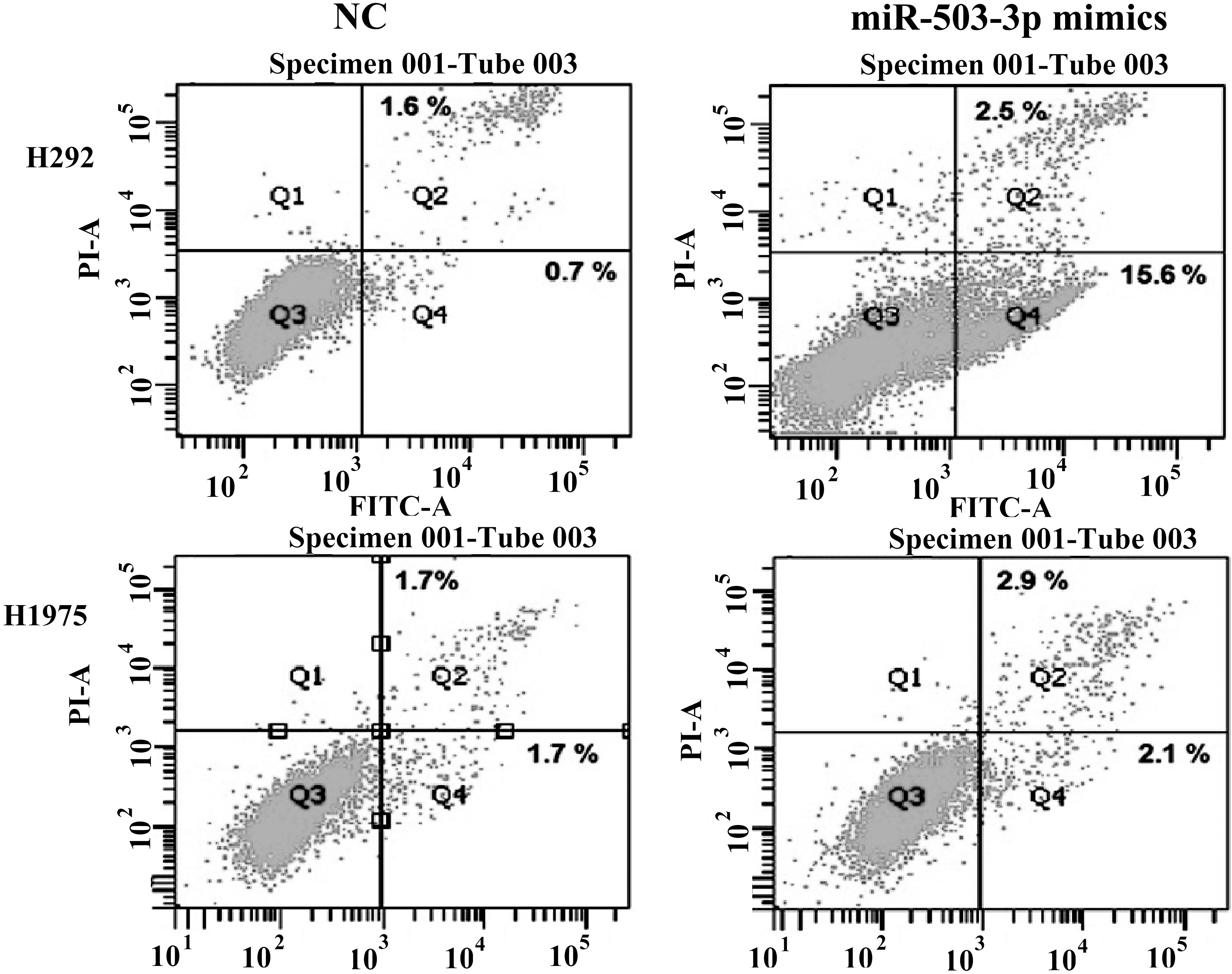
To investigate the effect of the miR-503-3p on cancer cell proliferation with the time increased, the H292 and H1975 cells were chosen for further study. H292 and H1975 cells were transfected with NC and miR-503-3p mimics. Then the cell proliferation was detected by MTT assay at 24, 48, 72 and 96 h time point after transfection, respectively. The cell proliferation of H292 cells was dramatically inhibited by the miR-503-3p mimics, compared to H1975 cells (Fig. 2A and B). In order to further confirm the effects of miR-503-3p on H292 cells proliferation, colony formation assay was performed. The survival fraction of H292 cells transfected with 10 nM and 20 nM miR-503-3p mimics were 47% and 5.38%, respectively (Fig. 2C), which are significantly lower than that of NC (20 nM) treatment group. Furthermore, the transfected H292 cells with miR-503-3p mimics ex-hibited atrophic morphology (Fig. 2D). Taken together, these results suggest that miR-503-3p inhibits cancer cell proliferation.
miR-503-3p suppressed the cell viability of lung cancer cells what we investigated. In many cases, cell proliferation and cell cycle progress are closely related. Thus, the cell cycle distribution of H292 and H1975 cells treated with/without miR-503-3p mimics were examined using flow cytometry. As shown in Fig. 3, the percentage of G1 phase cells of NC and miR-503-3p treated groups were 71.43%, 64.13% in 786-O cell, and 58.55%, 50.69% in H1975 cell, respectively. The G2/M phase percentage of H292 cells is 7.98%, 13.63% in NC and miR-503-3p mimics treated group, respectively. In H1975 cells, the G2/M phase per figure of NC group is 14.88%, and that of miR-503-3p mimics group is 21.83% (Fig. 3). These results suggest that treatment with miR-503-3p slightly decrease the percentage of G1 phase cells, whereas the G2/M phase percentage is increased in both H292 and H1975 cells.
A. Hierarchical clustering of the significantly differentially expressed gene in the two comparisons, using the RNA-seq data derived from four samples (NC1, NC2, miRNA 1, miRNA 2) based on log10 (FPKM+1) values. The red bands identify high gene expression quantity, and the green bands represent the low gene expression quantity. B. The transcript levels of cell cycle-related gene were tested by PCR array, experiment was repeated three times.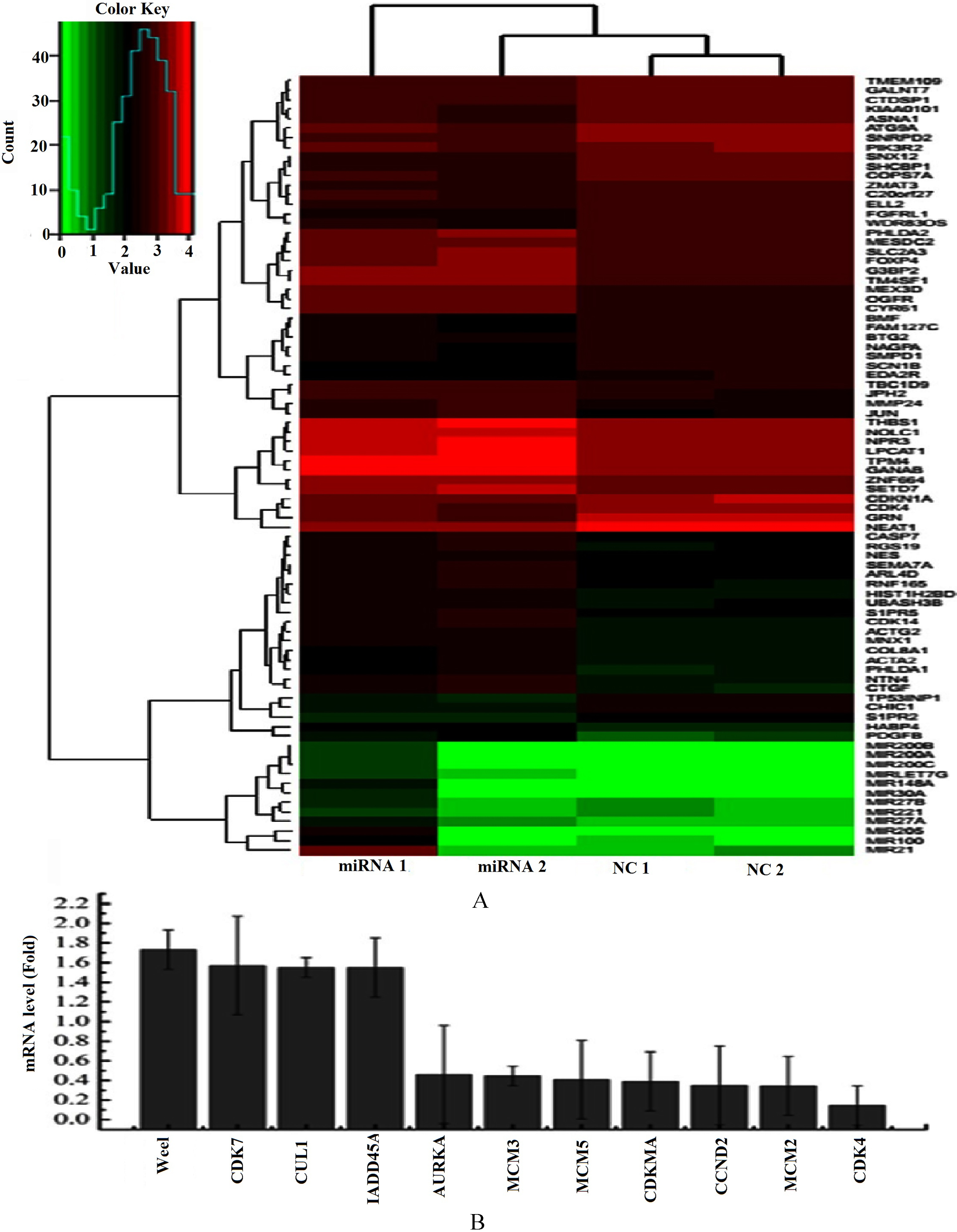
To better understand the mechanisms underlying the miR-503-3p dramatically suppressing the cancer cell viability. The H292 cells transfected with miR-503-3p were harvested at 48 h and cell apoptosis was analyzed using Annexin V-FITC Apoptosis Detec-tion kit. As shown in Fig. 4, H292 cells treated with miR-503-3p mimics significantly increased the early apoptotic cell populations compared to the NC group (15.6% vs. 0.7%). In H1975 cells, the early apoptotic cell populations are 1.7% and 2.1% in NC and miR-503-3p mimics treated grou-ps, respectively. These results indic-ate that miR-503-3p induced the cancer cell apoptosis.
Analysis of differential expression gene using RNA-seq and PCR array
To better elucidate the global interaction of miR-503-3p with other genes, RNA-seq analysis was performed. Through RNA-seq, we found some differentially expressed genes, such as p21 (CDK-N1A), JUN, CTGF, GRN, CDK4 etc, between miR-503-3p mimics tra-nsfected cells and the NC tre-atment. In addition, PCR array was performed to further investigate the transcript levels of differen-tial expression genes related to cell cycle, Our result demonstrated that the transcript of Wee1,CDK7, CUL1 and GADD45A were upregulated, and AURKA, MCM3, MCM5, p21, CCND2, MCM2, CDK4 were downregul-ated in miR-503-3p mimics transfected cells, compared to NC group (Fig. 5B). Interestingly, the transcript level of p21 and CDK4 detected by both PCR array and RNA-seq were decreased obviously.According to the RNA-seq data, we integrated a variety of proteins interaction database and established a protein-protein interaction map of gene collections from H292 cells after NC and miR-503-3p transfected (Fig. 6A). On the basis of protein interaction network, we estimated the number of each protein connect with others (Fig. 6B). The more number of other proteins interact with target protein, the more important role the target protein plays. In the diagram of protein interaction network, p21 and JUN interact with six, CTGF with five and CDK4, PDGFB, S1PR2, THBS with four proteins, respectively.
p21 is a direct target of miR-503-3p
We tried to predict the target genes of miR-503-3p using the target gene prediction website what we have known. Regrettably, no one target gene was found. From above, we know that p21 and CDK4 are down regulated by miR-503-3p either in PCR array or in RNA-seq. NCBI blast analysis indicated that p21 might be one of the target genes of miR-503-3p and there was one putative binding site in its 3’-UTR (Fig. 7A). Thus we synthesized p21 3’-UTR and p21 3’-UTR-mut oligonucleotide pairs containing miR-503-3p targeting site or mutating site, and then the dualluciferase reporter vector was constructed. Cells were cotransfected with miR-503-3p mimics and reporter vector for the dualluciferase reporter assay to validate the target prediction. Our results showed that p21 wild type 3’-UTR luciferase activity in H292 cells was significantly reduced after miR-503-3p transfection, whereas p21 3’-UTR-mut activity was not affected (Fig. 7B). These results indicate that miR-503-3p directly regulates the expression of p21 by targeting 3’-UTR of its mRNA.
Discussion
Dysregulation of microRNAs (miRNAs) is involved in the initiation and progression of several human cancers, including lungcancer, as strong evidence has been found that miRNAs can act as oncogenes or tumor suppressor genes [25, 26, 27]. Aberrant expression of miRNA-503 has been observed in various types of human cancer [28, 29]. In current work, we found that the upregulation of miR-503-3p inhibited lung cancer cell viability and induced cell apoptosis, particularly inhibited H292 cell viability dramatically, indicating that miR-503-3p may play an anti-oncogenic role in lung cancer cells. Furthermore, analysis of cell cycle and cell apoptosis indicated that miR-503-3p alters the distribution of cell cycle and induces the cancer cell apoptosis. Modulating the activities of some genes through molecular methods has already been used in cancer therapy. Hence, elucidate the mechanism of miR-503-3p in regulating cell viability may provide an opportunity for some carcinoma therapy.
RNAseq is a recently developed approach to investigate the transcriptome profiling using deep-sequencing technology. The data analysis provided a clue that various molecules in organisms complete many complex functions through interacting with each other. It will help us to understand the whole internal mechanisms of various biological processes. Through RNAseq, 51 genes differentially expressed between miR-503-3p treatment 1 (miRNA 1) and NC1 group (among the upregulated gene is 30, down-is 21), and 66 differential expression gene were found between miR-503-3p2 (miRNA 2) and NC2 group (upregul-ated gene is 39, down-is 27) (Fig. 5A). The functions of some differentially expressed genes, such as p21, JUN, CTGF, GRN, CDK4 etc, have been widely investigated in previous study. C-jun has effect on the cell proliferation, apoptosis, and invasion [30, 31, 32]. CTGF promotes malignant cell growth and inhibits apoptosis [33, 34, 35, 36]. Inhibitor of CDK4 impairs cell proliferation and induced apoptosis [37, 38, 39].
Construction of protein interaction network in H292 cells after transfecting with NC and miR-503-3p mimics. A. Red pie chart indicated the expression of protein is up-regulated. Yellow pie chart represent the expression of protein is down-regulated. B. Number represents the number of the protein interaction with other proteins in the network.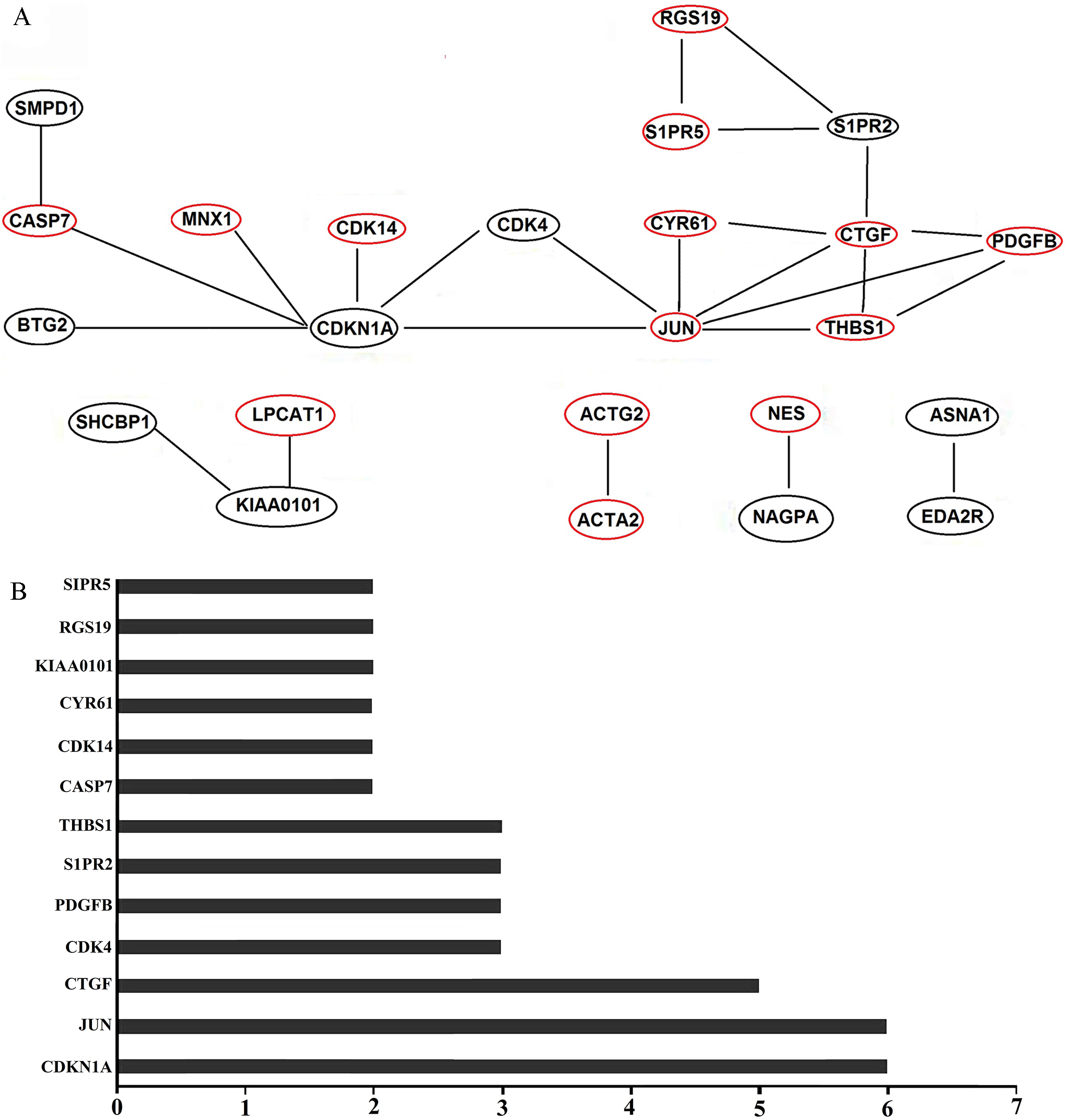
MiR-503-3p suppresses p21 expression by targeting the 3’-UTR sequences of p21. A. Putative miR-503-3p binding site within the human p21 3’-UTR are shown at the top. Sequences of mature miR-503-3p aligned to target site and the UTR mutated in the miR-503-3p seed-pairing sequence are shown below. B. Luciferase reporter assay were performed at 24 h following co-transfection in H292 cells with Wt p21 or Mut p21 vectors together with miR-503-3p mimics or nonsense small RNA oligonucleotides as the negative control. Significance was determined by Student’s t-test. 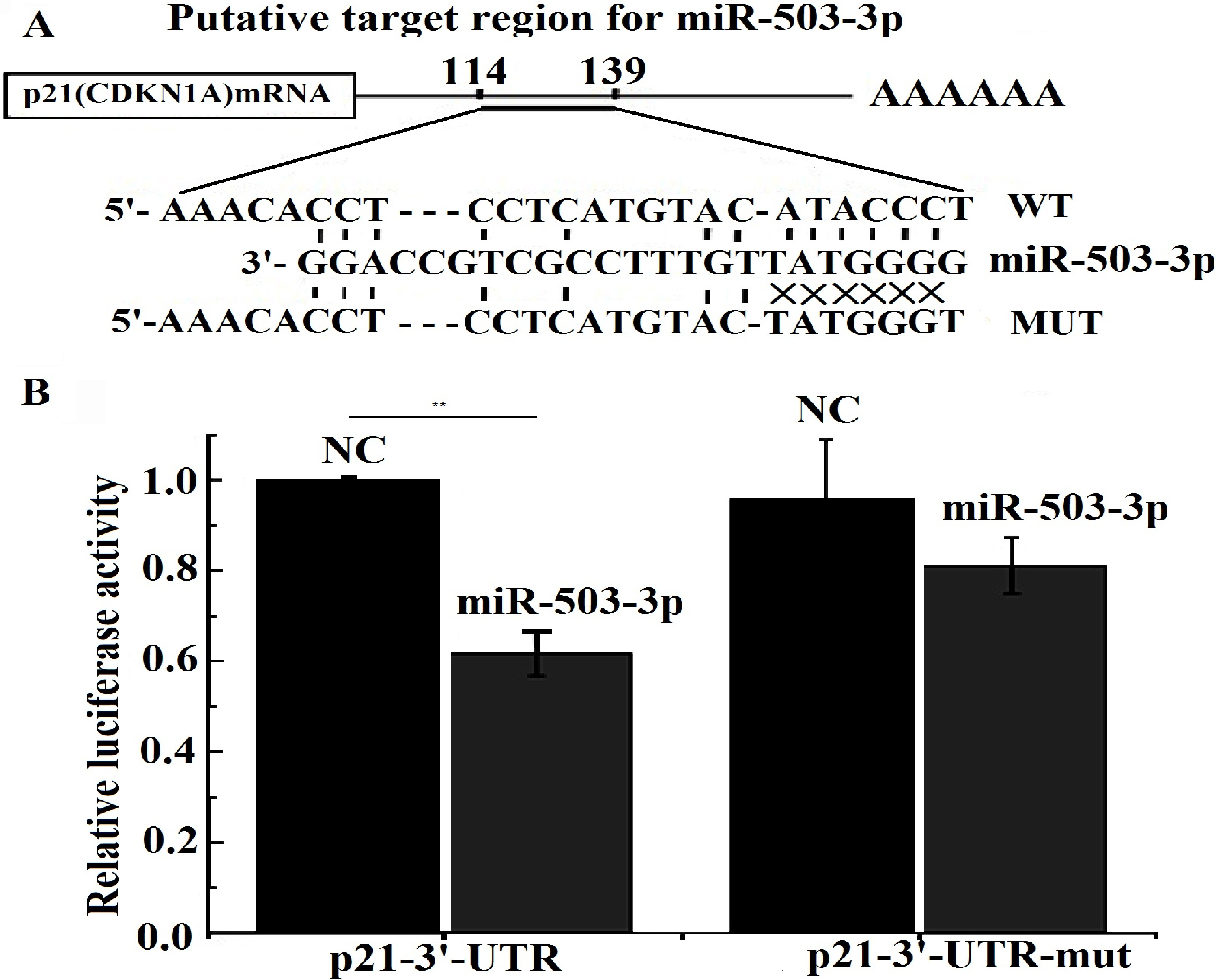
Our reports support the assumption that miR-503-3p induced tumor cell viability may be mediated by regulating the expression of these differential expressed genes. Biological interaction network is a system that various molecules in organisms complete many complex functions through interacting with each other. It will help us to understand the whole internal mechanisms of various biological processes. According to biological interaction network (Fig. 6), p21 was regarded as the most important protein that mediates miR-503-3p signal pathway, and subsequently it was proved to be a target gene of miR-503-3p. p21 was originally identified as a cyclin-dependent kinase (CDK) inhibitor that play an important role in regulating cell growth and differentiation, and can serve as an assembly factor for Cyclin D-CDK4 complex formation [40, 41].
Repressing p21 transcription and cyclin D1-Cdk4-p21WAF1/Cip1 complex formation induce apoptosis in vascular smooth muscle cells have been previously reported [42]. CDK4 and p21 proteins were simultaneously suppressed by miR-503-3p in H292 cells (Fig. 5), in keeping with the results of biological interaction network that miR-503-3p induces apoptotic cell death has emerged via p21 cooperate with CDK4 (Fig. 6). In the light of some recent studies, p21 can exert an opposite role being anti-oncogenic or oncogenic gene, depending on the cancer type and on the drug treatment. The current study supports the idea that p21 acts as an oncogene preventing cell apoptosis . Another study showed that increasing in p21 levels prevent cell apoptosis by sustaining an arrest of the cell cycle at the G2 phase [43, 44], consistent with miR-503-3p increase the percentage of G2/M phase cells in lung cancer cells. BTG2 is member of a novel family of anti-proliferative genes that prevents apoptosis of terminally differentiated rat pheochromocytoma PC12 cells [45]. p21 and BTG2 can be transcriptionally induced by p73 [46], suggesting p21 may also cooperate with BTG2 to regulate cell apoptosis.
In conclusion, the present study provide the evidence of miR-503-3p inhibiting lung cancer cell proliferation and apoptosis, suggesting that miR-503-3p may be a promising targets in lung cancer cell therapeutics. Some differential expression genes were discovered and the mRNA level of p21 and CDK4 were down-regulated in miR-503-3p overexpression cells, indicating that miR-503-3p inhibiting cell proliferation is partially mediated by these two proteins. The transcript of p21 is down-regulated by miR-503-3p, and dual-luciferase reporter assay showed miR-503-3p down-regulated p21 expression by directly targeting the 3’-UTR of p21 mRNA.
Conflict of interest
None.
