Abstract
BACKGROUND:
Although O(6)-methylguanine DNA methyltransferase (MGMT) promoter methylation status is an important marker for glioblastoma multiforme (GBM), there is considerable variability in the clinical outcome of patients with similar methylation profles.
OBJECTIVE:
We examined whether a MicroRNA (miRNA) signature can be identified for predicting clinical outcomes and helping in treatment decisions.
METHODS:
The differentially expressed miRNAs were evaluated in 6 pairs of short- (
RESULTS:
Six miRNAs were upregulated in the long-term survival group (fold change
CONCLUSIONS:
The 4-miRNA signature was identified as an independent prognostic biomarker that identified patients who have a favorable outcome.
Keywords
Introduction
Glioblastoma multiforme (GBM) is the most frequent primary brain tumor of adults and is highly malignant with a median overall survival (OS) of only 12–14 months [1]. The standard treatment for GBM patients include maximal safe neurosurgical resection and temozolomide (TMZ) chemotherapy with concomitant radiotherapy, followed by cycles of adjuvant TMZ [2]. However, evidence from two randomized clinical trials (the NOA-08 study and the Nordic Glioma study) shows not all patients are sensitive to this therapy [3, 4]. Although the short- and long-term surviving patients with GBMs have similar histological characteristics, biological and molecular characteristics of these tumors vary significantly [5].
Previous studies showed that molecular classification more reliably discriminates glioma and better predicts patient outcome than histological classifica- tion [6]. The 2016 WHO central nervous system (CNS) tumor classification also focuses on molecular markers. It builds on molecular and genetic basis of tumorigenesis and aims to incorporate these characteristics alongside histological findings in tumor classification [7]. There has been a great increase in the understanding of molecular alterations, both genetic and epigenetic, in GBMs. Major clinically relevant genetic and epigenetic abnormalities in GBM-such as mutations in IDH1/2, EGFR, PDGFRA, NF1 and hTERT were found [8]. The molecular profiles of patients affect therapeutic options. There is thus a critical need to implement clinical molecular markers tests. However, the clinically relevant molecular markers for GBM prognostication doesn’t meet the need of routine practice due to limited number and remain debatable [9]. Therefore, it is necessary to identify reliable molecular markers that can be used in prognosis and prediction of therapeutic response in GBM patients.
MicroRNAs (miRNAs) are highly conserved, 18–25 nucleotides in length, and non-protein-coding small RNAs that play key roles in post-transcriptional regulators of gene expression by silencing their mRNA targets [10]. Previous study showed that classification of multiple cancers based on miRNA expression signatures is more accurate than mRNA based signatures [11]. miRNAs have shown clear diagnostic, prognostic, and therapeutic potential in GBM. Previous studies showed that the expression of miR-21, miR-128a, miR-181b, miR-181c, miR-221, and miR-222 significantly altered in GBM. Moreover, miR-181b and miR-181c were significantly downregulated in patients who responded to RT/TMZ compared to patients with progressive disease [12]. miR-195, miR-455-3p and miR-10a were the three most up-regulated miRNAs in TMZ resistant cells [13]. Compare to anaplastic astrocytomas and normal brain tissues, GBM has higher miR-196a and miR-196b expression levels. Furthermore, high miR-196 expression levels is a predictor of worse prognosis in GBM patients [14].
Previous studies showed that the expression of O6-methylguanine-DNA methyltransferase (MGMT) is mainly regulated at the epigenetic level by promoter methylation [15]. As an acknowledged predictive marker, MGMT gene promoter methylation status was associated with prolonged OS in newly diagnosed GBM treated with the standard treatment [16]. Detecting MGMT promoter methylation level is crucial to indicate the option of alkylating agents or to select di- rectly a second line targeted therapy [17].
In this study, we identified a miRNAs-expression signature in GBM patients utilizing a qRT-PCR-based miRNA array and determine MGMT promoter methylation status by means of pyrosequencing (PSQ). Furthermore, a bioinformatics approach based on miRNA-target databases was used to identify relevant mRNA targets and biological pathways. The objective of this study was to obtain new prognostic and predictive biomarkers for GBM patients with the standard treatment.
Materials and methods
Patients
The retrospective study included 48 patients with primary GBM who were surgically treated at the Department of Neurosurgery, the Second Hospital, Lanzhou University. Written informed consent was signed by each patient before the treatment. After resection of the tumor, patients underwent adjuvant therapy including radiation (2 Gy per fraction for 6 weeks, total dose of 60 Gy) and chemotherapy with TMZ (75 mg/m
Patients characteristics
Patients characteristics
In this study, the median survival within the EORTC-NCIC trial 26981 of nearly 450 days as a survival cutoff-value that separate long- and short-term survivors [18]. Median OS was 827 days for long-term and 300 days for short-term survivors (
The GBM tissues samples were surgically resected and immediately fixed in formalin and embedded in paraffin (FFPE). Histopathological diagnosis of GBM was performed independently by two experienced neuropathologists. Total RNA with enriched fraction of small RNAs was extracted from, FFPE samples using TRIzol reagent (Invitrogen life technologies) by xylene deparaffinization and miRCURY
Bisulfite conversion of DNA and PSQ analysis of MGMT promoter
100 ng total DNA was subjected to bisulfite conversion by Epitect Bisulfite kit (Qiagen, Germany). A total of 40 ng bisulfite-treated DNA was carried on for PCR using the PyroMark PCR kit (Qiagen, Germany). The PCR conditions for MGMT gene were 95
We performed PSQ methylation assay to evaluate 4 CpG sites including CpGs 76–79 in exon 1 of the MGMT promoter region (genomic sequence on chromosome 10 from 131,265,519 to 131,265,537: CGACGCCCGCAGGTCCTCG) using MGMT Pyro Kit (Qiagen, Germany) on the PyroMark Q96 pyrosequencer (Qiagen, Germany). PSQ analysis was performed with PyroMarker Q24 software (Qiagen, Germany). PSQ yields a quantitative result giving the percentage of methylated alleles for each of the four investigated CpG sites.
miRNA array analysis
Twelve FFPE GBM tumor tissues (6 short- and long-term survivors each) were next analyzed using microarray to screen different expression of miRNAs. In brief, 1.5
Validation of the miRNA profile by qRT-PCR
Candidate miRNAs were validated by a second quantification method using qRT-PCR, in the validation cohort (
Functional interpretation of miRNA profile
A bioinformatics approach based on miRNA-target databases was used to identify relevant mRNA targets and biological pathways. We used the list of miRNAs that were significant with
Representative examples of MGMT promoter methylation status in GBM patients. A, Methylated MGMT promoter of a patient. B, Unmethylated MGMT promoter of a patient. The sequence analyzed in the study after bisulfite conversion is YGAYGTTYGTAGGTTTTYGT. Y indicates the potentially methylated sites and boxes indicate the analyzed CpG sites. Each C peak in a black background represents an internal control for bisulfte treatment.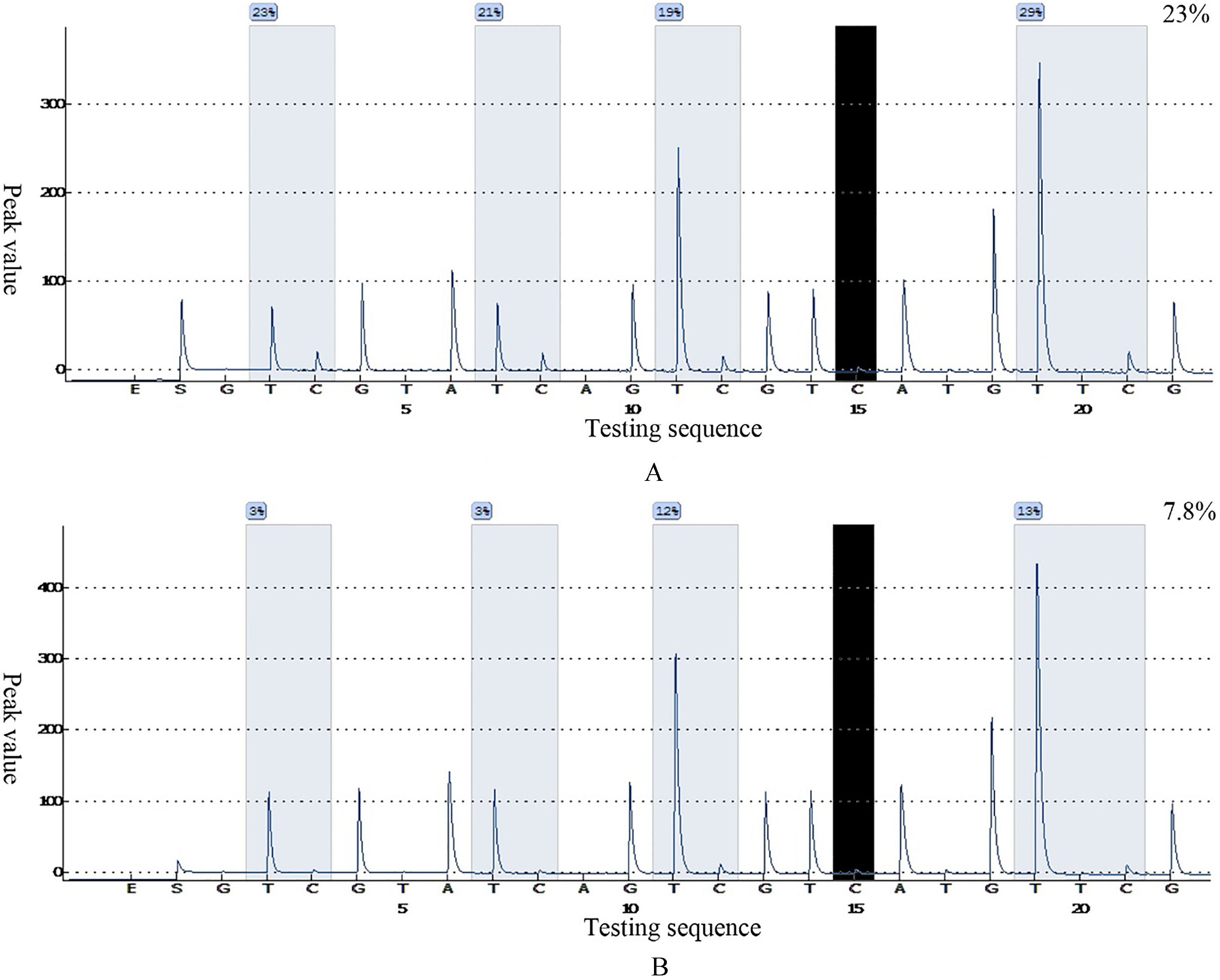
Survival data were calculated from initiation of each chemotherapy line to the date of death. OS rates, as related to the expression of the validated four-miRNAs signature, were evaluated using the Kaplan Meier method. The significance of the survival differences was determined by the log-rank test. The non-parametric Mann-Whitney U-test was used to identify significant differences in miRNA expression profiles between long- and shorter-term survivors. miRNAs with a
Kaplan-Meier overall survival curves for MGMT methylated (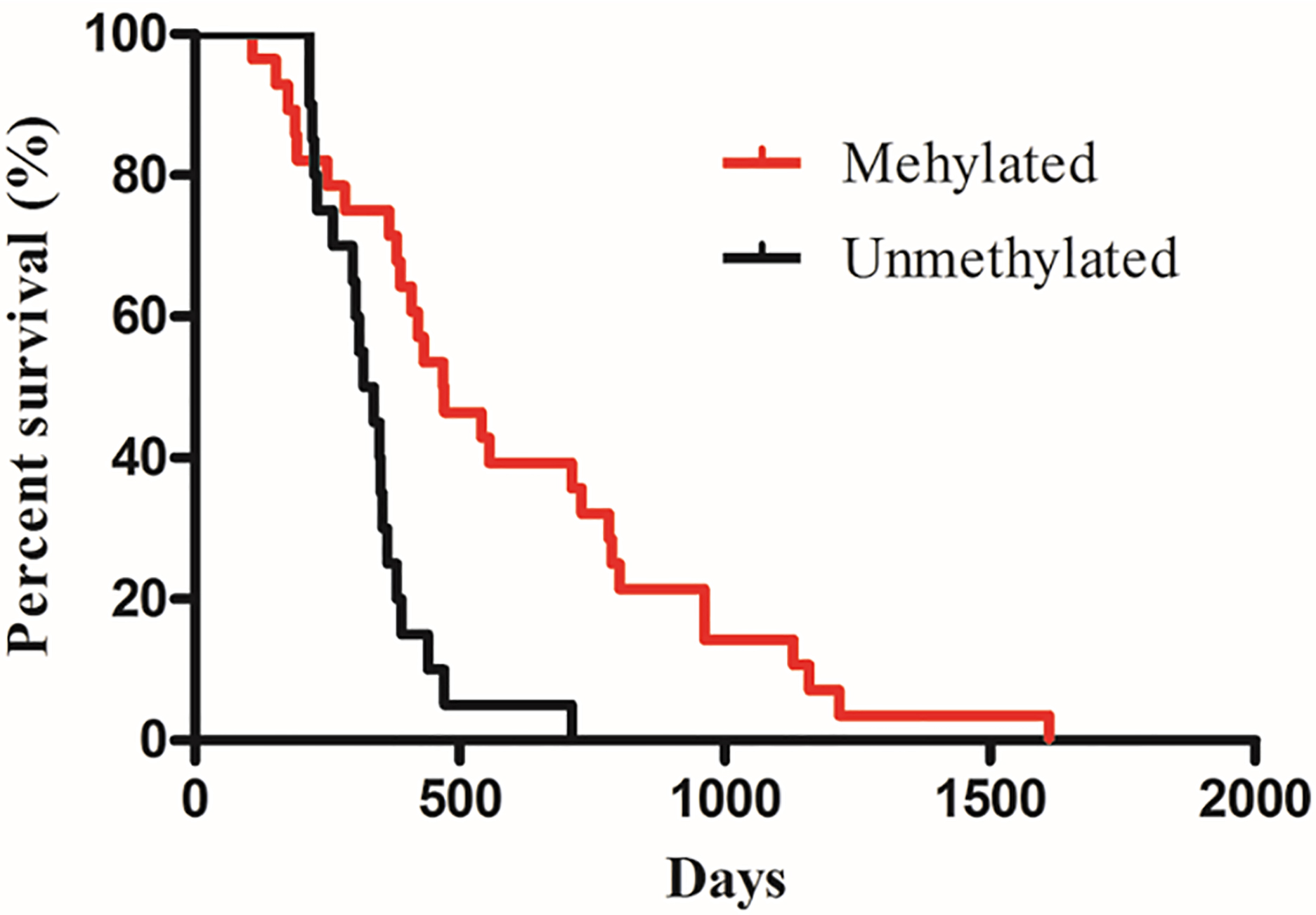
Screening by miRStarTM Human Cancer Focus miRNA PCR Array (AS-MR-0033, Arraysta). A, miRNAs expression difference between smples from long- and short-term survivors on a volcano plot. B, Heatmap showed expression pattern of six miRNAs differentially expressed between MGMT-methylation and MGMT-unmethylation GBM samples.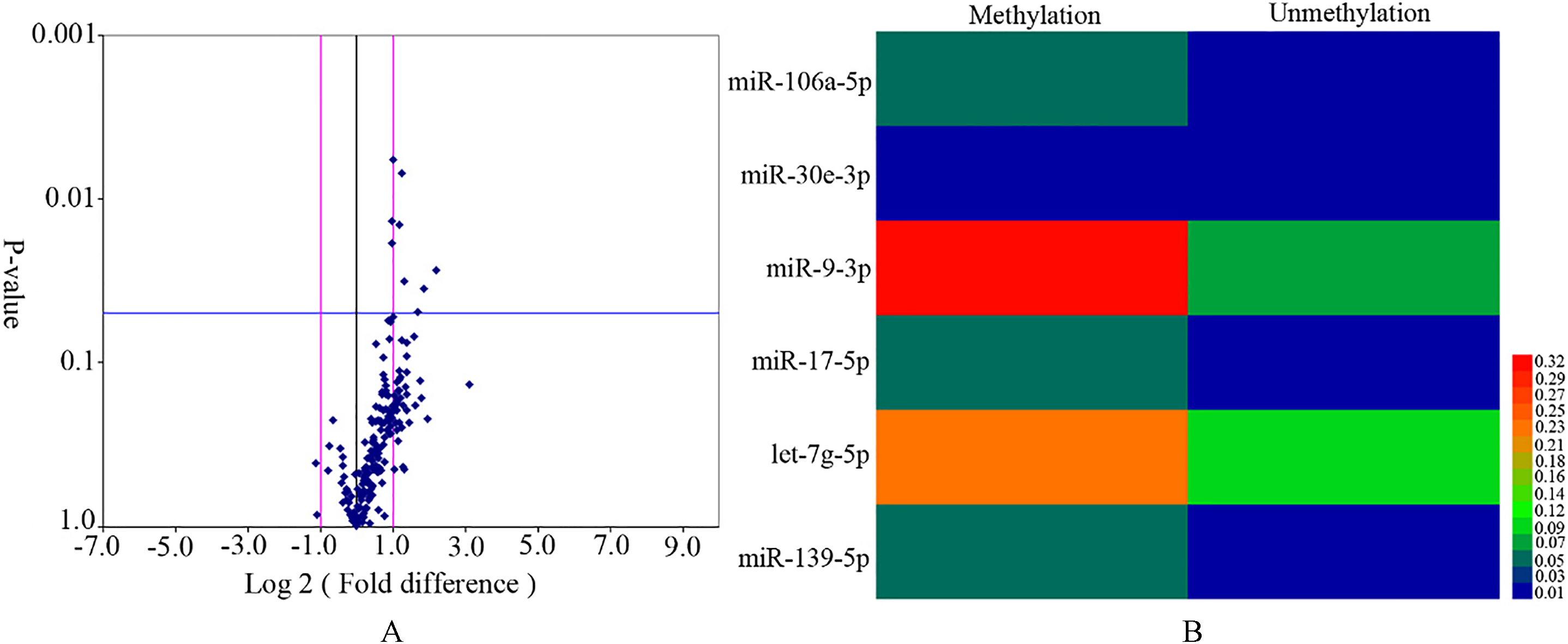
Expression of six differentially expressed miRNAs in 48 patients who constitute the validation cohort using RT-qPCR (long-term survivors 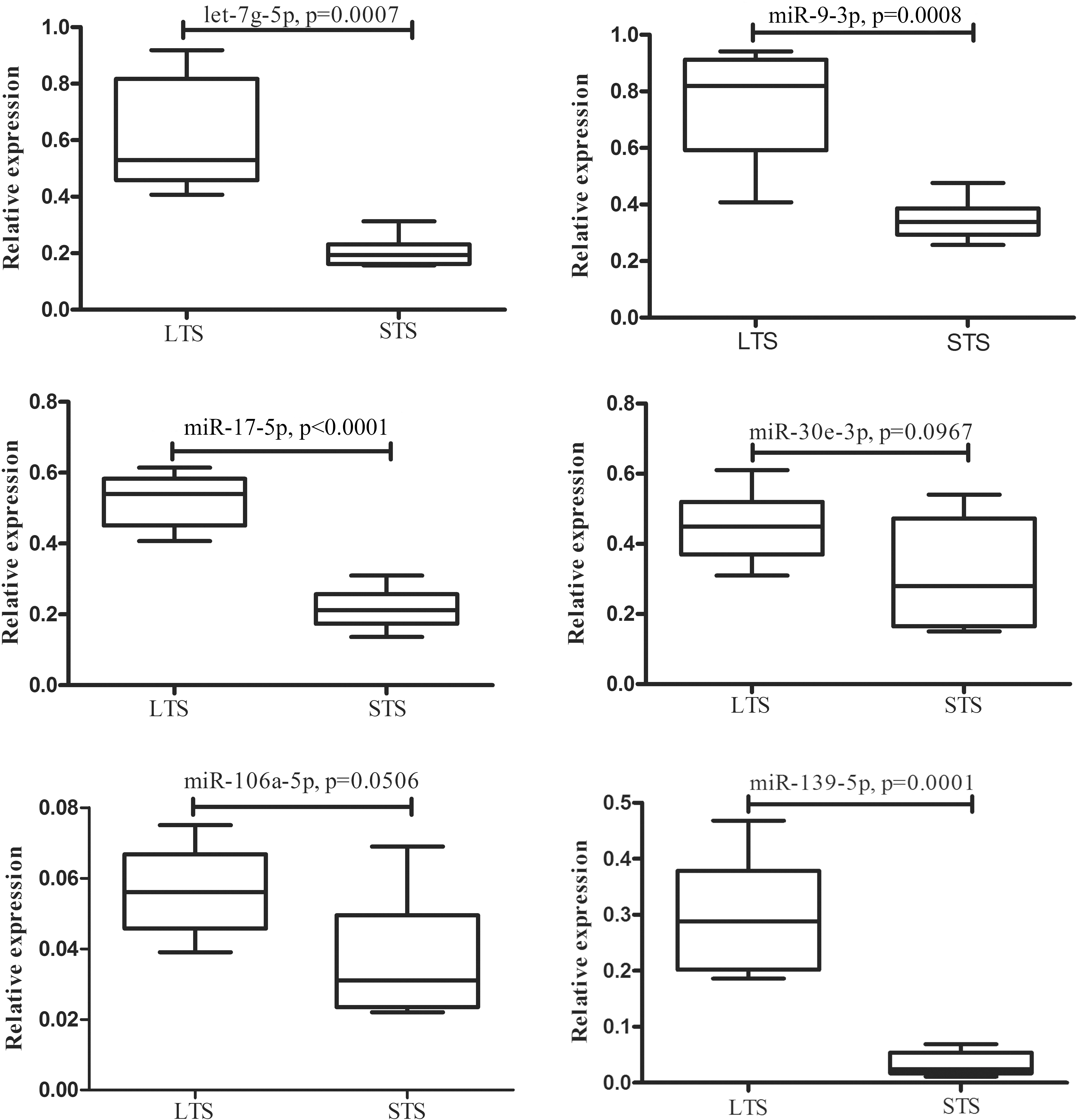
Representative two miRNAs (miR-139-5p and let-7g-5p) interact with their owen molecular targets in the cell cycle and signaling pathway including Calcium, MAPK, ErbB and mTOR associated with carcinogenesis from glial progenitor cell to primary GBM.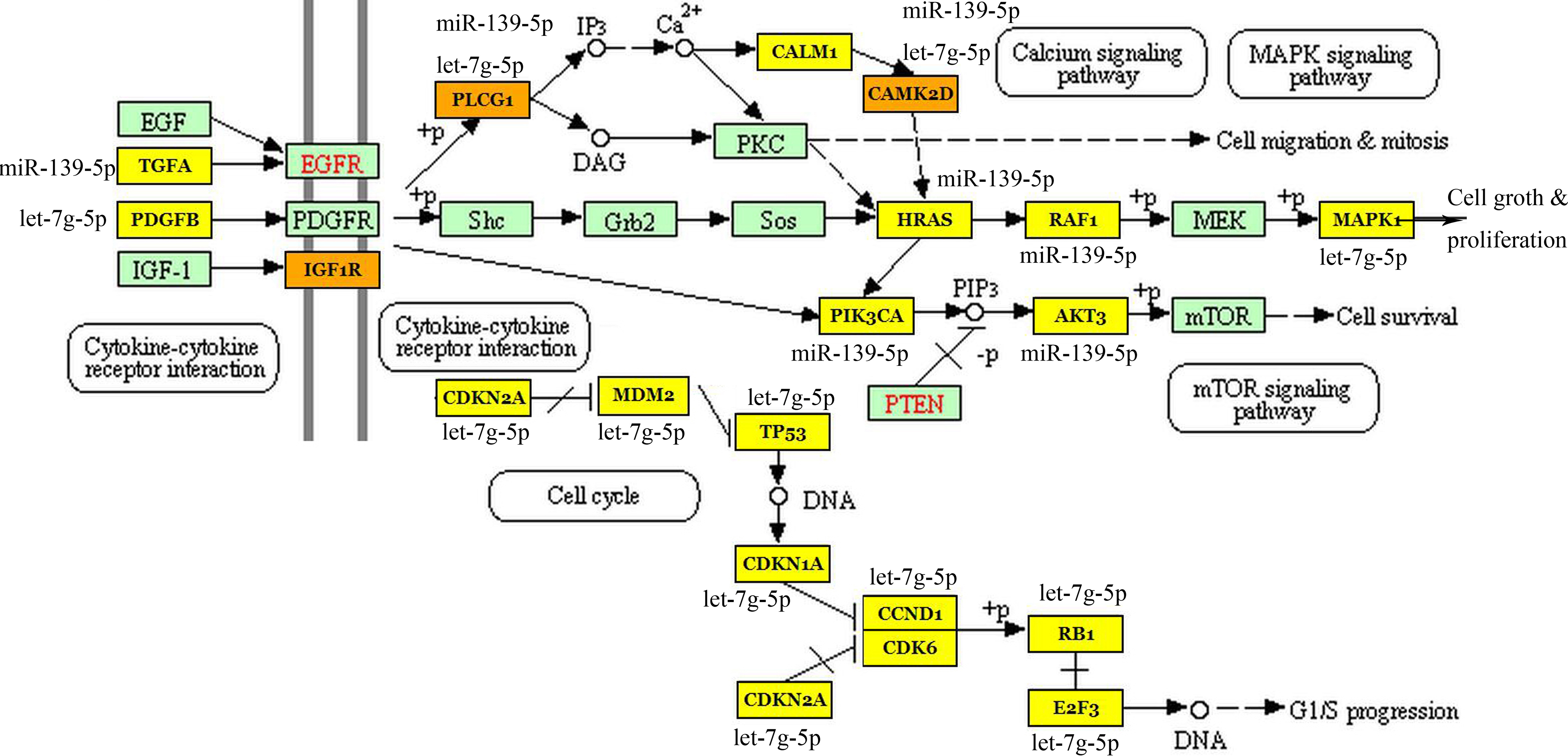
MGMT methylation status analysis
Twenty-eight of 48 samples (58.3%) showing a mean methylation level of all the invested four CpGs above 12.5%, were classified as methylated. And the other 20 samples (41.7%) were regarded as unmethylated ones. Average methylation level in methylated and unmethylated cases was 32.72%
Comparison of miRNA expression levels in long- and shorter-term survival tissue samples
The miRNA profile was analyzed between long-(
Validation by qRT-PCR
In order to technically validate the above-screened 6 miRNAs and to support potential applicability in clinical routine diagnostics, we assessed them in 17 long-term survivors and 31 shorter-term samples by qRT-PCR. The miRNAs were considered significantly altered only when they exhibited a mean fold-change
miRNA targets and signaling pathways associated with clinical response
To identify miRNA-mRNA regulatory signaling pathways in GBM patients, a novel bioinformatics strategy was used by means of KEGG-mirPathv.3. The selected mRNAs were considered reliable miRNA target genes due to the date were experimentally validated. With this approach, we identified 21 unique mRNA targets. Integrating the miRNA-mRNA target information, we found that most significant biological pathways were involved in carcinogenesis from glial progenitor cell to primary GBM. Figure 5 illustrates some of the most significant signaling pathways (Calcium, MAPK, ErbB and mTOR pathways) and molecules regulating cell cycle associated with tumor progression.
Factors associated with overall survival
In multivariate analysis, we found that additional prognostic factors such as MGMT methylation status, gender, age, extent of resection and KPS were shown in Table 2. In this regard, gender, age, extent of resection, type of resection and KPS were non-significant factors, whereas both MGMT promoter methylation status and the four-microRNA signature independently associated with OS.
Univariate analysis on potential prognostic factors for overall survival after primary diagnosis
Univariate analysis on potential prognostic factors for overall survival after primary diagnosis
Abbreviations: KPS, Karnofsky Performance Scale; HR, hazard ratio; CI, confidence interval; OS, overall survival; GTR, gross total resection; PR, partial resection; Meth, methylation; Unmehy, unmehylation.
Kaplan-Meier overall survival analysis for patients with miRNA signature high (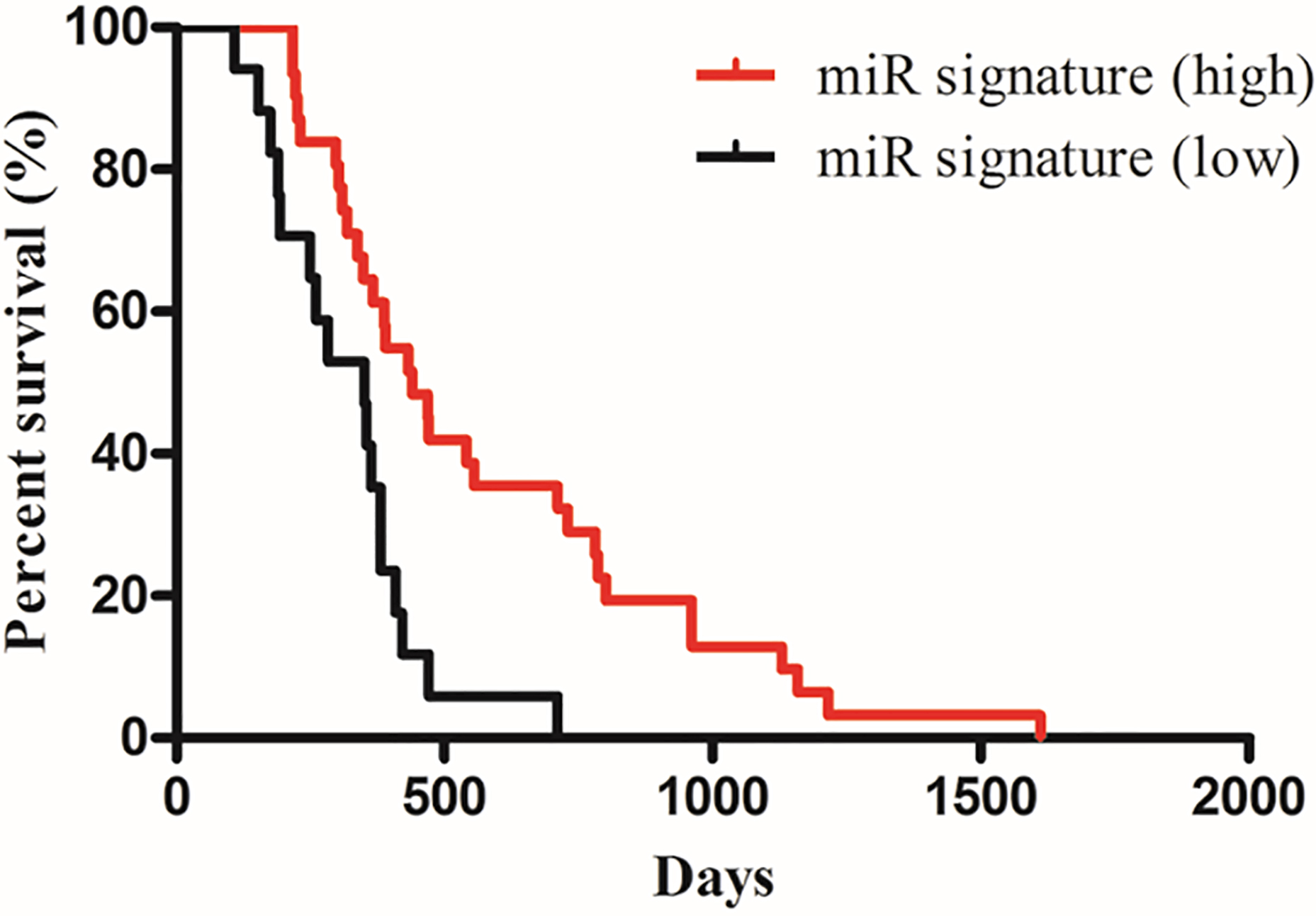
In order to examine the real influence of the four-microRNA signature on OS, Kaplan-Meier curves for two groups with high- and low-expression levels using the median value for each miRNA as the cutoff were plotted (Fig. 6). It could be shown that the four-microRNA signature was associated with OS,
MGMT promoter region includes 97 individual CpG sites spanning the proximal promoter region and the first exon [17]. Proper selection seems crucial due to methylation status varies for different CpG sites. In this study, the biological or clinical relevance of CpGs 76–79 are analyzed assessed. The reason to choose CpGs 76–79 is that these sites significantly attenuates promoter activity of MGMT in a luciferase reporter assay [20]. We confirmed MGMT promoter methylation could be a predictive factor for benefit from alkylating agent therapy in GBM. Patients with MGMT promoter methylation showed significantly longer OS (
Identification of biomarkers that predict GBM response to therapy is a critical step in the development of individual treatments that produce optimal clinical outcome. Recent clinical trials on patients with GBM revealed that only MGMT promoter methylation status significantly predicts patient’s response to alkylating agents and helps determine the patient population who likely benefit from such treatment [25, 26]. Noteworthy, MGMT promoter methylation is not a unique way to regulate the expression of MGMT protein and the response to therapy with alkylating agents [27]. The expression of MGMT protein is heterogeneous within the core of GBM tissue due to the transcriptional activation of expression mediated by hypoxia and activation of HIF-1
Increasing evidence shows that some miRNAs play critical roles in tumorigenesis of GBM and have a potential clinical value in diagnosis, treatment and prognosis evaluation for GBM [30, 31, 32]. In this study, miRNA microarray (Arraysta) was applied as a tool to screen expression of a larger number of miRNAs at a time. This approach allows identification of differential expression of miRNA not previously described for a separation of different prognostic groups in patients with GBM. Validation of the miRNA profile by RT-qPCR confirmed the microarray data results.
We found four miRNAs including let-7g-5p, miR-9-3p, miR-17-3p and miR-139-5p for the first time that accurately classified early long- and shorter-term survival (split at 450 days) in a significant way and this prediction was independent of the MGMT status.
In our study, we identified two miRNAs that were previously associated with resistance to radiation and chemotherapy. For instance, let-7g overexpression increase the radiation sensitivity of cancer cells [33]. let-7g-5p inhibits mesenchymal-epithelial transition (MET), invasion and migration by targeting VSIG4 in GBM cells [34]. let-7g-5p in the peripheral blood of patients could be regarded as a new biomarker for GBM diagnosis [35]. In the miRNA PCR array experiments used to obtain the global expression profile of the miRNAs, let-7g-5p was up-expressed 2.45 fold in long-term survival GBM patients (
In addition, we found that miR-139-5p could be used as a potential molecular marker to distinguish between short- and long-term survivors with significant p values (
The miR-17-92 cluster, located in chromosome 13 transcribes to 7 mature miRNAs (miR-17-5p, miR-17-3p, miR-18a, miR-19a, miR-20a, miR-19b, and miR-92-1) and widely shown to be important in tumor regulation, has been demonstrated by a variety of studies to be significant in GBM [40, 41]. Among miRs encoded in the miR-17-92 cluster, miR-17-5p is uniquely upregulated in secondary GBM but not primary GBM when compared to expression in a normal brain. Peripheral CD4+ T-cells derived from patients with GBM demonstrated decreased miR-17-5p expression compared to healthy controls [42]. These results support the potential role of miR-17-5p in GBM.
Conclusion
In the present study, we validated a four-miRNA signature that could serve as potential new prognostic and/or predictive marker allowing for GBM patient stratification. The mean level of CpGs 76–79 methylation within MGMT gene exon 1 can be used as dichotomization of quantitative MGMT PSQ results into “MGMT methylated” and “MGMT unmethylated” patient subgroups as a basis for further treatment decisions. The integration between a four-miRNA signature testing and MGMT promoter methylation analysis into GBM classification greatly improves diagnostic accuracy but also has practical implications concerning establishment and quality control of novel techniques, increased costs and prolonged time to diagnosis.
Conflict of interest
All authors declare that we have no competing interests.
Footnotes
Acknowledgments
We want to thank Dr. Kejun Ma from the Experiment Center of the Second Hospital, Lanzhou University for granting us access to the Pyromark Q96. Work was supported by Research Project of Chinese Society of Neuro-oncology (grant no. CSNO-2013-MS008), The Project of Healty and Famliy Planing Commission of Gansu (grant no. GSWSKY-2015-58/-2014-31), the Lanzhou Science and Technology Bureau Project (grant no. 2013-3-27/2015-3-86), and the Doctoral Research Fund of Lanzhou University Second Hospital (grant no. ynbskyjj2015-1-02/2015-2-11/2015-2-5).
