Abstract
BACKGROUND:
Remethylation of homocysteine is catalyzed by B12 dependent methionine synthase (MTR) in all types of cells and by B12 non-dependent betaine homocysteine methyltransferase (BHMT) in liver and kidney cells. Of many etiologies of cancer, an unexplored area is the variations of genes implicated in methylation reaction.
OBJECTIVE:
The study evaluated the association of BHMT (rs3733890) with acute lymphoblastic leukemia (ALL), followed by in-silico characterization of variations in BHMT gene.
METHODS:
BHMT [rs3733890; c.742G
RESULTS:
BHMT (rs3733890) showed an insignificant association with both childhood and adult ALL. Bioinformatics analysis showed that 18 nsSNPs are deleterious, 3 SNPs in 3
CONCLUSIONS:
BHMT (rs3733890) polymorphism showed no association with ALL. Hence this investigation needs further evaluation in larger sample size and effect of other SNPs, CNVs and miRNA’s is required to elucidate the role of BHMT gene in ALL development.
Introduction
Betaine-homocysteine methyltransferase (BHMT), one of the zinc-dependent cytosolic enzymes, is involved in the remethylation of homocysteine by B12 non-dependent manner especially in liver and kidney cells [1]. It has been predicted that BHMT protein can perform at least half of the total homocysteine remethylation reaction [2]. Though there were many pieces of evidence of elevated homocysteine in neural tube defect, cardiovascular disorders and cancers like acute lymphoblastic leukemia [3, 4, 5]; surprisingly there is little effort to evaluate the influence of BHMT gene variations on ALL. It has been observed that variations in other folate metabolism genes involved in the methylation reaction such as methylene tetrahydrofolate reductase (MTHFR), 5-methyltetrahydrofolate homocysteine methyltransferase (MTR) and 5-methylte- trahydrofolate homocysteine methyltransferase reductase (MTRR) can majorly contribute to the pathophysiology of ALL and other cancers [6, 7, 8, 9]. Scientific attention to exploring the possible role of BHMT in ALL development has been somewhat lacking. The human BHMT gene resides on chromosome 5q13.1-q15 [10]. BHMT gene has a common single nucleotide polymorphism (c.742G
ALL is a hematological malignancy with 80% estimates of total childhood leukemia and 20% of all adult leukemia [17, 18]. The knowledge about the most prominent etiology for ALL has been improved, but the molecular mechanism behind the disease development is not entirely known except for the role of ionizing radiations and chemicals like benzene [19, 20, 21]. It has been shown that the combination of many genetic variations and environmental elements can play a significant part in the ALL development as well as in the therapy [22, 23]. There has been an intense scientific focus on folate metabolism gene variations on ALL, but its unsustainable association with ALL in different populations [8, 9, 11] has impeded its establishment as a specific and prominent etiology for ALL. Furthermore, repeated analysis and new studies on population-specific genetic association may defiantly reveal the idea of the actual role of folate metabolism gene polymorphisms in the ALL development and therapy, which can be implemented to prevent and cure the disease in the future. Along with this the functional characterization of polymorphism by genetic expression analysis may add additional support for the disease management.
In India, existing data on ALL predisposing and drug response markers is limited and insufficient to draw a remarkable conclusion. Also, considering the heterogeneity in the populations, ALL predispositions and its therapy have always remained a challenge. Hence, it becomes necessary to evaluate the association of genetic polymorphisms with ALL. To decipher the associated risk of BHMT (rs3733890) with ALL, we conducted a case-control association analysis in South Indian population. An in-silico evaluation of other BHMT gene variants was also performed to predict the possible mechanisms of gene regulation.
Materials and methods
Case-control association analysis
Subjects
The study included 199 ALL individuals and 245 normal healthy individuals from South Indian population. The ALL was diagnosed through a morphological and immune phenotypic approach and subjects were characterized by World Health Organization (WHO) classification. The study protocols were approved by Kasturba Hospital institutional ethics committee, Manipal. The study conformed to The Code of Ethics of the World Medical Association. The written consent form was taken from both ALL individuals and normal healthy individuals. The written consent was acquired from parent or guardian in case of childhood ALL. Blood samples were collected from 125 childhood ALL individuals and 74 adults ALL individuals who were enrolled in the study from Kasturba Hospital, Manipal and Kasturba Hospital, Mangalore, Karnataka, India. The mean age of childhood and adult ALL participants were 6.57 years and 40.58 years respectively. Healthy normal subjects comprised of 191 young adults and 54 adults with a mean age 19.15 years and 47.78 years respectively.
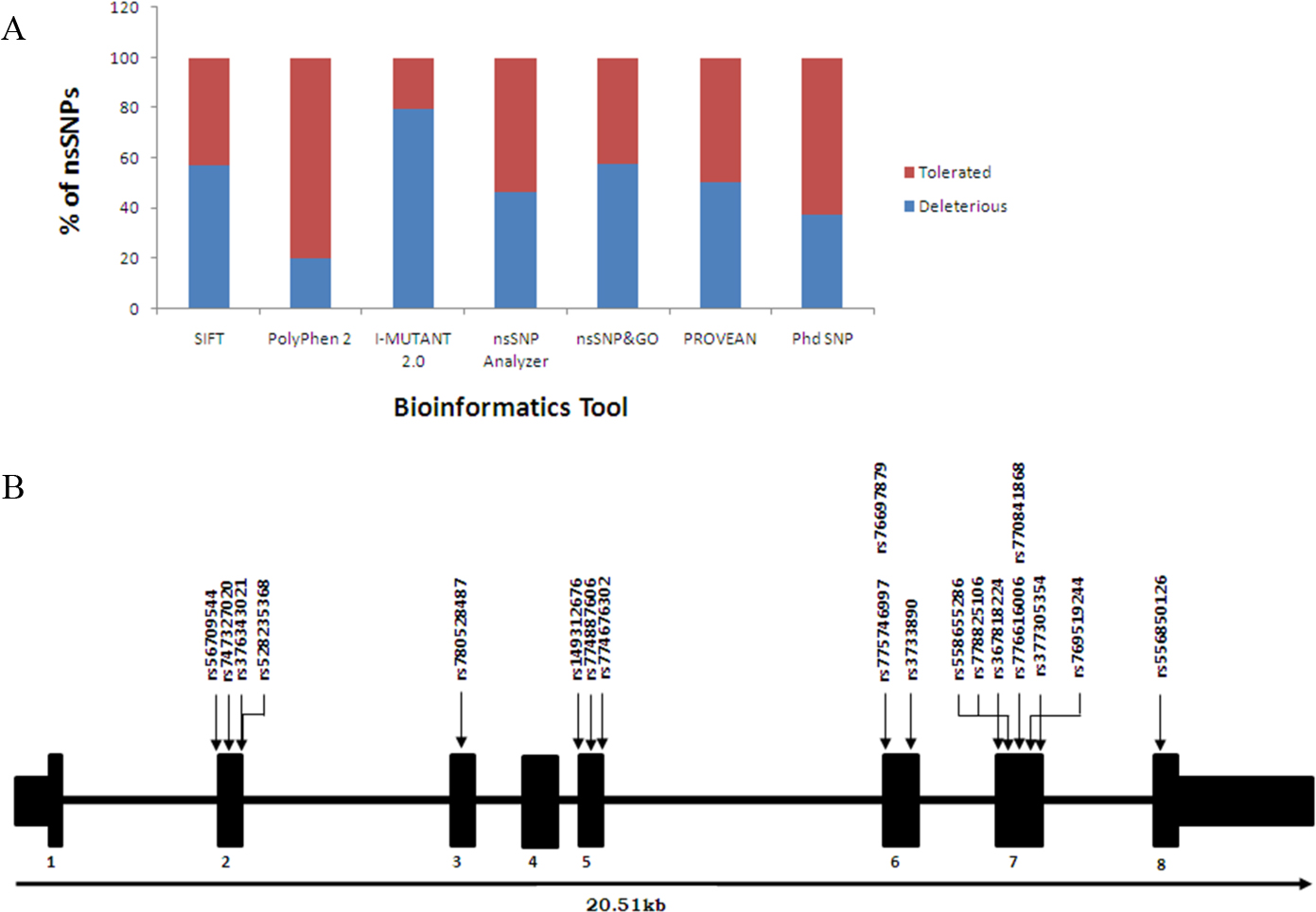
Bioinformatic analysis of nsSNPs in BHMT gene. (A) Percentage of deleterious and tolerated nsSNPs of BHMT gene predicted by 7 different bioinformatics tools. (B) Schematic representation of BHMT gene structure displaying the position of 18 deleterious nsSNPs along with rs3733890. Abbreviations: SIFT, Sorting Intolerant from Tolerant; PolyPhen-2, Polymorphism Phenotyping v2; nsSNPanalyzer, non-synonymous protein coding Single Nucleotide Polymorphisms analyzer; PROVEAN v1.1, Protein Variation Effect Analyzer; PhD-SNP; Predictor of human Deleterious Single Nucleotide Polymorphisms.
The genotyping of BHMT (rs3733890) in ALL individuals and normal healthy individuals were performed by tetra-primer amplification refractory mutation system-polymerase chain reaction (T-ARMS-PCR). The primers were designed using PRIMER1 web tool [24, 25]. The 25
Validation of genotyping by Sanger assay
The genotyping results of case-control populations captured by the T-ARMS-PCR were confirmed by gold standard Sanger DNA sequencing method by using ABI Prism 3130 genetic analyzer automated DNA sequencer (Applied Biosystems, USA) using ABI Prism BigDye Terminator v3.1 cycle sequencing kit. A total of 3 samples with three different genotypes of BHMT (rs3733890) were validated, and the T-ARMS-PCR genotype results were confirmed. The results of Sanger DNA sequencing were analyzed by Finch TV tool (
Statistical analysis
Distribution of genotype and allele frequencies of BHMT (rs3733890) in individuals with childhood ALL and normal healthy individuals
Distribution of genotype and allele frequencies of BHMT (rs3733890) in individuals with childhood ALL and normal healthy individuals
BHMT, BetaineHomocysteineMethyltransferase; ALL, Acute Lymphoblastic Leukemia; SNP, Single Nucleotide Polymorphism; OR, odds ratio; CI, confidence interval; *OR and 95% CI were calculated with BHMT 742GG genotype as reference group.
Distribution of genotype and allele frequencies of BHMT (rs3733890) polymorphism in individuals with adult ALL and normal healthy individuals
BHMT, BetaineHomocysteineMethyltransferase; ALL, Acute Lymphoblastic Leukemia; SNP, Single Nucleotide Polymorphism; OR, odds ratio; CI, confidence interval; *OR and 95% CI were calculated with BHMT 742GG genotype as reference group.
Pearson’s Goodness of fit chi-square test was used to calculate Hardy–Weinberg equilibrium (HWE) for observed genotypes in normal healthy subjects and ALL individuals. The risk association of BHMT (rs3733890) with ALL individuals was estimated by calculating odds ratio (OR) and 95% CI through chi-square test using Graph Pad Intent 3. Software.
Selection of SNPs for in silico analysis
For in silico analysis we have obtained all the information such as rsID, mRNA accession number, transcript ID, protein accession number, protein ID, protein sequences and Swiss-prot ID from National Centre for Biotechnology Information (NCBI), dbSNP database (
Prediction of nsSNPs, miRNASNPs, and CNVs by in-silico tools
In silico examination of non-synonymous SNPS (nsSNPs) were performed to predict the role of SNPs on protein structure and stability using bioinformatics tools such as Sorting Intolerant from Tolerant v. 1.03 (SIFT v. 1.03) [26], Polymorphism Phenotyping v2 (PolyPhen-2) [27], I-MUTANT 2.0 [28], Protein Variation Effect Analyzer (PROVEAN v1.1) [29], SNP&GO [30], nsSNPs protein coding single nucleo- tide polymorphisms analyzer (nsSNPanalyzer) [31], and Predictor of human Deleterious Single Nucleotide Polymorphisms (PhD-SNP) [32].
Furthermore, we analyzed the SNPs in the 3
List of polymorphisms which alter the binding of miRNA in 3
UTR region of BHMT gene and free energy change predicted using RNAhybrid
List of polymorphisms which alter the binding of miRNA in 3
MAF, Minor Allele Frequency.
The available three-dimensional (3D) structure of BHMT protein (4M3P) was retrieved from PDB (Protein Data Bank) and modeled for mutation at 239 position from arginine to glutamine (rs3733890; MAF: A
Bioinformatic network analysis
A network analysis to evaluate the closely related functional genes of BHMT were performed by using STRING 10.0 a functional protein association database by using seven different active prediction methods which include neighborhood, co-occurrence, co-expression, gene fusion, databases, experiments and text mining [39].
Results
BHMT (rs3733890) association analyses
The validation of genotypes by Sanger DNA sequencing showed 100% similarity with the genotype results by T-ARMS-PCR technique. The allele frequencies and genotype frequencies of normal healthy individuals, childhood ALL individuals and adult ALL individuals are represented in Tables 1 and 2. The genotype frequency of BHMT (rs3733890) followed HWE in childhood ALL individuals, adults ALL individuals and normal healthy individuals. BHMT (rs3733890) showed an insignificant association with ALL in South Indian population (Tables 1 and 2).
Bioinformatic analysis
Non-synonymous SNPs prediction
Details of copy number variations in BHMT gene from DGV database
Details of copy number variations in BHMT gene from DGV database
Note: nsv (NCBI; dbVar) andesv (EBI; DGVa) are accession IDs assigned to structural variations.
We observed 174 nsSNPs in BHMT gene from Ensembl database (
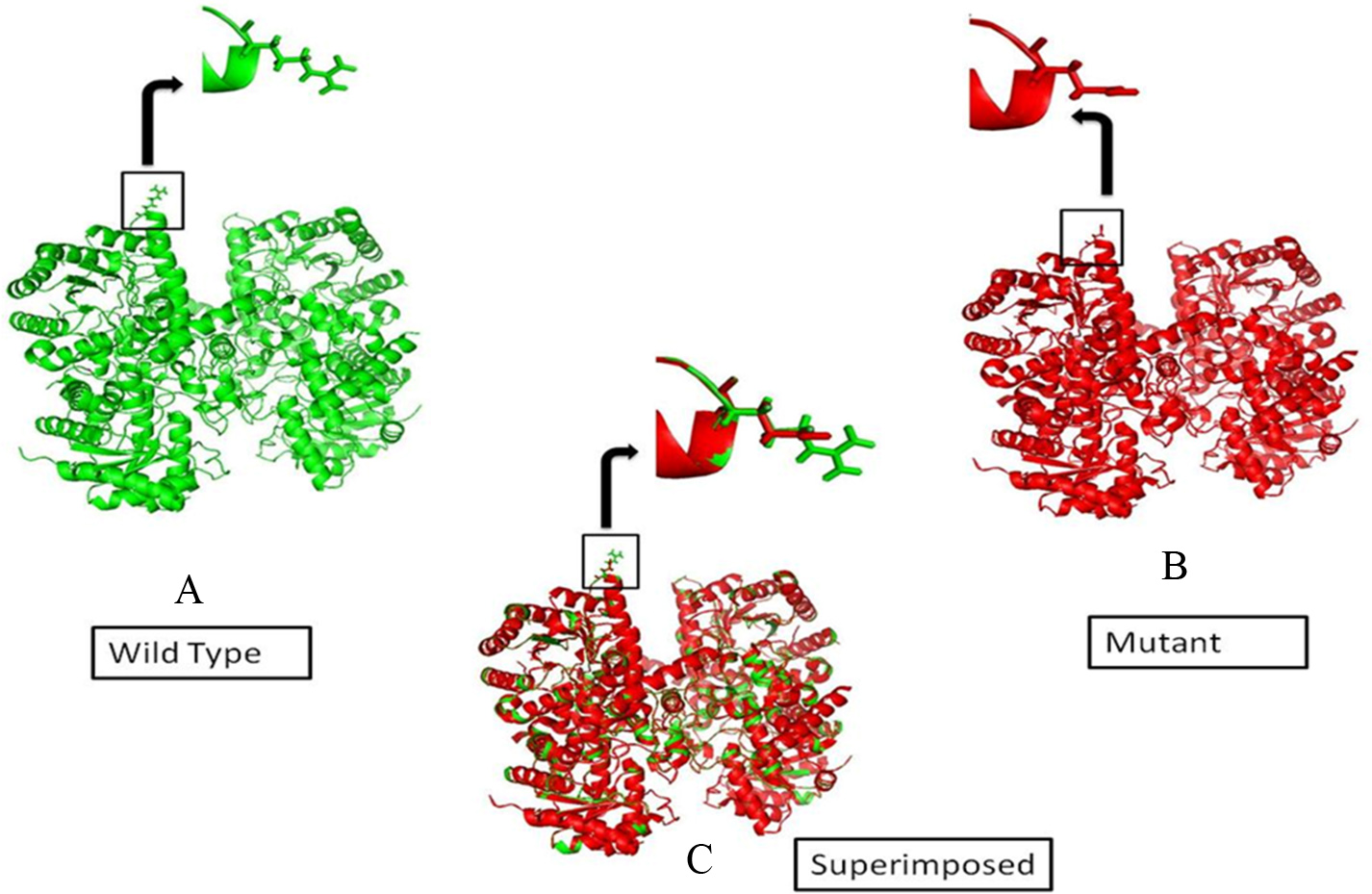
Modeled structures (cartoon shape) of BHMT (rs3733890). Native (A), Mutant (B) and Superimposition of native and mutant (C) using PyMOL 1.3.
The database search by using Ensembl showed 57 SNPs in 3
Analysis by using DGV and UCSC databases revealed that BHMT gene has 11 copy number variations (CNVs) (Table 4). A total of 6 copy number gains (esv2762527, nsv519156, nsv1022839, nsv1019727, nsv525398, esv1010320) were observed. Further, there were four insertions (nsv4892, esv3308186, esv34089 55 and nsv512853) and one duplication (esv3442711) (Supplementary Fig. 2).
Protein modeling
The impact of nsSNP rs3733890 on BHMT protein was assessed by performing mutagenesis on BHMT protein structure at position 239, where glutamine was introduced in place of arginine as a consequence of rs3733890 (Fig. 2) using PyMOL. It was observed that the total energy of the native and mutant type structure is
Bioinformatic network analysis
The network analysis of BHMT gene by STRING 10.0 web tool showed eleven functional partners Cys- tathionine-beta-synthase [CBS], Methylene tetrahydrofolatereductase [MTHFR], Methionine adenosyltransferase 1 A [MATIA], Adenosylhomocysteinase [AHCY], Methionine adenosyltransferase 2 A [MAT IIA], Dimethylglycinedehydrogenase [DMGDH], Sarcosinedehydrogenase [SARDH], Adenosylhomocysteinaselike 2 [AHCYL2], Adenosylhomocysteinaselike
Discussion
Confidence score of interactions between BHMT and its closely related functional genes by STRING 10.0 tool
Confidence score of interactions between BHMT and its closely related functional genes by STRING 10.0 tool
Abbreviations: CBS, Cystathionine-beta-synthase; MTHFR, Methylenetetrahydrofolatereductase; MATIA, Methionine Adenosyltransferase 1A; AHCY, Adenosylhomocysteinase; MATIIA, Methionine Adenosyltransferase 2A; DMGDH, Dimethylglycine Dehydrogenase; SARDH, Sarcosine Dehydrogenase; AHCYL2, Adenosylhomocysteinase Like 2; AHCYL1, Adenosylhomocysteinase Like 1; CTH, Cystathionine Gamma-Lyase; MATIIB, Methionine Adenosyltransferase 2B. “–”: represents no interaction.
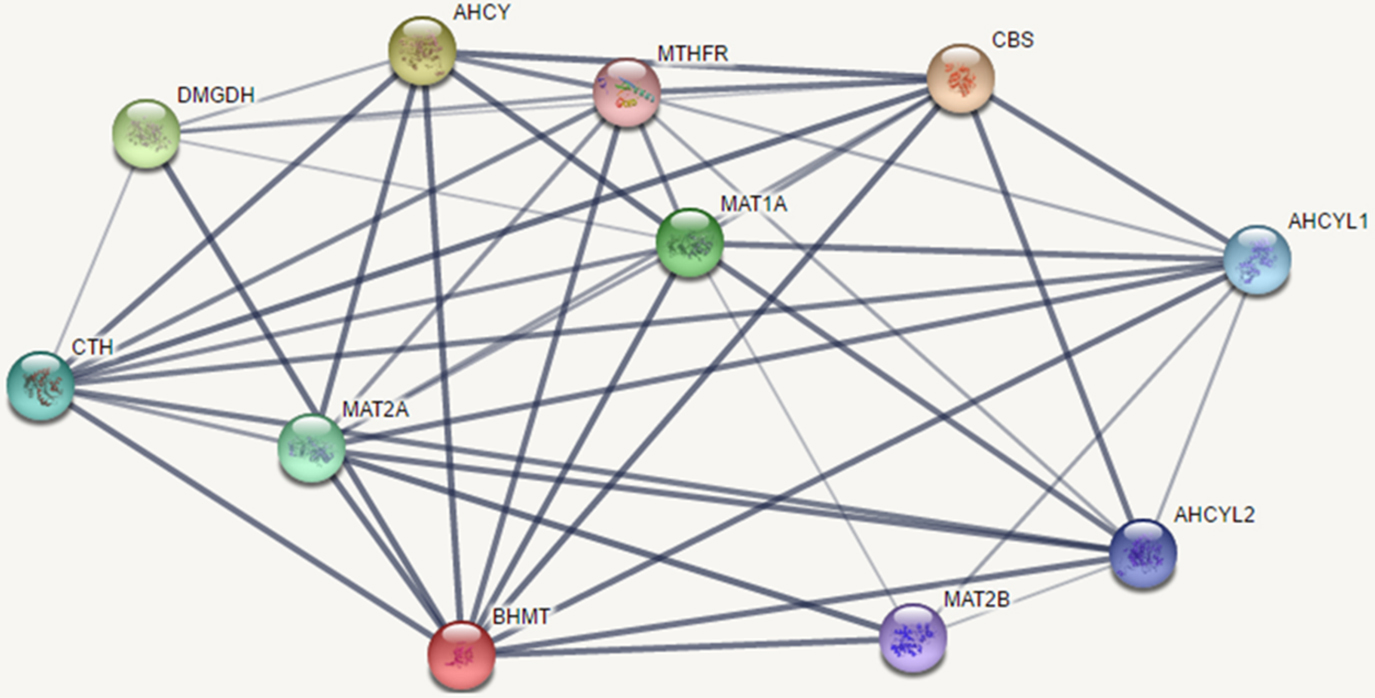
Network analysis of BHMT gene by STRING 10.0 tool. The network image represents the interaction between BHMT protein with closely related protein. Abbreviations: cystathionine-beta-synthase [CBS], Methylenetetrahydrofolatereductase [MTHFR], Methionine Adenosyltransferase1A [MATIA], Adenosylhomocysteinase [AHCY], Methionine Adenosyltransferase2A [MATIIA], Dimethylglycine Dehydrogenase [DMGDH], Sarcosine Dehydrogenase [SARDH], Adenosylhomocysteinase Like 2 [AHCYL2], AdenosylhomocysteinaseLike 1 [AHCYL1], Cystathionine Gamma-Lyase [CTH], and Methionine Adenosyltransferase 2B [MATIIB].
The folate and choline metabolisms are closely coupled pathways for homocysteine remethylation resulting in cellular methionine production. The remethylation of homocysteine to methionine by using 5-methyltetrahydrofolate as a methyl donor is catalyzed by methionine synthase and remethylation reaction by using betaine as a methyl donor is catalyzed by zinc-dependent cytosolic betaine-homocysteine methyltr- ansferase. However, the less availability of 5-methy- ltetrahydrofolate and betaine can cause the abnormal DNA methylation due to an imbalance in the development of SAM (S-adenosylmethionine) from methionine. There is no consistent association of polymorphisms in genes of folate metabolism including 5-methyltetrahydrofolate homocysteine methyltransferase (MTR) rs1805087 and methylene tetrahydrofolate reductase gene (MTHFR) rs1801133, rs1801131 in the ALL individuals [40, 41, 42, 43]. Hence, it is mandatory to focus on other genes involved in the homocysteine remethylation to draw a conclusion for the link between the ALL development and homocysteine remethylation.
The observations of the present study include an insignificant association of BHMT (rs3733890) with the risk of ALL in South Indian population and effect of other variations in the BHMT gene (including SNPs in the coding region, 3
In recent past, few evidence of association of BHMT (rs3733890) with few other cancers has been highlighted. In the case of uterine cervical cancer, BHMT(rs3733890) polymorphism acts as protective fac-tor [14], in the event of breast cancer this polymorphism is associated with reduced mortality [15], and in the event of head and neck cancer this polymorphism along with tobacco consumption increases the risk of head and neck squamous cell carcinoma [16]. BHMT (rs3733890) perhaps has also shown association with many other diseases. In the case of Down’s syndrome BHMT 742AA genotype reduces the risk in Brazilian population [11], in the case of neural tube defect of Caucasian American population the rs3733890 polymorphism acts as a significant risk factor [12], in regard to orofacial clefts of Polish population this polymorphism works in a protective fashion [13]. The pattern of association of this rs3733890 polymorphism conceivably different for different diseases but however it has been focused that the choline supplement can reduce the mortality of breast cancer thus it explored a hopeful strategy to adopt new lifestyle after cancer is diagnosed [15]. In our population, both MTHFR (rs1801133 and rs1801131) and BHMT (rs3733890) gene polymorphisms showed insignificant association with ALL [43]. This outcome may also be attributed to small sample size and distribution of minor allele in the study population.
Further, it may be necessary to evaluate the combined effect, and different haplotypes of pathogenic nsSNPs reported to be associated with the ALL and other cancers. The expansion of informatics into the field of biology has opened the windows for much graphical stimulation, one of which includes the prediction of disturbances in protein stability or structure caused due to changes in the sequence. Since the association analysis of the present study failed to reveal a significant association of BHMT (rs3733890) with ALL, we performed in-silico search for other non-synonymous SNPs and SNPs in 3
In summary, BHMT (rs3733890) showed insignificant association with ALL in South Indian population and the molecular mechanisms underlying the modulation in folate levels and ALL development in South Indian population is still unresolved. We further propose that the effective SNPs including rs59109725, rs116634518, rs138578732 and 11 CNVs which are present BHMT gene needs to be characterized in an attempt to better interpret the molecular mechanism of folate and cobalamin-dependent association of ALL for effective treatment and diagnosis in the future.
Supplementary data
The supplementary files are available to download from
Footnotes
Acknowledgments
This study was supported by Department of Biotechnology (DBT), Indian Council of Medical Research (ICMR), Technology Information Forecasting and Assessment Council-Centre of Relevance and Excellence (TIFAC-CORE) in Pharmacogenomics under Department of Science and Technology (DST), Government of India and Dr TMA Pai Endowment Chair for Pharmacogenomics, Manipal University.
