Abstract
BACKGROUND:
Breast cancer is the most common cancer among females, with an incidence of 6,41,000 cases annually. The genetic makeup of the individuals, ethnicity, geographical location, lifestyle, and BMI are some well-described factors associated with breast cancer. It is well known that pathogenic variants in BRCA1 and BRCA2 are associated with a majority of hereditary breast cancer. Genome-wide association studies (GWAS) have identified more than 80 germline susceptibility loci responsible for hereditary breast cancer.
METHODS:
In the present study, analysis of 94 genes associated with hereditary cancer was performed using next generation sequencing (NGS) in twelve patients having breast cancer and suspected with hereditary association.
RESULTS:
Four out of twelve (33%) patients harbored pathogenic mutation of the BRCA1 gene. Two patients was identified p. E23Vfs*17 mutation in BRCA1, one patient had p.Glu1580Gln in BRCA1, and a novel frameshift variant p.T1456Ifs*9(c.4367Cdel) in one patient.
CONCLUSION:
In the present study, out of four detected mutations in the BRCA1 gene, three were known and one was a novel BRCA1 mutation. It is advised to perform NGS-based genome sequencing to identify the genetic predisposition in breast cancer patients.
Keywords
Introduction
Breast cancer is the most common cancer among females, with an incidence of 6,41,000 cases annually. It has become one of the most common causes of cancer-related death among females accounting for 9–15% of all cancer deaths [1,2]. Studies have shown that among all breast cancer cases, 25% are of the familial type and 10% are of the hereditary type [3]. Patients with a hereditary type of breast cancer harbor inherited known pathogenic genetic variants and they may transmit to offspring or biologically related individuals. However, in the familial type, the cause of susceptibility of family members and its association with a causative genetic mutation may be unclear [4]. Studies have reported that globally 25–50% of hereditary breast cancer patients may harbor BRCA1 or BRCA2 mutation and its frequency depends on the ethnicity [5]. Genome-wide association studies (GWAS) helped to identify more than 80 genes associated with germline susceptibility for breast cancer other than BRCA1/BRCA2 [6]. Influence of patient’s ethnicity is associated with the type of genetic alteration and its penetrance, like in Ashkenazi Jewish women, BRCA1 or BRCA2 harbor up to 20% with a risk of 10- to 15-fold higher than non-Jewish [7]. In a study from India, nine known and six novel pathogenic mutations in BRCA1/BRCA2 and two mutations in CHEK2 were identified to be associated with Hereditary Breast-Ovarian Cancer Syndrome (HBOC) [8]. A better understanding of the predisposition of hereditary and familial cancer requires ethnic population-based GWAS studies.
In the field of genetics, Next-generation sequencing (NGS) has brought about a revolution by simplifying the process involved in multiple-gene sequencing. The NGS technology is being used extensively in cancer research and diagnosis [9]. The genetic bases in more than 20% of familial cancer are unknown to date hence identification of cancer-associated new variants associated is needed [10]. Therefore, in the present study, we have profiled hot spot regions of 94 genes using NGS in breast cancer patients with familial disease clustering.
Materials and methods
Patients and samples
Patients referred to the genetic counselling clinic of the Malabar Cancer Center in the year 2019 were included in the study. One hundred ten patients were registered in the cancer genetics clinic. Among the 110 patients assessed, 70 individuals were diagnosed with breast cancer. Out of these, 44 patients exhibited the following characteristics either individually or in combination: age under 40, had triple negative breast cancer, and had one or more first degree relatives affected by cancer. Out of which 12 gave informed consent for genetic testing. The study was performed in accordance with the Good Clinical Practice guidelines and approved by the MCC Ethics Committee (IEC meeting dated 13 Aug 2019 Agenda 1). Information about the age at diagnosis, histological type, grade, stage, and ER/PR/Her-2 receptor status was collected along with other clinical details. 2 ml of blood from the selected cases was collected for DNA isolation.
Sequencing
Genomic DNA was extracted from blood using the QIAamp DNA mini kit (Qiagen) and quantified using the QubitdsDNA BR Assay Kit (Thermo Fisher Scientific). Sequencing libraries were prepared with 50 ng of genomic DNA using the enrichment protocol “TruSight Cancer (Illumina)” for parallel sequencing of a panel of 94 genes (Supplementary Table ). The sequencing was performed using the MiSeq platform (Illumina) with MiSeq Reagent Kit v2, 2 × 150 cycles, according to the manufacturer’s instructions.
Data analysis and variant calling
NGS data analysis was performed using IlluminaBasespace Sequence hub, a cloud-based server for storing and analyzing NGS data. To ensure good call quality and to reduce the number of false positives, samples underwent proper quality analysis and filtering. NGS data alignment onto reference human genome (UCSC hg19) was performed with BWA Aligner application using BWA-MEM aligner. The BWA Enrichment app was used to analyze target DNA sequences with alignment and GATK pipeline for variant calling; variant analysis was performed for just the target regions. The Enrichment application used the rapid alignment and variant detection tool for targeted sequencing data. The Enrichment workflow maps reads using the Isaac aligner, performs small, structural, and copy number variant calling, annotates variants, and calculates enrichment metrics.
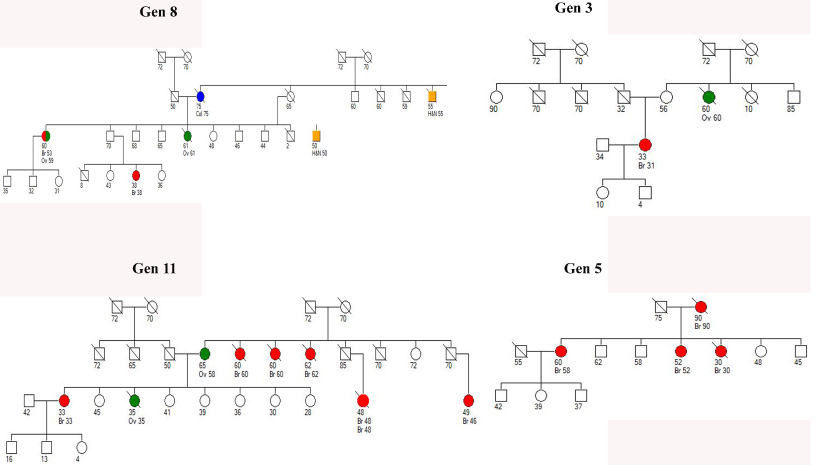
Pedigree of patients harbouring BRCA1.
Table showing patient’s details
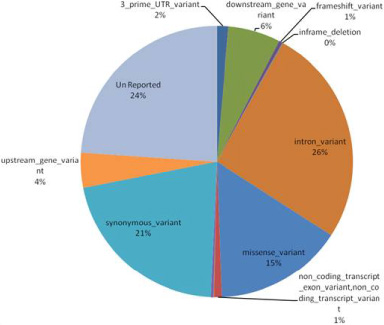
Graph showing frequency of different type of mutation detected in 12 cases.
IARC recommendations were used to classify the identified variants; they were classified into pathogenic, benign, and Variant of Unknown Significance (VUS) [11]. BRCA Share for identifying the pathogenic variants in BRCA1/2 was used [12]. dbSNP, ClinVar, and Leiden Open Variation Database (LOVD) were used to annotate the variants [13,14]. Variants not present in any of these databases were classified based on their characteristics. Only mutations introducing a premature stop codon (frame-shift and nonsense) and gross deletions were considered pathogenic/likely-pathogenic and classified in accordance with the guidelines of the American College of Medical Genetics (ACMG) [11].
Results
Clinicopathological and variants details of patients
We performed NGS analysis using 94 genes panel, covering all the hotspot regions (Supplementary Table S1). 12 cases out of 110 were selected based on the total number of first and second-degree relatives affected with cancer, age of index case or triple-negative breast cancer (TNBC) and willingness to perform NGS analysis (Fig. 1). 5 out of 12 patients (Table 1) had age below 40 years, one had bilateral breast cancer, and 4 patients had more than two first degrees relatives with history of cancer. Pathologically three patients had TNBC.
Samples harboring BRCA1 mutation
Samples harboring BRCA1 mutation
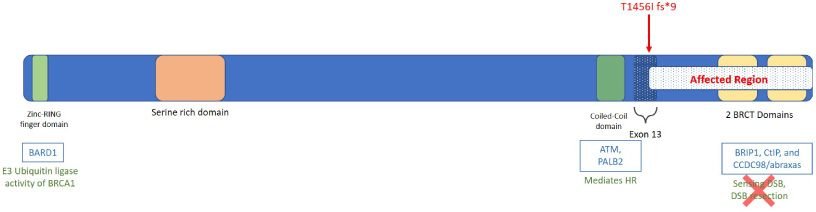
(a) BRCA1 novel variant p.T1456Ifs*9 location at genome and protein level, Schematic protein structure BRCA1, mutation of p.T1456Ifs*9 mutation is shown with lost region and altered pathway.
All the samples evaluated for NGS testing had Q30 above 94%, with 100x coverage above 1,90,000 bases (75%). An average of 88 variants per sample (range 39–127) was identified. All variants were categorized into synonymous or missense variants. Synonymous variants were detected in 68 genes; missense variants in 65 genes and frame-shift variants in 11 genes (Fig. 2). Eight pathogenic variants were detected in a total of six samples. Out of these, only three variants were known to be associated with breast cancer and one was a novel variant predicted as pathogenic.
In the present study, pathogenic/likely-pathogenic mutations were detected in four cases, out-off which three were known BRCA1/BRCA2 mutations, and in one case a novel mutation in BRCA1 was identified, which was predicted as pathogenic (Table 2). The patient with TNBC harbored a novel BRCA1 mutation and had two first-degree and three second-degree relatives affected with cancer (Fig. 1). Two out of four patients with BRCA1 pathogenic/likely-pathogenic mutation had one or more first-degree relatives having cancer. One 30-year-old patient without any affected first-degree relative also harbored BRCA 1 mutation however histologically the tumor was a triple-negative subtype.
In this study, three different types of BRCA1 mutations were identified: p.E23Vfs*17, p.Glu1580Gln and p.T1456Ifs*9. Patient 1 had the p.E23Vfs*17 mutation and was diagnosed with TNBC at age 30. There was no known cancer history in their immediate family. Patient 2 also had the p.E23Vfs*17 mutation and was diagnosed with breast cancer at age 52. She had one first-degree relative with cancer. The p.E23Vfs*17 (c.68_69CTdel) mutation is located in the ring finger+NES1 domain and results in a non-functional truncated protein, making it a well-known pathogenic variant associated with hereditary breast cancer. Patient 3 had the p.Glu1580Gln mutation and was diagnosed with TNBC at age 58. She had three first-degree relatives affected by cancer. Patient 4 had the novel p.T1456Ifs*9 (c.4367Cdel) frame-shift variant in the BRCA1 gene and was diagnosed with TNBC at age 33. She had two first-degree relatives with cancer (Fig. 3).
Samples harboring mutation other than BRAC1
Samples harboring mutation other than BRAC1
One pathogenic/likely-pathogenic mutation was identified (Table 1) in both the EHBP1 and PTEN. Both EHBP1 and PTEN mutation were present in a patient having metaplastic spindle cell carcinoma of breast carcinoma (Table 3).
Other variants
Non-pathogenic and Variants of Unknown significance (VUS):- 545 non-pathogenic variants and 300 variants having no information about the clinical significance (VUS) were identified in the twelve patients.
Discussion
In the present study, 94 genes were analyzed by NGS in 12 breast cancer patients from the northern part of Kerala, India. Pathogenic/likely-pathogenic mutations were detected in 5/12 (41.2%) patients, four patients harbored BRCA1 mutation. The two non-BRCA mutations were present in one patient and she had wild-type BRCA genes. The association between breast cancer predisposition and mutation identified in the present study in EHBP1 or PTEN is not known. It has been reported that patients with mutated BRCA1/BRCA2 develop breast cancer at a younger age and have affected first-degree relatives (Fig. 1). In the present study, 2/4 (50%) patients with BRCA gene mutation had ages below 35 years, and 3/4 (75%) patients had at least one first-degree relative previously diagnosed with cancer. The studies have shown that among TNBCs, BRCA1 mutation is more common than BRCA2 mutation [16]. This study had four cases of TNBC out of which, three cases had pathogenic mutations in BRCA1 gene.
One patient had a bilateral tumor and her two first-degree relatives were affected with cancer, hence it was expected that she may be harboring the carrier pathogenic mutation. However, we could not find any pathogenic mutation in that patient. Studies have shown that up to 50% of breast cancer patients with familial history may harbors BRCA mutation [17]. Both tumors were histopathologically IDC type grade 2 with 10--20% Ki-67 staining. However, poorly differentiated tumors and higher Ki-67 expression are reported as characteristic features of breast cancer cases with BRCA mutation [18,19]. This indicates that extensive GWAS are needed to understand the genetic factors associated with bilateral breast cancer.
The novel BRCA1 mutation (p.T1456Ifs*9) may affect the C-terminus of the protein altering the BRCT1/2 domains. In hereditary breast and ovarian cancers, the most altered domain of BRCA1 gene is BRCT1 domain. It is mainly involved in the interaction with CCDC98, CtIP, ABRAXAS, and BRIP1 via phosphopeptide-binding domains [20–22]. The C- terminal region of BRCA1 is known to be involved in the DNA repair mechanism, hence the loss of the C-terminal may affect the DNA repair system [23,24]. The patient harboring novel frame-shift mutation was presented with TNBC disease at the age of 33, and two of her first and three second-degree relatives were affected with cancer indicating the strong hereditary association of disease and high penetrance of the causative mutation, hence now the family members can be enrolled for the screening process.
Other variants in EHBP1 and PTEN genes are also known to be pathogenic and associated with an increased risk of other cancer. One study has shown that EHBP1c.1185+30064G >A(rs721048) is generally present in 31% and 4% of the general population as heterozygous and homozygous. In prostate cancer presence ofrs721048 SNP is found to be associated with aggressive prostate cancer [25]. PTEN variant p.Tyr178del (rs587780711) was documented as likely pathogenic. One study has shown that variants in the nearing location as benign [26]. In another study, the mutations in PTEN are found to be associated with the Bannayan-Riley-Ruvalcaba phenotype [27]. However, extensive genetic studies are needed to conclude the genetic association of such gene mutation with cancer. These results underscore the importance of using multi-gene approach for identifying the genetic cause of hereditary breast cancer than only BRCA1/2 genes testing. A population-based genetic database is also needed for predicting the pathogenicity of rare or unrelated variants.
It is very unlikely for breast cancer patients to harbor germline BRCA1 pathogenic mutation without a strong family history. However, few studies have shown that breast cancer patients aged below 50 and having the TNBC subtype may have germline BRCA1 mutation [28,29]. There are a few guidelines like NICE (National Institute for Health and Care Excellence), that have suggested genetic testing in this group of patients. Hence, it will be advisable to have genetic testing for breast cancer patients with TNBC type at a younger age, especially below 50 years.
Our observation showed that 50% of the cases, 6 out of 12, did not have any identified pathogenic mutations. Despite this, all patients had at least one first-degree relative who had previously been diagnosed with cancer. This highlights the necessity for more extensive and improved genome-wide studies that could potentially reveal any concealed pathogenic mutations.
NGS-based studies lead to identification of many variants that are not easy to classify as pathogenic or non-pathogenic. Several techniques can now be used to determine pathogenicity of such hard to classify mutations [30], yet quick, efficient and accurate methods for classifying variants are needed for translating this information into clinical practice. However, the present study strongly suggests a structured genetic testing protocol should be implemented for detecting the genetic alteration in breast cancer patients.
Footnotes
Acknowledgements
We thank all investigators and Mr KK Ragesh (RajyaSabha MP, Govt of India) for providing (MP LAID fund) funding support for setting up the genomic facility.
Conflict of interest
There is no confilict of interest.
