Abstract
BACKGROUND:
Benign and malignant breast tumors are the most commonly diagnosed tumor in females. Early and accurate diagnosis of malignancy is essential for effective breast cancer treatment. Human anterior gradient 3 (AGR3) has been suggested as a potential biomarker for the early detection and prognostic determination of breast cancer.
OBJECTIVE:
This study profiles AGR3 mRNA expression and serum protein levels in patients with benign and malignant breast tumors.
METHODS:
A case-control study was conducted on 40 benign and 40 malignant breast tumor patients in Makassar, Indonesia. AGR3 mRNA and protein were detected using qRT-PCR and ELISA, respectively.
RESULTS:
This study found significantly higher AGR3 mRNA expression in benign than malignant breast tumors using qRT-PCR (p < 0.001). In contrast, ELISA revealed no significant difference between AGR3 serum protein levels in benign and malignant breast tumors (p = 0.507).
CONCLUSIONS:
AGR3 is associated with non-aggressive tumors and could be used as a marker for less aggressive breast tumors.
Introduction
The most commonly diagnosed tumors in females are benign and malignant breast tumors, with approximately 10–20% confirmed by biopsy and approximately 50% by autopsy. As breast cancer is the leading cause of cancer deaths in female populations worldwide, the early detection and accurate diagnosis of malignant breast tumors are critical for successful treatment [1]. Breast cancer survival rates currently vary from 40–80%, with an average estimated survival period of 5 years [2,3]. In 2018 alone, an estimated 2,088,849 new breast cancer cases were diagnosed, and 626,679 breast-cancer-related deaths were reported, 60% of which occurred in developing countries.
Breast cancer can be detected by physical examination, radiological examinations (mammography, ultrasonography, and MRI), tissue biopsy, and testing for the expression of diagnostic markers including estrogen and progesterone receptors, human epidermal growth factor receptor 2 (HER2), TP53 tumor suppressor, and BRCA1 and BRCA2 mRNA. When used in conjunction, these screening methods are effective for the early diagnosis of breast cancer [3,4]. Advancements in technology have allowed the continued development of diagnostic methods for breast cancer detection at the level of organs to biomolecules. For example, Garczyk et al. identified human anterior gradient 3 (AGR3) as a potential biomarker for the early detection and prognostic determination of breast cancer [5,6]. This study seeks to determine the relationship between AGR3 mRNA expression and serum levels in patients with benign and malignant breast tumors and establish the diagnostic and prognostic utility of AGR3 in breast cancer.
Methods
Subjects
This research was performed in Makassar, Indonesia, from March–November 2019. Female patients with benign and malignant breast tumors were included following written informed consent. Inclusion criteria involved the histopathological confirmation of benign and malignant breast tumors. Subjects who had previously received chemotherapy or targeting or hormonal therapy were excluded. Following acceptance into the study, 5 ml venous blood was collected from each patient for use in the ELISA assay. Tumor tissue was biopsied from each patient and used for the qRT-PCR analysis of AGR3 mRNA expression. Tumor characteristics were determined through standardized methods.
Human AGR3 ELISA examination
Materials
Standard solution (24 ng/mL), pre-coated ELISA plate, standard diluent, streptavidin-horseradish peroxidase (HRP), stop solution, substrate solution A, substrate solution B, wash buffer concentrate (25×), biotinylated human AGR3 antibody (anti-AGR3), plate sealer, and zipper bag.
Assay procedure
The reagents, standard solution, and samples were prepared, and the number of strips required for the assay was determined. Standard (50 μL) and sample (40 μL) were added to each well of the ELISA plate, followed by the addition of anti-AGR3 (10 μL) and streptavidin-HRP (50 μL). The microtiter plate was covered with a clean adhesive strip and incubated for one h at 37 °C. The seal was then removed, and the plate was washed 5 × 30 s. Substrate solutions A and B (50 μL) were added to each well, and the plate was incubated in the dark for 10 min at 37 °C. Stop solution (50 μL) was added to each well, resulting in a color change from blue to yellow. Within 10 min of adding the stop solution, the OD at 450 nm was measured using a microplate reader.
Human AGR3 qRT-PCR examination
Materials
CFX96 Touch TM Real-Time PCR Detection System (Bio-Rad Laboratories, Inc., Beijing, China), refrigerated centrifuge (Thermo Fisher Scientific, Waltham, MA, USA), vortex (Bio-Rad Laboratories, Inc.), mini-centrifuge (Bio-Rad Laboratories, Inc.), laminar flow PCR cabinet (Esco Micro Pte Ltd., Singapore, Singapore), micropipettes and tips (10, 100, 1000 μL; Bio-Rad Laboratories, Inc.), RNA isolation kit (RNAprep Pure Tissue Kit; TIANGEN Biotech (Beijing) Co., Ltd., Beijing, China), cDNA kit (iScript TM cDNA Synthesis Kit; Bio-Rad Laboratories, Inc.), qPCR master mix (SsoFast EvaGreen supermix; Bio-Rad Laboratories, Inc.), nuclease-free water (Norgen Biotech Corp., Thorold, ON, Canada), Eppendorf tubes (0.2 and 1.5 mL), and qPCR plates/strip tubes. qPCR primers: IGF-1 (NM_184052) = Forward: GTGGATGCTCTTCAGTTCGTGTG, Reverse: TCCAGTCTCCTCAGATCACAGC; GLUT-2 (NM_031197) = Forward: GTTGGAAGAGGAAGTCAGGGCA, Reverse: ATCACGGAGACCTTCTGCTCAG; GCK (NM_010292) = Forward: GCATCTCTGACTTCCTGGACAAG, Reverse: CTTGGTCCAGTTGAGCAGGATG; and GADPH Mouse (NM_008084) = Forward: CATCACTGCCACCCAGAAGACTG, Reverse: ATGCCAGTGAGCTTCCCGTTCAG.
RNA extraction using the RNAprep pure tissue kit
Tissue (20 mg) and RL buffer (300 μL) were combined in a 1.5 mL tube. The tissue was macerated with a pestle, centrifuged for 5 min at 12 000 rpm, and the supernatant was transferred to a sterile 1.5 mL tube. Ethanol (150 μL) was added to the supernatant and mixed by pipetting. The lysate was transferred to a CR3 spin column and centrifuged for 1 min at 12 000 rpm. After removing the supernatant, RW1 buffer (350 μL) was added to the spin-column, which was then centrifuged for 1 min at 12 000 rpm, treated with 80 μL DNase I, and incubated at room temperature for 15 min. The spin-column was treated with RW1 buffer (350 μL), centrifuged for 1 min at 12 000 rpm, and isolated from the supernatant. The spin-column was then treated with RW buffer (500 μL), centrifuged for 1 min at 12,000 rpm, isolated from the supernatant, and centrifuged for 2 min at 12,000 rpm. After removing the supernatant, the spin column was transferred to a sterile 1.5 mL tube, layered with RNase-free water (50 μL), incubated at room temperature for 2 min, and centrifuged for 2 min at 12 000 rpm. The eluted RNA was then stored at −70 °C or directly analyzed by qRT-PCR using GCK, GLUT-2, and IGF-1 primers.
Data analysis
The clinicopathological characteristics of the subjects were analyzed using SPSS software (IBM Corp., Armonk, NY, USA). Data were analyzed using the non-parametric Mann–Whitney U test for comparative analysis between groups, with p < 0.05 considered statistically significant.
Ethical approval
Ethical approval was obtained from the Commission for Research Ethics, Medical Faculty, Universitas Hasanuddin, Makassar, Indonesia (Ethics Registration Number: 1007/UN4.6.4.5.31/PP36/2019).
Results
Subject characteristics
A total of 80 patients – 40 with benign and 40 with malignant tumors – were recruited for this study. The clinicopathological characteristics of the participants were recorded (Table 1). Within the benign group, most subjects were younger than 40 and displayed fibroadenoma histopathology. Within the malignant group, most subjects were older than 40 and displayed invasive breast carcinoma histopathology (80%), with the remaining malignant histopathologies identified as mixed invasive carcinoma (12.5%), intraductal papillary carcinoma (5%), and mucinous carcinoma (2.5%). The Nottingham histological grade categorized the malignant tumors as low (30%), moderate (47.5%), and high (22.5%) grade. Immunohistochemistry described the subtypes as luminal A (35%), luminal B (27.5%), HER2 (15%), and triple-negative (22.5%).
Clinicopathological characteristics of participants
Clinicopathological characteristics of participants
AGR3 mRNA expression as measured by qRT-PCR was significantly different between benign and malignant breast tumors (p < 0.001) (Table 2 and Fig. 1).
AGR3 mRNA expression by qRT-PCR
AGR3 mRNA expression by qRT-PCR
∗ Mann–Whitney U. IQR = Interquartile range.
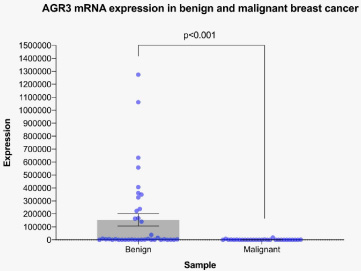
AGR3 mRNA expression in benign and malignant breast tumors by qRT-PCR. The bar graph shows mean ± standard error of the mean of n = 39 benign and n = 29 malignant samples. p = 0.001 by the Mann–Whitney U test.
AGR3 serum protein levels as determined by ELISA showed no significant difference between patients with benign and malignant breast tumors (p = 0.507) (Table 3 and Fig. 2).
AGR3 serum levels by ELISA
AGR3 serum levels by ELISA
∗ Mann–Whitney U. IQR = Interquartile range.
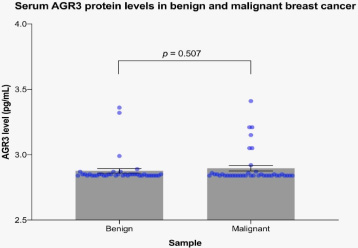
AGR3 serum protein levels in patients with benign and malignant breast tumors as measured by ELISA. The bar graph shows mean ± standard error of the mean of n = 40 samples/group. p = 0.507 by the Mann–Whitney U test.
Breast cancer has attracted substantial attention as a major cause of cancer-related deaths in women. The identification of potential biomarkers is essential for diagnostics can aid in selecting the course of treatment and can function in the prognosis of breast cancer [7]. In clinical practice, breast cancer markers have been identified based on four molecular subtypes: the estrogen receptor, progesterone receptor, Ki-67, and HER2 [7,8]. However, the anterior gradient protein complex (AGR) has also been reported to be present in numerous tumor variants, including breast, ovary, and prostate tumors.
The AGR family has three members: AGR1, AGR2, and AGR3. Although AGR1 lacks in higher vertebrates, likely due to decreased regeneration capacity, AGR2 and AGR3 are still expressed. Only AGR2 has been reported to function as a biomarker, with the role of AGR3 yet to be determined. Although the function of AGR3 in cancer is currently unknown, it has been associated with tumor diagnosis [9]. Xu et al. reported that AGR3 was expressed in patients with invasive ductal carcinoma with a luminal subtype histological class I–II and could predict a poor prognosis for the luminal B subtype [9]. They suggested that AGR3 misexpression could disrupt cell function and promote the proliferation and metastasis of cancer cells, accelerating the development of breast cancer [10]. AGR3 has also been found in ciliated cells of cyst epithelial inclusions by double immunofluorescence staining. Its expression was also reported in serous borderline ovarian tumors, consistent with an association with reproductive hormones [11].
AGR3 was initially identified as a membrane protein by proteomics of breast cancer cell lines and is also known as breast cancer membrane protein 11 (BCMP11). AGR3 is an estrogen receptor–resident protein required for regulating the frequency of ciliary movements and mucociliary cleaning in the airway epithelium [12]. It contains a catalytically active thioredoxin domain that is thought to influence growth and metastasis in breast cancer. In recent decades, isomerase disulfide protein derivatives, such as AGR2 and AGR3, have attracted researchers’ attention because of their involvement in carcinogenesis. AGR2 is known to be a pro-oncogene, and AGR3 is its homolog with 71% sequence identity.
This study found that AGR3 expression was higher in benign tumors than in malignant tumors, consistent with work by Obacz et al. which concluded that AGR3 was more readily detected in less aggressive tumors with better prognoses [13]. AGR2 and AGR3 are overexpressed in various adenocarcinomas and can be present in intra- and extracellular compartments, as evidenced by their secretion in specimens collected from cancer patients. In a study investigating Src kinase signaling, Obacz et al. showed that extracellular AGR3 (eAGR3) plays a role in regulating tumor cell adhesion and migration [14]. The authors also concluded that eAGR3 could function as a molecular signal in the tumor microenvironment [14].
This study found no significant results distinguishing benign and malignant breast tumors by ELISA than the qRT-PCR results. This could be related to the sensitivity and specificity of each test. Existing research suggests that a combined ELISA-PCR method provides good accuracy. Sue et al. found that ELISA-PCR performed better than ELISA, conventional PCR, and even qPCR, both in cost and diagnostic ability [15]. However, the non-statistically significant results obtained in this study prevent a conclusive recommendation of any particular method.
Conclusions
This study found that AGR3 was more highly expressed in benign tumors by qRT-PCR. The detection of high levels of AGR3 in patients could thus suggest a therapeutic route targeted for benign cases. As AGR3 is associated with non-aggressive tumors, it could be used as a novel biomarker in breast tumor patients. Further research is necessary to identify the optimal diagnostic test.
Footnotes
Acknowledgements
The authors would like to thank Dr. Gita Vita Soraya for her advice on statistical analysis and all the breast cancer patients who participated in this study.
Conflict of interest
The authors declare that there are no conflicts of interest.
Funding
This work was supported by a Block Grant Research for Lecturer from the Faculty of Medicine, Hasanuddin University.
