Abstract
INTRODUCTION:
Losses of genetic material from chromosomes 5q and 16q commonly occur in sub-sets of breast cancer. Their significance from a pathophysiologic point of view is not well-defined.
METHODS:
This study uses publicly available genomic data from extensive breast cancer datasets to define the landscape of losses in chromosomal arms 5q and 16q in the two sub-types of breast cancer they most commonly occur, basal-like and luminal A cancers, respectively.
RESULTS:
It is shown that dozens of genes from these chromosomal arms are putatively hemi-deleted in few samples each. No individual gene from either 5q or 16q shows an incidence of deep deletion above 10% in the cohorts with basal-like and luminal A cancers or in the whole cohorts. A few tumor suppressor genes are deleted in a small number of samples, less than 5% in each cohort. Losses of 5q or 16q confer no survival advantage in either the basal-like or the luminal A cohorts from TCGA that harbor them.
CONCLUSION:
Results suggest that there are no individual genes in chromosomes 5q and 16q whose loss can be implicated in a dominant pathophysiologic sequence of events in breast cancer or its sub-sets.
Introduction
Identification and characterization of molecular alterations at the level of the whole genome has classified breast cancers in several intrinsic sub-types which guide treatment based on surrogate markers used in the clinic [1]. Luminal cancers are characterized by their expression of the Estrogen Receptor (ER) and are negative for expression or amplification of the HER2 receptor. They are further divided into luminal A which are more indolent and show a lower proliferation index and luminal B which are more aggressive and display higher proliferation. HER2-enriched cancers are characterized by over-expression or amplification of HER2 receptor. Basal-like cancers are negative for expression of hormone receptors, ER and Progesterone Receptor (PR), and negative for HER2, and hence they are referred to as triple negative. Basal-like cancers represent the most aggressive phenotype of breast cancer, for which there are currently fewer therapeutic options.
Other molecular classifications based on patterns of copy number alterations (CNAs) have been proposed and are only partially overlapping with the intrinsic sub-types [2]. A classification based on patterns of CNAs across the genome assigns breast cancers to 11 so-called integrative clusters [3]. These 11 integrative clusters have been identified based on the pattern of chromosome segment duplications or deletions and the most common chromosomes that present these patterns, using representational oligonucleotide microarrays. Four main patterns of copy number changes in different chromosome segments have been revealed with these analyses, including a simplex pattern with broad amplified and deleted segments, two complex patterns (complex I with generalized narrow areas of duplications and deletions giving a sawtooth appearance and complex II with one or more localized areas of high level of amplifications interspersed with deletions called a firestorm pattern) and lastly a flat pattern with no clear gains or losses of chromosomal regions. The simplex pattern characterizes mostly luminal A breast cancers and is observed commonly in integrative clusters 7 and 8 that are comprised of luminal A cancers. The complex I pattern is observed most characteristically in the integrative cluster group 10 which mostly consists of basal-like breast cancers. Several clusters containing luminal B and HER2-positive cancers display a complex II pattern. In addition to this type of classifications based on copy number alterations, next generation genomic studies have the ability to determine copy numbers of individual genes and chromosome arms based on pre-specified algorithms [4]. Gains and losses of genetic material from different chromosome arm areas are variably prevalent in the breast cancer sub-types and this observation has also been incorporated in the integrative cluster classification. Loss of genetic material from the long arm of chromosome 5 (5q), for example, is very common in basal-like cancers while is rarely observed in luminal A cancers [5]. Loss of 5q arm defines a specific sub-type of a hematologic malignancy, myelodysplastic syndromes (MDS). This MDS subtype responds to therapy with immunomodulatory drugs (IMiDs) such as lenalidomide and genes hemi-deleted in the arm 5q have been postulated to be targets of IMiDs [6]. In contrast, luminal A breast cancers have more commonly losses in the long arm of chromosome 16 (16q). 16q loss defines ER+ breast cancers and it has been suggested that it may harbor tumor suppressors involved in carcinogenesis of these cancers [7].
This paper evaluates the copy numbers of genes in chromosome arms 5q and 16q in breast cancers as established by published genomic studies to determine the implications of chromosome arm copy number alterations as called in these studies and the corresponding alterations of genes located in the same arms. An analysis of the role of hemi-deleted genes from these chromosomal locations is performed and prognostic implications are investigated.
Methods
Genomic and clinical data from The Cancer Genome Atlas (TCGA) breast cancer genomic study cohort [4] and the METABRIC (Molecular Taxonomy of Breast Cancer International Consortium) study cohort [8,9] were analyzed using the cBioCancer Genomics Portal (cBioportal, http://www.cbioportal.org) platform [10,11]. The cBioportal is an open platform that provides data from genomic studies from various sources such as TCGA. It allows for interrogation of each study included in the platform for genetic lesions (mutations, copy number alterations) and mRNA expression modifications in any gene of interest that is included in the original study and links these data with clinical information including age of the patients, tumor grade and stage as well as survival outcomes.
TCGA breast cancer cohort contains 1084 patients of whom 1070 were assayed successfully for CNAs. Among these, 499 samples are of the luminal A sub-type and 171 samples are basal-like. The METABRIC cohort includes 2509 patients with 2173 patient samples included In the CNAs analysis. In METABRIC, 700 samples are luminal A and 209 samples are basal. For the sub-type and integrative cluster restricted analyses, the samples of interest were selected in the respective pie graphs provided in cBioportal. Similarly, for the analyses of samples with or without the specific amplifications of arms of interest (5q and 16q), samples were highlighted and selected from the arm-level CNA pie graphs generated under the charts tab in cBioportal. Putative CNAs are called in TCGA based on GISTIC (Genomic Identification of Significant Targets in Cancer) algorithm and are recorded and presented in the oncoprint section of cBioportal. Putative deep deletions correlating with hemi-deletion of a certain gene are defined by a score of −2.
For the TCGA and METABRIC breast cancer studies included in this report, cBioportal provides information on the breast cancer sub-type of each sample, based on the expression of estrogen receptor (ER) and HER2, and on genomic profile. Other information of each sample such as chromosomal instability as measured by the Aneuploidy Score (AS) and total Fraction of Genome Altered as well as clinical characteristics and outcomes data are also included in the cBioportal platform. AS is a measure of chromosomal instability at the chromosome arms level. It is calculated as the sum of the number of chromosome arms in each sample that have copy number alterations (gains or losses). A chromosome arm is considered copy number altered if the aggregate somatic copy number alterations of segments derived from this arm cover more than 80% of the length of the arm based on Affymetrix 6.0 SNP arrays [12]. The list of genes with deletions in 5q and 16q were scanned for putative cancer-associated genes as annotated in the OncoKB database [13].
For analyses that could not be directly performed in the cBioportal platform, the primary clinical and genomic data of the breast cancer TCGA and METABRIC studies were transferred to an Excel sheet (Microsoft Corp., Redmond, WA) for further analysis and calculations. Survival data (length of observation or time to event and survival status) were downloaded from cBioportal with the “download clinical data of selected cases” function into Excel, groups were annotated, and data were up-loaded to an R-based online application enabling construction of Kaplan–Meier survival curves. For CNAs genomic data, cBioportal includes samples that were not called (i.e. could not be categorized as gained, lost or unaltered) as a separate group. In the current study these samples were excluded and percentages were recalculated based only on samples for which information was available. For example, the total number of samples informative for the status of chromosome 16q in TCGA is 888, as 196 samples could not be categorized in one of the three informative categories (gain, loss or not altered).
Statistical comparisons of categorical and continuous data were performed with the Fisher’s exact test or the x2 test and the t test, respectively. For continuous data, the Kolmogorov–Smirnov test for normalcy was performed to confirm a normal distribution of all populations compared and normalcy was confirmed (Kolmogorov–Smirnov test statistic p > 0.05 for all comparisons). Kaplan–Meier survival curves were compared using the Log Rank test. All statistical comparisons were considered significant if p < 0.05. Calculations were performed with the use of open online statistical calculators (www.socialstatistics.com and https://merser.shinyapps.io/survival) and the graphpad statistics site (www.graphpad.com/quickcalcs).
Results
Deletion of genetic material from chromosome arm 5q characterizes basal type breast cancers, being present in 89.1% of them in TCGA breast cancer study, while it is significantly rarer in other sub-types (31% in HER2 positive, 22.1% in luminal B breast cancers and 4.8% in luminal A breast cancers) (Fig. 1).
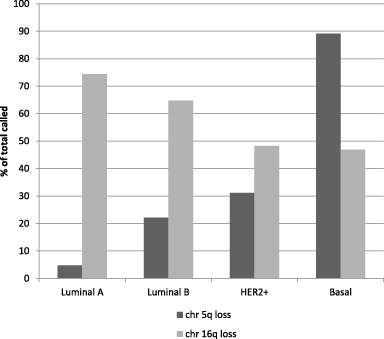
Loss of chromosome arms 5q and 16q in the sub-types of breast cancer as a percentage of total numbers of cases with data for the respective arms. Data are from TCGA breast cancer study.
Deletion of genetic material from chromosome arm 16q is common in luminal A breast cancers being observed in 74.4% of this sub-type in TCGA. The three other sub-types have lower rates of deletions of genetic material of arm 16q (64.6%, 48.3% and 46.9% in luminal B, HER2 positive and basal-like cancers, respectively) (Fig. 1). 16q deletions are the most frequent arm loss in luminal A cancers and is the only loss that is significant more frequent in this sub-type of breast cancer than in basal-like sub-type (Fig. 2). In contrast, 5q and several other chromosome arms including 17p, 14q, 15q, 4p, 4q and 8p are more frequently deleted in basal-like cancers (Fig. 2).
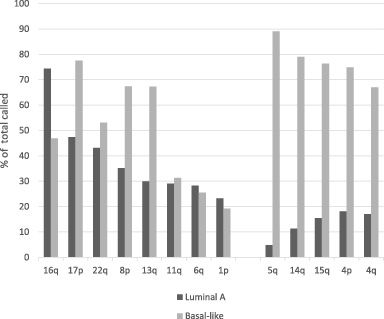
Left. Most frequently hemideleted chromosome arms in luminal A breast cancers and corresponding prevalence of these losses in basal-like cancers. For several frequent lost arms in luminal A cancers the frequencies of losses are even higher in basal-like cancers. Right: Additional prevalent arm losses (>60% of cases called) in basal-like cancers and corresponding frequency in luminal A cancers.
Through this paper “hemi-deleted” and “deleted” as well as “hemi-deletion and “deletion” are used interchangeably and are defined as putative loss of one gene copy. In luminal A cancers several dozens of 5q genes are putatively hemi-deleted in isolated samples or in two or three samples in TCGA cohort (Fig. 3). No individual 5q gene is hemi-deleted in more than three luminal A samples. Several genes from chromosome arm 16q are deleted in seven to fifteen luminal A cancers (1.3%–3%) with a peak at ten samples (2%) (Fig. 3). In basal-like cancers, genes from both arms 5q and 16q are commonly hemi-deleted in isolated and up to three cases (1.7%) (Fig. 4). A higher number of 5q genes (n = 554) are hemi-deleted in isolated cases or in two to three samples than genes of 16q (n = 257) in basal-like cancers. Moreover, hemi-deletions are observed in genes of 5q in 4–17 samples (2.3–9.9%) in 274 additional 5q genes (Fig. 4). However, among those genes, only nineteen had been deleted in more than 5% of cases (9–17 samples).
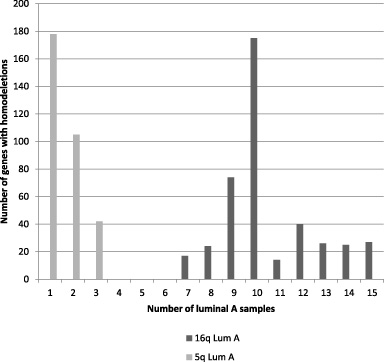
Distribution of number of genes from 5q or 16q with hemi-deletions in luminal A samples.
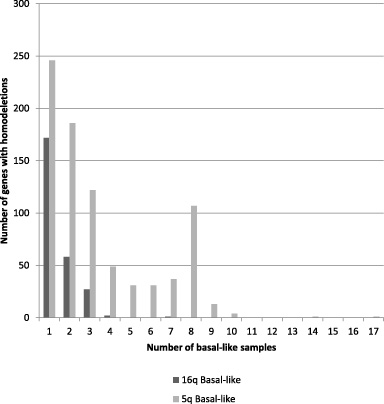
Distribution of number of genes from 5q or 16q with hemi-deletions in basal-like samples.
Focusing on putative cancer-related genes as described by the OncoKB database, among the 171 basal-like carcinomas in TCGA breast cancer study, 23 genes located in the 5q chromosomal arm and annotated in the OncoKB cancer gene database had a deep deletion in one or more samples (Table 1). The luminal A subtype of breast cancer with 499 samples in TCGA had a smaller number of seven genes from 5q included in the OncoKB cancer gene database with deletion. Six of these seven genes were among the 23 genes of the basal cancers and only one (SNCAIP) was present in a single luminal A case and was not observed in the basal cohort (Table 1). Among the 23 genes, decreased mRNA expression was rarely observed in TCGA samples and they were not associated with mutations of the same gene that might suggest complete loss of function (Fig. 5, upper panel). In the METABRIC study, 15 OncoKB-listed genes located in 5q display deep deletions. Those were mostly overlapping with the genes from TCGA (Table 2). In the integrative cluster 10, which is the main group where basal-like carcinomas segregate and contains a more homogeneous population of basal-like carcinomas, eight of these genes are putatively hemi-deleted in one or more samples (Table 2). Six of these eight are included in the 5q hemi-deleted genes of the basal-like sub-type from TCGA. Other integrative clusters have smaller numbers of 5q deleted OncoKB-listed genes ranging from none in cluster 2 and 6 to four in cluster 5 which is a cluster that contains mainly HER2 positive and some luminal B cases. A total of nine basal-like cases in METABRIC (0.4%) had clusters of two or more 5q hemi-deleted genes.
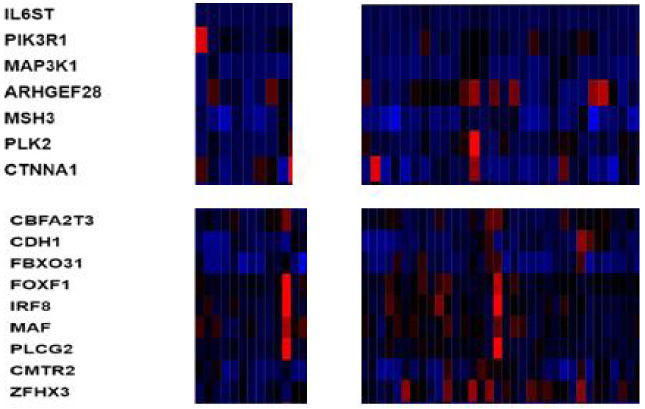
Upper panel: mRNA expression of representative hemi-deleted genes from 16q in luminal A breast cancer samples with hemi-deletions of these genes (left grid) and without hemi-deletions of these genes (right grid). Lower panel: mRNA expression of representative hemi-deleted genes from 5q in basal-like breast cancer samples with hemi-deletions of these genes (left grid) and without hemi-deletions of these genes (right grid). Data are from TCGA breast cancer study.
Genes of chromosome arm 5q hemi-deleted in basal-like and luminal A breast cancers. Data are from TCGA cohort
OG: oncogene, TS: tumor suppressor, ?: not designated. Genes that are also hemi-deleted in the METABRIC cohort are in bold.
Genes from arm 5q hemi-deleted in all samples and in integrative cluster 10 of METABRIC study. Deletions of all but 3 genes (AFF4, SNCAIP, IRF1) are also observed in TCGA basal-like cohort
The 499 luminal A breast cancers from TCGA contain 19 genes from 16q arm included in the OncoKB cancer gene database with hemi-deletion in one or more samples (Table 3). Fourteen of these genes are also hemi-deleted in one or more samples of the basal-like samples. Twenty-four cases (4.8%) contain deep deletions of a cluster of three or more of these genes, including ten cases (2%) with concomitant deletion of the whole cluster of 19 genes. Thirteen additional cases have deletion in one or two of these genes. Similar with the genes of 5q, decreased mRNA expression was only rarely observed in TCGA samples compared with samples possessing no deletions in these genes (Fig. 5, lower panel) and in only a single sample there was decreased mRNA expression associated with a concomitant mutation of the same gene (CBFB). In the METABRIC study, 19 genes included in the OncoKB cancer gene database located in 16q display deletions and those are completely overlapping with the 16q genes with hemi-deletions from TCGA (Table 4). All these 18 genes show deletion in one or more samples of the integrative clusters 7 and 8 which are clusters mostly comprised by luminal A cancers. Integrative cluster 10 show rare hemi-deletions in five of the 18 genes. No additional genes are deleted (Table 4). Overlapping hemi-deletion of more than one 16q genes in the same sample was observed in 27 cases (1.2%) in METABRIC. These results suggest that, despite loss of genetic material, coding chromosome areas are usually preserved and only a small minority of genes are deleted in any individual sample, independently of breast cancer sub-type. Deletion of individual genes that is observed in few samples is unlikely to be involved in pathogenesis of breast cancer but may be part of an underlying mechanism producing chromosomal instability.
OncoKB database listed genes hemi-deleted in 16q in basal-like and luminal A breast cancers. Data are from TCGA cohort
OG: oncogene, TS: tumor suppressor, ?: not designated as an oncogene or tumor suppressor.
Genes from 16q annotated in the OncoKB cancer gene database and hemi-deleted in all samples and in integrative clusters 7/8 and 10 of METABRIC study
A survey of deleted genes of arm 5q shows that nine genes are labeled in the KB database as tumor suppressors and six as oncogenes (Table 1). Similarly, regarding 16q arm genes, nine are listed as tumor suppressors in the KB database and two as oncogenes (Table 3). Interestingly, several tumor suppressors are amplified in a similar number of samples as they are hemi-deleted, including CYLD in 5 samples, ZFHX3 in 2 samples, NUP93 in 1 sample and EXOSC6 in 1 sample.
A survey of genes from 5q that are not listed in the KB database and are putatively deleted in 8 or more samples (≥4.7%) in basal-like cancers reveals a few additional candidate tumor suppressors or genes that are involved in cancer-related processes. Those include CCNB1 encoding for cyclin B1 and the genes encoding for centromere proteins CENPH and CENPK.
Genes belonging to the minimal deleted area of the 5q arm loss in the 5q- type of MDS (CSNK1A1, TNFAIP3, MIR-145/3P, MIR-146/146A, G3BP1, CD74, IL17B, CSF1R, SPARC, SLC26A2, FAT2, RPS14, RBM22, TCOF1, NDST1, CDC25C, EGR1) are hemi-deleted as a group in a single case of basal-like breast cancers and 2 additional samples include deletions of some of these genes. Interestingly, two other samples have amplification of this group of genes (besides TNFAIP3) and one additional sample has amplification of the whole group besides CDC25C and EGR1.
The gene encoding for casein kinase 1𝛼, CSNK1A1 at 5q has been suggested to be the target of lenalidomide in 5q- MDS. Since loss of individual genes in 5q including CSNK1A1 is uncommon in breast cancer, breast cancer would be unlikely to be sensitive to this drug through down-regulation of casein kinase 1𝛼. Other targets of lenalidomide and similar drugs include the Ikaros family transcription factors IKZF1 and IKZF3, the karyopherin sub-unit gene KPNA2, the translation termination factor GSPT1, transcription factors ZFP91 and ZNF692 (also known as AREBP) and the embryogenesis transcription factor SALL4. Besides isolated hemi-deletions in CSNK1A1, as mentioned above, and isolated hemi-deletions in ZFP91 (3 cases) and IKZF3 (1 case), no other hemi-deleted samples are observed in these genes in the breast cancer TCGA cohort. In contrast about 30% of cases have amplifications in one or more of immunomodulatory drugs target genes (Fig. 6). Most frequently amplified genes are IKZF3 (12%), KPNA2 (7%) and ZNF692 (10%). The two former genes are located at chromosome 17q and their amplifications are enriched in HER2-amplified cancers. The genes encoding for the CRL4CRBN ubiquitin ligase components (CUL4, DDB1, RBX1 and CRBN) and regulators of the ligase function (UBE2M, UBE2D3, UBE2G1), which form the endogenous ubiquitin ligase cellular machinery modulated by lenalidomide, pomalidomide and other immunomodulatory drugs, are amplified in 0.3%–2.5% of samples and even more rarely deleted (0–6 samples each) in breast cancer.
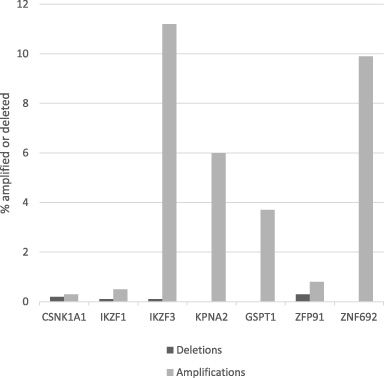
Prevalence of amplifications and deletions in the breast cancer TCGA cohort of immunomodulatory genes that have been proposed as targets of immunomodulatory drugs.
Patients with luminal A breast cancers and chromosome arm 16q loss tend to be older and have earlier stage disease than luminal A breast cancer patients with no 16q loss (Table 5). Luminal A cancers with 16q loss tend also to be more commonly (34%) lobular compared to luminal A cancers without 16q loss which display a lobular histology in 8.8% of cases. No significant differences in genomic instability as measured by the Aneuploidy Score (AS) and the Fraction of Genome Altered (FGA) between luminal A cancers with loss and no loss of 16q were observed. Luminal A cancers with 5q loss tended to occur in younger patients, to be more commonly of advanced stage and of ductal histology compared with luminal A cancers without 5q loss. 5q loss was also associated with higher AS and FGA scores (Table 5). In basal like carcinomas, loss of chromosome arm 5q as well as loss of 16q were both associated with higher AS and FGA scores, suggesting increased global genomic instability in these cancers (Table 6). Comparison of the Overall Survival (OS) of patients with luminal A breast cancer from TCGA and loss of chromosome arm 16q (n = 332) versus patients with no loss of 16q in whom 16q was either gained or not called (not gained or lost, n = 114) disclosed no significant differences (Log-Rank p = 0.8) (Fig. 7). In basal-like breast cancer patients with 5q loss (n = 115) OS was not different compared with patients with no loss of genetic material of 5q (n = 14, Log-Rank p = 0.7) (Fig. 8). In addition, loss of 5q did not alter OS prognosis in luminal A cancers nor loss of 16p altered OS in basal-like cancers (not shown). Similarly, no differences in PFS were detected in either luminal A or basal-like cancers with losses in 5q or 16q compared with the respective cancers without such losses (not shown).
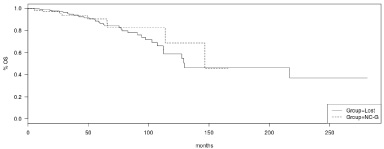
OS of Luminal A breast cancers with 16q loss (Group = Lost) versus luminal A breast cancers with the 16q arm not called or gained (Group = NC-G).
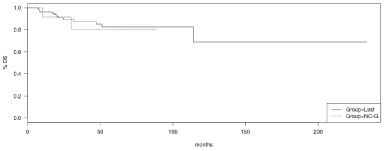
OS of basal-like breast cancers with 5q loss (Group = Lost) versus basal-like breast cancers with the 5q arm not called or gained (Group = NC-G).
Clinical and genomic characteristics of luminal A cancers with or without putative 5q and 16q hemi-deletions
5q-: 5q loss, 16q-: 16q loss, GNC: gained or not called. IDC: invasive ductal carcinoma. The histologic “other” category includes invasive lobular carcinoma, breast cancer not otherwise specified and invasive mixed mucinous carcinoma, AS: aneuploidy score, FGA: fraction genome altered. Comparisons for categorical values (stage and histology) were performed with the Fisher’s exact test and comparisons for continuous values (age, AS, and FGA) were performed with the t test. In some instances the sum of the categories does not amount to the total number of patients in each category due to missing data.
Clinical characteristics of basal-like cancers with or without putative 5q and 16q hemi-deletions
IDC: invasive ductal carcinoma. The histologic “other” category includes invasive lobular carcinoma, breast cancer not otherwise specified and invasive mixed mucinous carcinoma, AS: aneuploidy score, FGA: fraction genome altered. Comparisons for categorical values (stage and histology) were performed with the Fisher’s exact test and comparisons for continuous values (age, AS, and FGA) were performed with the t test. In some instances the sum of the categories does not amount to the total number of patients in each category due to missing data.
Sub-types of breast cancer based on genomics have different prognoses, patterns of recurrence and are treated with different drugs. The clinical characterization of breast cancer subtypes is based on immunohistochemistry for expression of hormonal and HER2 receptors which correlates reasonably well with genomic subtypes as defined by next generation genomic studies. Heterogeneity of breast cancers even within sub-types exists and is better characterized in triple negative cancers (negative for expression of ER, PR and HER2) which have been divided into five genomic categories (basal-like, immunomodulatory, luminal AR, mesenchymal and mesenchymal stem-like) [14]. Other studies have attempted to categorize breast cancers based on patterns of copy number alterations [15]. These patterns may be due to underlying molecular defects that lead to specific signatures of whole genome alterations such as those due to APOBEC deaminases defects or the tandem duplicator phenotype associated with mutations of CDK12 [16–18]. Genomic studies have made possible a parallel overview of the mutation and copy number status of the whole genome of cancers and open access initiatives, in association with user friendly platforms, have made the analysis of these data accessible to the broad oncology community. The current study takes advantage of the cBioportal platform to perform an analysis of putative loss/hemi-deletion of chromosomal arms 5q and 16q in luminal A and basal-like breast cancers. Publicly available data from two extensive breast cancer studies the TCGA and METABRIC are used in the analyses [4,8]. Chromosome arms 5q and 16q have been chosen for these analyses because they represent the most commonly lost arms in basal-like and luminal A cancers respectively. In contrast 5q loss is rare in luminal A cancers and 16q loss is significantly less prevalent in basal-like cancers (Fisher’s exact test p < 0.0001 for both comparisons, Fig. 1). Detailed investigation of copy number alterations of genes residing in chromosomes 5q and 16q shows that no individual gene or clusters of genes are deleted in high frequency in either luminal A or basal-like cancers. In addition, only a comparatively small number of cancer-related genes from 5q and 16q are deleted with a low prevalence in these cancers. These results strongly suggest that the loss of genomic material in 5q and 16q in these breast cancers is due to breaks that lead to loss of short sequences of DNA containing few genes each. However, isolated cases could contain longer losses as suggested by co-ordinated losses in a few occasions. Cancers with loss of genomic material from 5q or 16q have no significant different survival outcomes than counterparts with those chromosome arms either gained or not called.
An additional point of interest for 5q loss is the association with a completely different type of malignancy, the 5q- type of MDS [6]. Common loss of genetic material from 5q in basal-like breast cancers could imply similarities in the pathophysiology of these diseases stemming from loss of genes residing in 5q arm. The loss of 5q genetic material makes 5q- MDS sensitive to therapies with immunomodulatory drugs such as lenalidomide. Indeed, in an in vitro model with the triple negative breast cancer cell line MDA-MB-231, lenalidomide enhanced the cytotoxic effect of cisplatin, despite having minimal effect as monotherapy [19]. Increased apoptosis and suppressed BCL-2 were observed with the combination. However, since none of the proposed targets of lenalidomide is consistently hemi-deleted or mutated in basal-like breast cancers, the mechanism of the putative synergism of lenalidomide with cisplatin in MDA-MB-231 cells is unclear and may relate to alternative targets. These would be important to discover if lenalidomide and other immunomodulatory agents were to be repurposed for breast cancer therapy.
Luminal A carcinomas commonly harbor losses in genetic material from chromosome arm 16q. 16q losses have been present in in situ carcinomas and thus have been proposed to provide an early tumor advantage in carcinogenesis of ER-dependent carcinomas [7]. Other investigations have defined a most commonly minimal deleted region in 16q in the vicinity of which tumor suppressors ADAMTS18 and CNTNAP4 resided [20]. Consistently, these genes are co-deleted in 1.6% of samples in TCGA. However, given that luminal A cases with 16q losses do not display areas with loss in a consistent manner there is no consensus tumor suppressors residing in this arm that are unequivocally involved in carcinogenesis [21]. This is also true for tumor suppressor CDH1, a gene from 16q encoding for E-cadherin, and associated with lobular carcinomas, which is mostly mutated (in about 12% of cancers) and only hemi-deleted in 2.6% of luminal A carcinomas.
Three of the top deleted genes from 5q in basal-like cancers are CCNB1, CENPH and CENPK involved in mitosis and the assembly of the kinetochore as an anchor of mitotic spindle. Centromere proteins H and K are part of the machinery targeting CENPA (Centromere protein A) to centromeres and participate structurally in the assembly of the inner kinetochore [22]. Decreased dose of these proteins may interfere with the correct spindle assembly and could affect chromosomal integrity.
In conclusion, data presented in this study imply that there is no dominant tumor suppressor associated with losses of 16q and 5q in the sub-types of breast cancer where these losses occur. A plausible hypothesis for the common occurrence of loss of genetic material from these chromosomal arms in luminal A and basal-like cancers respectively, could be that another underlying defect in a gene that is part of the pathogenesis of these cancers favors the deletion pattern. Alternatively, these deletions could be part of more generalized structural aneuploidy, as suggested by the higher AS in cancers with both 5q and 16q losses (Tables 5 and 6). Randomly produced structural aneuploidy of different chromosome arm may then be favored in various sub-types with differing pathogenic molecular defects. Elucidation of these underlying mechanisms will be expected to advance understanding of breast cancer pathogenesis and advance new therapeutic opportunities.
Ethics approval and consent to participate
Not applicable.
Competing interests
The author declares that he has no competing interests.
Funding
No funding was received to perform this study.
Statement of contribution
Ioannis A. Voutsadakis is the only contributor in this paper.
