Abstract
BACKGROUND AND OBJECTIVES:
The role of human cytomegalovirus (HCMV) and Epstein-Barr virus (EBV) infections in breast cancer pathology is not well understood. Our study aimed to investigate the association of HCMV and EBV infections with breast cancer and distinguish the types of positive EBV and LMP-1 samples in Iranian patients.
METHODS:
Seventy-two formalin-fixed paraffin-embedded (FFPE) breast cancer tissues were analyzed between December 2014 and April 2016. Samples were analyzed for HCMV and EBV using nested-PCR and conventional PCR assays, respectively. Statistical analysis was performed using SPSS software version 18.
RESULTS:
Overall, HCMV and EBV genomes were detected in 6.9% and 16.7% of FFPE breast cancer tissues, respectively. Clinical factors were not statistically associated with the presence of HCMV and EBV.
CONCLUSION:
In this study, we reported EBV and LMP-1 typing in breast carcinoma cases for the first time in Iran. Our findings indicate that HCMV and EBV infections are not associated with the development of breast cancer.
Keywords
Introduction
Worldwide, breast cancer is the deadliest malignancy in women, causing more than half a million deaths annually. According to the last report of the World Health Organization (WHO), 2.1 million females were newly diagnosed with breast cancer, and 627,000 females died from the disease around the world in 2018 [1,2]. In Iran, breast cancer is considered the most common malignancy among women, associated with 7000 new cases and 4000 related deaths each year [3,4]. The etiology of breast cancer remains poorly understood, but it can be related to several conventional risk factors such as genetic predisposition, family history, obesity, physical inactivity, alcohol consumption, old age, nulliparity, and hormonal factors [5].
Environmental factors also play important roles during the initiation, development, and progression of various malignancies. Recently, the role of infectious diseases during carcinogenesis in different kinds of oncological diseases was well-established and described [6]. The International Agency for Research on Cancer (IARC) reports that biological carcinogens such as viral infections contribute to approximately 18–20% of all human malignancies [7–9]. To date, there are seven known human oncogenic viruses, including EBV, human papillomavirus (HPV), human T-lymphotropic virus 1 (HTLV-1), hepatitis B virus (HBV), hepatitis C virus (HCV), Merkel cell polyomavirus (MCPyV), and Kaposi sarcoma-associated herpesvirus (KSHV) [10–14].
HCMV is a member of the β-herpesvirus subfamily within the Herpesviridae family with a double-stranded DNA genome and a common human pathogen infecting 70–90% of the world’s population [15,16]. Nucleic acids and proteins of HCMV have been detected in several cancers, including breast, colon, and prostate cancers, as well as glioblastoma, medulloblastoma, mucoepidermoid cancer of the salivary gland, and rhabdomyosarcomas [13,17]. The molecular mechanism of HCMV in cancer pathology is not yet well understood but exhibits both oncogenic and oncomodulatory properties by expressing HCMV proteins that can interfere with cellular processes [17].
EBV is a gamma-1 herpes virus with a double-stranded DNA genome. Until now, the involvement of EBV in Burkitt’s lymphoma, Hodgkin’s lymphoma, and nasopharyngeal carcinoma has been well established [13,18]. The role of EBV in breast cancer is under investigation. It has been reported that EBV has an indirect role in breast cancer development by interfering with the immune response to HPV-transformed cells via the expression viral BCRF1 gene. This collaboration between EBV and HPV may increase their oncogenic potential [19,20].
Although various factors such as oncogenic viruses have been reported to contribute to the pathogenesis of human cancer, its molecular mechanism has not been recognized yet. It is difficult to determine which of these viruses is associated with breast cancer [13]. Not only the association of HCMV and EBV with breast cancer is important but also understanding of the cause of breast cancer for the early diagnosis, prevention, and treatment of breast cancer is necessary. In the light of contradictory and/or controversial findings from previous works concerning the role of HCMV and EBV in breast carcinoma development, this study aimed to investigate the molecular prevalence of these two infections in patients with breast cancer using an appropriate sample size in this study.
Materials and methods
Patients and histopathological features
In this study, during the period between December 2014 and April 2016, 72 formalin-fixed paraffin-embedded (FFPE) breast cancer specimens with histologically verified were collected from all hospitals affiliated to the Iran University of Medical Sciences (IUMS). A self-made questionnaire containing the following information filled for each patient: age, marriage status, cancer grade/type/malignancy, the estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (Her2). Breast cancer patients undergoing antiviral therapy such as ganciclovir and valganciclovir were excluded from the study. The institutional ethics committee approved the study protocol under approval number IR.IUMS.REC.1396.30553, and all procedures contributing to this work complied with the ethical standards of the relevant national and institutional committees on human experimentation and the Declaration of Helsinki.
Preparation of samples
Ten-μm-thin sections were cut from each FFPE tissue block with a standard microtome blade and were collected in sterile 2 ml microtubes. Because of contamination prevention of the specimens, microtome blades were changed between each block (Accu-Edge low-profile microtome blades; Co. SAKURA, Osaka, Japan). Furthermore, the sample holder of the microtome was washed with bleaching between cuttings. FFPE sections were treated with 600 μL xylene at 70 °C for 15 minutes, centrifuged at 13,000 rpm. The supernatant containing xylene was discarded, and this step was repeated two times. In the next step, FFPE sections were washed and rehydrated with 600 μL absolute ethanol, centrifuged at 13,000 rpm for 3 minutes, and this step was repeated three times. Finally, the specimens were allowed to dry at room temperature to remove remained ethanol.
List of sequences and other characteristics of primers used in this study
List of sequences and other characteristics of primers used in this study
DNA was extracted from FFPE tissues using the FavorPrepTM Tissue Genomic DNA Extraction Kit (Favorgen, Taiwan), following the manufacturer’s instructions. NanoDrop ND-1000® (Thermo Fisher Scientific Inc. Waltham, MA, USA) spectrophotometry and PCR amplification of a 167 bp fragment in the human β-globin gene as an internal control was performed to evaluate quantify and integrity of the extracted DNA. After extraction, DNA was eluted in 100 μL of elution buffer and stored at −20 °C for subsequent analysis.
HCMV detection by nested PCR assay
For detection of HCMV DNA, PCR reactions using a nested-PCR assay for amplifying regions of the UL55 gene were used. To perform the first round of PCR in a total volume of 20 μL of a reaction mixture of 0.5 μL of each outer primer (10 pmol), 4 μL extracted DNA as the template, 5 μL dH2O, and 10 μl of 2× PCR Master Mix (TaKaRa, Otsu, Japan) was used. The position and sequences of PCR primers are listed in Table 1. Amplification by PCR assay was performed in the Bio-Rad T100TM Thermal cycler under the following cycling program: one cycle, 95 °C for 5 min; 40 cycles, 95 °C for 1 min, 56 °C for 1 min (for external primers) and 52 °C for 30 s (for internal primers), and 72 °C for 1 min. To evaluate the amplification, PCR products in each round were visualized by a gel electrophoresis system that emitted UV radiation by E-Box Vilber version 15.03 (Vilber Lourmat, ZAC de Lamirault, France). Staining was performed using DNA Safe Stain in 1.5% agarose gel.
EBV typing and LMP-1 deletion analysis by PCR assay
For detection and typing of EBV, PCR assay with the specific primers amplifying the EBNA3C gene was used. The sequences and additional details of these primers are listed in Table 1.
PCR reactions were carried out in a 20 μL reaction mixture containing 0.5 μL of 10 pM of each primer, 10 μL 2× PCR Master Mix (TaKaRa, Otsu, Japan), 4 μL genomic DNA, and 5 μL dH2O. The PRC reaction was carried out under the following temperature conditions: an initial denaturation step of 5 minutes at 95 °C, followed by amplification for 40 cycles at the following conditions: 30 seconds at 94 °C, 60 seconds at 55 °C, 72 °C for 90 seconds, and finally a cycle of 4 °C for 8 minutes.
Distribution of HCMV and EBV according to the demographic and histopathological data
Distribution of HCMV and EBV according to the demographic and histopathological data
†Estrogen receptor; ‡Progesterone receptor; *Human epidermal growth factor receptor 2.
In the next step, all EBV-positive samples (EBV-I and EBV-II) were examined for the presence of the mutation in the EBV-LMP1 gene with the oligonucleotide primers flanking the clustered LMP-1 30 bp deletion segment (Table 1). Previous studies have shown that this region of the LMP-1 gene to be deleted in several malignancies caused by EBV, such as Hodgkin lymphoma. The PRC reaction was carried out in a total volume of 20 μL, comprising 0.5 μL of 10 pM of each primer, 10 μL 2× PCR Master Mix (TaKaRa, Otsu, Japan), 4 μL genomic DNA, and 5 μL dH2O. After an initial denaturation for 5 min at 95 °C, 44 cycles were performed as follows: 95 °C for 30 sec, 60 °C for 30 sec, and 72 °C for 30 sec. A final extension at 72 °C for 5 min completed the PCR amplification. In the next step, PCR products in each step visualized by a gel electrophoresis system that emitted UV radiation by E-Box Vilber version 15.03 (Vilber Lourmat, ZAC de Lamirault, France). Staining was performed using DNA Safe Stain in 2% agarose gel.
All data processing and statistical analysis were performed using SPSS software version 18 for Windows (SPSS Inc., Chicago, IL, USA), and P-values of less than 0.05 were considered statistically significant.
Result
A total of 72 patients with malignant breast tumors were enrolled in the study that seventy (97.2%) of them were married and two (2.8%) were single. The demographic data of the patients showed that the mean age was 52.15 ± 10.63 years (range 35–81 years). The type of breast tumor in all patients in this study was malignant breast tumor. Table 2 demonstrated the relationship between the age, HCMV and EBV status, and cancer grade in our study.
Out of a total of 72 participants, one had invasive medullary carcinoma, one had an invasive metaplastic carcinoma, 13 had an invasive lobular carcinoma (ILC), and 56 had invasive ductal carcinoma (IDC). The positive rate of ER, PR, and Her2 expression were 43.1%, 44.4%, and 18.1%, respectively (Table 2).
To evaluate the presence of HCMV DNA, nested PCR analyses of the UL55 gene were performed on 72 FFPE breast tumor specimens, and HCMV DNA was detected in 5 of 72 (6.9%) samples from breast tumor specimens (Fig. 1). Out of five HCMV-positive patients, all of them (100%) were married (P = 0.6). Regarding age, two HCMV-positive patients were under 50 years of age, and three patients were more than 50 years old (P = 0.8) (Table 2).
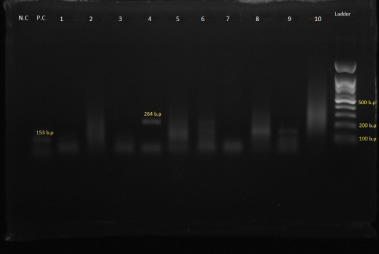
Electrophoresis image of amplification of HCMV UL-44 gene (532-bp PCR product) on 1.5% agarose gel stained with DNA Safe Stain.
The presence of EBV DNA in 12 of 72 (16.7%) samples of breast tumor specimens was identified. Of the twelve EBV-positive cases, two cases (16.7%) were single, and ten cases (83.3%) were married, and the difference was statistically significant (P = 0.001). Concerning the distribution of cases according to age group, seven EBV-positive patients were less than 50, and five were more than 50 years old (P = 0.2) (Table 2). Among the twelve EBV-positive cases, eight cases were type 1, two cases were type 2, and two cases had a dual infection by types 1 and 2 (Fig. 2).
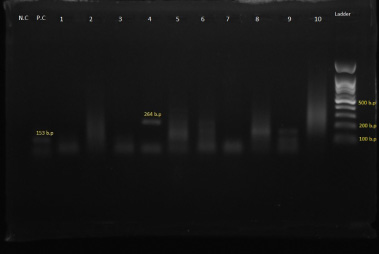
Electrophoresis image of PCR analysis of the EBNA3C gene of EBV (153-bp PCR product for EBV-I and 246-bp PCR product for EBV-II).
To identify the strains of the LMP-1 gene, PCR amplification was also performed with primers flanking the clustered LMP-1 deletion segment. In ten of the twelve EBV-positive samples (83.3%), a 161-bp product was seen, which was consistent with wild-type LMP-1. While in two cases out of twelve (16.7%), PCR amplification generated 131-bp products as deletion mutants.
Recent studies have shown viruses in cell lines, in addition to their contribution to some breast tumors. Hsu et al. study showed that HPV, EBV, CMV, HSV-1, and HHV-8 may be contributing to breast cancer [21]. In this regard in our study, HCMV and EBV genomes were investigated in 72 samples of FFPE breast cancer tissue, including 56 cases (77.8%) of IDC, 13 cases (18.1%) of ILC, 1 case (1.4%) of Invasive medullary carcinoma, and 1 case (1.4%) of Invasive metaplastic carcinoma by nested and conventional PCR, respectively, which all samples belonged to females.
Previous reports of PCR results of CMV and/or EBV in breast cancer have been inconsistent. There are fewer molecular investigations of CMV infection in breast cancer, but all that used IE-gene primers have been positive. Of seven studies, six (86%) found positive results, ranging from 7.4% to 100% of specimens tested, whereas found EBV in 10% of the samples, but one study did not detect CMV in any of 54 samples [15,22–26]. In our study, the HCMV genome was detected in 6.9% of samples of FFPE breast cancer tissue, and HCMV positivity was not associated with grade, receptor status, or disease stage.
There are conflicting reports about the role of EBV in breast cancer, but another possibility is that co-infection with multiple viruses increases the risk of breast cancer [13]. Previous studies showed that infection with EBV significantly increased the risk of breast cancer, indicates that EBV infection can play a key role in breast carcinogenesis. The results of other studies demonstrated an overall prevalence of 26.37% (95% CI: 22–31%) for EBV infection among patients with breast tumors in the world [27].
In our study, the EBV genome was detected in 16.7% of FFPE samples of malignant breast cancer by amplifying the EBNA-3 gene. EBNA-3 proteins (3A, 3B, and 3C) also cause the silencing tumor suppressor genes and involved in the formation and maintenance of cancer cells. EBV positivity was not associated with grade, receptor status, or disease stage but associated with marriage status so that all singles in the study were EBV positive.
The LMP-1 protein encoded by the EBV is known as an oncoprotein. Our findings have also shown that the LMP-1 30-bp deletion variant was found in 16.7% of the EBV-positive cases. In relation to the effects of this protein on cellular signaling pathways reported that LMP-1 protein activates NF-κB, MAPK, phosphatidylinositol 3-kinase/AKT, and c-Jun N-terminal kinase. Presence the 30-bp deletion in the C-terminal region of the LMP-1 gene involved in the progression from a non-oncogenic to an oncogenic state and has been found in different types of malignancies caused by EBV such as Hodgkin’s lymphoma, Burkitt’s lymphoma, nasopharyngeal carcinoma, and nasal T/natural killer cell lymphoma [28,29]. The molecular analysis of the EBV LMP-1 was also performed on breast cancer specimens for the first time in our study.
This study identifies HCMV and EBV genomes in malignant breast tumors, but their rates were lower than in other studies. Geographically, the molecular prevalence of these viruses in breast cancer varies. Another point is that biopsy specimens show more positive results than FFPE samples, indicating that biopsy samples (fresh or frozen) are more appropriate and sensitive than other specimens for the detection of HCMV and EBV. Also, the limited quantity of extracted genomic DNA and fragmentation of DNA are the problems of working with FFPE samples [30]. It can be concluded that the heterogeneity in results of different studies could be attributed to several variables, including geographic location, methods used to detect EBV infection, type of samples, detection regions or genes in HCMV and EBV genomes, and the histological types of breast cancer.
Conclusion
Overall, we found a low percentage of HCMV and EBV infections in this study. There was no causal relationship between HCMV and EBV infections and the development of breast cancer. It should be noted that conventional PCR and nested PCR assays used in this study have lower sensitivity than the real-time PCR or quantitative PCR (qPCR) that to be overcome. On the other hand, the small sample size of this study is another drawback, and so we suggest performing further independent studies with a larger sample size.
Footnotes
Acknowledgements
We would like to thank the Department of Pathology of Khatam-Al-Anbia Hospital for collecting the FFPE breast cancer tissue samples. This study has been funded and supported by Iran University of Medical Sciences, grant no. 1396.30553.
Conflicts of interest
All contributing authors declare no potential conflict of interest relevant to this article.
