Abstract
BACKGROUND:
Ki-67 immunohistochemistry (IHC) staining is a widely used cancer proliferation assay; however, its limitations could be improved with automated scoring. The OncotypeDX TM Recurrence Score (ORS), which primarily evaluates cancer proliferation genes, is a prognostic indicator for breast cancer chemotherapy response; however, it is more expensive and slower than Ki-67.
OBJECTIVE:
To compare manual Ki-67 (mKi-67) with automated Ki-67 (aKi-67) algorithm results based on manually selected Ki-67 “hot spots” in breast cancer, and correlate both with ORS.
METHODS:
105 invasive breast carcinoma cases from 100 patients at our institution (2011–2013) with available ORS were evaluated. Concordance was assessed via Cohen’s Kappa (κ).
RESULTS:
57/105 cases showed agreement between mKi-67 and aKi-67 (κ 0.31, 95% CI 0.18–0.45), with 41 cases overestimated by aKi-67. Concordance was higher when estimated on the same image (κ 0.53, 95% CI 0.37–0.69). Concordance between mKi-67 score and ORS was fair (κ 0.27, 95% CI 0.11–0.42), and concordance between aKi-67 and ORS was poor (κ 0.10, 95% CI −0.03–0.23).
CONCLUSIONS:
These results highlight the limits of Ki-67 algorithms that use manual “hot spot” selection. Due to suboptimal concordance, Ki-67 is likely most useful as a complement to, rather than a surrogate for ORS, regardless of scoring method.
Introduction
The immunohistochemical (IHC) marker Ki-67, which is a marker of cellular proliferation, has become routinely used in clinical practice as part of the pathologic diagnosis of breast cancer. Potential uses for Ki-67 include prognosis; use as a predictive marker of relative response to chemotherapy or endocrine therapy; as a biomarker for response to therapy before, during, and after neoadjuvant treatment, with a larger decrease in Ki-67 corresponding to a better response to therapy; and as an estimate of residual risk in patients after therapy [21,49,53,60,72]. However, there remain important concerns for use of Ki-67 as a clinical marker, including the lack of standardization of staining techniques, poor reproducibility, and great variability in cut-off ranges for risk categories [6,12,21,29,31,36,49]. Further development of automated scoring algorithms has been proposed by the International Ki67 in Breast Cancer Working Group (IKWG) as a potential solution to overcome current limitations of Ki-67 [49].
The use of automated algorithms based on artificial intelligence in pathology has dramatically increased in the past decade [3,17,48,59,63], with digital pathology applications spanning a wide range of both benign [13,34,74] and malignant conditions [16,27,32,50,64,65,71]. Automated algorithms specific to breast cancer have been the subject of extensive research, including but not limited to scoring of nuclear pleomorphism [19,20,25], analysis of tumor mitotic counts [9,10,44,46], and assessment of tumor infiltrating lymphocytes [5,8,37,42,45]. Automated scoring algorithms specific to the Ki-67 index have also been investigated, and many have shown promising results in the literature [1,6,26,31,36,38,55,56,62,75], suggesting the potential for automated scoring to improve the consistency, reproducibility, and accuracy of the Ki-67 assay.
Commercial scoring algorithms for Ki-67 have been developed, often as part of an “end-to-end” user experience – combining IHC platforms, whole slide scanners, high resolution hardware, and/or proprietary image analysis software – including the VENTANA Companion Algorithm image analysis software system (Roche) and the Aperio Nuclear Algorithm (Leica Biosystems). Standalone commercial scoring algorithms for Ki-67 on previously digitized slides have also been developed by companies including Visiopharm (10173 – Ki-67, Breast Cancer, AI) and Applied Spectral Imaging (GenASIs HiPath). Evidence suggests that many of these algorithms show high reliability and high concordance with manual assessment [23,76], and they have begun to be used as part of clinical diagnosis by some practicing pathologists. However, their high cost limits their overall availability, as many pathologists might be affiliated with laboratories that have existing significant investments in instruments such as IHC platforms and slide scanners that may not be compatible with proprietary image analysis software. Other algorithms have been built on top of open source software platforms, such as QuPath [11]. Many of these algorithms have shown high reproducibility and similar performance to proprietary algorithms [2,40,55]. However, while the QuPath platform is an extremely powerful research tool, its user interface for performing tasks such as automated Ki-67 scoring may be less intuitive to clinicians without technical backgrounds.
ImmunoRatio is an example of a freely available software program that can perform automated scoring of Ki-67 and easily be used in an “off the shelf” fashion [68]. Initially it was available as a web application, and as of publication of this article, is available as a plug-in for the also free ImageJ software platform [58]. The ImmunoRatio program has shown promising performance in initial studies [24,70,75], and has key advantages of accessibility in terms of both cost and user interface that could make it widely available in clinical practice, including the ability to use JPEG images captured from a standard microscope camera rather than requiring whole-slide digitized images. However, most of the initial research on ImmunoRatio involved comparing performance of the algorithm to exhaustive manual counting of digitally scanned images [24,75], rather than to visual estimates of Ki-67 IHC glass slides, as is commonly performed in clinical practice. Better evaluation of how algorithms such as ImmunoRatio compare to real-world estimates of Ki-67 proliferation for clinical diagnosis can help inform their clinical utility relative to proprietary solutions for practicing pathologists.
The OncotypeDX TM Recurrence Score (ORS) is a widely used molecular test used to predict response to chemotherapy in breast cancer patients with ER positive, HER-2 negative, and axillary lymph node negative or limited nodal disease. The test calculates a recurrence score based on a 21-gene RT-PCR assay that quantifies gene expression of 16 known cancer genes associated with prognosis and prediction, 5 of which are proliferative indices/markers, and 5 reference genes [52]. Based on the model coefficients involved in calculating the recurrence score, nearly half of the overall ORS is based on expression of the cancer proliferation genes alone. The ORS has emerged as a useful tool for evaluation of patients with low grade breast cancers and their likely risk of 10-year recurrence. It also aims to predict potential benefit of adjuvant therapy, thus reducing the use of unnecessary chemotherapy in many patients [12,14,30,47,66].
It is well established that ER positive breast cancers have a higher rate of late distant recurrence than ER negative breast cancers. Unlike in triple-negative breast cancer patients, who lack a well-defined therapeutic target such as ER or HER2 [35], the rate of distant recurrence in ER positive breast cancer often can be reduced effectively by endocrine therapy. Nonetheless, a portion of these patients will still recur, largely due to resistance to endocrine therapy [22]. Recent clinical trial data have shown that in patients with a mid-range recurrence score, chemotherapy was no better than endocrine therapy in terms of rate of recurrence [61]. Thus, the routine use of OncotypeDX TM in a subset of patients may identify a large proportion of women with endocrine receptor positive breast cancers who can be spared adjuvant chemotherapy [61,73]. However, OncotypeDX TM is a very expensive test with a long turnaround time. As both ORS and Ki-67 are markers of proliferation, if there is good correlation between semi-quantitative Ki-67 immunohistochemistry with the ORS, then Ki-67 may be a less expensive, more efficient, and more practical alternative that could help pathologists work up cases more efficiently without sacrificing accuracy. Indeed, prior researchers have identified Ki-67 as a potential surrogate marker for the ORS, with varying levels of accuracy [7,28,51,67]. Assessment of Ki-67 “hot spots,” in particular, may be more strongly associated with ORS [51,67], and at least some authors have shown that it may be better correlated with clinical outcomes [33,56], although it may suffer from decreased inter-observer reliability than “global” measurements of Ki-67 [51]. In this study, our goal was to compare manually reported Ki-67 values with results generated from the ImmunoRatio automated Ki-67 counter of Ki-67 hot spots, and correlate these results to ORS.
Materials and methods
With institutional review board approval, our institution’s diagnostic surgical pathology archives were searched for patients diagnosed with invasive carcinoma of the breast who also had analysis with OncotypeDX TM . 105 cases from 100 patients diagnosed with invasive ductal and/or lobular carcinoma (mean age = 55.6, range 37–77) between 2011 and 2013 and who had their samples analyzed using OncotypeDX TM were evaluated. Pathologic features including grade, size, stage, manually determined semi-quantitative IHC tumor marker results (ER, PR, HER-2, Ki-67, p53), and OncotypeDX TM Recurrence Score were recorded. The manual semi-quantitative Ki-67 score (mKi-67) was recorded as the Ki-67 score reported at the time of diagnosis. If slides for Ki-67 were unavailable for a particular case, the Ki-67 IHC was repeated on the most representative tumor block. Initial and repeat immunohistochemical staining for Ki-67 was performed using anti-Ki-67 rabbit monoclonal antibody (30-9, Ventana), according to the manufacturer protocol.
To recreate the workflow most likely to be available to a practicing pathologist, JPEG digital images of the immunohistochemical slide that was initially used to manually estimate Ki-67 in the area of highest Ki-67 staining were manually captured for each case using an Olympus BX40 microscope, Olympus DP71 camera, and OLYMPUS cellSens Entry digital software. To ensure quality and representativeness of high-power field selection, these images were reviewed in a third of the cases by a senior pathologist with specialty training in breast pathology and compared to the original glass slide prior to their use in automated analysis. Finalized images were used to interpret an automated quantitative Ki-67 score (aKi-67) using ImmunoRatio. ImmunoRatio calculates the percentage of positively stained nuclear area (labeling index) by using a color deconvolution algorithm for separating the staining components (diaminobenzidine and hematoxylin) and adaptive thresholding for nuclear area segmentation [68]. A dual image report including the aKi-67 result (as a percentage) was generated for each case (Fig. 1). As part of a secondary analysis, the specific JPEG images included in the automated analysis were re-reviewed by a pathologist in a large subset of cases (N = 73), blinded to the original and automated values, to provide a manual visual estimate of Ki-67 index corresponding to the exact field analyzed by the ImmunoRatio software.
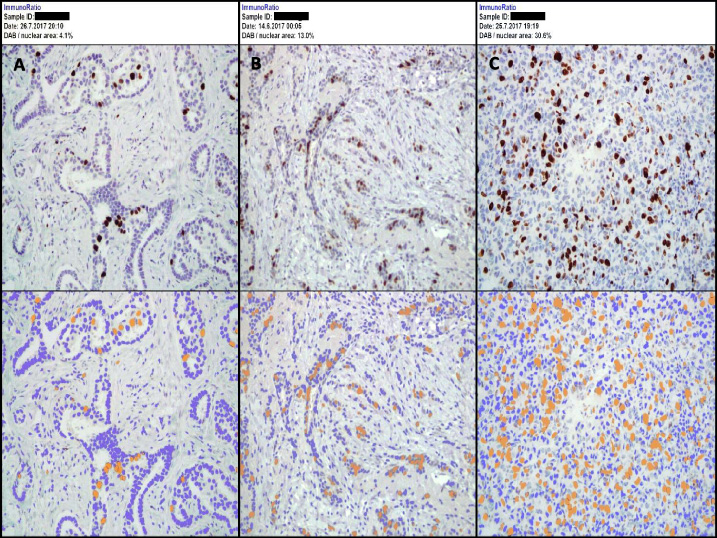
Example images obtained via ImmunoRatio digital imaging software. A. Low (Favorable) Ki-67 Score: ≤10%; B. Intermediate Ki-67 Score: 11–20%; C. High (Unfavorable) Ki-67 Score: >20%.
For both mKi-67 and aKi-67, results were reported as a percentage and as a semi-quantitative category according to our institution’s criteria (≤10% = low/favorable, 11–20% = intermediate, and >20% = high/unfavorable). Reported mKi-67 (percentage and category) was compared to the calculated aKi-67 (percentage and category) for each case. In a secondary analysis, an alternative binary cut-off system was used to compare mKi-67 and aKi-67, with potential cut-offs of >5%, >10%, >15%, >20%, and >25% assessed. Too few cases >30% were present to accurately assess this cut-off. Automated and reported Ki-67 category for each case was compared to reported OncotypeDX TM Recurrence Score category (Score <18 = low risk, 18–30 = intermediate risk, ≥31 = high risk). The categories for each reported Ki-67 value and OncotypeDX TM Recurrence Score are summarized in Table 1.
Ki-67 and OncotypeDXTM Recurrence Score categories
Percent agreement between the manual and automated Ki-67 categories, as well as between these categories and the ORS category, was assessed via cross-tabulation. Overall concordance between methods was calculated using an unweighted Cohen’s Kappa statistic (κ), which is a measure of agreement that accounts for the expected agreement due to chance [15]. Although there are no universal cut-offs for values of κ that constitute “good” concordance, a commonly used scheme characterizes κ values of 0–0.20 as slight, 0.21–0.40 as fair, 0.41–0.60 as moderate, 0.61–0.80 as substantial, and 0.81–1 as almost perfect agreement [41]. Additionally, the correlation between the actual percentage values of the manual and automated Ki-67 indices was assessed using the Spearman correlation coefficient to account for the skewed distribution of the data. Significant differences in percentage values of the manual and automated Ki-67 indices were tested via the paired Wilcoxon rank sum test. P-values <0.05 were considered statistically significant. All analyses were performed using R 3.4.1 [54].
A total of 100 participants, including 105 total cases of invasive carcinoma, had available data for analysis. Basic clinical and histopathologic characteristics of the cases are shown in Table 2. Raw mKi-67, aKi-67, and ORS results for each case are reported in Supplementary Table S1.
Clinical and histopathologic characteristics of the cases (N = 105). All data presented as N (%) unless otherwise specified
Clinical and histopathologic characteristics of the cases (N = 105). All data presented as N (%) unless otherwise specified
∗One case was primarily DCIS with a small focus of microinvasive carcinoma and could not be graded. †One case was recurrent breast carcinoma, so staging was not applicable.
Agreement in manual and automated Ki-67 index categories was seen in 57 of 105 cases (54%, Table 3), with κ = 0.31 (95% CI 0.18–0.45). Agreement was highest in mKi-67 “high” cases at 80%, compared to 57% and 47% agreement among mKi-67 “intermediate” and “low” cases, respectively. Just over two-thirds of discordant cases occurred between the mKi-67 “low” score and aKi-67 “intermediate” and “high” scores. When looking at differences between actual reported automated and manual percentages, the median difference of aKi-67 minus mKi-67 indices was 3% (IQR −0.5%–11%), and the median absolute difference between the aKi-67 and mKi-67 indices was 4.9% (IQR 2.5%–12.1%). This increase in aKi-67 percentage relative to mKi-67 percentage was statistically significant (P < 0.001). The Spearman correlation coefficient for the mKi-67 and aKi-67 index percentages was 0.43 (P < 0.001).
Cross-table of manual vs automated Ki-67 index categories
Cross-table of manual vs automated Ki-67 index categories
As a secondary analysis, we compared manual to automated estimates of the Ki-67 index in a subset of cases using the same high-power field (same JPEG image) used to calculate aKi-67. In this analysis, agreement in mKi-67 and aKi-67 index categories was seen in 50 of 73 cases (68%) overall, with κ = 0.53 (95% CI 0.37–0.69). When looking at differences between actual reported automated and manual percentages, the median difference of aKi-67 minus mKi-67 indices was 0.5% (IQR −1.5%–3.9%), and the median absolute difference between the aKi-67 and mKi-67 indices was 2.9% (IQR 1.1%–6.3%). The difference between the aKi-67 and mKi-67 percentages using the same high-power field was not statistically significant (P = 0.14). The Spearman correlation coefficient for the mKi-67 and aKi-67 index percentages based on the same high-power field was 0.83 (P < 0.001).
In another secondary analysis, we compared concordance between mKi-67 and aKi-67 using a binary cut-off system, rather than the three-category system used at our institution. Results for different cut-offs of >5%, >10%, >15%, >20%, and >25% are shown in Table 4. Overall, concordance was higher at higher cut-offs, ranging from κ = 0.25 (95% CI 0.12–0.38) at a cut-off of >5% to κ = 0.44 (95% CI 0.22–0.67) at a cut-off of >25%. When comparing manual and automated Ki-67 values using the same high-power field, concordance was highest at the extremes, with κ = 0.80 (95% CI 0.62–0.99) at a cut-off of >5% and κ = 0.85 (95% CI 0.67–1.00) at a cut-off of >25%. If the cut-offs of >5% and >25% are combined into a three-category system, then κ = 0.28 (95% CI 0.15–0.41) for reported mKi-67 vs aKi-67 and κ = 0.80 (95% CI 0.67–0.94) for manual vs automated Ki-67 scoring in the same high-power field.
Comparison of reported manual, manual high-power field (HPF), and automated Ki-67 index concordance rates at different binary cut-off values. Note that manual HPF Ki-67 was estimated using the same JPEG image of a single high-power field as was used in the automated analysis, and was assessed in a subset (73/105) of cases
Table 5 shows the cross-tabulation for the mKi-67 index and ORS categories. Agreement was observed in 62 of 105 cases (59%), and κ = 0.27 (95% CI 0.11–0.42). Agreement rates were higher for the manual Ki-67 “low” group than for the “intermediate” and “high” groups (73% agreement vs 36% and 47%, respectively). Cross-tabulation for the aKi-67 index and ORS categories is shown in Table 6. Agreement was observed in 42 of 105 cases (40%), with κ = 0.10 (95% CI −0.03–0.23). Similarly, the highest agreement was observed in the “low” aKi-67 group compared to the “intermediate” and “high” groups (71% agreement vs 23% and 26%, respectively).
Cross-table of manual Ki-67 index vs OncotypeDXTM risk score (ORS) categories
Cross-table of manual Ki-67 index vs OncotypeDXTM risk score (ORS) categories
Cross-table of automated Ki-67 index vs OncotypeDXTM risk score (ORS) categories
To the best of our knowledge, this study is one of the largest to date to compare the “real-world” performance of manual Ki-67 scores and automated Ki-67 scores using the ImmunoRatio software platform, as well as compare these scores to the OncotypeDX TM Recurrence Score. Although aKi-67 and mKi-67 scores showed fair to moderate overall correlation and concordance (κ = 0.31; 95% CI 0.18–0.45), we found that aKi-67 scores consistently overestimated manual scores, and tended to show worse concordance with ORS than mKi-67. Although no gold standard exists for the evaluation of Ki-67, the better correlation with ORS suggests that in our study, mKi-67 was more accurate as a measure of cancer proliferation than aKi-67. In clinical practice, a bias toward overestimation could result in breast cancer patients’ inappropriately receiving adjuvant or neoadjuvant chemotherapy, and being at risk of many toxic side effects, when they are less likely to benefit. Our findings are in contrast to those of some prior authors that have shown better correlation between Ki-67 hot spots than average proliferation with ORS [51,67], and may reflect the greater variability of hot spot vs global based scoring methods, as has been reported by the IKWG [43,49]. Future researchers studying Ki-67 should be extremely cautious when aggregating results across multiple studies that incorporate analysis of Ki-67 hot spots, paying close attention to methodologic details of how the hot spots are selected to avoid problems with poor reproducibility across studies.
Based on the results of our secondary analysis, which showed much higher correlation between manual and automated Ki-67 scores when performed on the exact same high-power field (correlation coefficient 0.83 vs 0.43 with reported values), it seems likely that most of the overestimation of mKi-67 by aKi-67 reflects human inputs rather than the algorithm itself, with the selection of the representative high-power field driving most of the discrepancy. Although manual post hoc inspection of images generated by the ImmunoRatio program reveals specific instances where the algorithm classified some non-tumor cells, such as lymphocytes and stromal cell, as either positive or negative-staining tumor cells, this kind of algorithmic misclassification error does not appear to have systematically affected the results in our study overall. Although this study was only performed with a single algorithm, the ImmunoRatio online software, our results suggest that any automated algorithm for Ki-67 that involves manual selection of high-power fields for analysis should be treated cautiously before being adopted into clinical use. Indeed, while institutional clinical use of any automated Ki-67 scoring algorithm would require site-specific validation to ensure algorithmic scoring accuracy and consistency, algorithms such as ImmunoRatio that rely on manual hot spot selection would likely need additional initial and ongoing personnel training in field selection itself to maintain reliability, as the “human element” of manual selection cannot be fixed with algorithmic fine-tuning alone.
When estimating the Ki-67 index manually, extreme care needs to be taken by the reviewing pathologist to choose the correct area of interest, with the hot spot with the highest level of Ki-67 staining present on a representative tumor slide for analysis with the imaging software [33]. A more systematic approach to assessing Ki-67 manually can be taken using a whole slide, assessing IHC staining and averaging the results. One study recommends counting at least 3 randomly selected high-power fields, scoring a minimum of 500 cells and averaging the results between the three areas [4,57]. The IKWG has proposed calculating a global score based on the weighted average of 100 nuclei scored in each of four separate fields representing negligible, low, medium, and high Ki-67 proliferation, with manual visual estimation of the overall proportion of tumor belonging to each category to determine the weights [49]. However, these methods are time consuming and are not widely used in practice, especially at high volume centers. Automated scoring algorithms based on whole-slide digital image analysis may be especially useful for performing global estimates of average Ki-67 proliferation without requiring a prohibitive increase in pathologist workload.
A limitation of this study includes the use of a single high-power field for digital analysis of Ki-67, as well as the fact that this field was not chosen by the original diagnostic pathologist for all cases, which would have been logistically impracticable. It is unclear from the available data in our current study whether use of additional high-power fields or personnel training would lead to a meaningful improvement in the observed overestimation of aKi-67 relative to mKi-67, or whether our results are more reflective of inherent limitations to hot spot based scoring methods. Future studies are thus needed to evaluate whether use of multiple high-power fields might yield more robust results with better concordance with both manual evaluation, molecular studies such as ORS, and clinical outcomes. Furthermore, manually capturing multiple images to upload into a software program could substantially increase the time required to use the scoring algorithm. As a result, if several high-power images are required to make freely available platforms such as ImmunoRatio more reliable in practice, then practicing pathologists might be more likely to either use higher cost but more robust solutions involving whole-slide scanned images or continue using entirely manual visual estimation methods.
Currently, there are no agreed upon standard cut-point definitions for Ki-67 proliferation index related to particular clinical outcomes, and the IKWG has recognized the importance of additional research into determining better cut-off definitions [49]. We chose the cut-offs of ≤10% = low/favorable, 11–20% = intermediate, and >20% = high/unfavorable, as these are the standard cut-offs used at our institution and would have been the cut-offs by which the practicing pathologists would have made their original Ki-67 assessments in the diagnostic pathology report. However, we also conducted a secondary analysis examining different potential binary cut-offs. We found that increasing the cut-off improved the concordance of reported mKi-67 and aKi-67, likely by reducing the impact of hot spot variability within the sample. When limiting the analysis to manual vs automated assessment of the same exact high-power field, concordance was highest at both the lowest and highest cut-off selection points, with κ = 0.80 (95% CI 0.62–0.99) at a cut-off of >5% and κ = 0.85 (95% CI 0.67–1.00) at a cut-off of >25%. If the cut-offs of >5% and >25% were combined into a three category system, then concordance remained excellent when estimated on the same high-power field (κ = 0.80; 95% CI 0.67–0.94), but was essentially unchanged from the original cut-offs when comparing reported manual vs automated values (κ = 0.28; 95% CI 0.15–0.41). These findings are consistent with the recent IKWG consensus that Ki-67 scores of ≤5% or ≥30% can be used reliably enough to estimate prognosis; however, they suggest that different cut-offs alone are insufficient to overcome the limitations of manual hot spot selection. Unfortunately, a large percentage of breast cancer cases will fall within the intermediate range in between these values, and meaningful prognostic information is likely held within this range as well. Better assessment of cut-offs in the intermediate range will likely require analysis using automated scoring systems based on whole-slide digital images to overcome the limitations of human visual assessment.
Our study also showed that mKi-67 exhibited fair concordance with ORS overall (κ = 0.27; 95% CI 0.11–0.42), and that aKi-67 was not significantly more concordant with ORS than would be expected by chance alone (κ = 0.10; 95% CI -0.03–0.23). Agreement was highest in the low Ki-67 category, both for manual and automated methods. These results suggest that the Ki-67 assay alone will be unlikely to be useful as a cheaper alternative to ORS for most patients, despite both assays being largely based on an assessment of cancer proliferation rate. Some researchers have examined whether combining Ki-67 with additional clinical information, such as tumor grade, size, and/or hormone receptor status into a statistical model, such as the modified Magee Equation or the IHC4 score, may better predict ORS [18,30,39,69]. These models overall seem accurate in predicting a high ORS, but they may be more likely to miss intermediate ORS individuals who may still have some benefit from adjuvant chemotherapy [30]. These models relied on Ki-67 assessment by manual scoring, and further research is needed to determine whether incorporation of automated Ki-67 analysis can improve the accuracy of ORS prediction with clinicopathologic models. Other research has suggested that presence of high Ki-67 may have complementary value to ORS by identifying individuals who may have a slightly higher recurrence risk despite having a low or intermediate ORS [28]. The IKWG has recommended that presence of a Ki-67 score of ≥30% or ≤5% can be used in certain breast cancer patients to recommend either for or against the use of adjuvant chemotherapy, respectively, without needing to use ORS. In our study, 15% of individuals (2/13) with a high ORS had an mKi-67 ≤5% and 0% had an aKi-67 ≤5%. By contrast, approximately 3% of individuals (2/62) with a low ORS had an mKi-67 >25% and 13% (8/62) had an aKi-67 >25%. Thus, it may be the case that hot spot and global assessment may have complementary uses clinically, with a lower Ki-67 assessed via hot spot methods better for excluding a high ORS and a higher Ki-67 assessed via global methods better for excluding a low ORS. Additional research is needed to help clarify the clinical utility of each approach in different clinical scenarios, as well as to develop automated methods for assessing Ki-67 hot spots that will be less vulnerable to human variability in field selection. More research is also needed to determine the clinical implications of discordance between ORS and Ki-67 index for patient treatment response and prognosis.
In summary, our study highlights the potential limits of Ki-67 quantification algorithms that rely on manual hot spot selection, as well as the suboptimal concordance of the Ki-67 index and the ORS. While automated quantification algorithms have the potential to improve the reproducibility of the Ki-67 index, it may be difficult for “off the shelf” algorithms to achieve the reliability required for clinical use without also utilizing methods that minimize the impact of human inputs, such as whole-slide digital imaging. Finally, while Ki-67 remains an important predictor of treatment response and prognosis in breast cancer, it will likely continue to primarily be useful as a complement to, rather than as a surrogate marker for ORS in cases where the latter is indicated clinically.
Footnotes
Acknowledgements
None.
Conflict of interest
None.
