Abstract
Background
Artificial intelligence (AI) persists as a focal subject within the realm of medical imaging, heralding a multitude of prospective applications that span the comprehensive imaging lifecycle. However, a key hurdle to the development and real-world application of AI algorithms is the necessity for large amounts of well-organized and carefully planned training data, including professional annotations (labelling). Modern supervised AI techniques require thorough data curation to efficiently train, validate, and test models.
Objective
The proper processing of medical images for use by AI-driven solutions is a critical component in the development of dependable and resilient AI algorithms. Currently, research organizations and corporate entities frequently confront data access limits, working with small amounts of data from restricted geographic locations.
Methods
This study provides an in-depth examination of the publicly accessible datasets in the field of medical imaging. This work also determines the methods required for preparing medical imaging data for the development of AI algorithms, emphasizes current limitations in dataset curation. Furthermore, this study explores inventive strategies to address the challenge of data availability, offering a detailed overview of data curation technologies.
Results
This study provides a comprehensive evaluation of medical imaging datasets emphasizes their vital significance in improving diagnostic accuracy and AI models, while also addressing key problems such as dataset diversity, labelling, and ethical implications.
Conclusions
The paper concludes with an insightful discussion and analysis of challenges in medical image analysis, along with potential future directions in the field.
Keywords
Introduction
Since the invention of medical imaging technology, the field of medicine had entered a new era. The beginning of medical imaging started with the adoption of x-rays. Artificial intelligence (AI) has emerged as an important topic in the field of health care during the last two decades, 1 especially for medical imaging.2,3 These imaging techniques have been useful to diagnose and treat a variety of diseases. Medical imaging is critical for providing useful information into the diagnosis and treatment of a wide range of diseases. The utilization of various imaging modalities captures scans of the human body by exploiting its diverse responses. Reflection and transmission are prevalent techniques in medical imaging, leveraging the distinct reflection or transmission ratios of various body tissues and substances. Many researchers turned their attention to AI in the field of medical imaging, believing that it might be one of the solutions to problems (such as medical resource shortages) and take use of technology advancements.4–6
A frequent challenge encountered in employing deep learning (DL) methods within a particular domain, especially in the context of medical image analysis, is the issue of insufficient data. In medical image analysis research, individuals typically utilizing DL methods are often computer scientists rather than having a background in the medical field. This data scarcity problem tends to be more pronounced in the realm of medical image analysis. Due to the absence of access to medical equipment and patients, these researchers are unable to independently gather data. Additionally, the lack of relevant medical knowledge prevents them from annotating the obtained data. Moreover, medical data ownership lies with institutions, which face challenges in making it publicly available due to privacy and ethical constraints. Consequently, when researchers assess their algorithms using private data, the comparability of their research results becomes compromised. 7
Nowadays, numerous research groups and organizations do not have their access to medical images, and a small number of datasets impede the ability to be generalized and accuracy of developed methods. The objectives of these challenges include the enhancement and creation of automatic or semi-automatic algorithms, thereby encouraging research in medical image processing through the utilization of computer-aided methodologies.7,8 Meanwhile, several scholars and organizations initiate efforts to collect and share medical datasets for research purposes. Furthermore, successful AI algorithms require effective curation, analysis, labelling, and clinical application.
In the present study, research publications were retrieved using several search keywords, including ‘ medical imaging datasets’, ‘dataset curation tools’, ‘medical imaging analysis’, and ‘DL/ML techniques for medical imaging’ from different academic databases. After accumulating articles from various databases, we used precise selection criteria to decide which ones to include or omit, as shown in Table 1.
Articles selection criteria.
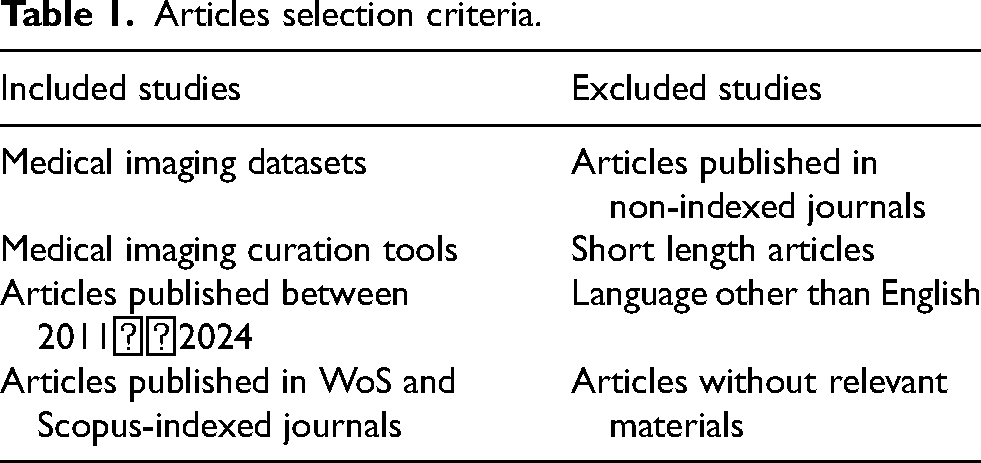

Table 1 shows that research publications were retrieved using several search keywords, including “short-length articles or articles without relevant facts,” which were disregarded. publications for additional evaluation were chosen based on their quality assessment standard marks.
This study aims to tackle these issues by offering a detailed review of available medical datasets and outlining several important steps for preparing a large volume of medical image data. It also explores new methods to solve the problem of limited data availability, giving a comprehensive look at how data can be effectively curated. Finally, the paper wraps up with a discussion on the challenges of processing medical images and the future possibilities in this area. Table 2 shows the overview of current review studies on medical imaging.
Overview of recent review publications on medical imaging.
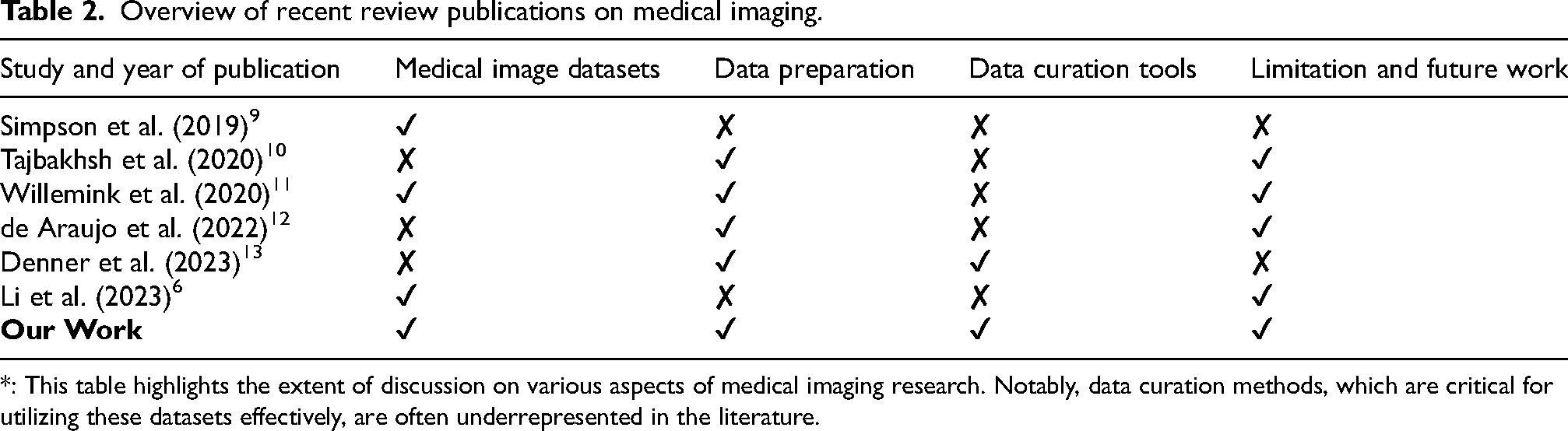
: This table highlights the extent of discussion on various aspects of medical imaging research. Notably, data curation methods, which are critical for utilizing these datasets effectively, are often underrepresented in the literature.
Table 2 summarizes key review publications that have addressed medical imaging datasets, data preparation techniques, and future directions. This work makes the following specific contributions: (1) Table 3 shows medical imaging datasets with details on modality types, bodily organs, purposes, and file formats; (2) Data preparation as shown in Figures 1 and 2 and Tables 4 and 5; (3) Data curation tools as shown in Table 6; and (4) Discussion and future works.
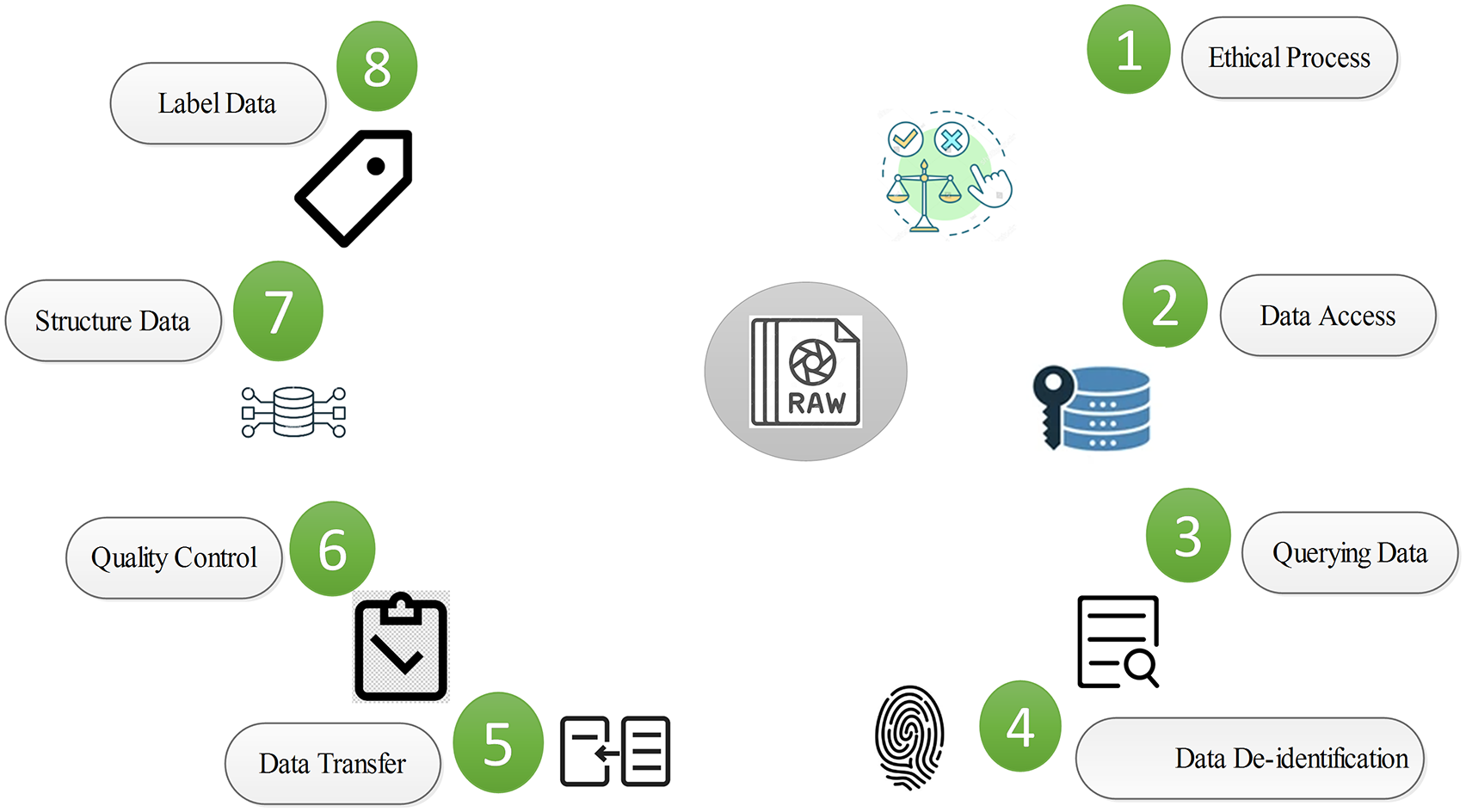
Medical image preparation process.
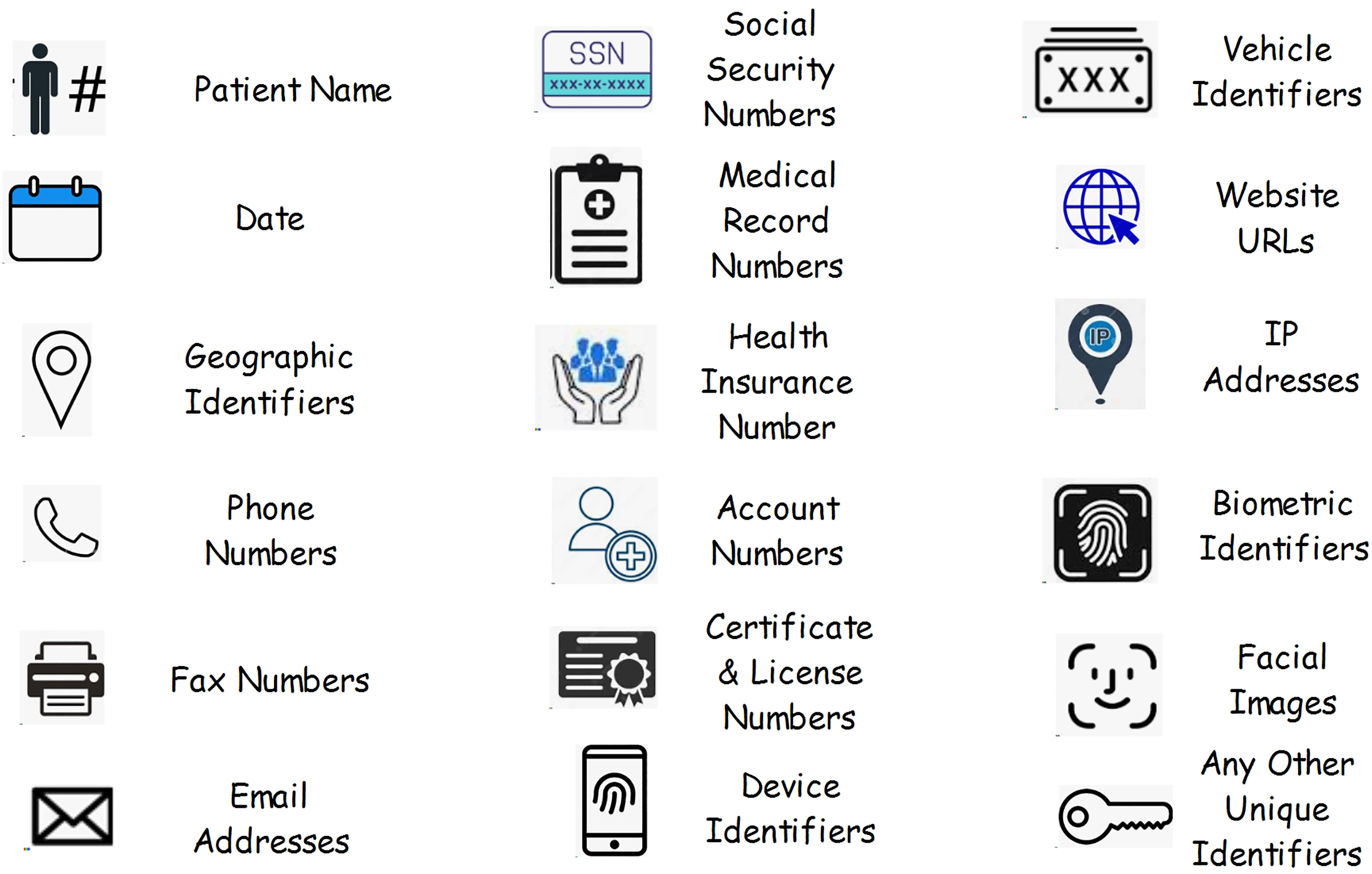

List of HIPAA identifiers (https://www.healthcarecompliancepros.com/blog/phi-and-you-the-basics-you-need-to-know).
Medical image datasets
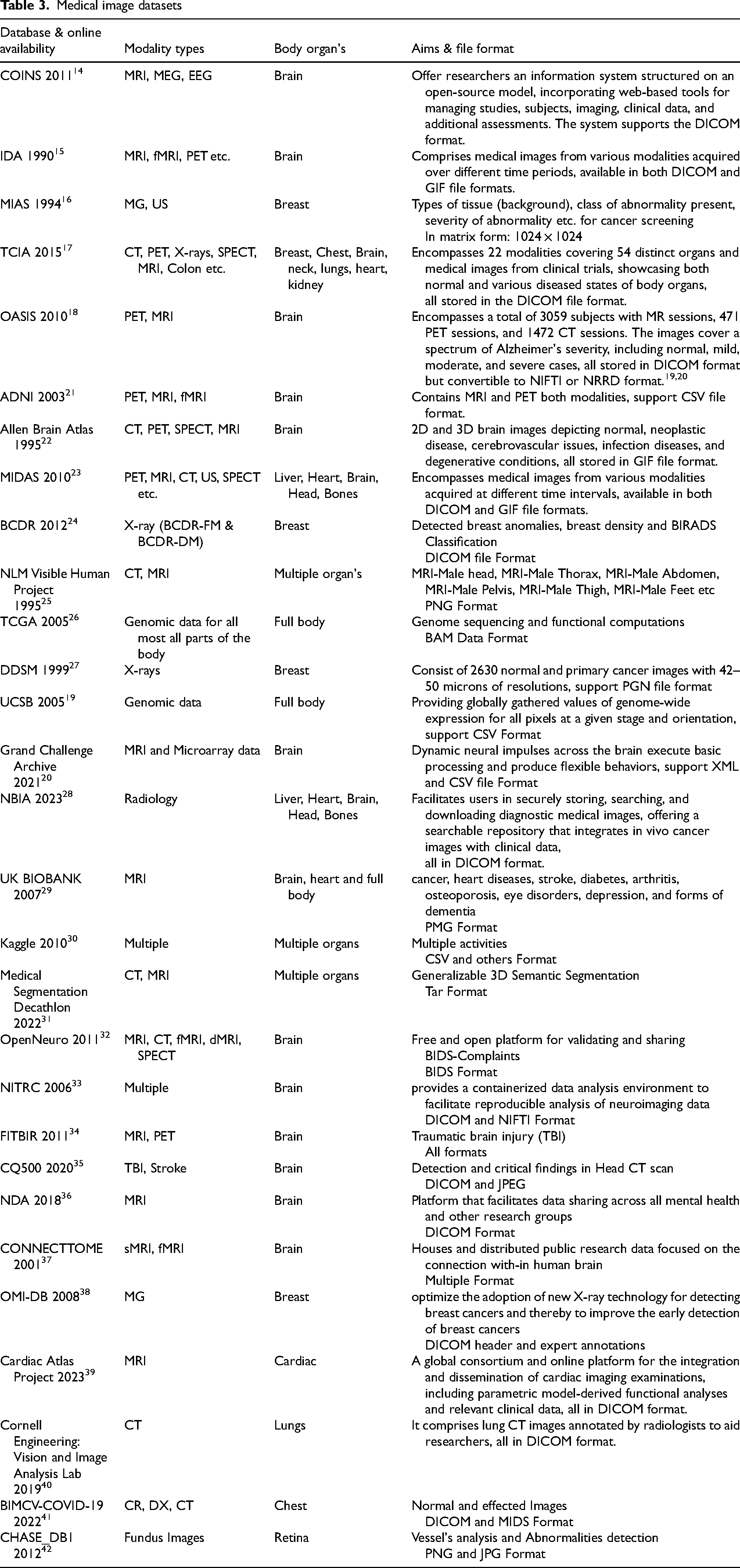
Some important medical image open-source solutions.
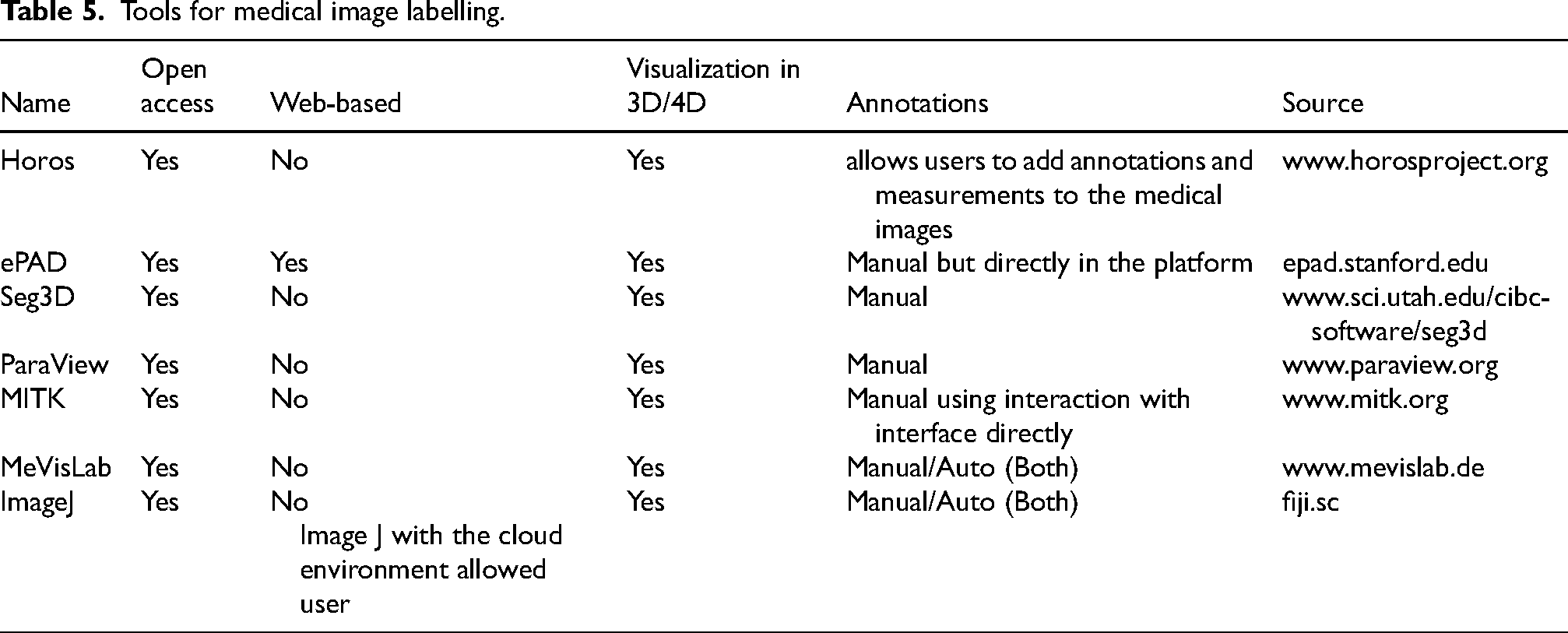
Tools for medical image labelling.

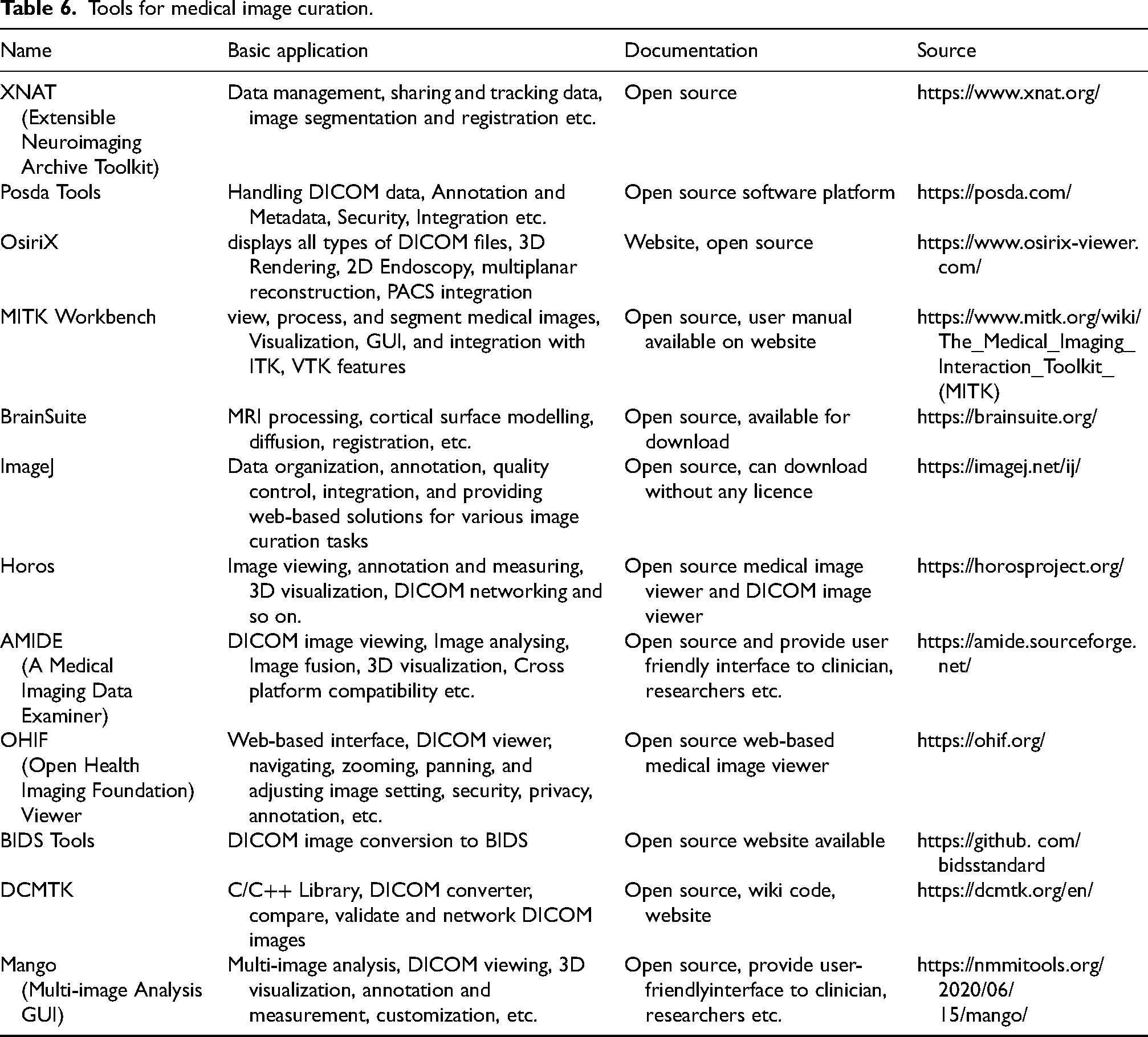
Tools for medical image curation.
The rest of the article is structured as follows:.
Section 2 provides an extensive overview of the medical imaging databases. Section 3 describes the steps for medical imaging data preparation. Section 4 introduces the data curation tools, while Section 5 examines existing issues and future directions. Section 6 concludes the review.
Medical image datasets
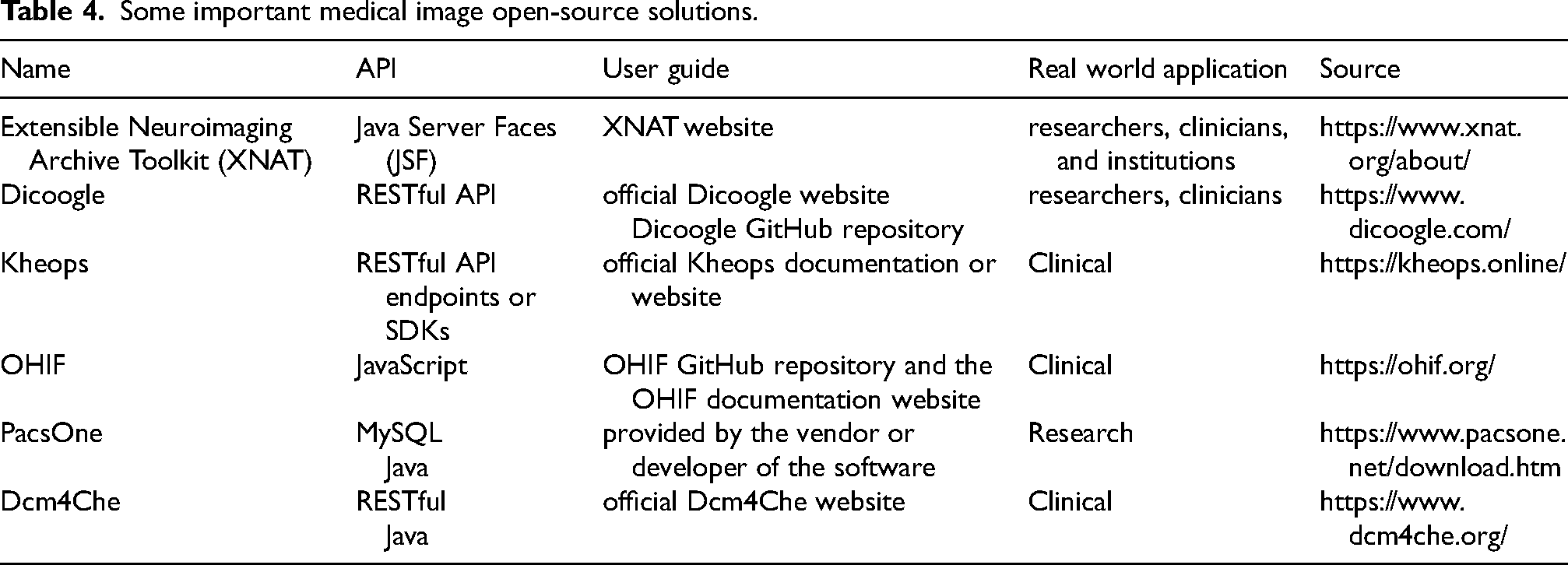
Obtaining imaging data is an essential element of developing AI algorithms for imaging diagnostics. These data sets are useful for training and testing of AI algorithms. Considering the importance of patient privacy, many market oriented AI models rely on private datasets or hospital datasets that are not publicly available. The main aim of this study is to explore the best accessible medical imaging datasets along with modality types, body organs, medical image classification and file format in depth. We believe that this succinct overview will help the scholars in an efficient and straightforward manner. Most of the datasets are open access; however, few of them require registration to view the data. Table 3 displays the details of the state-of-the-art medical imaging datasets.
There are many challenges to working with medical imaging datasets. Many of them were explored at MICCAI (a well-known conference—https://miccai.org/), and researchers/organizations attempted to address these difficulties by providing specialized datasets. Starting in 2018, MICCAI additionally developed an online platform for sharing and addressing these challenges. Table 2 presents many well-known databases, such as TCIA data repository provides curated imaging sets for several organs, focusing on cancer imaging. 17 The database expanded to include x-ray and computed tomography (CT) images for COVID-19 patients. Images from this site can be downloaded in collections based on a common condition or imaging modality. Similarly, the UK Biobank is another important resource in the field of medical data collection and research. 29 In addition to a wide range of clinical data, such as electronic health records (EHR), it has imaging collections from over 100,000 individuals, including scans of the abdomen, brain, heart, carotid artery, and bones.
The imaging repositories mentioned earlier contain datasets spanning multiple organs and diverse medical conditions. Nonetheless, there are additional initiatives for data collection that concentrate on specific organs. For instance, substantial collections of neuroimaging datasets are available from repositories such as IDA, OASIS, NITRC, CQ500, and so on. These repositories encompass imaging data for healthy individuals across different age groups, as well as patient data pertaining to various neurological disorders. Medical imaging databases are essential for clinical AI applications. The accuracy, reliability, and impartiality of diagnostic methods are determined by the datasets used to train and evaluate the models.
Data preparation
Data preparation is the most critical stage before using medical images for developing AI techniques. 11 One of the primary objectives of this research work is to offer a comprehensive overview of the medical image data preparation, which can be employed before and during the development, execution, and validation of AI algorithms. Next, we highlight the necessary steps when working with medical images.
Ethical process
Normally, obtaining approval from a local ethical committee is a prerequisite before utilizing medical data for the development of AI techniques. 11 A review committee is tasked with evaluating the risks and benefits of the study for patients. 11 In clinical studies, individual principal researchers may be required to grant permission for the disclosure of data concerning their patients. 11 After completing all ethical formalities, the relevant data should be made accessible, systematically searched, accurately de-identified, and securely stored. Any confidential patient data must be omitted from each of the DICOM metadata and the image files. 11 Figure 1 summarizes the process of preparing medical images for AI development.
Accessing data
AI algorithm developers often lack direct access to medical imaging data through PACS (Picture Archiving and Communication System), particularly for commercial purposes. PACS is a comprehensive solution for the storage, management, and retrieval of medical images. This approach transformed old film-based procedures by automating radiological images like MRI, CT, PET, and X-rays, enabling for computer storage. Enabling data access for AI developers is a challenging task that involves multiple processes, one of which is data de-identification (discussed below). After data is available to researchers, there are several ways for searching for medical images and clinical information.
Querying data
After data becomes accessible to AI developers, many methods exist for searching medical images and clinical information/data. 11 Researchers may use customized search commands to access medical data. Custom search commands may include strings, globally disease categorization codes, and modern medical terminology codes. PACS or radiology information system search engines can be used to conduct a systematic search and retrieval of data from hospital PACS and digital health records. Data must be regularly examined and extracted from both PACS and digital health records. Many PACS providers, for example, provide users with access to metadata like the annotations, the source, sequence, and image numbers, as well as unique target injury name and relationship. Researchers can access this data in some PACS and can further organize and control it by other systems such as digital cancer repositories, medical repositories, and other databases. 43 Alternatively, software tools exist to facilitate the process of data querying.43,44
Image de-identification
Image de-identification is the process of removing or anonymizing personal information, such as name, address, and medical record number, from medical images to protect the privacy of those concerned. 6 De-identification is crucial when sharing or using medical images for research, education, or other purposes outside of direct patient care. The de-identification process typically involves the various steps such as removal of patient name, date of birth, medical record number from metadata, anonymization of dates, overlay removal and so on.6,11 These identification data is normally available in DICOM format, and many tools are available to remove this information. The goal of image de-identification is to balance the utility of medical images for research and educational purposes while safeguarding patient privacy and complying with ethical and legal standards, such as the Health Insurance Portability and Accountability Act (HIPAA) in the United States or similar regulations in other regions. 11 Figure 2 shows the list of 18 HIPAA identifiers.
Data storage
Medical image storage is the secure and effective archiving of medical images created by x-rays, CT scans, MRIs, ultrasounds, and other imaging modalities. Proper storage is vital for maintaining, retrieving, and preserving these valuable diagnostic and patient care assets. Here are the main characteristics of medical image storage:
Aside from commercial alternatives such as PACS, Vendor Natural Archives (VNA), there are various open-source solutions that require little commitment from academics and physicians to implement.45,46 All these open-source alternatives offer platforms for creating your own server, but they do not provide a free storage facility, which would be expensive to operate. They are, however, extendable and include several plugins, allowing you to save medical images on the cloud with an individual provider that complies with data protection rules. Table 4 represents open-source medical image storage solutions.
In summary, medical image storage involves the careful management of digital images, ensuring their security, accessibility, and compliance with regulatory standards, to support effective patient care and medical research.
Quality control
Quality control in medical image processing is an important step in ensuring that the images obtained are of good quality, free of artifacts, and suitable for accurate diagnosis and analysis.47,48 It is a systematic process of checking and evaluating images to identify any issues that may impact their quality. The overall accuracy, precision, and durability of images acquired throughout the acquisition stage. It includes a variety of parameters that influence the quality and usefulness of the collected images for their final usage.49,50 The main parameters to improve the image quality are noise reduction, color accuracy, sharpness, and so on. Artifact identification and removal is the second step in quality control. It is the process of detecting and removing undesired features in images known as artifacts. Artifacts are distortions, abnormalities, or inconsistencies that occur during image acquisition, processing, or transmission.51,52 They may affect the quality of images, hide essential details, and restrict proper interpretation and analysis. Similarly,
Physicians, technicians, and other medical professionals need to work together to ensure top-notch medical image preparation. Regular inspections and ongoing training promote an environment of quality and constant advancement in the use of medical imaging. Furthermore, compliance with laws and regulations is required to achieve quality assurance standards.6,11
Structure data
It is the process of arranging and classifying medical information in a uniform way that enables retrieval, storage, analysis, and exchange. It is essential for ensuring stability, interoperability, and effectiveness across multiple healthcare networks. 6 Organizing medical imaging datasets is essential for efficient storage, retrieval, and analysis in health care organizations. The DICOM format is widely used for storing and exchanging data associated with medical imaging. DICOM images are frequently stored in a file format that follows the DICOM standard. These files may contain extensions like.dcm or.dicom. it includes metadata such as patient history, imaging type, acquisition settings, and so on. This metadata is essential to properly order and analyze the photos.11,13
In addition, medical images must be organized systematically at the patient, study, series, and occurrence levels. Likewise, employ a uniform file name approach to make it easy to find and handle image files. PACS systems have been developed to efficiently organize and make available DICOM images.6,46,54 Medical personnel can successfully handle medical images by following prescribed guidelines, arranging data hierarchically, and incorporating metadata. This structured strategy optimizes medical procedures, research activities, and cross-institutional collaboration. 46
Labelling data
Labelling medical images means annotating or classifying regions of interest in the image to offer ground truth information for AI algorithms training/testing or research purposes.6,46 Image annotation is one of the basic problems in image labelling.
It consists of regions of interest, landmarks, and label classification. Selecting appropriate annotation tools for each annotation class is an additional vital component of data labelling. These tools might vary from simple sketching tools for identifying regions to more advanced tools for 3D volumetric annotations. 6
Likewise, uniformity in labelling and multidimensional annotations are challenging aspects of data labelling. To produce reliable and consistent information, annotators must employ a uniform labelling mechanism. Define annotation criteria or utilize standardized taxonomies when appropriate, and experiment with annotating medical images from various imaging types, such as MRI, x-ray, or CT scans, to create large datasets for algorithm training/testing or research studies. Another challenging area of data labelling is incorporating metadata annotations such as patient history, imaging parameters, and acquisition information into the labelled dataset. This information enhances the contextualization of the images. Furthermore, documentation of the labelling process, including guidelines, definitions, and any obstacles encountered, is essential throughout the data labelling process. This documentation is useful for future use and sharing the dataset with others.
Labelling medical image data is an important step in testing and validating AI algorithms for analyzing medical images. Whether for training deep learning models or supporting clinical workflows, accurate and standardized annotations contribute to the reliability and effectiveness of medical image datasets. A comprehensive list of data annotation tools has also been provided by research studies.55,56 Nevertheless, Table 5 also offers a variety of open-source platforms and applications focused on medical images.
Data curation tools for medical imaging
Medical image data curation tools are advanced software applications or platforms designed to assist in the organization, management, integrity, annotation, verification, extraction, and quality control of medical image datasets. These tools play a crucial role in preparing medical imaging data for research, training, and clinical applications. Without proper data curation techniques, AI algorithms may exhibit low efficiency and accuracy, resulting in unsatisfactory outcomes and, in some situations, failure. To present a comprehensive and informative collection of curation tools, we focus on those with general uses that address common use cases in medical imaging. For example, Table 6 illustrates widely used curation tools. Data curation tools are essential for reviewing, detecting, avoiding, and addressing shortcomings in datasets. 6 As a result, without data curation techniques, possible issues such as errors from untrustworthy data, bias introduction, and doubt about the validity of prediction outcomes may arise later in the AI development process.
Table 5 highlights some prominent data curation tools available to researchers, clinicians, and healthcare professionals. Choosing the right data is critical for training and evaluating AI algorithms.
An effective data curation tool must be able to clean, arrange, and locate relevant data for particular tasks. This requires handling huge datasets and filtering data based on specific criteria. Likewise, comprehending and interpreting large datasets needs unique data visualization.
An effective tool should be able to provide data in a variety of formats, such as graphs, tables, and images, and allow the user to modify these visualizations to meet their specific needs. A data curation system should be able to deal with a wide range of data types, including images, DICOM, and BIDS, as well as all labelling standards such as boundaries, grouping, and polyline.
Furthermore, data curation tools should have a simple and user-friendly interface, as they are frequently used by both technical and non-technical stakeholders. According to the individual demands and requirements of a project or study, multiple technologies may be employed to successfully organize and curate medical data for a range of clinical and scientific applications. Some important free medical imaging software are discussed below.
Open-source software for medical imaging
The research community has shown a strong interest in open-source software, making substantial contributions to the development of publicly accessible software for a wide range of applications. This includes image processing tasks essential to AI research, such as anonymization, curation, categorization, and labelling of medical images.
For example, ‘ImageJ’ is a multiplatform, Java-based image processing and analysis tool. 52 It is freely available in the public domain and does not require a license. It supports various file formats using freely accessible plugins. ImageJ offers a wide range of capabilities for manipulating images, such as image filtering, edge recognition, sharpening, and morphological processing. Additionally, it provides analysis tools for computing regions, it also includes analysis tools for analyzing regions, boundaries, and angles on specified regions. In addition, ImageJ can natively manage multidimensional data, such as image stacks obtained from MRI scans.
Medical Image Interaction Toolkit (MITK) is an open-source program developed using the Insight Toolkit (ITK). 52 The software offers intuitive tools for both manual and computer-assisted image classification. It is compatible with all commonly used file formats in medical imaging and has a multi-window interface that makes it easy to view and interact with the images. The integrated semi-automatic tool employs active contour techniques. In addition, the latest iterations of the latest version of MITK known as ITK-Snap, now incorporate registration functionality, which improves the handling of multimodal images. In addition, it provides a decentralized segmentation solution that allows for cloud-based segmentation using algorithms given by the web developer community.
The Open Health Imaging Foundation (OHIF) (https://ohif.org/) Viewer is a web-based open-source platform of medical imaging. 57 The objective is to provide a basic framework for constructing advanced imaging applications. The purpose of this open-source software is to quickly load huge radiology trials by retrieving information in advance and streaming the necessary imaging pixel data as needed. The OHIF enables users to develop web-based imaging applications without the need to repeatedly build essential viewer functionality for every new application.
Several open-source applications and libraries have been developed specifically for medical image processing. Notable examples include DIPlib (https://diplib.org) and Icy (https://icy.bioimageanalysis.org/). However, for the sake of conciseness, they are not elaborated upon in this paper.
Discussion and future directions
According to various healthcare professionals,47,58 AI is profoundly changing the field of medicine, with a particularly significant impact on medical imaging. AI has shown tremendous potential to outperform humans in certain operations, such as image segmentation. Furthermore, AI offers vital insights into the medical process of decision-making. Without AI, it would have been exceedingly difficult, almost impractical, to optimally combine and extract this information. The advancement of AI owes much to the growing availability of (publicly available) medical imaging databases. These images serve as critical inputs for AI models that retrieve the most relevant attributes, which aid in the identification of anatomical structure boundaries and the prediction of disease. Nevertheless, before this stage, it is imperative to adequately prepare medical images to ensure best utilization and maximize their abilities in developing AI and assessment.
As presented in this study, during the last decade, various publicly available medical image databases and open-source tools have evolved to encourage established standards for preparing data for clinical imaging. Nonetheless, significant issues remain, requiring ongoing attention and research, as detailed below.
Image de-identification is critical for protecting patient privacy. In recent years, many acts, such as GDPR and HIPPA, have been revised, necessitating the regular alignment of image de-identification tools with these changes. As a result, techniques for automating this process (which is now done semi-automatically) are required, as is validation that these de-identification tools effectively satisfy regulatory guidelines. For example, when developing a 3D reconstruction of the head, it is critical to avoid revealing the individual's identification. As a result, certain spatial information, such as facial features, should be erased. Nevertheless, the challenge lies in removing identifiable facial features while maintaining essential scientific knowledge without modification. This offers a dilemma, especially in conditions such as neck cancer and radiation therapy planning, where essential data is currently compromised. Data curation is a crucial phase that ensures the data is well-organized and managed. After defining the data collection procedure objectively, we should focus on improving the data quality. This can be accomplished by creating standards and rules across the entire process of medical image preparation, spanning from the de-identification phase to the data annotation stage, with a special emphasis on data curation. Furthermore, data collection operations are critical for studies on AI in clinical imaging since they enable the formation of standards to assess AI across numerous centres and scanners. There is an urgent need for automated tools and standards to evaluate image quality, especially for quantitative studies. Additionally, the development of methods for automatically identifying and rectifying image artifacts will be required to ensure a consistent level of quality in images used to train AI algorithms. Encouraging such initiatives to promote quality standards and tools is critical for assuring the dependability of image labelling, annotation, and attributes. Image annotations play a crucial role in ensuring the accurate training of AI algorithms and should be conducted meticulously. Nevertheless, achieving precise delineations or annotations poses significant challenges and is exceedingly time-consuming, particularly in 3D imaging modalities. For clinicians, annotating the thousands of images to train AI algorithms could be a challenging and impractical task. Both existing and future public and collaborative annotation technologies have tremendous value in capturing the diversity of annotations generated by multiple physicians.
Aside from diving into data preparation, the major focus of this study, it is important to anticipate the possible future trends of AI for clinical imaging, which include: (a) data augmentation, (b) ethical considerations regarding AI, (c) federated learning, and (d) uncertainties estimation.
Data augmentation has emerged as a promising approach within AI to enhance the data preparation phase. Cutting-edge data augmentation methods span from fundamental strategies employing practical geometric transformations, color adjustments, cropping, flipping, and noise injection, 48 to more sophisticated techniques. 59 Federated Learthe futurening is a cutting-edge technique that has been promoted in medical research to protect patient privacy while simultaneously improving the imaging datasets utilized by AI algorithms. 60 In the conventional approach, de-identified data are moved from the hospital (or silo) to a central storage system. However, with federated learning, the data remain within the hospital while the algorithm can be trained locally at multiple locations. Figure 3 shows the layout of existing versus federated learning mechanisms.
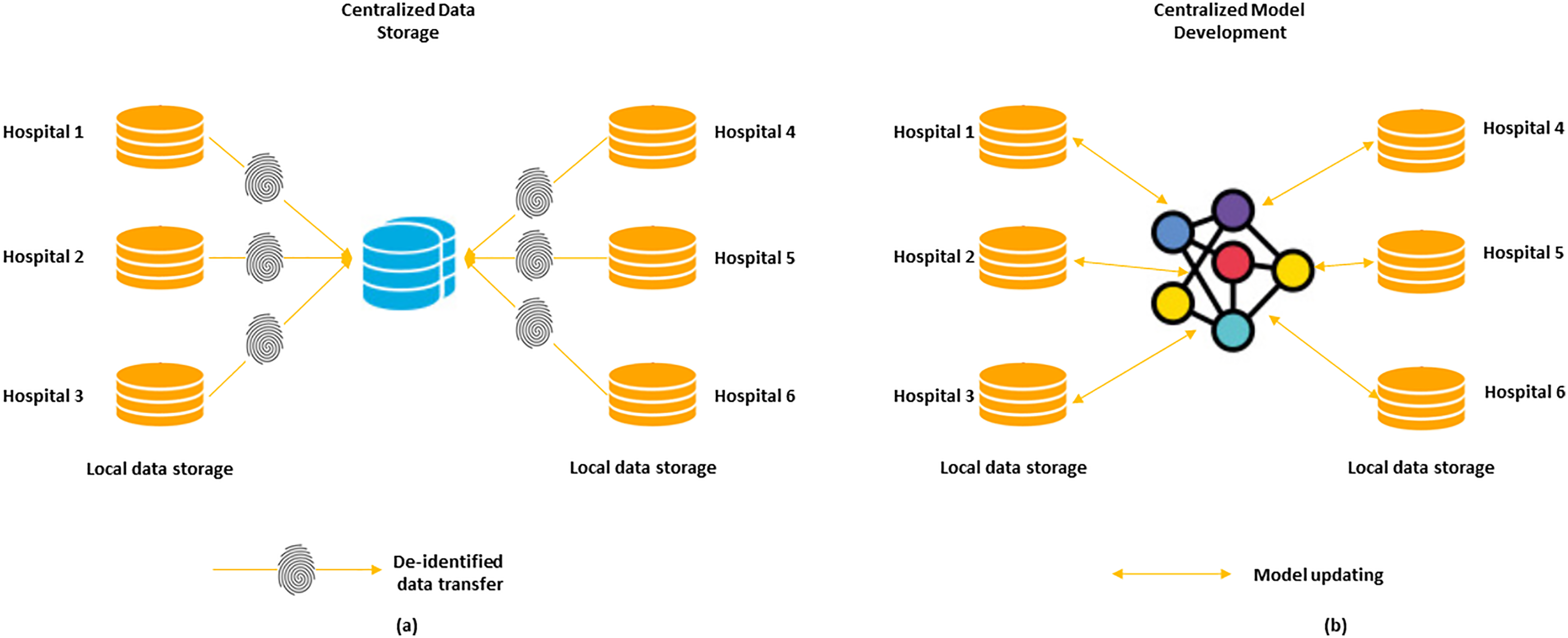
Existing versus federated learning (a). Today's AI model development involves transferring de-identified data to a centralized storage system. (b). Federated learning may be used in the future.
Another key component in medical sciences is the usage of AI tools. The study presented in61,62 addresses a significant concern, emphasizing that the ethical application of these tools in the medical sciences should aim to improve well-being and reduce suffering. As discussed earlier, numerous factors influence the data preparation process and its quality during training. As a result, alongside reliability and accuracy, the prediction confidence level of AI algorithms must be evaluated for image analysis. Uncertainty estimation holds particular significance given the imperfect nature of data preparation. Healthcare professionals should be notified of high uncertainty values so that they can incorporate this data in their final decisions. We are hopeful that this emerging research area will increase the applicability of AI in real-world scenarios by improving the reliability of methods that are currently seen as black boxes.
Conclusion
In this paper, we reviewed an in-depth examination of the publicly accessible datasets in the field of medical imaging. This work also determined the methods required for preparing medical imaging data for the development of AI algorithms and emphasized current limitations in dataset curation. This phase must be completed before starting the design or deployment of any AI algorithm. Furthermore, this study explored inventive strategies to address the challenge of data availability, offered a detailed overview of data curation tools. The provided organized explanation provides researchers and clinicians with an in-depth guide to selecting from the many currently accessible tools for preparing clinical images prior to implementing AI methods. The paper concluded with an insightful discussion and analysis of challenges in medical image analysis, along with potential future directions in the field.
In addition to the primary focus of this work, which was data preparation, it is imperative to anticipate emerging trends in AI for clinical imaging, such as data augmentation, ethical issues surrounding AI, federated learning, and uncertainty estimation.
Footnotes
Acknowledgments
This study is supported via funding from Prince Sattam bin Abdulaziz University, Alkharj, Saudi Arabia.
Authors contribution
Abdulrahman Alabduljabbar (Data curation; Writing – original draft; Writing – review & editing); Sajid Khan (Conceptualization; Data curation; Writing – original draft); Anas Alsuhaibani (Formal analysis; Investigation; Project administration; Resources; Writing – review & editing); Fahdah Almarshad (Data curation; Formal analysis; Methodology; Validation); Youssef N Altherwy (Formal analysis; Funding acquisition; Project administration; Software; Supervision; Validation; Visualization).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Thanks to Prince Sattam University for funding via project number ((PSAU/2024/01/29710).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
Data may be provided by request to corresponding author.
