Abstract
Background:
The investigation of mitophagy in Alzheimer’s disease (AD) remains relatively underexplored in bibliometric analysis.
Objective:
To delve into the progress of mitophagy, offering a comprehensive overview of research trends and frontiers for researchers.
Methods:
Basic bibliometric information, targets, and target-drug-clinical trial-disease extracted from publications identified in the Web of Science Core Collection from 2007 to 2022 were assessed using bibliometric software.
Results:
The study encompassed 5,146 publications, displaying a consistent 16-year upward trajectory. The United States emerged as the foremost contributor in publications, with the Journal of Alzheimer’s Disease being the most prolific journal. P. Hemachandra Reddy, George Perry, and Xiongwei Zhu are the top 3 most prolific authors. PINK1 and Parkin exhibited an upward trend in the last 6 years. Keywords (e.g., insulin, aging, epilepsy, tauopathy, and mitochondrial quality control) have recently emerged as focal points of interest within the past 3 years. “Mitochondrial dysfunction” is among the top terms in disease clustering. The top 10 drugs/molecules (e.g., curcumin, insulin, and melatonin) were summarized, accompanied by their clinical trials and related targets.
Conclusions:
This study presents a comprehensive overview of the mitophagy research landscape in AD over the past 16 years, underscoring mitophagy as an emerging molecular mechanism and a crucial focal point for potential drug in AD. This study pioneers the inclusion of targets and their correlations with drugs, clinical trials, and diseases in bibliometric analysis, providing valuable insights and inspiration for scholars and readers of JADR interested in understanding the potential mechanisms and clinical trials in AD.
INTRODUCTION
Alzheimer’s disease (AD), a prevalent and progressive neurodegenerative disorder, was initially identified by German psychiatrist Alois Alzheimer in 1906. AD is primarily characterized by a decline in cognitive function. AD imposes a remarkable burden and causes tremendous loss not only for patients and their families but also for society [1]. The escalating number of AD cases has led to extensive research aimed at a deeper understanding of the disease.
Thus far, only two categories of drugs have received approval from the Food and Drug Administration (FDA) for the clinical treatment of AD. The first category focuses on alleviating AD symptoms, including drugs such as donepezil, rivastigmine, galantamine, and memantine, and controlling behavioral and psychological symptoms, such as orexin receptor antagonists and atypical antipsychotics [2]. The second category aims to modify the underlying biology of AD, exemplified by monoclonal antibodies such as aducanumab [3–5] and lecanemab [6]. Recent findings from an ongoing clinical trial demonstrated that another monoclonal antibody, donanemab, notably slowed cognitive and functional decline in early symptomatic patients with AD (either mild cognitive impairment or mild dementia) [7]. Although these antibodies remove amyloid-β (Aβ) from the brain, the mechanisms of action of these treatments differ. However, considerable attention has been devoted to other AD drugs for asymptomatic patients or those with preclinical AD, such as solanezumab [8] and gantenerumab [9]. Despite the outcomes of clinical trials for these drugs (solanezumab, NCT02008357, completion; gantenerumab, NCT04339413 and NCT04374253, termination) not meeting expected cognition rescue outcomes, there is anticipation surrounding the A4 study (anti-amyloid treatment in asymptomatic Alzheimer’s disease) targeting cognition improvement in asymptomatic elders [10, 11].
Several widely recognized hypotheses have been postulated regarding AD pathogenesis. These hypotheses encompass the Aβ cascade [12–14], abnormal phosphorylation of tau protein [15, 16], neuroinflammation [17, 18], APOE allele mutations [19, 20], oxidative stress [21], mitochondrial dysfunction, cholinergic dysfunction [22], and imbalances in neurotransmitter and metal homeostasis [23–25]. Despite multiple hypotheses attempting to elucidate AD pathogenesis, the precise mechanisms underlying the disease remain elusive.
Neurons, being highly energetically demanding cells in the brain, rely on normal mitochondria for their active and crucial functions. Mitochondria, dynamic organelles with a distinctive balance between fission and fusion in their life cycle, are distributed from the soma into dendrites, axons, and synaptic terminals [26]. Mitochondrial dynamics play a critical role in maintaining the function and energy status of neurons [27, 28]. Altered mitochondrial morphology and dysfunction might be related to neuronal metabolic disorders and energy deficiencies in AD [29, 30]. Aβ-induced mitochondrial dysfunction manifests as a defective citric acid cycle, decreased respiratory chain complex III/IV activity, and increased generation of reactive oxygen species (ROS) [28, 31].
Autophagy serves a crucial role in safeguarding cells by eliminating damaged mitochondria and preserving healthy mitochondria. Over the past decade, research into autophagy and its relation to mitochondria has garnered significant attention in the context of AD. Initially proposed by Lemasters JJ [32], “mitophagy” was identified to potentially play a pivotal role in halting age-related accumulation of mutations in mitochondrial DNA. Subsequently, mitophagy has been recognized as a focal point in the pathological progression of multisystem diseases. Parkin, a critical protein in mitophagy, mediates the removal of damaged mitochondria into autophagosomes and facilitates their degradation through selective autophagy [33]. In AD brains, the extent of mitochondrial autophagic degradation is increased, accompanied by mitochondrial dysfunction and oxidative damage [34]. Fang EF has highlighted that impaired clearance of dysfunctional mitochondria is a key event in the pathological mechanism of AD. Enhanced mitophagy has the potential to reduce Aβ plaques and tau pathology, thus mitigating cognitive impairment [35]. In summary, the aforementioned findings suggest that mitophagy might serve as a potential target in AD treatment, which is increasingly prevalent but lacks effective clinical therapeutic drugs.
For decades, bibliometric analysis has been widely used by applying statistical and mathematical methods to comprehensively examine publications with summaries and prospects of countries, journals, articles, authors, citations, keywords, and other information in various fields [36, 37], including autoimmune diseases and tumors [38, 39]. Since 2006, bibliometric analysis has been used in AD research. Ansari MA was the first to conduct a bibliometric analysis of AD publications from 2003 to 2004, focusing on publication types, languages, countries, authors, and collaborations [40]. Subsequently, several researchers successively analyzed publication data, co-citations, keywords, and topics in AD [41–44]. While a few studies have explored mitophagy in other fields [45, 46], previous bibliometric analyses have seldom addressed targets, drugs, or clinical trials in AD. Notably, neither mitophagy nor mitochondrial dysfunction as a pathogenic factor in AD has been encompassed in past bibliometric analyses.
This study aimed to bridge the gap in the application of bibliometric analysis in understanding mitophagy and AD. Leveraging data from 5,146 articles in the Web of Science Core Collection (WoSCC) database, VOSviewer, Citespace, Pajek, Scimago Graphica, GenDoma, and other analytical methods were employed to scrutinize basic bibliometric data, keywords, targets, drug-target associations, clinical trials, and their relationship with mitophagy and AD publications over the past 16 years. This research endeavors to offer a comprehensive overview of research trends in mitophagy and AD for the benefit of researchers in this field.
MATERIALS AND METHODS
Data source, extraction, collection, and search strategy
The Web of Science Core Collection (WoSCC, https://www.webofscience.com/wos, Clarivate Analytics, USA) is a citation-based database facilitating the index and analysis of publications. This comprehensive resource includes crucial information such as titles, authors, affiliations, countries/regions, and author keywords. Its reference data empowers researchers to unlock its full potential, making WoSCC the ideal database for bibliometric studies, extensively employed in numerous research endeavors [47, 48]. Hence, the present study on AD and mitophagy utilized data from this database.
To mitigate potential researcher bias, two authors executed the search strategy concurrently. Employing the search formula #1: Ts = (“alzheimer’s disease” OR “alzheimer disease”); #2: Ts = (“mitophagy” OR “mitochondrial autophagy”); Final dataset: #1 AND #2 (Supplementary Table 1), pertinent publications were systematically collected. Adhering rigorously to the search formula, detailed in Supplementary Table 1, all papers were retrieved utilizing the Topics (Ts) search strategy within WoSCC. Ts search records encompassed titles, abstracts, author keywords, and keywords plus. Thus, both terms “mitophagy” and AD, as Ts, were required in the title, abstracts, author keywords, or keywords plus to meet our criteria for bibliometric analysis. Subsequently, publications from 2007 to 2022 were selected, limited to review and original articles only. Specific inclusion criteria are outlined in Fig. 1, resulting in a total of 5,146 publications considered for analysis. These publications were downloaded in text format, including “full records and cited references” for bibliometric analysis. Supplementary bibliometric details such as Impact Factor (IF) were sourced from the 2023 Journal Citation Report (JCR, Clarivate Analytics, Clarivate PLC, USA) using Clarivate Analytics. Additionally, data such as the number of citations and average citations per year were obtained from WoSCC’s citation report.
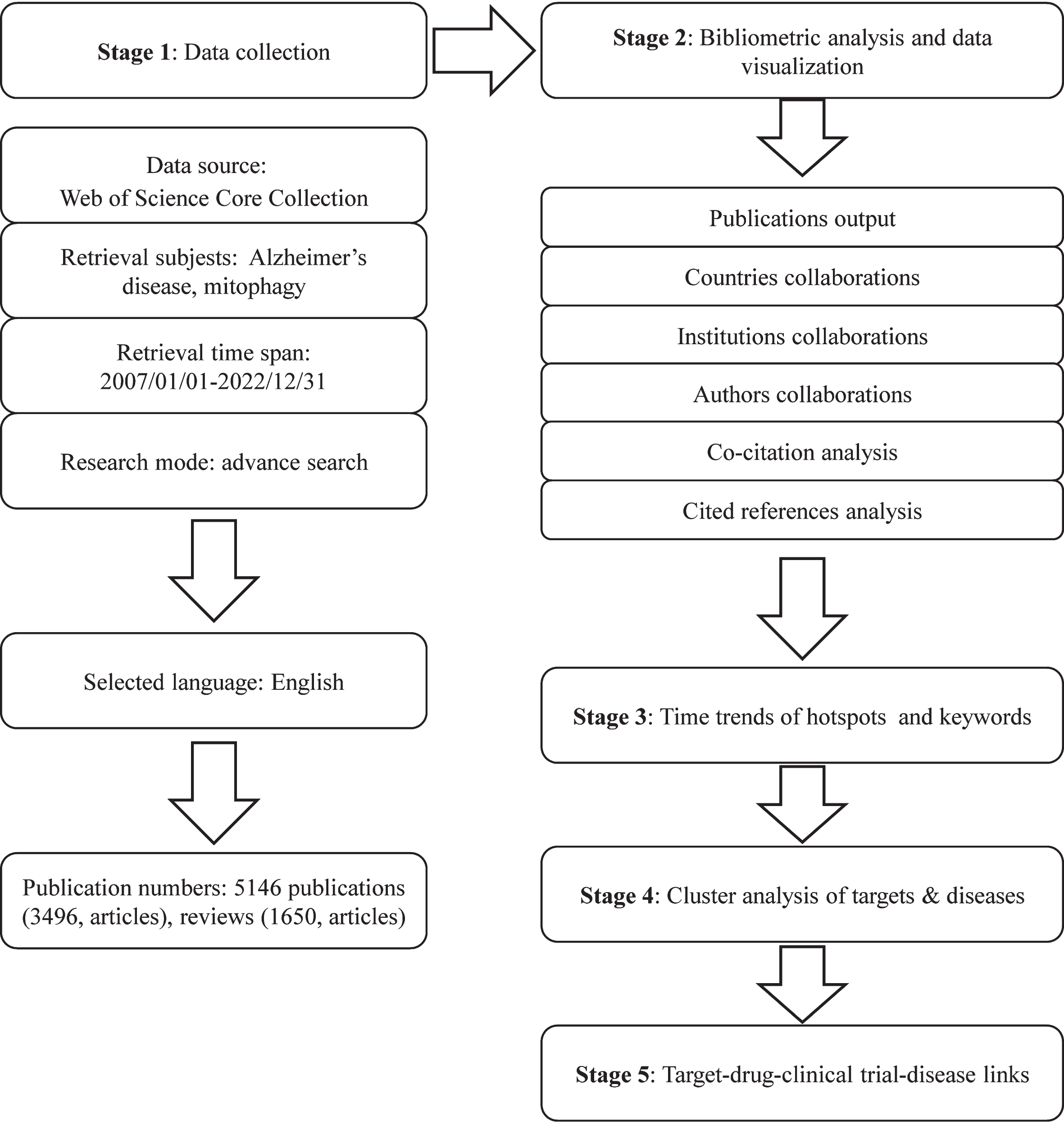
Flowchart showing the search strategy and analysis of publications.
Bibliometric analysis
To conduct a comprehensive data analysis, various bibliometric software tools were utilized, namely VOSviewer, Pajek, CiteSpace, and Scimago Graphica.
VOSviewer (v.1.6.18, https://www.vosviewer.com/, Centre for Science and Technology Studies, Leiden University, The Netherlands), a well-established bibliometric analysis software application, is widely recognized for clustering and mapping crucial information from publications [49, 50]. Utilizing VOSviewer, collaborations among countries, institutions, journals, and high-frequency keywords were visualized. In the generated visual maps, nodes represented different entities such as countries, institutions, authors, or keywords, with node size reflecting their frequency of co-occurrence in titles, abstracts, and keywords. The links between nodes symbolized cooperation and relationships, distinguished by different colors representing distinct clusters.
In tandem with VOSviewer, Pajek (v.5.18, http://vlado.fmf.uni-lj.si/pub/networks/pajek/, Andrej Mrvar and Vladimir Batagelj, Faculty of Computer and Information Science, University of Ljubljana, Ljubljana) was employed to elucidate relationships and connections. Pajek serves as a proficient network analytic tool designed to visualize and streamline large networks. Furthermore, in this study, graph data and correlation networks were computed and refined using Pajek to enhance clarity and comprehension.
CiteSpace (v.6.1.6, https://citespace.podia.com, Mountain View, CA, USA), a free Java-based application developed by Professor Chaomei Chen in 2004 [51], serves as a globally recognized tool for visualizing and analyzing scientific literature trends and citations. Its capabilities extend to presenting the structure, laws, and distribution of scientific knowledge, offering a comprehensive overview of research fields and structured bibliometric networks. In this study, CiteSpace was used for clustering and visualizing the co-citation patterns among 5,146 publications.
In addition to CiteSpace, Scimago Graphica (v.1.0.34, https://graphica.app/, SCImago Lab, SRG S.L. company, Spain) was employed to visualize collaborative relationships among countries and to track time trends in keywords and research areas. The global distribution map of national publications and the cooperation network of countries were depicted, with each node representing a country/region and its size determined by the total publications. Lines connecting nodes indicated collaboration. The time trend map of keywords was segmented into 4-, 3-, 3-, 3-, and 3-year intervals, after which they were ranked and correlated using Scimago Graphica [52].
Cluster analysis
For keyword analysis, a custom algorithm in CiteSpace generated a clustering network diagram based on keyword co-occurrence. Utilizing the LLR algorithm, this software calculated modularity and mean silhouette values, identified themes and occurrence times of keywords, and highlighted core keywords within the 5,146 publications. The resulting co-occurrence network diagram displayed clustering results reflecting similarity and co-occurrence features. In the realm of target and disease analysis, VOSviewer employed its algorithm to cluster institutions, authors, keywords, and targets within the 5,146 publications. This method classified co-occurrence phenomena into clusters representing specific themes, depicted as distinct-colored nodes. Node size indicated occurrence or co-occurrence, while relative node positions conveyed topic similarity. Edge thickness represented co-occurrence frequency, and network density indicated connection strength.
Drug interactions and clinical trials
To explore drugs or molecules in literature, a preliminary definition was established before conducting the analysis. Using Bidirectional Encoder Representations from Transformers for Biomedical Text Mining (BioBERT, https://github.com/naver/biobert-pretrained, https://github.com/dmis-lab/biobert) [53], all terms related to drugs or molecules were identified within the abstracts of relevant literature. Subsequently, these drugs or molecules were extracted and subjected to statistical analysis. Following this, a keyword co-occurrence analysis was performed to establish connections between the identified drugs and their gene-related targets. Identification of drug or molecules associated with diseases was accomplished through the GenDoma web server (https://ai.citexs.com). GenDoma refined the retrieved data into drug or molecules-related terms, arranging them based on their frequency of occurrence. To enrich the target-drug-disease information, data from online resources such as the PubChem database and Clinical Trial website were integrated.
PubChem (https://pubchem.ncbi.nlm.nih.gov), an online chemical database, facilitated the analysis of drug or chemical interactions with targets. The Clinical Trial website (https://clinicaltrials.gov/) supplied information on phases, statuses, interventions, or treatments for diseases. A collection of 28 drugs/molecules, notably the top 10 representative drugs/molecules, was collated, including the number of their representative clinical trials, phase, status, associated diseases, and mechanisms of action.
Statistical analysis
The general bibliometric analysis encompassed WoSCC, VOSviewer, CiteSpace, and Scimago Graphica, as previously mentioned. GraphPad Prism (v.7.0) was used for analyzing line charts, while ggplot2 generated circular heat maps. Sankey figures were analyzed using ggplot2 (v.3.3.6) and ggalluvial (v.0.12.3), and cloud map was created using ggwordcloud (v.0.6.0). Targets obtained from GenDoma were clustered using VOSviewer, and heat maps of clustered targets were generated using GraphPad Prism (v.7.0, GraphPad Software, Boston, USA). Pathway analyses of targets were conducted using the Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO). Protein-protein interaction (PPI) analysis of targets involved tools such as STRING (https://string-db.org) and GeneMANIA (http://www.genemania.org). Diseases, also obtained from GenDoma, were clustered using VOSviewer.
RESULTS
Publication outputs and trends
After following the screening strategy flowchart and conducting a publication search process (Fig. 1), a total of 5,146 publications were identified, comprising 3,496 articles and 1,650 reviews. Notably, a rising focus on mitophagy in AD is evident annually (Fig. 2). The average annual publications within this research realm showed a steady increase from 2007 to 2022, growing at a rate of 12.84%, despite minor fluctuations. To assess the trend of AD and mitophagy studies further, a growth curve was fitted using the index function (Y = 108.29e0.1128x, R2 = 0.985) for annual publication trends. The years 2007 to 2011 represented an early stage, with fewer than 250 publications per year on the link between AD and mitophagy. However, from 2012 to 2019, publications steadily increased, exceeding 300 per year after 2016 and continuing to increase annually. Notably, the publications surged from 2019 to 2020, with nearly 21.8% (n = 97) of all publications occurring in the latter year. These findings carry significant implications for exploring and expanding the field of mitophagy in AD.
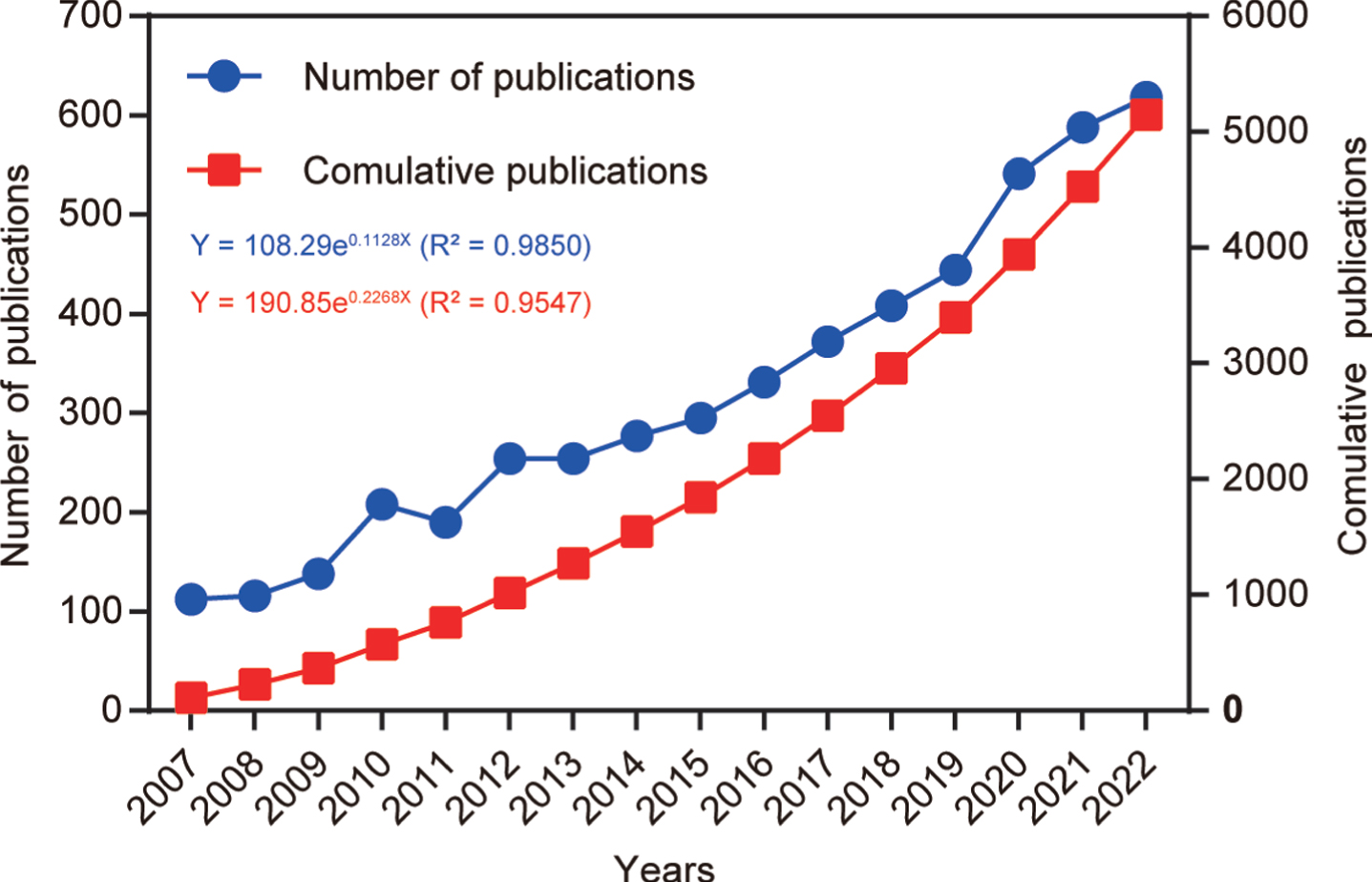
Number of annual publications from 2007 to 2022. The line chart displays the annual number of publications (red square point) and the cumulative number of publications (blue round point) from 2007 to 2022. The X-axis represents the year, and the Y-axis represents the annual number of publications.
Analysis of countries/regions and cooperation relationships
Subsequently, an analysis of publication numbers and collaborative relationships among countries or regions was conducted. A total of 90 countries or regions contributed publications on mitophagy in AD, with 48 of them producing more than 10 publications over the past 16 years. Using Scimago Graphica and VOSviewer software tools, a visual cooperation relationship network map was generated for these 48 countries (Fig. 3A). The USA maintained the highest number of publications (n = 1,692, 24.38%), closely followed by China (n = 1,204, 17.35%). Additionally, the USA exhibited a strong inclination for collaborative efforts with other countries, with the United Kingdom (UK) and China following suit. Further, these 48 countries were clustered based on the amounts of publications, weight value (cooperation links among countries), and score of citations, and displayed them as a heat map (Fig. 3B). Both the USA and China contribute to most publications on mitophagy in AD while fostering cooperative relationships, consistent with the findings of the visual network map (Fig. 3A). Detailed breakdowns of each country were presented in the sankey diagram and line chart (Fig. 3C, D). Notably, countries such as Colombia contributed fewer publications (n = 15) but garnered higher citation scores (116.8), a pattern akin to Denmark (80.62), Hungary (79.19), and Switzerland (78.48). Despite anomalies, most countries/regions showcased a positive correlation between publication numbers and citation scores.
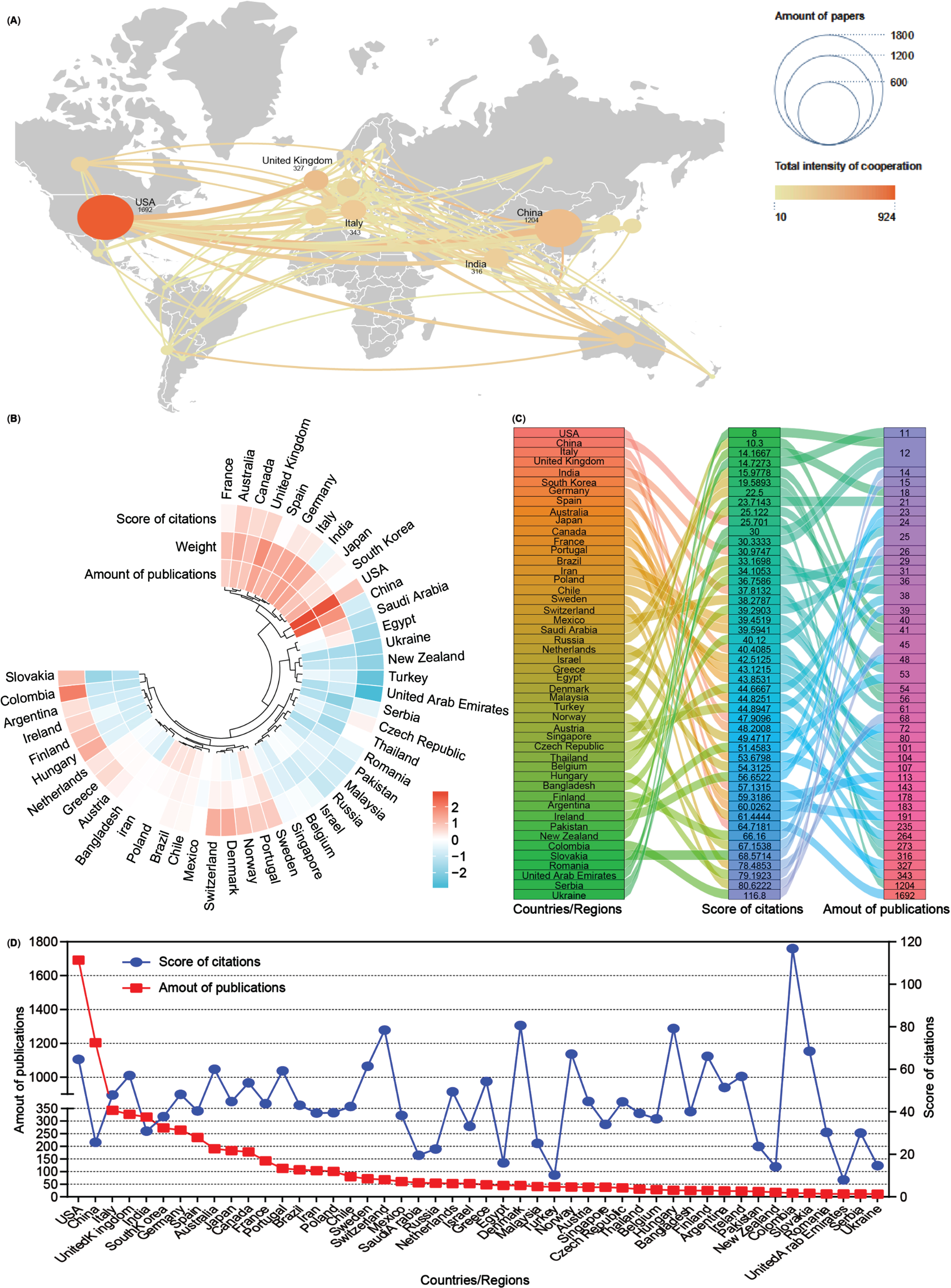
Co-operation analysis of countries/regions. A) Overlay world map of country analysis. The nodes in the map represent countries/regions, and the lines between the nodes represent cooperation relationships. According to the amount of publications and color gradient in the right corner, different nodes were allotted various sizes and colors. B) A circulus heat map shows the relationships among the publications, citations, and cooperation weight among 48 countries/regions. C) Sankey diagram shows the score of citations and the total amounts of publications in 48 countries/regions. D) Line chart displays the distribution of the amounts of publications (red square point) and the score of citations (blue round point) in 48 countries/regions.
Analysis of funding agencies and institutional cooperations
Drawing from WoSCC data, the top 10 funding agencies from the 5,146 publications were identified (Table 1). The National Institutes of Health (NIH) emerged as the most frequently cited funding agency (1,409 times), followed by the United States Department of Health (1,040 times) and Human Services and the National Natural Science Foundation of China (675 times). Among the top 10 funding agencies, three are based in the USA, with subsequent representation from the UK and Japan.
Top 10 funding agencies and frequency of funding
Subsequently, an analysis of the number of publications and collaborative relationships among institutions in the field of mitophagy in AD was conducted. A total of 4,355 institutions have contributed 5,146 publications related to this subject. To visualize these relationships, Scimago Graphica and VOSviewer software were used to generate an institutional network map that distinctly displays publications with at least 28 publications and collaborations among different institutions through lines (Fig. 4A).
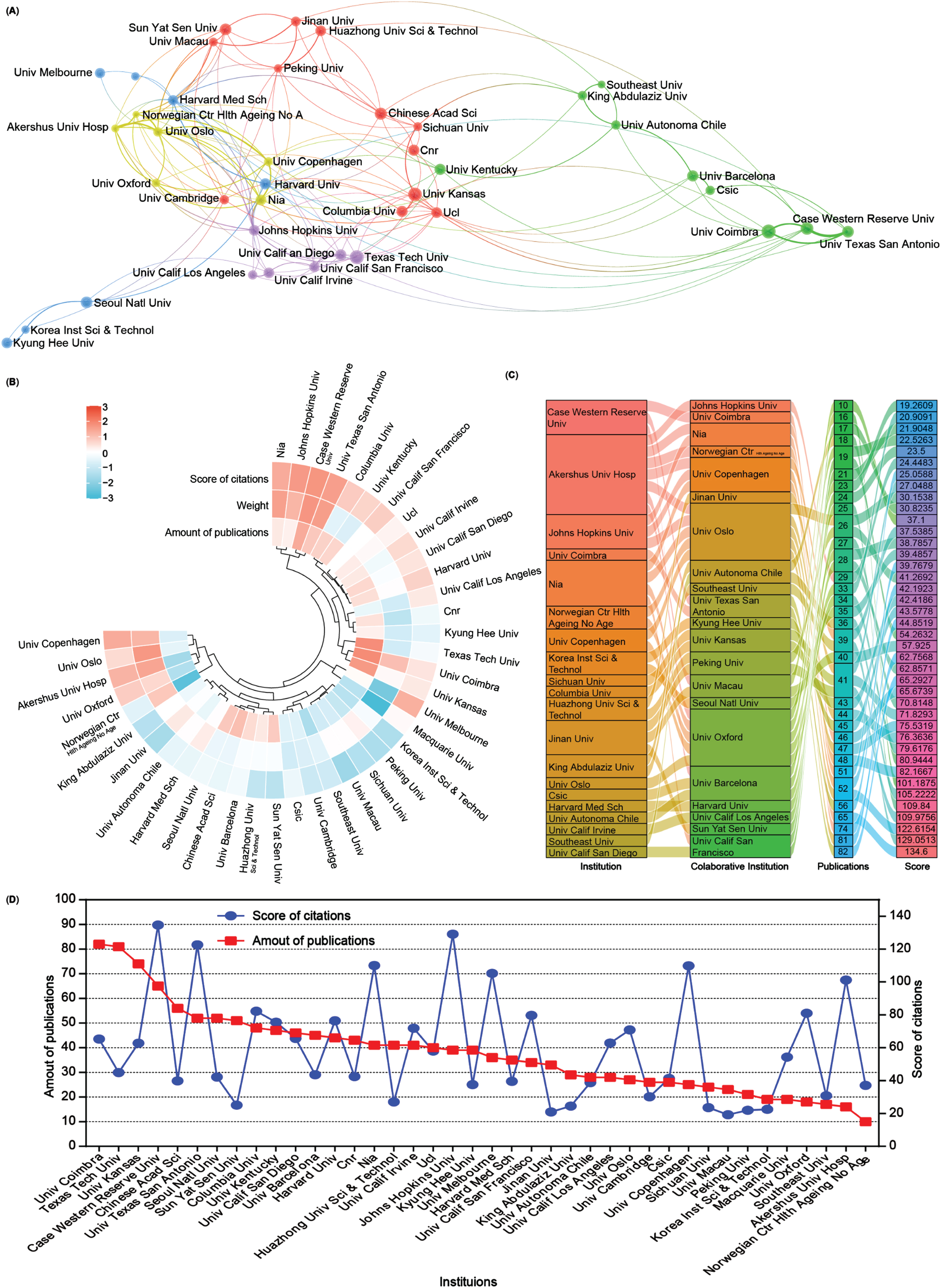
Collaboration analysis of institutions. A) Network of institutional collaborations. A node represents an institution, and the size of each node represents its quantity of publications. Lines represent the cooperation relationship between two institutions. B) A circulus heat map shows the relationships among the publications, citations, and cooperation weight in the top 40 institutions. C) Sankey diagram shows the cooperation relationships among institutions as well as the collaborative institutions, amounts of publications, and score of the average citations. D) Line chart displays the number of publications (red square) and the score of average citations (blue round) in the top 40 institutions.
A heat map and a sankey diagram were constructed to illustrate the publication quantities, citation scores, and institutional relationships (weight value). Following clustering, both publications and the total link strength weight value collectively affected the average citation rates. For instance, institutions such as Case Western Reserve University and the University of Texas at San Antonio exhibit high productivity, relatively high citations, and cooperation weight value in AD and mitophagy. Conversely, higher production and collaboration weight value at institutions such as Texas Tech University did not necessarily translate into higher citation scores. Moreover, institutions with fewer publications but higher link strength weight, such as Akershus University Hospital and the University of Oxford, could still contribute to achieving higher citation scores (Fig. 4B, C). Further details about each institution are presented in the line chart (Fig. 4D). Overall, the top 5 productive institutions in this field are the University of Coimbra, Texas Tech University, University of Kansas, Case Western Reserve University, and the Chinese Academy of Science. However, there seems to be no significant correlation between the number of publications and citation scores (Fig. 4D).
Analysis of the authors
This study further clarified the relationship between authors’ publications and the degree of collaboration. A total of 23,777 authors contributed to the 5,146 publications related to AD and mitophagy. The author’s collaboration is visually represented in the author’s collaborative network diagram (Fig. 5A), where each cluster (the same color) comprises at least 20 authors. The circle size correlates with the number of an author’s publications, and the lines linking circles indicate the intensity of collaboration among different authors. Furthermore, a line chart was generated to delve into the number of publications and the collaboration weight value among the top 20 authors. Our analysis results show that George Perry ranks first in the collaborative weight score, and P. Hemachandra Reddy ranks first in publishing output. Overall, these analysis graphs highlighted a pattern: higher collaboration intensity leads to increased publication outputs, particularly for authors such as P. Hemachandra Reddy, George Perry, Xiongwei Zhu, and Paula I Moreira (Fig. 5A, B). Additionally, this study presented information on the top 15 highly productive authors, including the number of publications, institutions, and countries (Table 2). Notably, 13 of these 15 authors were from the USA, accounting for 86.7%, aligning with the analysis results (Fig. 3). Further observation reveals that five authors were from Texas Tech University and three were from Case Western Reserve University, consistent with the earlier findings (Fig. 4). Additionally, the other two high-yielding authors are from Portugal and China (one each).
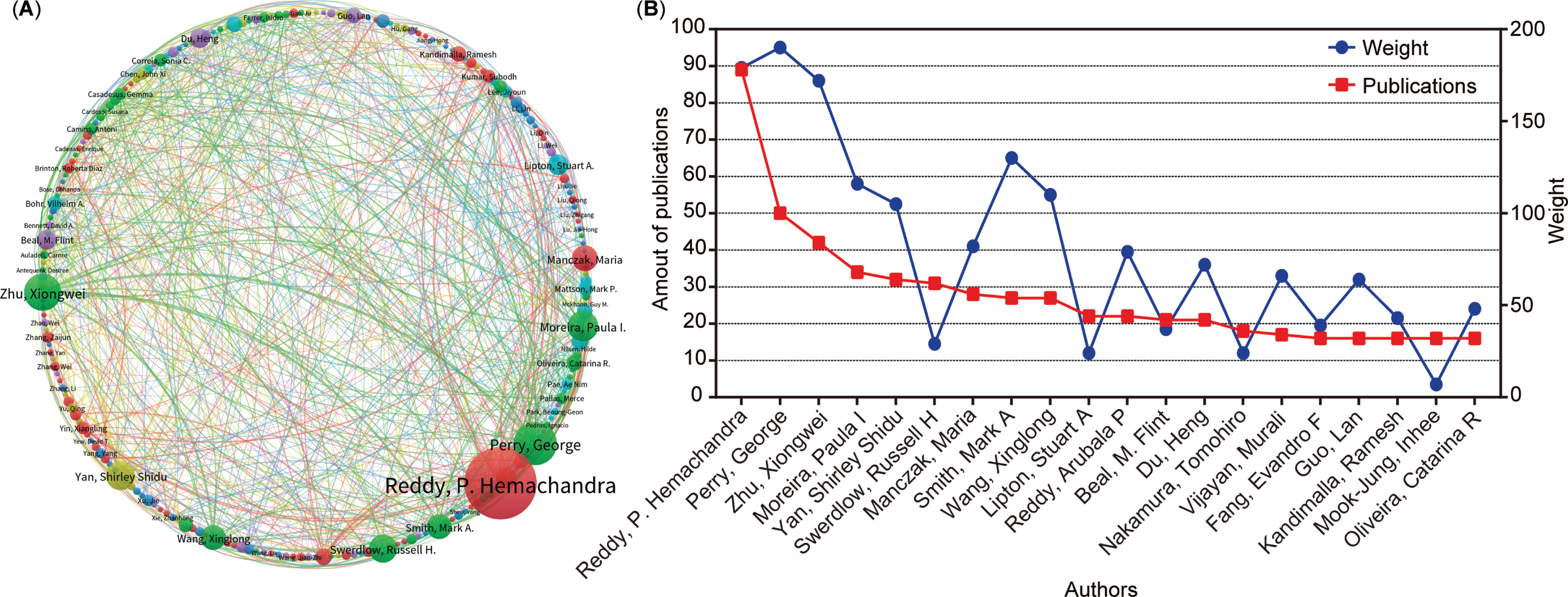
Collaboration analysis of authors. A) Map of collaboration within authors. The nodes represent authors, and the lines between the nodes represent collaboration relationships. The color represents clusters. B) Chart of the top 20 productive authors’ collaboration weight (blue round) and amount of publications (red square).
Top 15 most productive authors in AD and mitophagy
NP, Number of publications; USA, United States of America.
The collective contribution from various countries/regions, institutions, and authors in the realm of mitophagy in AD aligns with the trend observed: higher collaboration intensity contributes to increased publication outputs overall (Figs. 3–5).
Analysis of journals and the top 10 publications with the strongest citation bursts
A detailed analysis of the correlation between publications and journals was conducted, outlining essential information about the top 10 journals that extensively cover AD and mitophagy. Table 3 presents the key data from our assessment. Notably, from 2007 to 2022, the Journal of Alzheimer’s Disease stood out as the most prolific, publishing 253 articles. It was followed by the International Journal of Molecular Science, contributing 140 publications, and Molecular Neurobiology with 105 publications. This ranking aligns with the latest June 2023 release of the Journal Citation Reports (JCR) by Clarivate. Among these journals, 5 of the top 10 were classified in the JCR 2 region, while only one was in the JCR 4 region. Furthermore, the three journals classified in the JCR 1 region are the International Journal of Molecular Science, Free Radical Biology and Medicine, and Biochimica et Biophysica Acta-Molecular Basis of Disease (Biochim Biophys Acta).
Top 10 most prolific journals in the research field of AD and mitophagy
NP, Number of publications; NC, Number of co-citations per publication; IF, Impact factor; JCR, Journal citation report; UK, United Kingdom; USA, United States of America; Biochim Biophys Acta, Biochimica et Biophysica Acta-Molecular Basis of Disease.
Following this, the top 10 highly cited publications were presented for an in-depth analysis of article types (articles/reviews), titles, citations, and journal details within the mitophagy in AD research field (Table 4). The journals containing the top ten publications are mainly distributed in the JCR Q1 and Q2 regions. These 10 publications primarily belong to the JCR Q1 and Q2 regions. Among them, six are reviews and four are articles, each cited more than 300 times. The most heavily cited publication (with the highest strength of citations 74.43) emerged from Natural Neuroscience, exploring “The role of mitophagy on Aβ, tau pathology, and cognitive defects in AD” [35], This was followed by two articles (54.48 and 54.09) focusing on “Mitochondrial fission/fusion in AD” [27, 54], a review (52.48) on “Oxidative stress and mitochondrial dysfunction in AD” [55], and another review (50.20) on the “Mitochondrial cascade hypothesis in AD” [56].
Top 10 publications ranked by number of citations in the AD and mitophagy field (the statistics on the number of citations stopped in June 2023)
NC, Number of citations; ACY, Average citations per year; IF, Impact factor; JCR, Journal citation report. Proc Natl Acad Sci U S A, Proceedings of the National Academy of Sciences of the United States of America; Biochim Biophys Acta, Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease.
To summarize, our findings suggest a growing scholarly interest in mitochondria and mitophagy concerning AD. The aforementioned journals serve as valuable references for research publications in the AD and mitophagy domains.
Analysis of keyword clustering
The keywords listed after abstracts from 5,146 publications were extracted, generating a word cloud map (Fig. 6A). The most frequently mentioned keywords included AD, mitochondria, oxidative stress, neurodegeneration, mitochondrial dysfunction, and mitophagy.
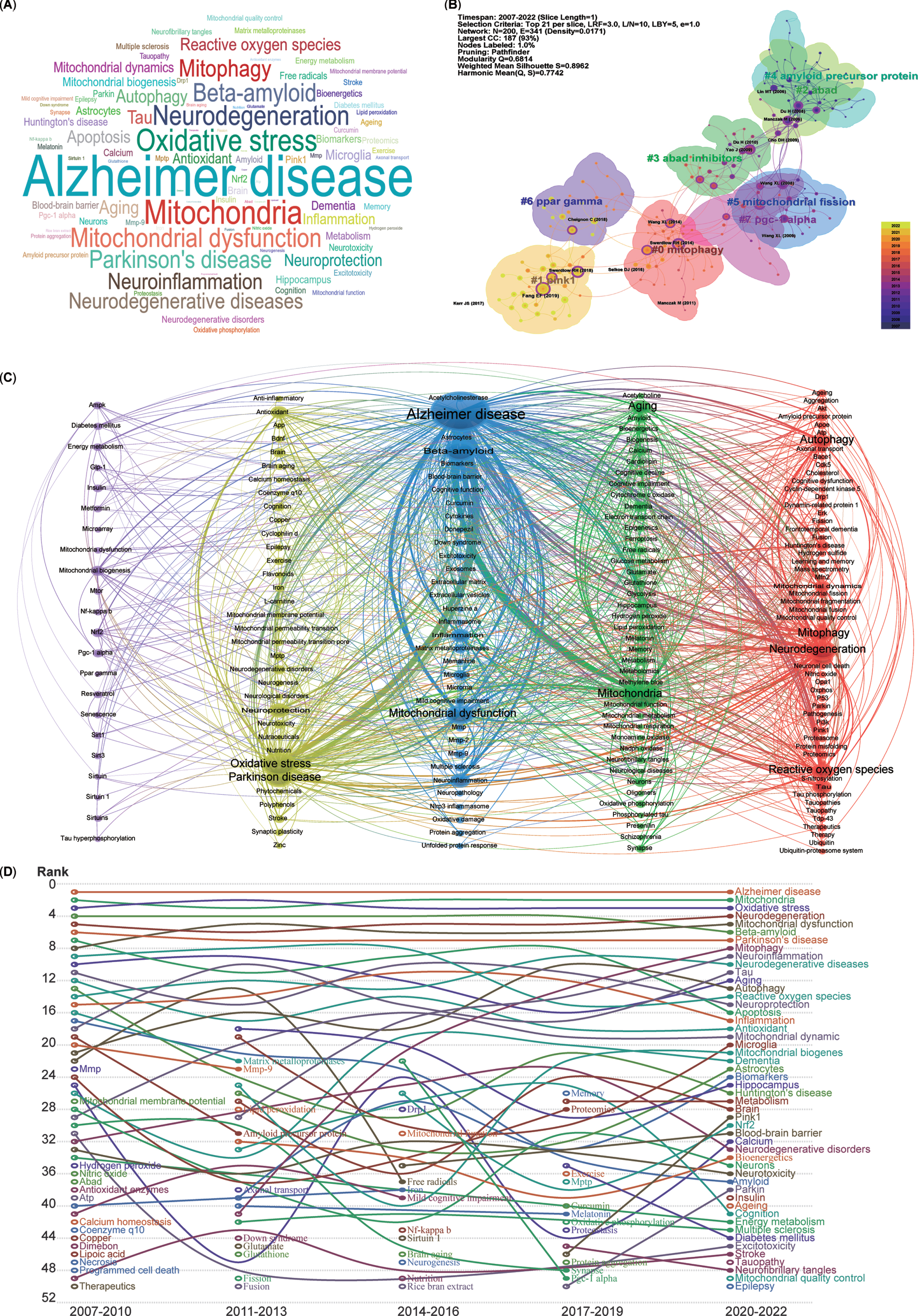
Cluster analysis and time trends of keywords. A) Word cloud map of the high co-occurrence of keywords and theme terms. B) Cluster map of co-citation analysis. Each circle represents an article, and the circle size represents the number of citations. The lines between the circles represent co-citation. Purple indicates that the article was cited relatively early, and yellow indicates that the citation time is relatively closer. C) A total of 191 keywords were selected for visual map construction, with each node consisting of a circle and a label. Nodes of different colors represent different clusters, with each color representing a different research direction. D) Time trend map shows the fluctuations in keywords from 2007 to 2022.
To further demonstrate the co-occurrence of the keywords in the 5,146 publications, the LLR algorithm clustering method was used to burst main keywords, and a network map of the co-occurrence of keywords clustered into eight research fields was generated using CiteSpace software (Fig. 6B). The cluster map indicated that the recently burst cluster (2022) can be categorized as mitophagy (Cluster 0). Similarly, the other co-citation publication clusters can also be categorized into the following research fields, which include mitophagy-related proteins (Cluster 1-PINK1, PTEN-induced kinase 1), mitochondrial proteins related Aβ production (Cluster 2-ABAD, Amyloid-binding alcohol dehydrogenase), inhibitors for mitochondrial Aβ production (Cluster 3-ABAD inhibitors), Aβ production (Cluster 4-Amyloid precursor protein, APP), mitochondrial dynamics (Cluster 5-Mitochondrial fission), critical regulators of mitochondrial biogenesis (Cluster 6-PPAR gamma, Peroxisome proliferator-activated receptor-gamma), and key factors for mitochondrial biogenesis and mitophagy (Cluster 7-PGC-1 alpha, PPAR-gamma co-activator-1 alpha).
Moreover, for a more precise depiction of keyword connections, a co-occurrence network of keywords was generated using VOSviewer, clustering them into five colors (Fig. 6C). The blue cluster, with the most connections and larger nodes, centers on the pathogenesis of AD (middle cluster), involving Aβ and mitochondrial dysfunction. The green cluster, adjacent to the blue one, relates to mitochondrial function in neurodegenerative diseases (NDDs), encompassing mitochondria, aging, dementia, and amyloid. The red cluster includes keywords such as neurodegeneration, autophagy, mitophagy, and ROS. The yellow cluster denotes oxidative stress and Parkinson’s disease (PD). Lastly, the purple cluster, with relatively smaller nodes, focuses on mitochondrial biogenesis, mitochondrial dysfunction, and key factors such as NRF2, PGC-1 alpha, AMPK, SIRT1, MTOR, and PPAR gamma. In essence, all extracted keywords from the 5,146 publications revolve closely around AD and mitophagy.
Analysis of time trends in keywords and research areas
To analyze the temporal trends of specific keywords, the frequency of all keywords was computed from a pool of 5,146 publications. Subsequently, the top 50 keywords with the highest occurrence were identified to create a time trend chart spanning from 2007 to 2022 (Fig. 6D). The top 5 keywords—AD, mitochondria, oxidative stress, neurodegeneration, and mitochondrial dysfunction—have remained relatively consistent over the past 16 years, signifying the primary focal points. Although mitophagy was initially proposed in 2005 and was in the early stages of AD research before 2010, it has exhibited a consistent upward trajectory year by year since 2011 (Fig. 6D). Furthermore, the pivotal regulators of mitophagy, namely PINK1 and Parkin, have shown an ascending trend in the last 6 years. Additionally, several emerging hotspots from the most recent 3 years’ topics have secured positions among the top 50 keywords, including insulin, aging, epilepsy, tauopathy, and mitochondrial quality control (MQC).
Overall, the research focus on mitochondria, mitophagy, and mitochondrial quality control has significantly solidified within the domain of AD studies.
Analysis of targets clustering and pathway enrichment
After scrutinizing the keywords, 213 targets encompassing genes and proteins were extracted from the abstracts of the aforementioned 5,146 publications. VOSviewer was used to visually represent these 213 targets as a visualization map organized into three clusters (Supplementary Figure 1). To present a more detailed and representative view of the targets, the top 30 targets within each cluster were selected and their relationships were compared based on occurrence and link weight (Fig. 7). The analysis revealed that these targets, named after the leading target in each cluster, could be categorized into three distinct clusters, namely APOE (Apolipoprotein E), MAPT (Microtubule-associated protein tau), and APP cluster. The APOE cluster encompassed factors associated with inflammation, such as TNF, IL6, NF-κB, along with insulin (INS) and AKT1, integral to insulin-pathway signaling (Fig. 7A). The composition of the MAPT cluster included elements related to mitophagy and mitochondrial function, incorporating OPA1, MFN1, MFN2, TOMM20, MAP1LC3A, SQSTM1, MTOR, DNM1L, VDAC1, FIS1, and the newly burst targets PINK1 and Parkin (Fig. 7B). Meanwhile, the APP cluster comprised APP, PSEN1, and other apoptotic targets such as CASP3, CASP9, BCL2, and BAX (Fig. 7C).
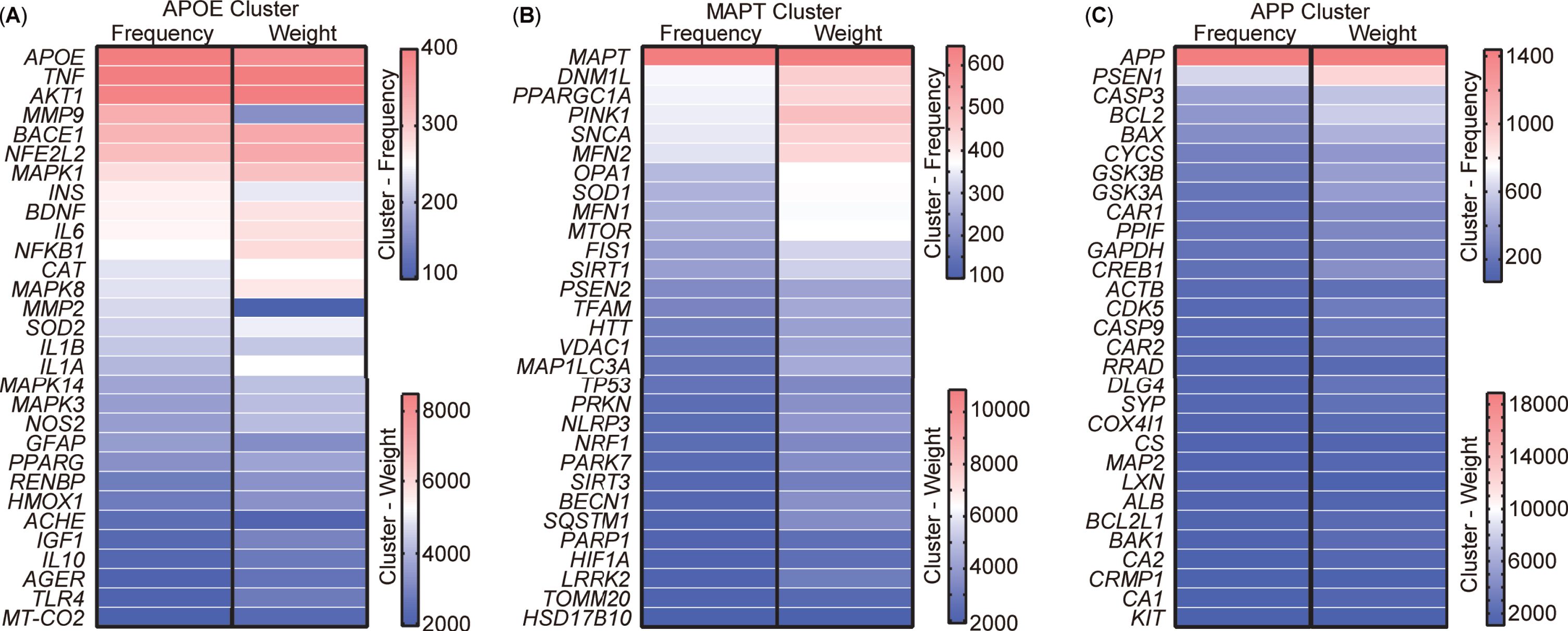
Cluster analysis of targets from the abstract of 5,146 publications. A) Top 30 most frequent targets, named the APOE cluster. B) Top 30 most frequent targets, named the MAPT cluster. C) Top 30 most frequent targets, named the APP cluster. Targets represent the gene or protein names appearing in the abstract.
For a more intuitive understanding of the functions associated with the clustered targets, enrichment analyses of KEGG and GO for these three clusters were conducted (Fig. 8). The KEGG analysis highlighted both similarities and differences among the clusters (Fig. 8A, C, E). Specifically, the APOE cluster included pathways associated with AD, lipid and atherosclerosis, and AGE-RAGE signaling pathway in diabetic complications. The MAPT cluster showed involvement in neurodegeneration, PD, and mitophagy pathways, while the APP cluster was implicated in neurodegeneration, AD, and amyotrophic lateral sclerosis pathways. Overall, the KEGG analysis indicated that the three clusters shared common pathways, notably AD. The GO enrichment analysis revealed three main modules: biological processes (BPs), cellular components (CCs), and molecular functions (MFs) (Fig. 8B, D, F). The APOE cluster was associated with biological processes such as cellular response to peptides, oxidative stress, lipopolysaccharide, nutrient levels, and extracellular stimulus. The MAPT cluster was linked to the regulation of autophagy, mitochondrial disassembly, and autophagy of mitochondrion. Conversely, the APP cluster focused on axon development, regulation of membrane potential, generation of precursor metabolites and energy, axonogenesis, neuron apoptotic process and death, and mitochondrial transport. Notably, the APOE and MAPT clusters shared BP related to mitochondria. In terms of CC, the APOE cluster involved the membrane raft, endoplasmic reticulum, vesicle lumen, and presynapse, whereas the MAPT cluster emphasized components such as the mitochondrial outer membrane, mitochondrial matrix, intrinsic component of mitochondrial membrane, and mitochondrial outer membrane, and presynapse. By contrast, the APP cluster predominantly concentrated on the presynapse, soma, distal axon, exocytic, and synaptic vesicle in neuron. Notably, all three clusters had associations with the presynapse in the CC. Regarding MFs, the APOE cluster centered on signaling receptor activity, Aβ binding, and cytokine activity, while the MAPT cluster focused on ubiquitin or ubiquitin-like protein ligase binding, and tubulin binding. However, the APP cluster was enriched in tau protein binding and protein serine/threonine/tyrosine kinase activity. Additionally, all three clusters exhibited participation in DNA-binding transcription factor binding. Both the KEGG and GO enrichment analyses highlighted the close association of our target enrichments with AD, as well as with mitochondria and mitophagy pathways.
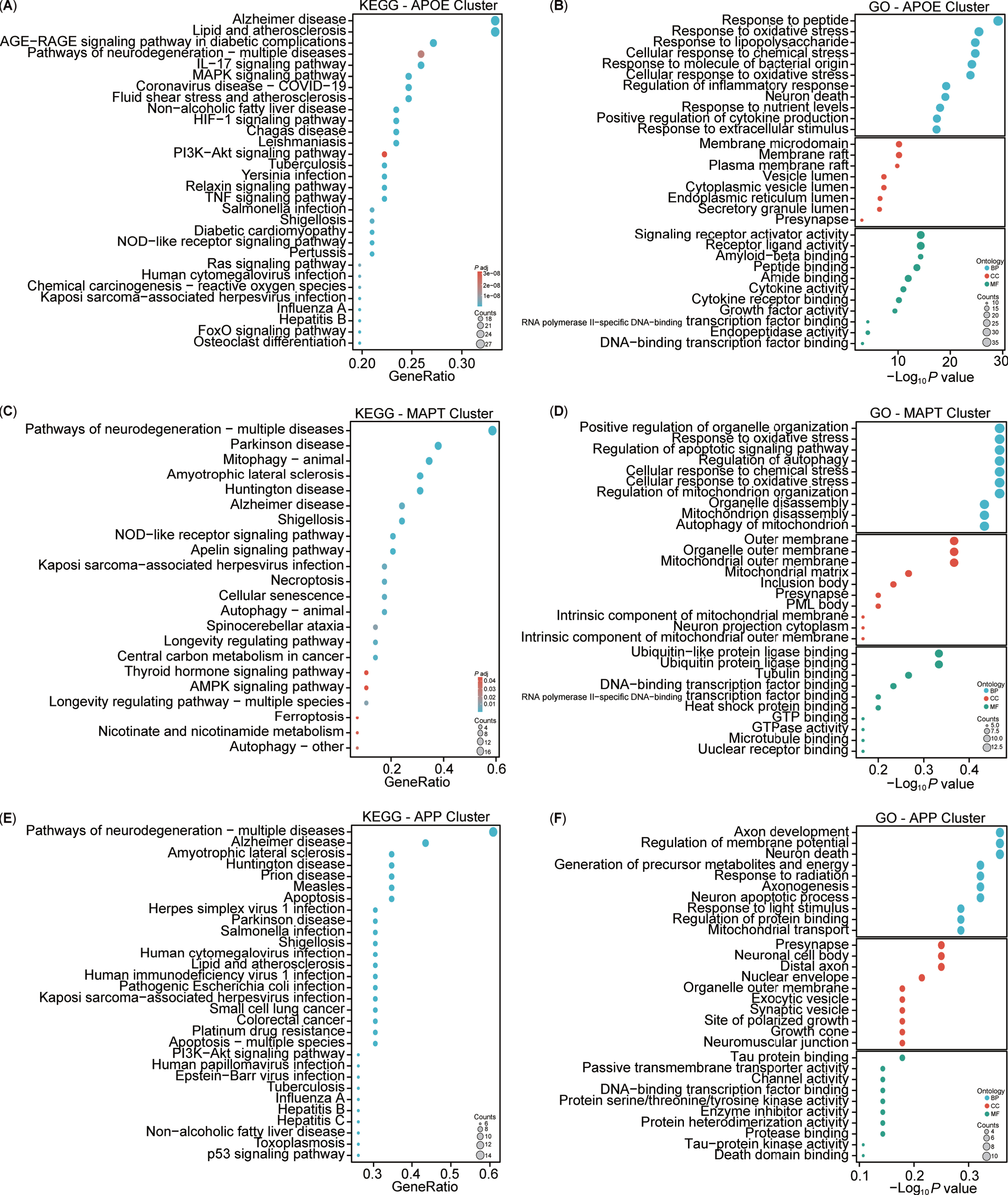
Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analysis of AD and mitophagy targets. A-F) KEGG and GO analyses of the three clusters (APOE, MAPT, and APP).
After conducting a protein-protein interaction (PPI) network analysis with 30 targets selected from the top 10 targets in each cluster, STRING and GeneMANIA were used to explore connections between these targets (Fig. 9A, B). The resultant networks revealed intricate yet comparable interactions associated with Aβ synthesis (APP, PSEN1, and BACE1), regulation of mitochondrial function (PPARGC1A and MAPK1), progression of mitophagy (OPA1, DNM1L, MFN1, MFN2, and PINK1), and energy metabolism (GSK3B, MTOR, and AKT1). Identifying consensus and potential targets in AD and mitophagy research could significantly benefit forthcoming drug interventions and potential clinical trials in future.
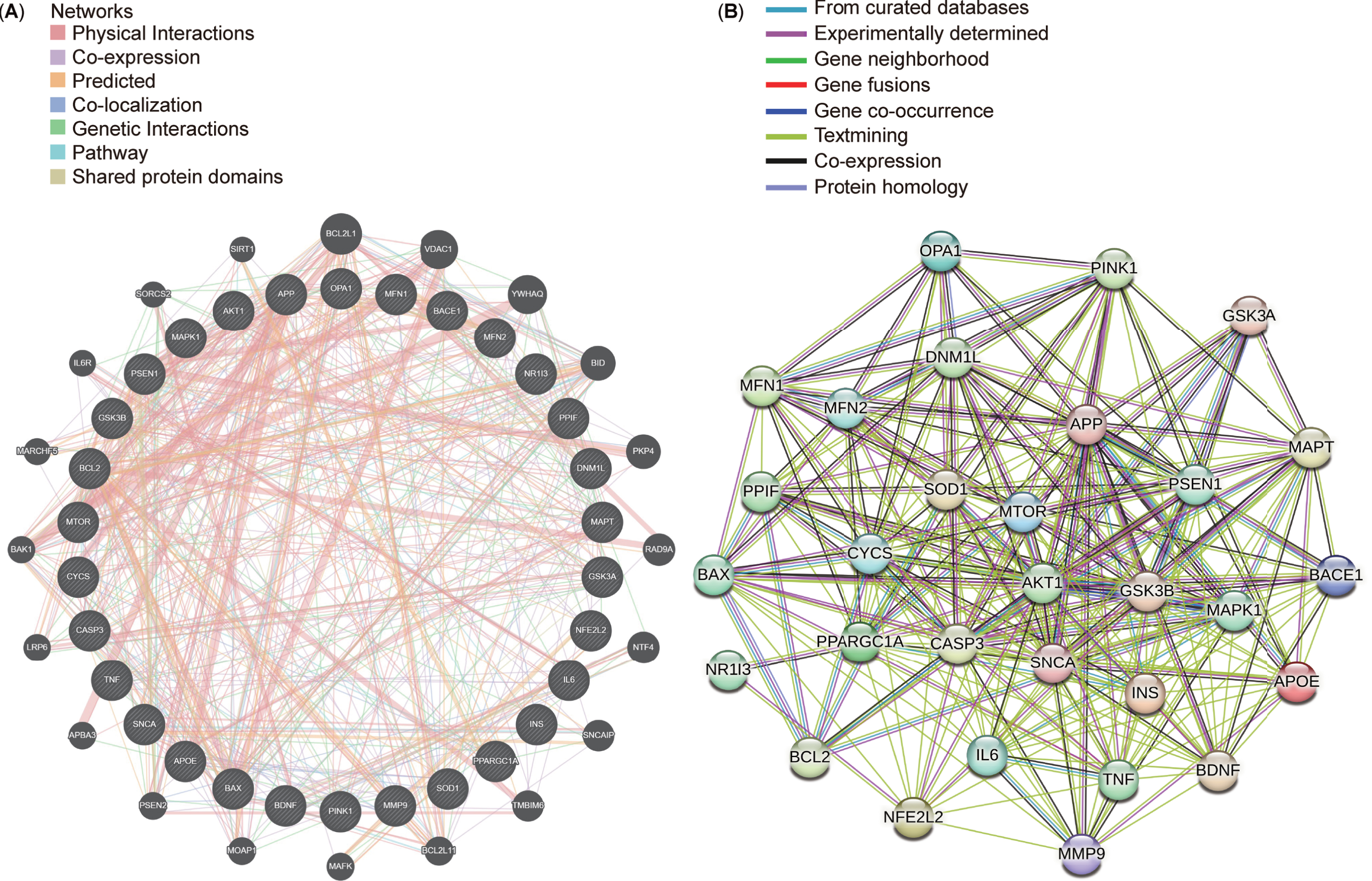
Top 30 targets in the protein-protein interaction (PPI) networks. A) Top 30 targets in the PPI network generated using GeneMANIA. B) Top 30 targets in the PPI network displayed using STRING.
Analysis of diseases with drug trials and targets involving mitophagy
After clustering analysis of the targets, we identified 200 diseases mentioned or recorded in the abstracts of the publications (each disease occurring at least 35 times) and conducted clustering and frequency analysis by VOSviewer. It is obvious that “AD” ranks first among the top ten diseases in terms of frequency, followed by “mitochondrial dysfunction” and “other NDDs” (Table 5 and Supplementary Table 2).
Top 10 most frequently occurring diseases from four clusters
The previous results demonstrated that the clustered keywords include diseases, targets, and drugs, indicating that the latter two may be involved in treating of the clustered diseases (Figs. 6 and 7). Consequently, we identified gaps in targets, drugs, and diseases, and attempted to fill these gaps. We extracted all drugs or molecules (total of 28) studied from 5,146 publications, such as donepezil, memantine, huperzine A, and NAD (+) (Supplementary Table 4). Further, we summarized the top 10 representative drugs or molecules, the mechanism of action of mitophagy, representative clinical trials with stages and completion status (Table 6). Our results showed that above ten drugs applied in various stages of AD-related clinical trials (e.g., curcumin, NCT00164749, NCT00099710; insulin, NCT05081219, NCT01595646, NCT00438568; melatonin, NCT04522960, NCT00000171, NCT00940589). Most of these drugs such as curcumin, insulin, melatonin, iron, glutamate, and donepezil regulate mitochondrial function and mitophagy as potential treatments (Table 6). Meanwhile, increasing publications reported different targets in these mechanisms (e.g., MFN2, PINK1, Parkin, OPTN, TOMM20), and these targets were consistent with our clustered results (Figs. 6–9), which indicates that mitophagy is an important target for potential drug in AD. Besides, we also summarized our drug-target link data from PubChem of the National Library of Medicine (Supplementary Figure 2). The results show that all drugs and targets of PubChem data are also consistent with bibliometric analysis results (Figs. 7–9, and Table 6).
Top 10 drugs or molecules extracted from publications with representative clinical trial information and mechanisms of action in AD
AD, Alzheimer’s disease; C, Completion; R, Recruiting; U, Unknown; AN, Active, not recruiting; W, Withdrawn or No funding; NA, Not Applicable.
In this part, we attempted to incorporate information such as targets and target-drug-clinical trial-disease into the scope of bibliometric analysis and summarized their relationship in mitophagy and AD research by this method to provide AD researchers and readers with an overview of mitophagy in AD candidate drugs.
DISCUSSION
Bibliometric analysis
This study focused on analyzing the research trend of mitophagy in AD during 2007-2022, presenting significant findings through bibliometric analysis.
The volume of scientific publications serves as a pivotal indicator of a field’s trajectory [57]. Our analysis of the global publications curve demonstrates a consistent and rapid increase in published works from 2007 to 2022. This upsurge indicates the promising trajectory of mitophagy research within the realm of AD for the foreseeable future.
The total volume of literature from a country stands as a significant indicator of its scholarly output. This study identifies the USA, China, and Europe as the primary contributors to AD and mitophagy research. Among the top 10 most productive countries, the USA exhibits the highest citation score, reflecting its substantial contributions and leading position of AD and mitophagy research. Furthermore, this study revealed that the strength of collaboration, number of publications, and citation scores maintained the general obedience to more vital collaboration, higher outputs, and more citations, respectively, in countries and institutions worldwide. Nevertheless, notable variations exist, particularly in Colombia. A highly cited collaborative study in 2008 reported the direct relationship between increased abundance of Aβ1–42 and noncoding RNAs in AD [58]. Another significant collaboration in 2014 highlighted that carriers of the PS1-E208A mutation exhibited elevated abnormal mitochondria in familial AD (FAD) and impaired ER/mitochondria tethering in neuronal cell lines [59]. Additionally, the NIH is the leading institute supporting AD research. From the perspective of funding agencies, the USA has indeed laid the groundwork for AD and mitophagy research.
However, while collaborative efforts primarily involve most local institutions in the USA and Europe, there is a lack of substantial collaborative connections with institutions in other countries or regions. This disparity suggests that limited academic collaboration might, to some extent, impede the progress of AD and mitophagy research. The findings obtained in this study strongly advocate for increased collaboration among institutions from diverse geographic locations to expedite the advancement of AD and mitophagy research.
Sorensen AA evaluated authors’ influence based on total publications [41]. Our study primarily emphasizes total publications and identifies the top three most prolific authors: P. Hemachandra Reddy, George Perry, and Xiongwei Zhu. The contributions of the last two authors align with Sorensen AA’s analysis. The absence of Reddy PH in Sorensen AA’s research ranking might be attributed to differences in data retrieval timelines. Reddy PH’s initial publication, “Mitochondrial dysfunction and oxidative damage in AD” [60], marked the beginning of his focused contributions to AD and mitophagy research, significantly affecting publication outputs and the advancement of this field.
Certain bibliometric analysts posit that high-impact journals signify leadership in a particular field [41]. In our investigation, publications featuring substantial co-citations were predominantly found in high-impact journals such as Nature Neuroscience, Trends in Neuroscience, and Proc Natl Acad Sci U S A. Interestingly, our analysis also highlighted the Journal of Alzheimer’s Disease as the most prolific journal. Additionally, among the top 10 publications with the strongest citation bursts, researchers not only contributed to specialized journals such as the Journal of Alzheimer’s Disease but also engaged with others such as the Journal of Neuroscience, and Biochim Biophys Acta. These findings suggest that while high-impact journals are crucial, classic journals within the professional sphere also hold considerable merit for publication.
Hotspot trends and research areas
In general, publications exhibiting the strongest citation bursts serve as significant milestones in the field. Our bibliometric analysis delineated a temporal trend of hotspots in AD, notably marking three citation bursts during 2007–2022.
The initial citation burst transpired during 2008-2009. In 2008, Wang X’s research introduced the concept that APP might differentially regulate mitochondrial dynamics through mitochondrial fission or fusion proteins in AD [54]. Subsequently, in 2009, their findings elucidated that abnormal mitochondrial dynamics played a pivotal role in Aβ-induced mitochondrial and synaptic dysfunction in AD. They suggested regulating mitochondrial fusion and fission proteins (DRP1 and OPA1) as a potential therapeutic strategy for AD [27].
The second citation burst occurred in 2014 when Swerdlow RH summarized the features of the mitochondrial cascade hypothesis, originally proposed by himself [61], along with discussions on AD biomarkers, diagnostics, and clinical trials in 2014 [56], announced that the mechanism of regulation of mitochondrial function and cellular biogenesis could assist in delaying aging and preventing the development and progression of AD. Simultaneously, Wang X’s review systematically emphasized that the interaction between mitochondrial dysfunction and oxidative stress might lead to the initiation and amplification of ROS, emphasizing their significant role in the pathogenesis of AD [55].
The third citation burst emerged in 2019. Fang EF’s team published an article discussing mitophagy’s role in AD progression, examining the effect of accumulated damaged mitochondria and impaired mitophagy on neurons and microglia in AD [35]. Their results suggested that maintaining mitochondrial mass in neurons or at the organismal level could potentially treat AD. This publication, being part of the strongest citation burst, played a pivotal role in driving deeper research on AD and mitophagy.
Keyword analysis, a widely used bibliometric tool, revealed emerging hotspots and themes in our gathered publications by keyword co-citation analysis. PINK1 was co-cited in studies published from 2017 to 2022 and was closely correlated with autophagy, mitophagy, and MQC from 2017 to 2022. Additionally, our time trend analysis unveiled intriguing patterns. Notably, AD, mitochondria, and mitochondrial dysfunction held central positions over the past 16 years. Moreover, Aβ and APP were prominent during the early stages (2007–2010), while mitophagy gained attention starting in 2011. Furthermore, PINK1 and Parkin emerged as key keywords in the last 6 years, accompanied by increasing interest in MQC, insulin, aging, tauopathy, and epilepsy in the past 3 years. In recent years, PINK1, a mitochondrial serine/threonine kinase, has been reported to play a major role in Parkin-dependent mitophagy [62]. Activating PINK1/Parkin dependent mitophagy can promote mitophagy, reduce their insolubility for Aβ, and synaptic dysfunction in AD mice [63, 64]. The MQC system is developed by mitochondria to respond to changes in NDDs, and its regulation partially relies on mitophagy [65]. Growing evidence suggests that AD is also called type III diabetes in recent study, because it has many similar pathological manifestations with diabetes [66]. The addition of brain insulin can prevent Aβ-induced dysfunction [67]. Overall, besides the prevailing theories of Aβ cascade and abnormal tau protein aggregation, researchers are exploring alternative mechanisms, notably mitophagy. Particularly noteworthy is the growing interest in PINK1 and Parkin, linked to the newly summarized theory of mitophagy, indicating their gradual emergence as hotspots in the field of mitophagy in AD.
Remarkably, our study analyzed targets in 5,146 publications. The results showed that the keywords APOE, MAPT, and APP ranked first in the frequency and weight values of the three clusters, respectively, which has also been supported in other research reports. APOE, as a sporadic risk factor for AD, especially APOE4, may cause mitochondrial dysfunction in AD patients, ultimately leading to mitophagy defects [68]. Fang EF also observed impaired mitophagy, characterized by decreased phosphorylation of mitochondrial initiators TBK1 and ULK1, as well as decreased expression of PINK1, LC3-II, and BCL1 in AD patients and animal AD models [35]. Several studies have reported that pathological phosphorylation of MAPT/tau can induce mitochondrial damage and is closely related to the mechanism of mitophagy [69, 70]. Previous studies have reported overexpression of the AD-associated mutant Aβ-APP leads to abnormal mitophagy [71]. Mitophagy enhancer, urolithin A, can exert protective effects against mutant APP and Aβ-induced mitochondrial and synaptic toxicity [72]. Other results indicate an increasing focus on mitophagy-related targets, such as OPA1, MFN1/2, and TOMM20, within the AD research field. Additionally, insulin-related metabolism targets, such as APOE, INS, and AKT1, may guide future research in AD.
Furthermore, the KEGG and GO enrichment results revealed significant clusters enriched in the mitophagy signaling pathway, particularly the MAPT cluster involved in mitochondrial regulation, disassembly, mitochondrial membrane and matrix components. Notably, Sorensen AA published a study ranking diseases based on the PubMed database [41]. This study identified the top 3 clustered diseases as cerebral vascular accident, AD, and dementia. In our study, the top 3 clustered diseases were AD, mitochondrial dysfunction, and NDDs. While our research shares some “disease” terminology with Sorensen AA’s work, the appearance of the term “mitochondrial dysfunction” suggests an emerging role of mitochondria in diseases beyond their cellular organelle function, which is consistent with literature reports that mitochondrial dysfunction, a hallmark of early pathophysiology [69], is closely related to the occurrence and development of AD [73].
In summary, our bibliometric analysis encompasses target and disease clustering analysis. It aims to broaden understanding regarding potential mechanisms and targets among AD researchers and readers.
Mitophagy, a penetrating point for AD candidate drug research
Despite numerous studies indicating that Aβ cascade, tau protein abnormal phosphorylation, neuroinflammation, APOE allele mutations, oxidative stress, cholinergic dysfunction, the hypothesis of neurotransmitter and metal homeostasis imbalance, and mitochondrial dysfunction are the pathological mechanisms of AD, more clinical trials of drugs focus more on Aβ, especially monoclonal antibodies [74].
In recent years, the emergence of numerous monoclonal antibodies (mAb) has expanded the treatment options for patients with AD at different disease stages and with diverse symptoms (Table 7). As an example, aducanumab and lecanemab, newly approved anti-amyloid drugs by the FDA, eliminate Aβ from the brain, reducing cognitive and functional impairment, despite associated side effects such as amyloid-related imaging abnormalities with edema or effusions, headaches, falls, seizures, nausea, dizziness, confusion, walking problems, and vision changes. While clinical trials of these drugs have focused on early-stage AD, FDA approval limits the use of the drugs to stages 3P/Fa and 4P/Fa. Insufficient data on their safety and efficacy in these stages hinder their current use.
Anti-Aβ monoclonal antibody (mAb) drugs from literature and clinical trial information and mechanisms of action in different AD
AD, Alzheimer’s disease; C, Completion; R, Recruiting; AN, Active, not recruiting; NA, Not Applicable; EAD, Early Alzheimer’s disease; NY, not yet recruiting; ARIA, amyloid-related imaging abnormalities; ARIA-E, amyloid-related imaging abnormalities with edema or effusions.
The literature documents that asymptomatic patients or those with preclinical AD exhibit no cognitive symptoms but might display underlying biological changes in the brain [10, 76]. Consequently, early detection of cerebrospinal fluid (CSF) Aβ and prompt diagnosis can facilitate timely medical intervention for the progression of asymptomatic patients or those with preclinical AD. Current research, follow-ups, and clinical trials for such patients predominantly emphasize anti-amyloid protein strategies. Lecanemab is being tested in asymptomatic patients through the AHEAD 3–45 study to evaluate its effectiveness in slowing down biomarkers and cognitive abilities [77]. Other drugs such as solanezumab and gantenerumab are undergoing further evaluation for efficacy (Table 7), despite solanezumab’s unsatisfactory results (NCT02008357) in preclinical AD [8] and gantenerumab’s unsuccessful phase III trials (NCT04339413 and NCT04374253) in early AD [9]. Additionally, the A4 study continues to assess solanezumab’s effect in a clinical trial (NCT02008357) aimed at rescuing cognitive function decline in older asymptomatic individuals.
In addition to the aforementioned anti-amyloid protein antibodies, non-selective oral administration of β-secretase inhibitors has been conducted over the past few decades to reduce Aβ levels. Notably, JNJ-54861911 has completed trials, demonstrating a 67–90% reduction in mean CSF Aβ1 - 40 in both Caucasian and Japanese patients with early-stage AD (NCT01978548 and NCT02360657) [78]. However, atabecestat’s trials (NCT02569398) were terminated due to treatment-associated dose-related cognitive worsening and neuropsychiatric adverse events in asymptomatic patients with AD [79]. In summary, the therapeutic effects of these monoclonal antibodies primarily focus on anti-amyloid protein.
Although all these drugs are trying to treat AD, there is still no drug that can fundamentally prevent and reverse the condition of AD patients, which also prompts basic and clinical researchers to explore new mechanisms of AD in order to find new strategies for prevention and treatment of AD.
Ever since Swerdlow RH proposed the “mitochondrial cascade hypothesis” in 2004 [61], increased attention has been directed towards fluctuations in mitochondrial physiology in AD. Abnormal mitochondria affect various aspects, including biogenesis, mitochondrial dynamics [80–82], axonal transport [83–85], ER-mitochondrial association [86–89], proteostasis [90–96], and mitophagy [34, 97–99]. The accumulation of damaged and dysfunctional mitochondria constitutes an early symptom of AD [100], worsening the effects of Aβ and tau pathologies, resulting in synaptic dysfunction, cognitive impairment, and memory loss [101, 102]. Neuronal mitochondrial dysfunction and impaired metabolism [103], along with glial mitochondrial dysfunctions and mtDNA abnormalities [104–106], contribute similarly to AD progression. Targeting mitochondria, be it in neurons or glial cells, for instance, through mitophagy enhancement, may be critical for effective retention and cognitive function [107, 108]. Mitochondrial dysfunction, mitophagy damage, and AD-related proteins such as Aβ, APP-derived fragments, tau, and APOE are contributors to mitophagy failure in patients with AD [109]. Mitophagy impairments play a role in AD pathology observed in animal models and human brain tissues [110]. Furthermore, Fang EF’s research highlighted the pivotal role of impaired defective mitochondrial removal in AD pathogenesis. Increased mitophagy has shown promise in reducing Aβ plaques and tau pathology, thereby preventing cognitive impairment and suggesting a potential therapeutic intervention for AD [35, 111].
In traditional bibliometric analysis, certain studies have indeed focused on analyzing trends and ranking drugs related to AD publications, along with the frequency of “treatment” keywords [43, 112]. However, these studies have given limited attention to understanding drug mechanisms, targets, and the ongoing clinical trial progress. This study specifically concentrates on identifying potential AD treatment drugs and screened 28 candidate drugs/molecules that have been featured in 5,146 publications. Additionally, it highlights the top 10 representative drugs/molecules, namely curcumin, insulin, melatonin, iron, glutamate, glutathione, resveratrol, nitric oxide, metformin, and donepezil. For instance, previous research highlighted curcumin’s inhibition of Aβ-induced abnormalities in mitochondrial metabolism and neuronal damage caused by oxidative stress [113]. Similarly, insulin has been reported to enhance mitochondrial function and regulate impaired insulin-signaling pathways [114–117]. Resveratrol, functioning as a mitophagy inducer, has shown positive effects on AD models by promoting mitophagy and reducing Aβ-induced mitochondrial damage, thereby improving cognitive function [118]. Moreover, metformin, recognized as an insulin sensitizer, has demonstrated its potential to ameliorate mitochondrial function to treat AD [119, 120]. Additionally, melatonin, iron, glutamate, glutathione, resveratrol, and nitric oxide have displayed promising functions in AD through various mechanisms, regulating mitophagy and mitochondrial quality control, thus reducing damaged mitochondria [118, 121–124]. In summary, there is a growing consensus on targeting mitophagy as a potential treatment for AD [125]. At the same time, the most urgent issue currently faced is how the discoveries and progress made in basic research on targeting mitophagy can be realized in clinical trials. Artificial intelligence (AI) has become a cost-effective and fast method for identifying and screening drugs or compounds, as well as discovering new targets [126, 127]. Therefore, future research can attempt to focus on candidate drugs for AD treatment, infer the best potential drugs and targets by AI models, and then conduct clinical trials.
Furthermore, detailed insights into the clinical trials of these drugs were presented. For instance, donepezil has completed phase IV clinical trials (NCT03073876, NCT02787746, NCT02097056), curcumin has completed phase II clinical trials for AD treatment (NCT00164749, NCT00099710), and insulin has shown similar clinical progress (NCT01595646, NCT00438568). Most other drugs are primarily in phase I or II clinical trials. However, whether the aforementioned drugs in these clinical trials are involved in stimulating mitophagy mechanisms in AD remains unclear. Therefore, future clinical drug research should aim to evaluate changes in mitophagy (proteins or genes) in patients with AD simultaneously.
In summary, although AD has a complex pathogenesis, researchers have made significant progress in the mechanism study and drug research of mitophagy in AD over the past 16 years. Therefore, we hope that this study can promote researchers to have a deeper understanding of the molecular mechanisms, targets, and drugs of mitophagy in AD, providing a penetrating point for its mechanism and anti-AD drug research.
Conclusion
To the best of our knowledge, this report represents the first examination of mitophagy in AD research trends using a bibliometric method. Our analysis reveals that mitophagy constitutes a novel molecular mechanism in AD research and could potentially guide AD candidate drug research.
Additionally, this study introduces targets, drug associations, clinical trials, and disease links into the domain of bibliometric analysis for the first time. This innovative approach might inspire subsequent researchers engaged in bibliometric analysis. Furthermore, our analysis aims to broaden the understanding of potential mechanisms and targets among AD researchers and readers, offering a theoretical overview for the development of new clinical trial drugs for AD treatment.
Nevertheless, the present study has several limitations. First, articles and reviews were solely analyzed, potentially missing out on other publication types that could reveal more significant research trends. Second, this study only objectively presented targets and drugs from 5,146 publications, as well as the latest monoclonal antibody drugs from literature or pharmaceutical websites, without leveraging AI models to predict the optimal candidate targets and drugs related to mitophagy in AD. Future research, based on existing studies, could focus on candidate drugs for AD treatment, employing AI models to infer the most promising drugs and targets.
In conclusion, the findings of the present study will provide researchers and readers with a comprehensive understanding of the evolving trend of mitophagy in AD. It also strives to offer new insights into basic research on mechanisms and clinical trials on drugs in AD.
CRediT AUTHOR STATEMENT
Hongqi Wang (Data curation; Formal analysis; Investigation; Methodology; Project administration; Software; Validation; Visualization; Writing – original draft); Xiaodong Yan (Funding acquisition; Methodology; Software; Supervision); Yiming Zhang (Validation; Visualization; Writing – original draft); Peifu Wang (Writing – review & editing; Assist in the major revision of the article); Jilai Li (Supervision; Validation; Writing – review & editing; Assist in the major revision of the article); Xia Zhang (Data curation; Formal analysis; Investigation; Methodology; Project administration; Supervision; Validation; Visualization; Writing – original draft; Writing – review & editing).
Footnotes
ACKNOWLEDGMENTS
We are grateful to the WOS, PubChem, Clinical Trial, GenDoma, STRING, and GeneMANIA that made the data available, and all data obtained from these datasets kept to the rules for their usage and publication.
FUNDING
This work was supported by the Natural Science Foundation of Shanxi Province (202103021223011).
CONFLICT OF INTEREST
The authors have no conflict of interest to report.
DATA AVAILABILITY
All data can be obtained from WOS (https://www.webofscience.com/wos), VOSviewer (https://www.vosviewer.com/), Pajek (http://vlado.fmf.uni-lj.si/pub/networks/pajek/), CiteSpace (https://citespace.podia.com), BioBERT (https://github.com/naver/biobert-pretrained, https://github.com/dmis-lab/biobert), PubChem (https://pubchem.ncbi.nlm.nih.gov), Clinical Trial (https://clinicaltrials.gov/), Gen-Doma (https://ai.citexs.com), STRING (https://string-db.org), and GeneMANIA (![]() ).
).
