Abstract
Thrips are important pests of avocados causing damage to the fruits, making them unsuitable for exportation. Many of them are quarantine species and their identification is necessary. Despite their importance, not many studies describe thrips diversity on avocado, especially in Colombia, partly because identification of some of the species is difficult using classical taxonomic keys. In this study, 93 adult thrips specimens were collected on avocado and adventitious dandelion plants from plantations in the department of Antioquia, Colombia. Morphological characters were used to classify 62 of them and all of them were analyzed by PCR amplification and sequencing of cytochrome c oxidase subunit I (COI) gene and internal transcribed spacer (ITS) region. The species identified by morphological features were Frankliniella gardeniae Moulton, F. gossypiana Hood, F. panamensis Hood, Liothrips perseae Watson, Thrips palmi Karny, T. simplex (Morison), T. trehernei Priesner, Microcephalothrips abdominalis (Crawford DL), and Scirtothrips hansoni Mound & Hoddle (Tubulifera), being F. panamensis, M. abdominalis, and S. hansoni reported for the first time on avocado in Colombia. Thrips simplex and T. trehernei were also found on adventitious dandelion plants, but not on avocado. Phylogenetic analysis of COI and ITS sequences showed that most species, except for F. gardeniae and F. gossypiana were mainly identified with the COI gene. Furthermore, FST values obtained between them suggest no genetic differentiation at COI level. The results obtained here contribute to the knowledge of the diversity of thrips species found on the avocado crop in Colombia and are meaningful for pest management.
Introduction
Colombia registered 39.172 hectares of avocado (Persea americana Mill.) harvested in 2017, ranking third worldwide in planted area and fourth in total production with 314.275 Tons (FAO, 2018). In recent years, avocado has been considered one of the most important crops for exportation in this country. The department of Antioquia stands out in Colombian avocado production with 11.321 hectares and an increase of 126 % in planted area in the last 5 years (Ministerio de Agricultura, 2018). However, some factors limit competitiveness such as pests which require constant management, increasing production costs (Bravo-Pérez et al. 2018).
Thrips (Thysanoptera) represent serious invader pests of avocado because they produce scars on the fruit, making it unsuitable for export markets (Cambero-Campos et al. 2011). They are difficult pests to control due to their small size, cosmopolitan distribution, high reproductive rate, polyphagous habits, invasive behavior (both in greenhouses and in the field), and a limited number of natural enemies (Brunner et al. 2002; Monje et al. 2012; Mouden et al. 2017). For these reasons, many thrips species have been considered quarantine species for many countries. The exaggerated use of pesticides has led to resistance, residue problems on marketable crops, toxicity towards beneficial non-target organisms, and contamination of the environment (Mouden et al. 2017).
One important problem is their taxonomical identification at the species level due to their imperceptible morphological differentiation, intraspecific polymorphisms, and sexual dimorphisms (Brunner et al. 2004; Tyagi et al. 2008; Iftikhar et al. 2016). The requirement of an expert in thrips morphology is essential for taxa identification. However, an alternative to morphological identification is the use of molecular techniques, which can be used together with morphological variation to identify species, as well as to improve the taxonomical status and reconstruct phylogenies (Brunner et al. 2004; Buckman et al. 2013; Iftikhar et al. 2016). Several molecular markers, including nuclear and mitochondrial loci, have been used for species identification and phylogenetic analysis (Asokan et al. 2007; Buckman et al. 2013; Iftikhar et al. 2016; Tyagi et al. 2017).
Despite their importance, there are not many studies reporting the thrips species affecting avocado and often they are recognized as belonging to a species complex (Echeverri-Florez et al. 2004; Cambero-Campos et al. 2011). In Colombia, a morphological characterization made from adults collected from avocado leaves and flowers at Caldas and Risaralda departments (West-Central Colombia) allowed the identification of the phytophagous species: Selenothrips rubrocinctus Giard, and Frankliniella gardeniae Moulton, and individuals of the genera: Leptothrips and Karnyothrips have predatory habits (Echeverri-Florez et al. 2004). A recent study based on the molecular identification of the genera Arorathrips, Caliothrips Frankliniella, Leptothrips, Neohydatothrips, and Scirtothrips, was conducted in Mexico using 44 specimens collected from avocado and sequencing of the mitochondrial COI (Cytochrome Oxidase I unit) gene (Bravo-Pérez et al. 2018).
Given the little knowledge on thrips species in Colombia and the Antioquia department, the aim of this study was to identify the species of thrips present on avocado trees from commercial plantations in Antioquia. In addition, we tested if the use of molecular techniques could aid in the identification of the species present in this crop easier. For this, 93 specimens were sampled, taxonomically identified by morphological characters, and analyzed by PCR amplification and sequencing of the Cytochrome Oxidase I (COI) gene and Internal transcribed spacer (ITS) region of each specimen. Results obtained here are useful since they widen the information on the trips species affecting avocado and could be used to improve an integrated pest management strategy based on COI sequencing as a simple and accurate method that facilitates the identification of thrips species (Rebijith et al. 2014).
Materials and methods
Specimen's collections
Thrips specimens (adults) were randomly collected from several commercial avocado trees during 2017 in 7 avocado orchards (approx. 1-2 ha area/orchard), in the department of Antioquia in Northeast Colombia (Supplementary Table 1). For molecular analysis, 93 adult individuals were collected from avocado, P. americana Mill. (Lauraceae) and dandelion, Taraxacum officinale (L.) Weber ex F.H.Wigg. (Asteraceae) plant inflorescences. The dandelion plants were considered in this study because they could be a reservoir of avocado thrips and because they are commonly found in avocado orchards in the region. Thrips were collected and transferred to a 50 mL Falcon tube with ethanol at 96%. Location was recorded with GPS on each sampling site. The tubes were kept at -20 °C at the laboratory of Molecular and Cell Biology at Universidad Nacional de Colombia in Medellín until processing. Of 93 thrips, morphological identification was carried out on 62 specimens based on various characters described by Mound and Marullo (1996). For taxonomical identification, the specimens were sent to Dr. Laurence Mound in Australia at the CSIRO Institute, Canberra, and followed the classification of Mound and Hoddle (2016). Species identification based on morphology was carried out for only 62 individuals as these samples were useful enough to determine that on avocado and dandelion plants different thrips species are associated. Photographs of three species of thrips were taken with a motorized Leica M250A Stereo microscope using the plan Apo 1X objective and a DMC 4500 camera with AX support at 5X magnification.
DNA extraction
Thrips stored in alcohol were dried for 2 min on rice paper. Total DNA from thrips specimens was performed using a “salting-out” protocol adapted used by Rugman-Jones et al. (2006). Adults were transferred in a 1.5 ml Eppendorf tube with 100 µl of TNES buffer (50 mM Tris, pH 7.5, 400 mM NaCl, 20 mM EDTA, and 0.5 % SDS) and 1.7 µl of proteinase K (10 mg/ml). Each specimen received an abdominal puncture for DNA extraction. Furthermore, some adults were stored for morphological identification. Tubes were incubated at 37 °C for 18h. Afterward, proteins were precipitated with 28 µl of 5M NaCl for 15s with vigorous agitation. Proteins and adult specimens were sedimented on a microcentrifuge at 13.000 rpm for 5 min. The supernatant was transferred into a new 1.5 ml Eppendorf tube. DNA was precipitated from supernatant through the addition of ice-cold ethanol (100%) and incubated for 1h at 20°C. DNA was further sedimented in a cold microcentrifuge at 4°C and 13.000 rpm for 5 min, ethanol was washed with ultra-pure sterile water-free with RNase and DNase and DNA was air-dried and finally dissolved in 30 µl of sterile water (Rugman-Jones et al. 2006).
COI gene and ITS region amplification
The barcode region COI-5' of 600 bp was made with primers LCO1490 and HCO2198 (Folmer et al. 1994) and amplified under the following conditions: 95°C (5 min), 35 cycles of 94 °C (30 s), 50.7 °C (45 s), 72 °C (1 min); and a final extension at 72 °C (10 min). The final 30 µl volume mix contained 1x PCR buffer, 1.5 mM MgCl2, 0.2 mM dNTP, 0.06 U recombinant Taq polymerase, 0.4 µM de of each primer, and 3 µL of DNA, complemented with ultra-pure water.
Amplification of the ITS region (1000-1200 bp) was performed with the 18SMP / 28SMP primers (De Grazia et al. 2016) and the polymerase chain reaction contained: 4 µl template; 3.0 µl of 10x PCR-buffer; 0.6 µl of 10 mM dNTP; 2.4 µl 25 mM MgCl2; 0.48 µl of each primer; 0.1 µl of Thermo scientific T™ recombinant Taq DNA polymerase and ultra-pure water to complete a final volume of 30 µl. Amplification was carried out in a T100 ™ Thermal Cycle Thermal Cycler (BIO RAD, CA, USA). DNA was denatured at 95 °C for 3 min, followed by 35 cycles of denaturation at 95 °C for 45 s. Alignment was done at 53 °C for 45 s and an extension cycle at 72 °C for 2 min. A final extension cycle for 5 min was done at 72 °C to complete amplification. PCR products were visualized on 1.2% agarose gels with Red gel® (Biotium, Hayward, CA, USA).
Amplified DNA fragments were sequenced (forward and reverse), assembled, aligned, and edited with CLC software v. 7.0.3 (QIAGEN, Hilden, Germany) and submitted to GenBank. Sequence codons were also checked with MEGA 7.0 (Kumar et al. 2016). All these sequences are available at the GenBank COI (MZ298133-MZ298225) and ITS (MZ331549-MZ331581).
Sequencing analysis phylogeny, barcoding gap, and haplotypes
All sequences obtained with both loci were compared to GenBank and BOLD databases using “BLASTn” (http://www.ncbi.nlm.nih.gov/blast/) and http://www.barcodinglife.org/index.php/IDS_OpenIdEngine, respectively (Supplementary Table 1). Two representatives per haplotype group were selected to perform the phylogeny of the COI gene. Sequences of the COI gene and ITS region were aligned using MUSCLE (Edgar, 2004). Two dendrograms were inferred separately for the COI gene and ITS region with the platform Beast 2.0 (Bouckaert et al. 2014). Data was analyzed in Beauti using a MCMC with 46 million generations. Each consensus tree obtained for the COI gene and ITS region was maintained every 4600 generations and preBurning was established for the first 10 million generations for the MCMC. The posterior probability was corroborated with Tracer v1.6 (Rambaut, 2014), the consensus trees were summarized with Treeannotator and graphed with Figtree (Rambaut, 2014).
The barcoding gap was calculated to determine the potential of the COI marker for the identification of the thrips species. For this, the ABGD (Automatic Barcode Gap Discovery) web software (Puillandre et al. 2012) was used with a K2P substitution model a maximum a priori intraspecific divergence value between 0.001 and 0.1, 20 recursive steps, and a relative gap width (X) of 3.0 as proposed by Iftikhar et al. (2016) for thrips.
An analysis of the genetic differences between the haplotypes of F. gardeniae and F. gossypiana was performed for COI gene and ITS region separately for both species, since phylogenetic analysis clustered both species within the same clade, suggesting no genetic differences between them. Seven additional COI GenBank sequences of Mexican origin were also used here. DnaSP v6 (Rozas et al. 2017) and Network v5 (Bandelt et al. 1999), were used to analyze COI and ITS haplotypes. Finally, an AMOVA (Analysis of Molecular Va riance) using Arlequin 3.1 software (Excoffier et al. 2005) was performed to determine whether sampled individuals of the species F. gardeniae and F. gossypiana were genetically different according to FST estimations using COI sequences.
Results
Morphological identification carried out in 62/93 specimens (collected in avocado and T. officinale = dandelion) by Dr. Laurence Mound, demonstrated the presence of the following species in avocado and dandelion: Frankliniella gardeniae Moulton, F. gossypiana Hood, F. panamensis Hood, Thrips trehernei Priesner, T. simplex (Morison), T. palmi Karny, Microcephalothrips abdominalis (Crawford DL), Scirtothrips hansoni Mound & Hoddle, and Liothrips perseae Watson (Tubulifera). In general, the most abundant species found were F. gardeniae with 55.65 % followed by S. hansoni 16.52 % and F. gossypiana 13.91 %. However, F. gardeniae and F. gossypiana were found on flowers, whereas S. hansoni was found on leaves (Fig. 1). To our knowledge, this is the first time that S. hansoni has been reported on avocado in Colombia, while M. abdominalis and F. panamensis have not been reported on avocado before this study. Thrips simplex and T. trehernei were only found on dandelion flowers, suggesting that dandelion plant is not a reservoir of avocado thrips. They were included in this study as many dandelion plants were surrounding the avocado trees and could serve as reservoirs of this insect.
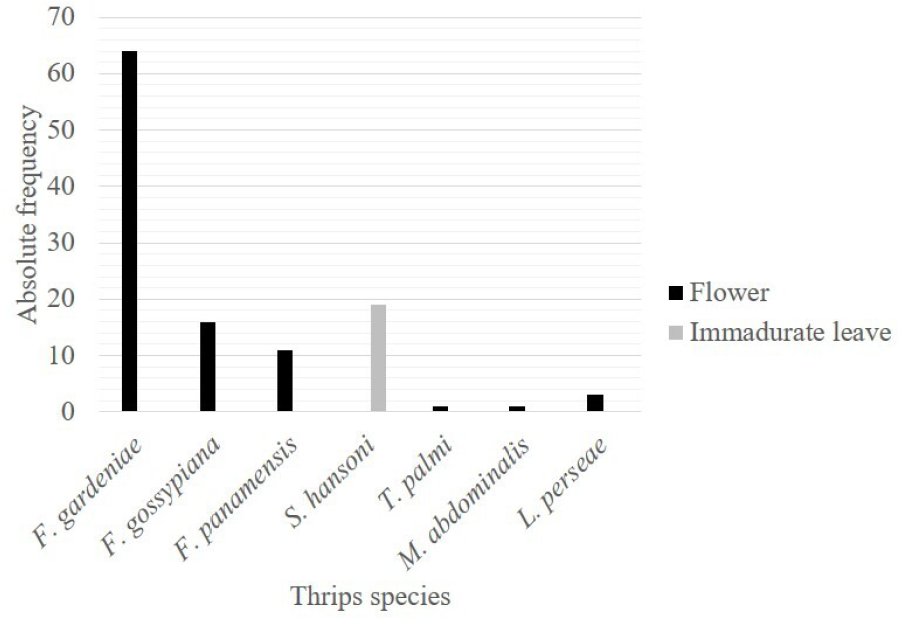
Percentage of the found thrips species, according to the different plant organs. Thrips were collected in avocado crops from Antioquia, North-West Colombia. Black = flower, grey = immature leaves.
The COI gene produced an amplification fragment of 600 bp in 93 individuals, but only 372/600 bp was used to perform the phylogenetic approximation. This was done to include another group of 137 sequences obtained from the GenBank to produce a Bayesian phylogeny (Fig. 2). Furthermore, the ITS sequences produced a 1242 bp segment from 61 samples that were also used to generate another Bayesian phylogeny (Fig. 3). The first dendrogram (Fig. 2) showed that the genus Frankliniella clustered all the species included here within this genus. The species F. gardeniae and F. gossypiana were clustered within the same group, demonstrating that no clear differentiation was obtained between these two species with this locus. Frankliniella was closely related to Microcephalothrips, which was only composed by the species M. abdominalis. Furthermore, the genus Thrips was also closely related to Frankliniella and Microcephalothrips and was composed by the species T. palmi, T. simplex, and T. trehernei. Scirtothrips were grouped separately from the other three genera in a group consisting of S. hansoni, S. persea, and S. dorsalis. The topology obtained on this dendrogram suggests that more COI sequences are needed to improve the phylogeny of the genera Microcephalothrips and Thrips. In the second dendrogram (Fig. 3), the genus Scirtothrips was also separated from the genera Frankliniella and Thrips, corroborating the results obtained with the COI phylogeny. However, the close relationship found between these two genera demonstrates that more sequences are needed for the ITS region as grouping between these two genera did not differentiate the species well.
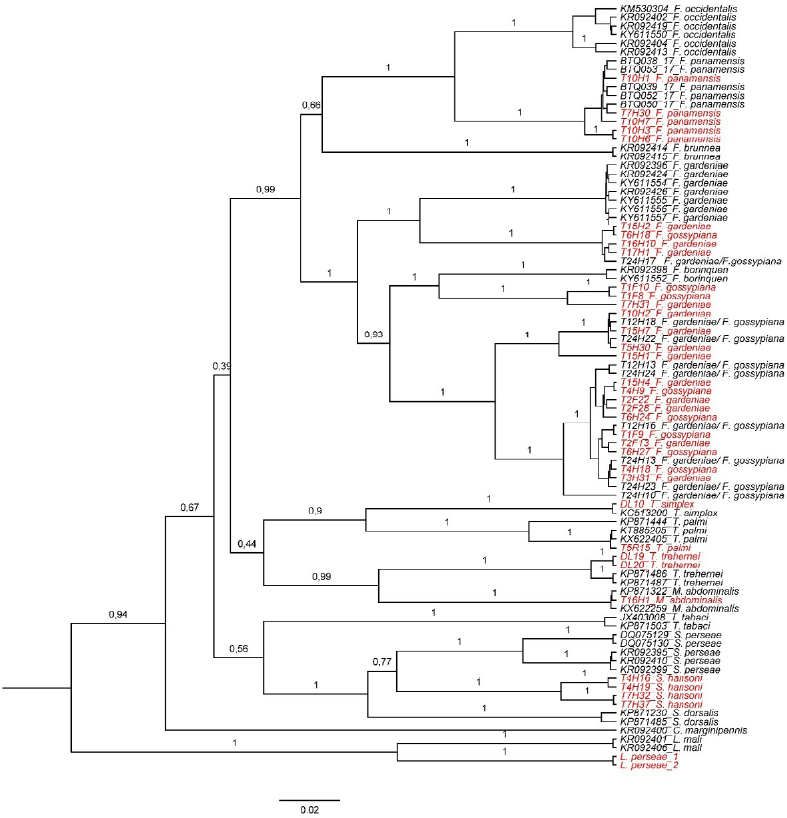
Bayesian phylogeny was obtained with COI sequences from avocado thrips. The sequences used as references are with their GenBank or Bold access codes. The trips species found in this work were: F. gardeniae, F. gossypiana, F. panamensis, T. palmi, T. simplex, T. trehernei, Microcephalotrips abdominalis, and S. hansoni. Out-group = L. perseae. In red, in red, thrips samples identified morphologically and molecularly.
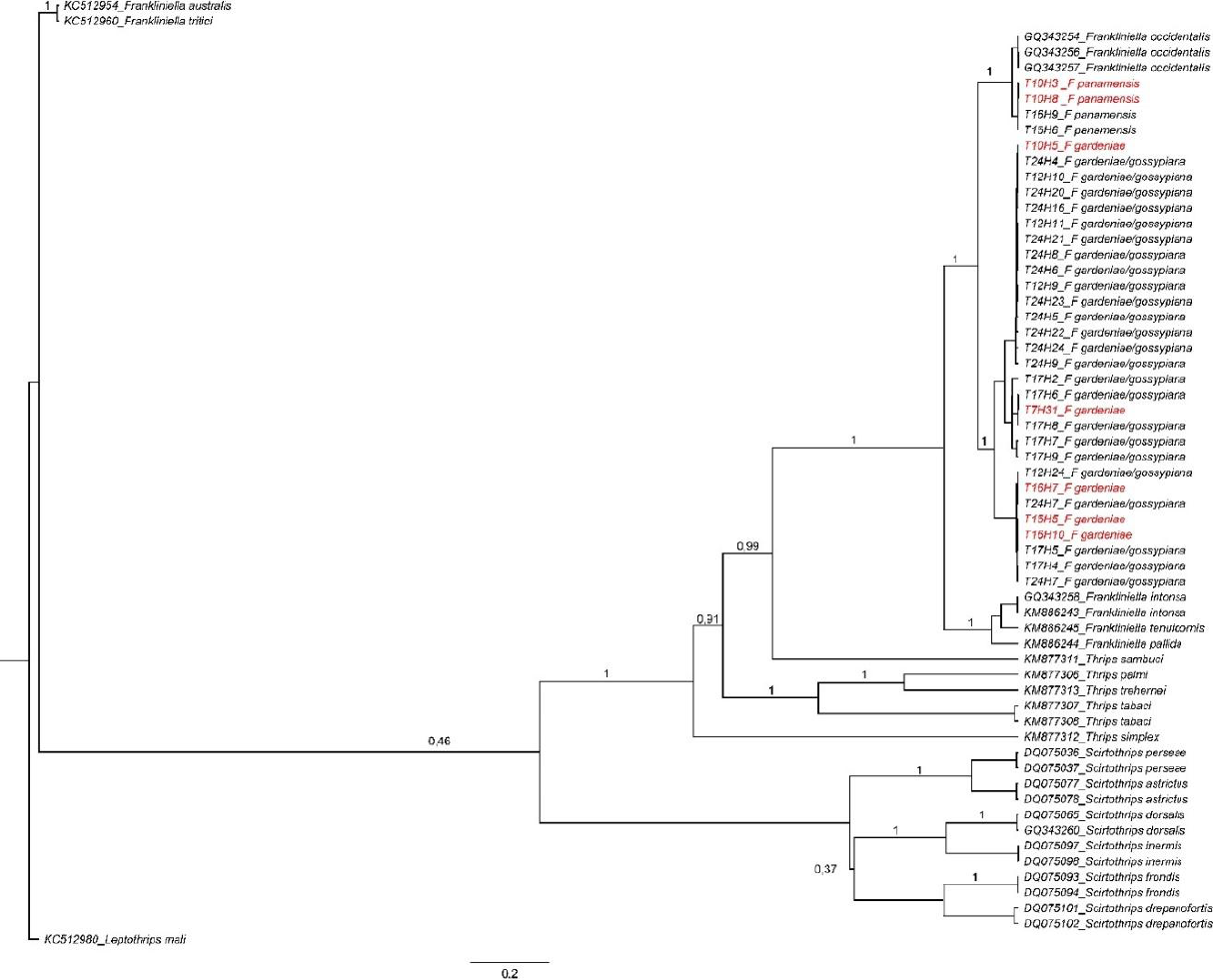
Bayesian phylogeny was obtained with ITS sequences from avocado thrips. The sequences modified as references are with their GenBank or Bold access codes. The trips species found in this work were: F. gardeniae, F. gossypiana, F. panamensis, T. palmi, T. simplex, T. trehernei, Microcephalotrips abdominalis, and S. hansoni. Out-group = L. mali. In red, in red, thrips samples were identified morphologically and molecularly.
The ABGD barcoding gap analysis showed a genetic gap between 8 to 11 % (Fig. 4). The presence of 42 groups was observed which were composed of the different species of thrips (Supplementary Table 2) and in general, each group contained a single species. However, some species such as F. panamensis and S. hansoni formed several groups indicating intra-species variation. Additionally, in agreement with the other results obtained in phylogenetic analysis, genetic divergence between F. gardeniae and F. gossypiana was not evidenced, suggesting that these two species belong to one genetic unit.
Haplotype network revealed the existence of 19 haplotypes (Fig 5a), being H2 the most common. This haplotype had thirty-one individuals where four of them corresponded to F. gossypiana, followed by haplotypes H1, H6, H7, H8, and H3. The rest of the haplotypes presented a single individual, including F. gardeniae and F. gossypiana. Haplotype identification carried out with Dnasp showed that the species F. gardeniae and F. gossypiana cannot be differentiated with the COI marker. Frankliniella gardeniae have more haplotypes than F. gossypiana suggesting that the origin of the latter species is more recent. Additionally, Mexican F. gardeniae produced a single haplotype (H1) unlike the Colombian haplotypes that were divided into the remaining eighteen haplotypes. Mexican haplotypes were far apart from haplotypes of the same genus from Colombia suggesting that F. gardeniae from Colombia is genetically distant from the same species from Mexico.
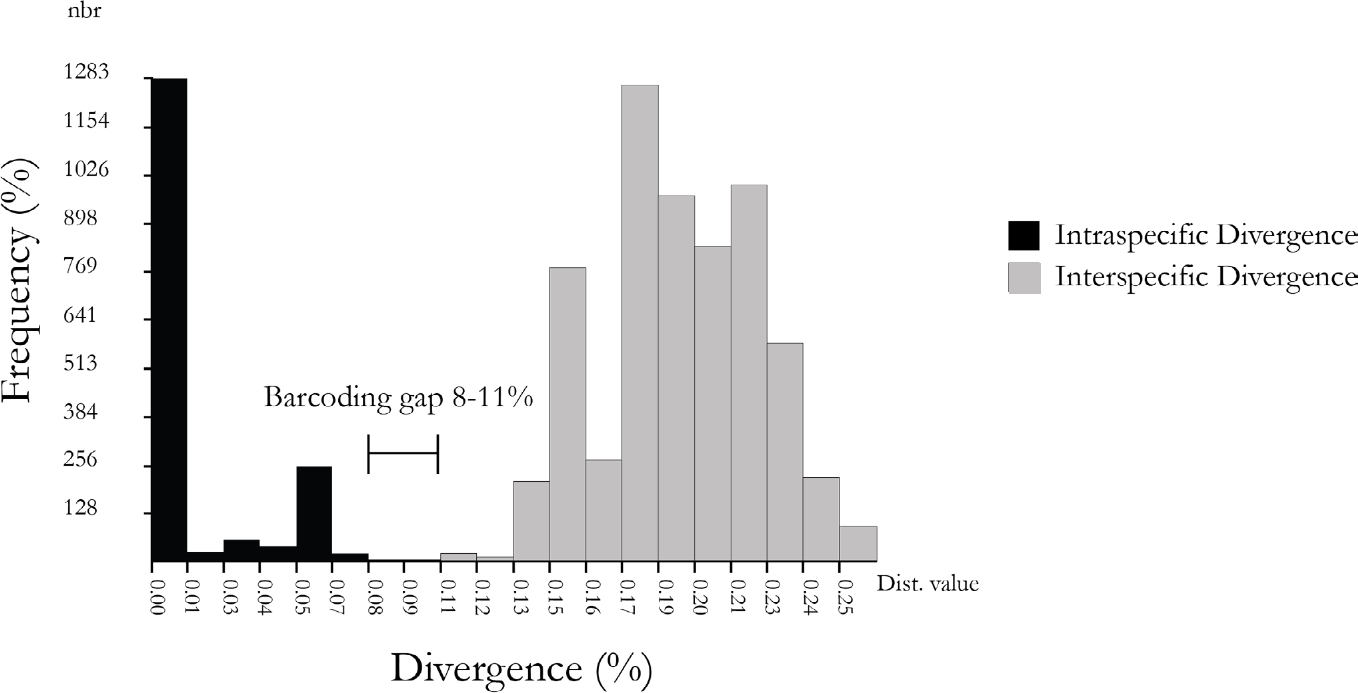
Analysis of the barcode gap of thrips species generated by automatic barcode gap detection (ABGD) (Puillandre et al. 2012). The histogram shows the K2P distance within and between each pair of samples of the COI gene.
The ITS haplotype network (Fig. 5b) showed that some of the specimens that were collected on avocado might belong to the genus Frankliniella given their genetic proximity to sequences obtained from the GeneBank for the genus. Since fewer sequences were used here to analyze the genetics of this group, these haplotypes mainly corroborate the results obtained for the COI gene. In this case, no analysis of genetic differentiation was made for F. gardeniae and F. gossypiana since more information is needed in the databases of these two species. The haplotype network showed that the genus Thrips is closely related to Frankliniella as sequences of T. simplex and T. sambuci were close to Frankliniella. On the other hand, the genus Scirtothrips is not closely related to the other mentioned genera suggesting that is genetically different from them, corroborating the results obtained for the COI gene network and phylogeny.
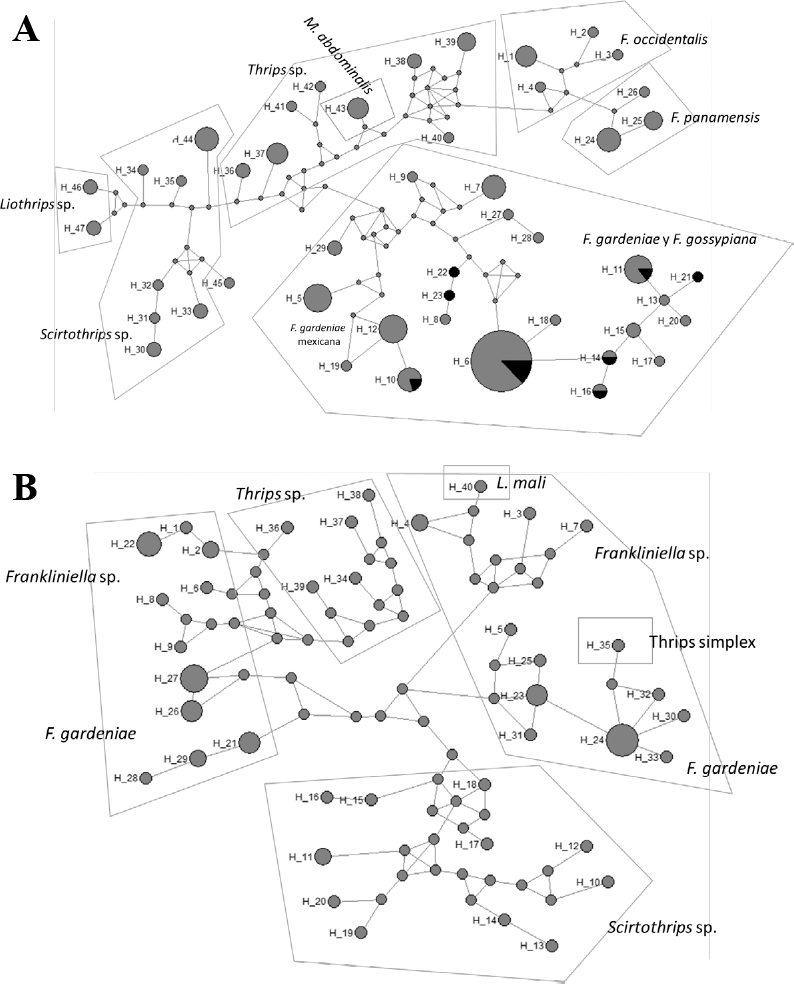
Haplotypes networks. a. Haplotype network obtained for COI sequences from Antioquia avocado and GeneBank and BOLD Databases. b. Haplotype network obtained for ITS sequences from Antioquia avocado and GeneBank
Finally, AMOVA results obtained from COI sequences of F. gardeniae and F. gossypiana showed a low FST value that was not significant (Table 1), suggesting no genetic differentiation between these two species.
AMOVA results were obtained from genetic comparisons between individuals of the species F. gardeniae and F. gossypiana collected from avocado.
Discussion
So far, no molecular genetic studies of thrips have been done before this work in Colombia. DNA barcoding was employed here as it represents an effective method for species-level identification. Based on the molecular data obtained, we consider that F. gardeniae and F. gossypiana are likely to be the same species. However, to obtain more robust results it would be important to have more morphologically-identified specimens from both species to be studied at the molecular level. Bravo-Pérez et al. (2018) used sequences of COI to identify several trips species on avocado from Mexico by using specimens collected from fruits and leaves. These authors mentioned that species identification was useful in differentiating F. borinquen, F. brunnea, F. rostrata, and F. insularis on avocado, demonstrating the efficacy of this gene for the identification of species belonging to the Frankliniella genus. A similar analysis, using the COI gene, was carried out by Brunner et al. (2004) with 10 thrips species. They found a high FST of 0.9713 between Thrips angusticeps and T. tabaci and low FST values such as 0.16 between Echinothrips americanus and T. palmi. The significant population differentiation between the 10 species involved showed that this gene successfully identified thrips species from different geographic origins (England, North America, Israel, amongst others).
When determining the percentage contribution of thrips species in crops per plant organ, F. gardeniae, F. gossypiana, and F. panamensis were found to be predominately associated with floral organs suggesting that this genus feed mainly on flowers. In contrast, S. hansoni was found mostly on young leaves. Mound and Teulon (1995) reported that members of the Frankliniella genus are mainly found on flowers and leaves, but Scirtothrips are present only on young leaves. In other studies, different thrips species have been reported to prefer different plant tissues. For example, Heliothrips haemorrhoidalis collected from citrus (Rutaceae) and Quercus (Fagaceae) prefer mostly old leaves (Marullo 2009). In a previous study conducted in Colombia on avocado, the following thrips species were found: Frankliniella gardeniae Moulton, Karnyothrip sp., Leptothrips sp., and Selenothrips rubrocinctus Giard, and on avocado (Echeverri-Florez et al. 2004). In our study, we found a higher number of thrips species but no potentially predatory thrips. Differences between these two studies carried out in Colombia might be due to the fact that our study was carried out in Antioquia, North-East Colombia whereas the study by Echeverri-Florez et al. (2004) was carried out in Risaralda and Caldas departments at the West-Center of Colombia. These differences may be due to environmental factors, management at the agrochemical level, or the presence of different adjacent crops. Another interesting aspect of our study is that the species found on dandelion: Thrips simplex and T. trehernei differed from the species that attack avocado, discarding the possibility that dandelion could be a reservoir of thrips that affect avocado.
Sequencing of the barcode COI gene was used for the identification and elucidation of all the species collected in this study, and its implementation in avocado was very useful. Comparisons between the sequences obtained in this work together with those obtained from GenBank and BOLD databases showed matches with F. panamensis, L. perseae, M. abdominalis, T. simplex, T. palmi, and T. trehernei, with a high percentage of similarities (98 %). Similar results were obtained with ITS sequences, suggesting that both markers can also be implemented in the integrated pest management of this group of cryptic species of avocado. However, the COI gene allows obtaining a better image of the phylogenetic relationship of thrips species.
Phylogeny found for the nine species with COI gene and ITS region showed a close relation between Frankliniella, Microcephalothrips, and Thrips and a distant relation of these three genera with Scirtothrips. The topology of the phylogeny obtained with COI in this study coincided with previous studies (Iftikhar et al. 2016) and (Liu et al. 2017) showing that the genera Scirtothrips and Thrips are closely related and share a common ancestor with Frankliniella, corroborating the phylogenetic approach obtained here. Similarly, Rebijith et al. (2014) also found the same topology.
In the present study, only two loci were used to carry out a phylogeny and only members of the family Thripidae were analyzed. The results obtained support a close relation of the genera Frankliniella, Microcephalothrips, and Thrips with some differentiation with the genus Scirtothrips. However, the phylogenetic relationship between F. gardeniae and F. gossypiana remains difficult to determine. A recent phylogeny obtained by Bravo-Pérez et al. (2018) on avocado from Mexico found that F. occidentalis specimens might also constitute a group of cryptic species of avocado. In our study, F. gardeniae and F. gossypiana were found clustered within the same group, suggesting the possible existence of cryptic species or biotypes within the same species as they differ in morphology but did not differ in their genetics. To our knowledge, this is the first time that information on the genetic variation of these species is recorded at the genetic level in the world. AMOVA results also showed no genetic differentiation between F. gardeniae and F. gossypiana (Fst = 0.0076, P > 0.05) supporting the hypothesis that these two species represent the same genetic unit, even though they differ morphologically.
In a population genetics study made with Thrips tabaci, a thrips species considered a relevant pest of onions in Australia, Westmore et al. (2013) showed that the species evolved into different biotypes, based on the analysis of differences of the COI gene (sequences) found between thrips sampling sites (Westmore et al. 2013). Two biotypes were found, one mainly associated with onion which does not transmit viruses, and the other with tobacco which transmits viruses, differing only in 6 nucleotide positions. In our case, F. gardeniae and F gossypiana differed in 41 nucleotide sites meaning that F. gossypiana might represent a biotype or a subspecies of F. gardeniae. However, according to Hebert et al. (2003), if two populations differ from 2 % K2 distances, they represent different species, and the results obtained here for F. gardeniae and F. gossypiana showed K2 = 6 % (data not shown).
Classical taxonomy using morphological characters cannot always be employed in the identification of thrips species in all life stages, as phenotypic variation can be insufficient (Brunner et al. 2004; Rebijith et al. 2014). The presence of unusual morphological forms on different hosts, small size, co-existence on the same host, complex life cycles, parthenogenetic lifestyle, amongst other traits, makes difficult the identification of species in this group (Rebijith et al. 2014; Iftikhar et al. 2016). As an alternative, the proposed concept of DNA barcoding could be a powerful tool to identify all metazoan species employing 658 bp fragment of the 5'end of the mitochondrial COI locus (Hajibabaei et al. 2007). DNA barcoding has been employed for discriminating cryptic species, biotypes, haplotypes, and host-associated pests. Therefore, sequencing this gene plays an important role for integrated pest management programs since it will allow to easily identify the species occurring in the crop thus generating better monitoring methods. Having an easy to implement identification method will help to identify the preferred areas of the insect on the plant as well as other possible hosts (Hajibabaei et al. 2007; Rebijith et al. 2014; Iftikhar et al. 2016; Liu et al. 2017). Implementing the use of sequences from the ITS region also might be useful to corroborate the identification of thrips in avocado and other crops. However, in our case, the ITS region was difficult to amplify and therefore it was not possible to have an adequate number of sequences. This might have been due to the quality and quantity of the DNA obtained from thrips, and to the fact that the primers used were generic for the genus. This difficulty was particularly seen with S. hansoni which is one of the smallest insects, so it may be related to the amount of hemolymph obtained from it. The fact that the COI gene amplified better might be due to better annealing of primers or because it is a mitochondrial gene, it might be present in higher copy numbers in the insect cells.
Conclusions
The use of the COI gene and ITS region allowed for the identification of avocado thrips species. In addition, a good hypothesis of the phylogenetic relationship of these species using the COI gene was obtained. Molecular analysis results suggest that F. gardeniae and F. gossypiana are the same genetic unit and that F. gardeniae from Colombia is genetically distant from the same species from Mexico. However, we obtained few sequences from the ITS region, perhaps due to the quantity and quality of the DNA extracted from thrips or because of the primers used in this work. Therefore, we suggest that future studies should use another extraction method or other set of primers better suited for the sampled species. This study showed that DNA sequencing is a fast and effective method to identify most thrips species in avocado and it could be implemented in integrated pest management programs of this group of insects in Colombia. Identification of pest species is relevant for the improvement of their control given that specific pesticides should be used for each species and in the case of avocado at least seven cryptic species of thrips were found in this study.
Origin and funding
The research reported in this publication was supported by the Universidad Nacional de Colombia under Project Code 202010013471, contract number FP44842-132-2017. This material is based on work supported by the Ministry of science, technology, and innovation (Minciencias) of Colombia Graduate Research Fellowship Program for Daniela Cano-Calle No.727 in 2015 under Grant No. 201010020475. Foundation for the Promotion of Research and Technology, Bank of the Republic of Colombia Project No 201423.
Author's contributions
Daniela Cano-Calle participated in experimentation processing, design, data processing, analysis, and discussion of the information in the manuscript and the writing of the manuscript.
Clara Inés Sadamando-Benjumea participated in the analysis, data processing, discussion of the information and the writing of the manuscript.
Claudia Ximena Moreno-Herrera participated in the design, project administration, discussion, supervision, review, and editing of the manuscript.
Rafael Eduardo Arango-Isaza participated in the design, data processing, analysis, and discussion of the information in the manuscript and the writing of the manuscript.
