Abstract
Ovarian cancer is the leading cause of gynecologic cancer death in the Western world and more than 70% of patients are diagnosed with advanced stage disease. The high mortality rate is due to the difficulty in the early detection of ovarian cancer. Current screening strategies lack the necessary sensitivity and specificity to reliably and accurately diagnose affected women, prompting investigators to seek alternative means of analysis found in protein pathways and networks. Proteomics seeks to advance the understanding of how proteins interact in cancer and may provide a mechanism for early stage diagnosis. The proteomic techniques of laser capture microdissection, mass spectrometry and tissue lysate arrays have led to the discovery of new biomarkers and the identification, development and approval of a number of targeted therapeutic agents. Following validation through clinical trials, the application of these techniques will contribute to the changing paradigm of cancer detection and treatment toward personalized medicine.
Keywords
Ovarian cancer remains the leading cause of death from gynecologic malignancy among women in the USA. In 2005, an estimated 22,200 women will be diagnosed with ovarian cancer and 16,210 will succumb to the disease [1]. Over 70% of women will be diagnosed with advanced stage disease. Despite appropriate surgical and chemotherapeutic interventions, the 5-year survival rate is 30–40% versus 90–95% with organ-confined cancer. Early detection of ovarian cancer itself could have a profound effect on successful interventions for this disease. Our current assessment methods of physical exam, cancer antigen (CA)-125 testing and transvaginal ultrasound lack the necessary sensitivity and specificity to provide accurate and cost-efficient screening for the general population, or from which to predict who will best benefit from which treatment. These challenges have led investigators to seek and discover new alternatives for the diagnosis and treatment of ovarian cancer.
Oncology is becoming closely intertwined with molecular medicine and moving beyond genomics to proteomics [2–4]. Proteomics characterizes the information flow within the cell and organism through protein pathways and networks. It seeks not only to catalog proteins, but also to advance the understanding of how these proteins interact in cancer and may provide a mechanism for early stage diagnosis [3]. The application of proteomic techniques is now underway in several cancers including colon, pancreatic and breast cancer, resulting in the discovery of new biomarkers and molecularly targeted therapeutics [5–7]. Cancer is fundamentally a genetic defect that confers survival advantages to malignant cells. These genetic defects are transcribed and translated into proteins that drive cell survival and/or apoptosis. The completion of the human genome sequencing initiative indicates the genetic diversity inherent in individuals, which translates to a much broader proteomic diversity that emphasizes the need to tailor therapy for individual patients, based on their unique genetic and proteomic constitutions [8].
Proteomic techniques
The field of proteomics is heavily dependent on technology, and technologic advances have been paralleled by enthusiastic translational programs. A number of approaches have been used to study proteins, employing techniques that range from disruption of the cellular environment for quantitative measurements to attempts at preserving the integrity of signaling networks and architecture contained within the cell. Proteins have traditionally been studied using immunohistochemical staining, in situ hybridization and western blotting for quantification [9], while two-dimensional gel electrophoresis (2D PAGE) has been used for protein identification [10]. Newer techniques include protein microarrays, of which there are two types: tissue lysate arrays (TLAs), also known as reverse phase arrays (RPAs), and antibody or forward phase arrays (FPAs). In addition, laser capture microdissection (LCM) or other sensitive selection techniques can be used for procurement of defined cell samples from tissue specimens [11]. Various applications of high-throughput mass spectrometry (MS) are used to analyze and sequence protein samples. Bioinformatics plays a critical role in analyzing the massive amounts of information yielded by these approaches and has proven to be essential for progress in proteomics [12].
2D PAGE separates proteins by isoelectric point and molecular weight. Initially crude, this technology has evolved over the last 30 years and is now capable of yielding high resolution, reproducible protein separation [13]. Protein spots are visualized by staining with coomassie blue, silver staining or using fluorescent dyes. Identification of protein spots was classically carried out by transferring the separated proteins to a membrane and sequencing from the N-terminus. However, this system was labor-intensive and lacked sensitivity. Therefore, newer methods were developed which relied upon chromatography separation techniques coupled to a MS platform. Within the last 10 years, significant improvements in sensitivity, throughput and automation, combined with databases of protein sequences, produced much faster methods of protein identification [14].
The different types of protein microarrays differ in the sample that is applied. Protein microarrays are composed of an immobilized protein sample bound to a nitrocellulose-coated glass slide. The sample contains a target molecule, such as an antibody or protein fragment, to which tagged probes bind and subsequently emit a signal in a relatively quantitative fashion [15]. FPAs contain multiple different probes applied to the array chip as individual spots; the most common FPA is an antibody array. The whole array is incubated with a labeled test sample, for example a cellular lysate, thus yielding an overview of target protein presence. Major limitations of this format include variability between runs and low sensitivity [16]. Conversely, in TLAs/RPAs, test samples are immobilized on the chip and then queried with a single probe, such that a single end point is determined for multiple samples [17]. TLAs utilize high-sensitivity detection probes and signal amplification techniques in order to obtain information from small amounts of tissue sample.
LCM is often used to obtain cell samples for protein microarrays as it is able to separate a relatively homogeneous population of cells from a heterogeneous sample [11]. For example, LCM enables the removal of an enriched sample of omental stroma or invasive ovarian carcinoma from a single biopsy that contains multiple cell types, allowing protein expression analysis in those cells. The combination of LCM and TLAs has led to multiple applications of this technology from basic research to clinical trials. It is a powerful technique that allows for the monitoring of changes in protein expression across multiple cell types over time.
High-throughput MS currently plays a large role in proteomics (Figure 1). With MS, samples are ionized and detected by an ion detector plate. The time required for the sample to reach the detector plate is a function of the mass-to-charge ratio (m/z) [18]. There are several methods by which to introduce the sample into the spectrometer. Matrix-assisted laser desorption and ionization with time-of-flight (MALDI-TOF) and surface-enhanced laser desorption and ionization with time-of-flight (SELDI-TOF) are most commonly used. MALDI-TOF employs the use of a matrix that traps a subset of proteins in the sample [19]. This method is particularly effective with low molecular weight proteins of less than 20 kDa [20]. In contrast, SELDI-TOF utilizes a commercial chip customized with specific bait molecules that either chemically bind protein samples using cationic or hydrophobic interactions, or employ an antibody to which samples bind followed by a matrix to facilitate ionization [21]. These are all amenable to mass throughput.

Proteomic patterns for diagnosis of disease.
The use of MS technology has given rise to multiple approaches applied to early ovarian cancer detection. These include the discovery of biomarkers and diagnostic proteomics where spectral patterns guide detection [22]. Analysis of spectral images requires software that is capable of segregating different classes of sample, for example, diseased and unaffected, based on the pattern of proteomic peak intensities rather than identification of the proteins responsible for the individual peaks. The use of high resolution MS can yield millions of data points from which to glean potential biomarker panels and/or to identify specific peaks for isolation and sequencing.
A recent advance in the area of MS proteomics research was the application of the high-resolution, high mass accuracy mass spectrometer, such as the hybrid quadrupole (Qq)TOF. Liotta and Petricoin used QqTOF and the lower resolution series PBS-II mass spectrometer to address the question of whether the high resolution instrument conferred an advantage [23]. They obtained mass spectra of 248 serum samples from unaffected women and women with ovarian cancer. They built and validated the diagnostic models produced by each instrument using independent training and validation sets. The results of their study demonstrate that the diagnostic models obtained from QqTOF were statistically more effective for testing patient samples. This suggests that high-resolution technology is preferable.
Applications of proteomic technologies
The artillery of new proteomic technologies has seemingly limitless applications in the battle against ovarian cancer. The capacity to analyze heterogeneous biologic samples and detect specific proteins is remarkably powerful. However, the utilization of these technologies in screening for ovarian cancer is dependant upon the discovery and validation of reliable serum biomarkers of ovarian cancer.
A useful ovarian cancer biomarker-based screening test must possess several key characteristics. First, the test must be easy to administer. Ideally, the biomarker should be present in an accessible body fluid such as blood, serum or urine. Furthermore, the test must be both sensitive – consistently positive in affected patients – and specific – consistently negative in unaffected patients. Moreover, an efficient ovarian cancer screening program must also have a strong positive predictive value (PPV). Currently, if a patient tests positive for a biomarker, costly and invasive surgery may be necessary to obtain a biopsy of the ovary. Researchers in the field agree that a PPV of at least 10% (i.e., one cancer discovered for every ten biopsy procedures) is acceptable [24]. A biomarker employed in such a test must have a specificity of 99.6% to yield the PPV of 10% [25].
CA-125, a high molecular-weight glycoprotein, is regarded as the gold standard biomarker, and is approved by the US Food and Drug Administration (FDA) to monitor progression and response of ovarian cancer to treatment. However, it does not fit the statistical profile of a specific presymptomatic screening biomarker. Its physiologic concentration is significantly increased in only 80–85% of late and 50–60% of early stage disease [26]. Moreover, CA-125 concentrations are increased in a number of conditions other than ovarian cancer, including the first trimester of pregnancy, breast cancer and endometriosis [27]. These studies demonstrate that CA-125 is not an effective biomarker for ovarian cancer screening. Current studies using the rate of change, risk of ovarian cancer algorithm (ROCA) model have improved the utility of CA-125 [25]. These studies demonstrate that a single value of CA-125 is neither sufficiently specific nor sensitive for ovarian cancer screening in a general population.
Over the past 5 years, much effort has been expended to discover new serum biomarkers to improve upon the predictive value of CA-125. The National Cancer Institute's Early Detection Research Network (EDRN) recently described a five-step system of biomarker discovery and evaluation:
Preclinical exploration to identify promising candidate biomarkers
A clinical assay to determine the ability of the test to detect the disease
A retrospective/longitudinal determination of a putative biomarker's ability to detect preclinical disease
Prospective screening to identify the extent and characteristics of disease detected by the test and the false positive rate
A definitive trial to determine the effectiveness of the putative biomarker in reducing the disease burden on the population [28]
Many promising markers have been identified and are currently in the first two steps of the EDRN discovery and evaluation system. While the EDRN's guidelines apply to the general process of biomarker discovery, they do not recommend specific techniques to employ. Scientists have utilized a wide range of technologies to discover biomarkers, from simple immunoblots to more complicated cDNA microarrays to proteomic technologies, including MALDI-TOF and SELDI-TOF MS.
Preproteomic techniques, including cDNA microarray and differential polymerase chain reaction (PCR), have been widely used to identify many potential ovarian cancer biomarkers for use in screening tests. An example of one such identified biomarker is a protein called HE4 [29,30]. This protein is a member of the whey-acidic secretory protein (WAP) family, which includes several protease-inhibiting proteins present in breast milk and seminal fluid [31]. Initially, cDNA microarrays have shown the HE4 gene, WFDC2, to be overexpressed in ovarian carcinoma, whereas its expression in normal tissues is uniformly low [21,32,33]. Furthermore, HE4 protein levels have been shown to be increased in ovarian carcinoma by both reverse transcription (RT)-PCR [33] and enzyme-linked immunosorbent assay (ELISA) techniques [21]. Based on these data, researchers developed a clinical assay using ELISA to measure serum HE4 levels. In addition, Drapkin and colleagues demonstrated that HE4 overexpression is subtype specific (serous and endometrioid) by immunohistochemistry and that the protein is secreted by tumor cells as a glycoprotein [33]. HE4 was found to be comparable with CA-125 in distinguishing women with both early and late-stage ovarian cancer from healthy patients, with 95% specificity and 94% sensitivity [21]. Furthermore, this study indicated that HE4 was more effective than CA-125 at distinguishing malignant ovarian disease from benign ovarian disease [21]. Importantly, it appears that HE4 is positive less frequently in patients with benign gynecologic disease, an issue that has affected the use of CA-125 for accurate and reliable screening.
MS has also played an important role in ovarian cancer biomarker discovery. A recent study used MS technology to confirm the utility of two subspecies of lysophosphatidic acid (LPA) already defined as putative ovarian cancer biomarkers. LPA has been shown to downregulate expression of the Fas-ligand in several ovarian cancer cell lines and thus, reduce apoptosis [34]. Furthermore, LPA has been shown to activate G-protein signaling pathways that lead to the activation of phosphoinositol-3 kinase and the downstream activation of the antiapoptotic protein kinase B (PKB/AKT) [35]. In addition, laminin-induced secretion of LPA-stimulated ovarian cancer cell migration [36]. It has long been known that LPA expression is increased in early and late-stage ovarian cancer [37]. Recently, MS was used to measure the concentrations of the dozens of subspecies of LPA simultaneously to clarify which of its many subspecies has the most potential as an ovarian cancer biomarker. The most significant changes were observed in two LPA subspecies: 16:0 LPA and 20:4 LPA. The two subspecies demonstrated sensitivity of 91.1% and specificity of 96.3% in clinical assays [38]. MS served in this fashion to refine the process and application of biomarker discovery.
Past research has focused on identifying a single optimal protein biomarker, a more specific and sensitive version of CA-125. However, recent studies support new approaches to biomarker-based proteomic ovarian cancer screening. Current data suggest that multiplexed, composite or multiple biomarker targets yield improved sensitivity and specificity of protein fingerprints generated by MS.
A composite biomarker of CA-125 and mesothelin has shown promise as a target of proteomic-based ovarian cancer screenings. The glycoprotein mesothelin is a differentiation antigen that is present on normal cells and highly overexpressed in a variety of cancers, including ovarian cancer [39]. Interestingly, Rump and colleagues have shown that mesothelin binds to CA-125 and that this interaction facilitates cell adhesion [40]. This adhesive phenotype may play a role in the metastatic implantation of ovarian cancer in the peritoneal cavity. Small quantities of mesothelin are detectible in the blood of patients with mesothelin-positive cancers [41]. When used in concert with CA-125 as a composite marker, it was found that the mesothelin–CA-125 combination had specificity equal to CA-125 and a greater sensitivity (98%). It was also shown that the composite marker was more successful in differentiating between benign and malignant ovarian disease than CA-125 alone [42].
The authors' group hypothesized that serum would contain diagnostic protein signatures with which to differentiate cancer patients from unaffected women. This approach focused on mining the MS datastreams for patterns of proteins that function as successful discriminants of disease [43]. A training set of defined samples was used in order to provide knowns for a bioinformatics algorithm to generate the discriminating pattern. An independent set was used to test the validity of the signature defined. The preliminary findings within the limited test cohorts demonstrated 100% sensitivity and 95% specificity. These results illustrated a proof of principal that proteomic pattern technology could be used to differentiate affected from unaffected samples. A prospective population-based study is warranted, in which this approach could be studied and validated in a larger cohort. It is not yet known if there are histology or stage-specific protein signatures and, if so, what value they may provide to the clinical community. If a specific signature for mucinous ovarian cancer were developed and found to distinguish between mucinous ovarian versus mucinous appendix or other gastrointestinal tumors, an important advance will have been made.
On a tissue level, these differences are evident when examining protein expression in invasive tumors compared with low malignant potential tumors. In 2002, Brown and colleagues examined the technique of LCM with epithelial tumor cells in human tissue specimens with 2D PAGE to identify proteins that may serve as invasive ovarian cancer-specific biomarkers for early detection and/or new therapeutic targets [14]. A total of 23 proteins were consistently differentially expressed between both the low malignant potential (LMP) and three invasive ovarian tumors in the limited study set. The direct comparison of LCM generated proteomic profiles of invasive versus LMP ovarian cancer that may more directly generate important markers for early detection and/or therapeutic targets unique to the invasive phenotype.
Since that time, there has been much discussion in the scientific, medical and lay press regarding the reliability and validity of the proteomics approaches. The consensus of the community is that the hypothesis and concept are valid. Multiple other investigators in many disease disciplines have used serum MS and higher order bioinformatics to mine diagnostic and prognostic data. The community agrees that this approach must be rigorously advanced using independent and blinded serum sets, preferably from a variety of sources, in order to reduce or minimize collection bias. Furthermore, there is a strong call to put raw data into the public domain so that the maximal amount of information may be gleaned by the most people. Other groups reported that, using proteomic patterns derived from one ovarian cancer data set, they were able to segregate affected and unaffected samples in a blinded subset [44]. However, Baggerly and colleagues reported other challenges inherent in proteomic profiling of the serum, including programming errors that confound reproducibility across these data sets and incomplete randomization of sample processing, that bias the results [45]. If ovarian cancer is suspected on the basis of physical examination and the results of transvaginal ultrasonography, an exploratory laparotomy is performed for histologic confirmation, staging and tumor debulking [46]. Clinical trials are underway through the Gynecologic Oncology Group and the European Institute for Cancer (Milan, Italy) to develop the serum resources from which to generate sensitive and specific proteomic signature(s) to discriminate malignant from nonmalignant pelvic masses. These are anticipated to be used ultimately as a presurgical secondary screen for ovarian disease [47].
An additional proteomic technique that can be used in combination with surgery is MALDI-TOF MS imaging. Sections of biologic tissues are introduced into a MALDI-TOF MS instrument, where the ultraviolet pulsed laser of the MALDI-TOF source is used to raster over a selected area while acquiring mass spectra of the ablated ions at every image point. Hundreds of analyte-specific images can be generated based on the selected masses from this array of spectra [48]. This technique may permit molecular assessment of tumor biopsies, with the potential to identify subpopulations of cells that are not evident based on the cellular phenotype determined microscopically. It may permit assessment of surgical margins at the molecular level [49]. This same technique can be used to detect the presence of pharmacologic agents in specific tissues after drug administration [50].
Personalized medicine
Recent evidence suggests that each patient's cancer may have a unique subset of pathogenic molecular derangements [3]. These derangements may be the result of alterations at different points in the information cascade and may account for the heterogeneous response to existing treatment regimens [51]. Over the past 30 years, advances in surgical technique, chemotherapy and supportive care all have contributed to the steady increase in the median survival of women with advanced stage ovarian cancer; however, there has been no improvement in the cure rate. The oncologist is challenged, particularly with advanced cancers, by the following [52]:
Predicting tumor response to a given drug or regimen
Determining which tumors of identical histology will remain indolent and which will be likely to progress
Determining the timing of the emergence of drug-resistant cancer cells and hence switch to appropriate therapy
In the future, proteomic technology may yield a molecular profile or pathway outline for an individual tumor. The incorporation of that information into clinical practice may be the beginning of an improvement in the cure rate.
The majority of patients with epithelial ovarian cancer will require postoperative adjuvant chemotherapy in an attempt to eradicate residual disease [46]. Protein activity is the current target of drug discovery. The evolution of proteomics techniques is thus of major importance to the drug development process [53]. Common applications of proteomics in ovarian cancer in academic and pharmaceutical industry research include target identification and validation, identification of efficacy and toxicity biomarkers from readily accessible biologic fluids and investigations into mechanisms of drug action or toxicity. Examples of these agents include imatinib mesylate (Gleevec™/Glivec™), which is targeted against the BCR—ABL fusion and c-kit kinases for treatment of both leukemias and gastrointestinal stromal tumors and is under investigation in ovarian cancer trials, and gefitinib (Iressa™, ZD1839) a tyrosine kinase inhibitor that targets the epidermal growth factor receptor, which is important in a number of epithelial cancers [54].
Proteomics is being applied specifically to ovarian cancer in two ways. The first is MS to identify either signatures and/or actual proteins that may be diagnostic or prognostic to screening or outcome. The protein array approach is being used in an exploratory fashion to assess what biochemical changes may occur in response to therapy. Following rebiopsy or molecular imaging, the effect of treatment can theoretically be monitored in real time. This may also allow proof of target in showing that the drug is altering its target as directed. Furthermore, this can allow recognition that a single target may not be sufficient for clinical benefit. In the authors' trial of imatinib for ovarian cancer, limited clinical benefit was observed, but downregulation of c-kit did occur [Posadas E, unpublished data].
Combined molecular targeted therapeutics with immunotherapy may, in future, become part of the standard care for ovarian cancer. The ability to identify suitable markers of disease would greatly benefit the application of immunotherapy by improving the therapeutic index and cost efficacy [55]. New blood-based analyses using proteomic TOF MS and high-content screening offer supporting evidence of the quantitative and predictive capacities of these strategies. They could serve as surrogates for disease and response, enabling for future advances in the development of novel therapies in the adjuvant and neoadjuvant setting if validated [55].
Executive summary
Ovarian cancer is commonly diagnosed at an advanced stage and is the most deadly gynecologic malignancy in the USA (30–40% 5-year survival rate). In contrast, early stage disease is curable in 80–90% of patients, emphasizing the need for earlier detection methods.
Over the past 30 years, advances in surgical techniques, chemotherapy and supportive care all have contributed to the steady increase in the median survival of women with advanced stage ovarian cancer; however, there has been no substantial improvement in cure rate.
Current screening methods lack the necessary sensitivity and specificity to provide accurate screening, prompting investigators to discover new alternatives to aid in the early diagnosis and treatment of ovarian cancer.
Proteomics characterizes the information flow within the cell and organism through protein pathways and networks.
Basic and clinical proteomics research has contributed to the advancement of proteomic techniques, including laser capture microdissection, mass spectrometry and protein microarrays.
Laser capture microdissection is a tool used to procure selected cell samples from tissue specimens, preserving the molecular composition and architecture of the cells and allowing for independent analysis of the tumor and its local microenvironment.
Mass spectrometry is a high-throughput tool yielding massive datastreams characterizing the targeted proteome. Mining this information can lead to the discovery of new biomarkers.
Protein microarrays examine protein expression and activation states, giving a functional view of protein networks. Two types of protein arrays are commonly used, antibody arrays and tissue lysate arrays.
Four key characteristics of biomarkers include: sufficient sensitivity, sufficient specificity, the availability of an effective intervention and ease of the test.
Limitations of the cancer antigen (CA)-125 test have led to the pursuit of better serum biomarkers for ovarian cancer detection, both for early diagnosis and monitoring of recurrence.
New approaches to biomarker-based proteomic ovarian cancer screening involve the use of composite biomarkers and protein fingerprints generated by mass spectrometry.
The ability of proteomic technology to obtain a molecular profile or signature of an individual tumor may be incorporated into current clinical practice and aid in the personalized selection of combinatorial therapeutics to tailor treatment to the individual patient.
The evolution of proteomics is of major importance to the drug development process and has led to the identification, development and approval of a number of targeted therapeutic agents.
There are several challenges to consider before clinical proteomics is incorporated into current clinical practice, including the current translational infrastructure, patent and regulatory barriers and public–private partnerships [56]. The discovery of new classes of therapeutic drugs, aimed at novel protein targets, has declined in recent years. Overcoming these barriers by providing funding for the performance of validation studies conducted in multidisciplinary research settings, improving communication and outreach efforts to other interested communities in the public policy arena, and bringing new protein tests to commercial feasibility and use has been recommended.
Conclusion
Despite significant surgical and chemotherapeutic advances, the 5-year survival rate for ovarian cancer has increased only slightly from 30 to 40% over the past 20 years [57]. This has led several investigators to seek new techniques to aid in the early detection of ovarian cancer and discover new treatment modalities. The novel proteomic technique of MS has yielded several biomarkers for ovarian cancer that may be of significant clinical importance, either alone or in conjunction with CA-125 based testing. As with any diagnostic test, however, well powered and executed trials following completion of quality control and assurance testing are required before introduction to clinical practice [58]. The raw data of MS databases and high-throughput screening results must be broadly available. Proteomic techniques are not solely limited to diagnostic testing. TLAs are being used in exploratory investigations in clinical trials to provide validation of several molecularly targeted therapeutic agents. The vision of personalized medicine has broadened our horizons and emphasized the uniqueness of our individual genetic diversity. The willingness to overcome the barriers to the advancement of the field of proteomics will ensure continued discovery and progress.
Future perspective
Within the next 5–10 years, significant advances in the field of proteomics will enable its application in clinical medicine to have a profound impact on the way in which patients with ovarian cancer are both diagnosed and treated. The ultimate goal of these changes is threefold:
To detect ovarian cancer before it has progressed to advanced stage where cure is rare
To analyze the unique proteomic profile in an individual ovarian cancer in order to select combinatorial molecular therapies targeted to that tumor
To use proteomic techniques for real-time monitoring of the efficiency of a therapy for its specific molecular pathway [8]
In addition, it will be possible to assess toxicity at the molecular level before damage has occurred (Figure 2)
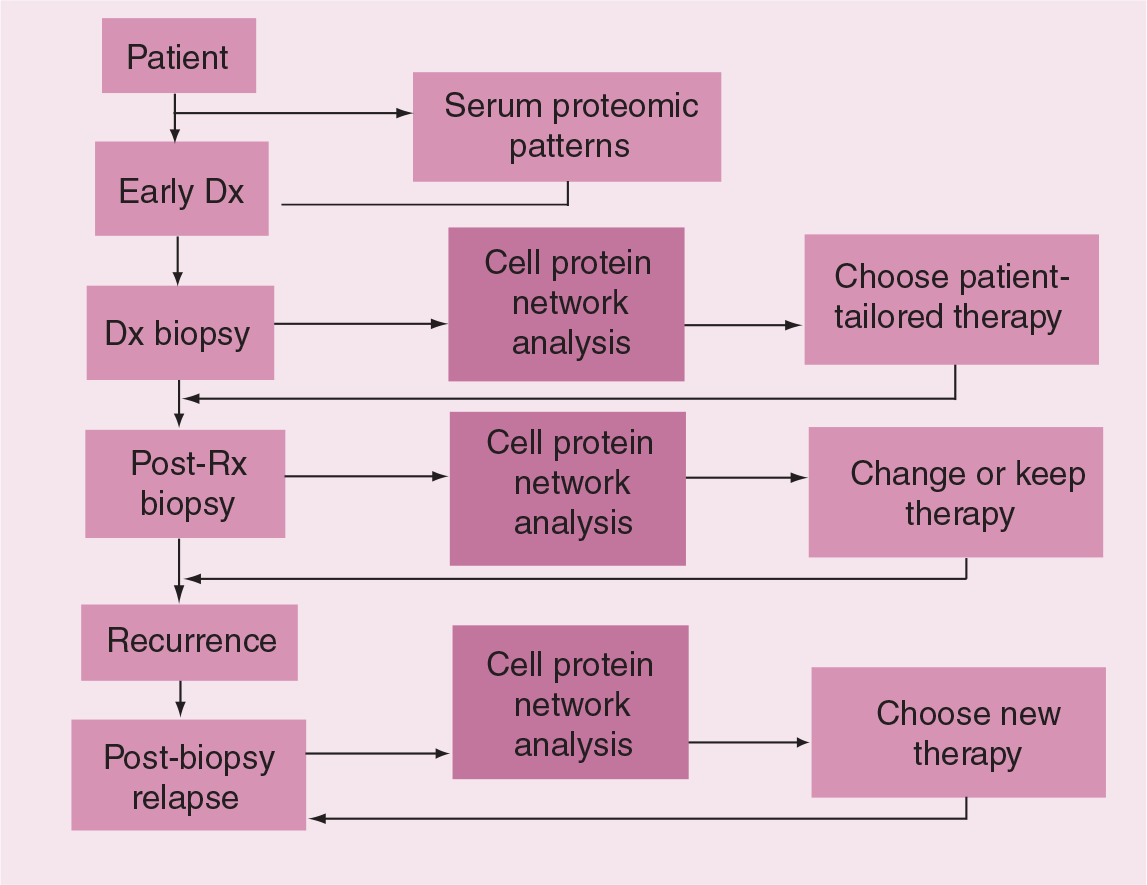
Schematic of personalized molecular therapy.
Proteomics has employed two techniques for disease detection: pattern recognition of unknown proteins and measurement of identified biomarkers. It is anticipated that within the next few years, these tests will reach the very high levels of specificity necessary for their application as a screening method [25]. Combinations of analytes have already yielded improvements in the ability to detect disease-free and cancer patients, reaching a sensitivity and specificity of 95% [59]. However, as past examples have illustrated, further advances are requisite upon standardization of how samples are handled, experiments are conducted and results analyzed between institutions before validation can occur. Universal accepted standards will need to be adopted by the field of proteomics for this technology to be properly used and optimally interpreted.
Proteomics will play a pivotal role in monitoring the efficacy of these drugs in patients in this age of molecularly targeted therapies. Currently, selected clinical trials incorporate TLAs to evaluate how therapies affect their purported molecular targets. This technique may be routinely used in clinics in the future to measure patient responsiveness to specific molecular interventions, as well as to assess early toxicity before damage occurs. In addition to helping physicians select the most effective therapy for a specific patient, cross-talk between molecular pathways will also be revealed, thus allowing for identification of new targets and increasing the repertoire of anticancer therapies.
