Abstract
CXCL12 (SDF-1), which binds CXCR4, is involved in several physiological and pathophysiological processes. In heart, this axis seems to play a key role in cardiogenesis and is involved in the neovascularization of ischemic tissues. Rats have three known CXCL12 mRNA isoforms, of which only α and γ are present in the normal heart. However, little is known about CXCL12 protein expression and localization. We investigated the pattern of protein expression and the localization of both CXCR4 and CXCL12 in the heart, using isolated cardiomyocytes and a rat myocardial infarction model. Western blots showed that cardiomyocytes contained a specific 67-kDa CXCR4 isoform and a 12-kDa CXCL12 isoform. Confocal and electron microscopy clearly showed that CXCR4 was present at the plasmalemma and CXCL12 in continuity of the Z-line, in the proximal part of T-tubules. In conclusion, we provide the first description of the expression and fine localization of CXCR4 and CXCL12 proteins in normal rat heart and cardiomyocytes. These results suggest that the CXCL12/CXCR4 axis may be involved in cardiomyocyte calcium homeostasis regulation. Our work and the well-known chemoattraction properties of the CXCL12/CXCR4 axis highlight the importance of deciphering the function of this axis in both normal and pathological hearts.
CXCL12, also known as stromal cell-derived factor-1 (SDF-1), interacts with two seven-transmembrane domain G protein-coupled receptors, CXCR4 and CXCR7, better known as the orphan receptor RDC1 (Balabanian et al. 2005; Burns et al. 2006). The physiological and/or physiopathological function of the CXCL12/CXCR7 interaction remains unclear, but CXCL12/CXCR4 interaction has clearly been shown to form a major CXC-chemokine/receptor axis. This axis plays a key role in many biological processes including hematopoiesis, cardiogenesis, vasculogenesis, neuronal development, and immune cell trafficking (Kucia et al. 2004). CXCL12 and CXCR4 have also been implicated in pathological conditions such as cancer, human immunodeficiency syndrome, inflammatory processes, and tissue ischemia (Juarez et al. 2004; Ceradini and Gurtner 2005). CXCR4 was recently identified as a new drug target for preventing HIV entry and cancer development (Rusconi et al. 2004; De Clercq 2005).
The rat CXCL12 gene encodes three splice variants, α, β, and γ. αCXCL12 transcripts are produced mostly in the liver, spleen, kidney, heart, brain, and lung; βCXCL12 transcripts are produced in the liver but also in the kidney, spleen, and thymus, whereas γCXCL12 mRNAs predominate in the heart, lung, brain, and peripheral nerve system (Gleichmann et al. 2000; Pillarisetti and Gupta 2001). These splice variants encode three protein isoforms with a common signal peptide sequence: αCXCL12 (89 amino acids, AA), βCXCL12 (93 AA), and γCXCL12 (119 AA). To our knowledge, detection of the γCXCL12 protein has never been reported in normal heart tissue. Regarding CXCR4, Sloane et al. (2005) showed that this receptor is a heterogeneous protein and that its expression pattern depends on cell type. This diversity was attributed to CXCR4 dimerization and/or posttranslational modifications such as N-glycosylation, tyrosine sulfation, or serine chondroitin sulfation.
Little is known about the localization of CXCL12 and CXCR4 proteins in pathological heart tissues. In cases of heart failure, immunohistochemical labeling has demonstrated the presence of CXCL12 and CXCR4 in the vessels and cells of the infarcted and border zones in both humans and mice (Damas et al. 2000; Abbott et al. 2004; Yamani et al. 2005). These studies have shown that the CXCL12/CXCR4 axis may facilitate progenitor cell therapies designed to improve cardiac function. In this therapeutic context, it would be particularly useful to determine whether this chemokine/receptor axis does exist in endogenous cardiac cells, to improve our understanding of the in situ functioning of this axis in the heart.
We aimed to determine, for both the CXCL12 and CXCR4 proteins, the pattern of isoform expression and fine subcellular localization in both normal rat heart and in cardiac myocytes. For this purpose, we combined Western blot analysis with high microscopic resolution methods (confocal laser and transmission electron microscopy) that allow the determination of bidimensional topographic distribution of a protein. We found that CXCR4 was present on the plasmalemma of cardiac myocytes. We also detected two CXCL12 proteins in heart and demonstrated, in cardiac myocytes, specific expression of the 12-kDa CXCL12 protein, most probably corresponding to the γ isoform, located in the continuity of the Z-line, in the proximal part of the T-tubules.
Materials and Methods
Experimental Animals and Isolation of Cardiac Myocytes
Animal investigations were performed according to institutional guidelines, in compliance with international law and policies (EEC Council Directive 86/609, OJ L358, 1 Dec. 12, 1987; Guide for the Care and Use of Laboratory Animals, US National Research Council, 1996).
Ten-week-old adult male Wistar rats (Janvier; Le Genest St Isle, France) were anesthetized by IP administration of pentobarbital (50 mg/kg; Abbott, Rungis Cedex, France). Fifteen hearts were either frozen in liquid nitrogen for immunoblotting and immunofluorescence analysis or fixed for transmission electron microscopy. Isolated myocytes were obtained from five hearts, as previously described (Ait-Mamar et al. 2005). Cells were used frozen for immunoblotting or fixed for immunofluorescence.
Myocardial infarction was induced by left coronary artery ligation in 10-week-old male Wistar rats, as previously described (Michel et al. 1988). Four months after surgery, this procedure infarcted ∼30% to 40% of the left ventricle (LV) and produced a LV dysfunction (Milliez et al. 2005). For each failing heart, the infarcted scar and the non-infarcted tissue of the left ventricle free wall were frozen separately in liquid nitrogen for immunoblotting and histological and immunofluorescence studies. One complete infarcted left ventricle free wall was frozen for transverse section to illustrate the infarct scar.
Protein Extraction and Immunoblotting
For total protein extraction, cardiac samples (n=10) or isolated myocytes were homogenized (10% w/v) in buffer 1 (50 mM Hepes/Tris (pH 7.4), 300 mM sucrose, 1 mM EGTA, 0.1 mM phenylmethylsulfonyl fluoride, 10−3 mM pepstatin, 10−2 mM leupeptin, 1 mM o-phenanthroline, 2.5 mM Na3VO4, and 10−2 mM NaF and stored at −80C.
For fractional protein extraction, frozen isolated myocytes were homogenized in buffer 2 (50 mM Hepes/Tris (pH 8.0), 5 mM MgCl2, 1 mM EDTA, 300 mM sucrose, 1 mM iodoacetamine, 0.2 mM AEBSF, 10−2 mM aprotinin, 10−2 mM leupeptin, 10−3 mM pepstatin, 2 mM phenylarsine oxide, 2.5 mM Na3VO4, and 2.5 mM o-phenanthroline). Homogenates were then centrifuged at 20,000 × g for 20 min at 4C, and the resulting supernatant (S1, cytosolic fraction) was stored at −80C. The membrane pellet (P1) was resuspended in buffer 2 supplemented with 150 mmol/liter NaCl and 0.1% Triton X-100 and centrifuged again. The S2 supernatant (Triton-soluble membrane proteins) and the P2 pellet (Triton-insoluble membrane proteins) were stored at −80C.
Samples were subjected to SDS-PAGE (15% and 10% acrylamide gels for CXCL12 and CXCR4, respectively) and transferred to PVDF membranes. CXCL12 was detected using a mouse monoclonal anti-CXCL12 antibody made by immunizing BALB/c mice with the SDF-1-derived peptide KPVSLSYRSPSRFFC (clone K15C, 1/500; generously provided by Dr. F. Arenzana) (Amara et al. 1999). CXCR4 was detected with a goat polyclonal anti-CXCR4 antibody raised against a peptide mapping at the C-terminus of fusin of human origin (1/5000; Santa-Cruz Biotechnology, Tebu, France). Blots were then treated with secondary peroxidase-conjugated goat anti-mouse (1/500,000; Sigma, St Quentin Fallavier, France) or rabbit anti-goat (1/500,000; Sigma) antibodies. Peroxidase activity was detected using the ECL-Advance kit and Hyperfilm ECL (Amersham; Orsay, France). Specificity of the primary antibodies used was checked by probing membranes with the secondary antibodies alone and by preincubating the primary antibodies with either immunizing peptide (CXCR4) or the protein (CXCL12).
Histological and Immunohisto/cytofluorescences Studies
Ventricles were rapidly excised and rinsed in PBS. To reveal the aspect and distribution of the different cell types present in tissue section, we performed a hematoxylin/eosin coloration. Interstitial collagen was stained with picrosirius red F3BA on three successive sections.
Indirect double-immunofluorescence staining was carried out on 8-μm frozen ventricular sections or on rat ventricular myocytes fixed in 4% formaldehyde (Rucker-Martin et al. 2002). Samples were incubated overnight with a solution containing goat anti-CXCR4 (1/50; Santa-Cruz Biotechnology) and mouse anti-sarcomeric α-actinin (clone EA-53, 1/200; Sigma) antibodies or rabbit anti-ryanodine receptor (RyRs, 1/50; generously provided by Dr. I. Marty) (Hatem et al. 1997) and mouse anti-CXCL12 raised against the 20-89 amino acid sequence of human SDF-1 alpha (clone 79018, 1/50; R&D Systems Europe, Lille, France) antibodies. Desmin was revealed using a rabbit anti-desmin antibody (1/20; MP Biomedicals Europ/Cappel, Vannes, France). The following secondary antibodies, anti-rabbit IgG coupled to Alexa Fluor 488, anti-goat IgG coupled to Alexa Fluor 488, and anti-mouse IgG coupled to Alexa Fluor 594, were purchased from Molecular Probes (Invitrogen; Cercy Pontoise, France) and used at a 1/100 dilution. F-actin was identified using Alexa Fluor 488-phalloidin (1/200; Molecular Probes, Invitrogen). For negative controls, we omitted incubation with primary antibodies.
Confocal Laser Microscopy
Images were acquired with a multitracking laser scanning confocal microscope (Zeiss LSM-510, Carl Zeiss SAS; Iena, Germany), as previously described (Ait-Mamar et al. 2005). Differential interference contrast images were collected under monochromatic light. Fluorochromes were detected sequentially at 488 nm and 543 nm. Images were coded green (Alexa Fluor 488) and red (Alexa Fluor 594). We used a X63 oil-immersion objective (numerical aperture 1.4), giving a resolution of 100 nm in the x,y plane and 300 nm along the z-axis (pinhole 72 μm). An optical section (0.8-μm thick) was observed every 0.38 μm. We determined the distribution of CXCR4, CXCL12, and F-actin by analyzing fluorescence intensity profiles with LSM Image Browser version 3 (Carl Zeiss).
Quantification of Fluorescence Overlap
We analyzed CXCL12 and RyRs colocalization by fluorescence intensity profiles (Bolte and Cordelières in press). We studied six individual cardiac myocytes and analyzed 20 ± 5 successive optical sections acquired in each image stack. Overlap in the distributions of CXCL12 and RyRs in the x,y plane was determined by analyzing fluorescence intensity profiles with LSM Image Browser version 3 (Carl Zeiss). Positive structures were evaluated by the Costes approach (Bolte and Cordelières in press) using threshold values of 1683 ± 189 (CXCL12) and 1627 ± 181 (RyR) pixel units on a grayscale of 0 to 4095. A scatterplot of the individual pixels was generated from each pair of images. Overlapping CXCL12 and RyR pixels were characterized by an overlap coefficient value of 1. The number of overlapping pixels was expressed as a proportion of the total number of positive pixels for each labeling (CXCL12 or RyR).
Transmission Electron Microscopy
Small blocks (1 mm3) of tissue were fixed by immersion in 4% paraformaldhehyde-0.2% glutaraldehyde in 0.1 mol/liter Sorensen phosphate buffer, pH 7.4, (1 hr), postfixed in 4% paraformaldehyde (2 hr), and then in 0.4% paraformaldehyde-0.2 mol/liter sucrose, overnight at 4C. Samples were dehydrated in a graded series of ethanol solutions and embedded in LR White resin. Thin sections (80 nm) were stained with mouse anti-CXCL12 antibody (clone 79018, 1/10) and then with anti-mouse IgG coupled to 10-nm gold particles (1/50; British Biocell International, Tebu, France). Sections were examined using a Philips CM12 electron microscope at 80 kV (Philips; Suresnes, France).
Results
Rat Cardiac Myocytes Express the CXCR4 Receptor
In rat atria and ventricles, the anti-CXCR4 antibody detected a major immunoreactive 67-kDa protein, together with two additional bands at 93 and 130 kDa (Figure 1A, Lanes 1 and 2). Only the 67-kDa band was recovered from isolated rat ventricular myocytes (Figure 1A, Lane 3), indicating that the 67-kDa isoform was the only isoform detectable in myocytes.
We localized CXCR4 on heart ventricular sections by immunofluorescence labeling (Figure 1B). To reveal the aspect and distribution of endothelial cells and cardiac myocytes present in the tissue section, we performed a hematoxylin/eosin coloration (Figure 1B, panel 1). A strong fluorescence labeling with the polyclonal anti-CXCR4 antibody was observed in both ventricular myocytes characterized by α-sarcomeric actin expression (Figure 1B, panels 2 and 3) and in the endothelial cells of the endocardium (Figure 1B, panel 3).
Confocal analysis after immunofluorescence staining identified CXCR4 at the periphery of rat ventricular myocytes along the plasmalemma (Figure 1C, panels 1-3). This localization was confirmed by an analysis of fluorescence intensity distribution. Along an axis perpendicular to the myocyte, fluorescence peaked regularly every 15.1 ± 2.0 mm, corresponding to the mean width of rat ventricular myocytes (Figure 1D).
Thus, cardiac myocytes express a specific 67-kDa CXCR4 receptor on the plasmalemma.
Rat Cardiac Myocytes Essentially Express a 12-kDa CXCL12 Protein
Western blot analyses on whole rat heart protein extract with the anti-CXCL12 antibody (clone K15C) identified two specific bands at 12 and 8 kDa in the atria and ventricles (Figure 2A, Lanes 1 and 2, respectively). We determined the subcellular distribution of these CXCL12 isoforms in rat ventricular myocytes by fractional protein extraction. The 12-kDa CXCL12 protein was detected in cytosolic fractions (S1) but not in Triton-soluble (S2) or Triton-insoluble (P2) membrane protein fractions (Figure 2A). The 8-kDa CXCL12 protein was not detected in any of the fractions. These data clearly show that, in cardiac myocytes, CXCL12 was present solely as a 12-kDa protein in the cytosol. This protein probably corresponds to the γCXCL12 isoform (Gleichmann et al. 2000; Yu et al. 2006), whereas the 8-kDa band probably corresponds to the α/βCXCL12 isoform (NCBI nucleotide accession numbers NM_022177 and NM_001033882) (Yu et al. 2006).
We localized CXCL12 on heart ventricular sections by immunofluorescence labeling to confirm that this protein was present in various heart cell types. Distribution of endothelial cells of the endocardium and ventricular myocytes present in the tissue section was first revealed by hematoxylin/eosin coloration (Figure 2B, panel 1). At low magnification, a fluorescence labeling with the monoclonal anti-CXCL12 antibody was observed at the plasma level of ventricular myocytes identified by desmin expression (Figure 2B, panel 2). Endothelial cells were strongly stained with the anti-CXCL12 antibody (Figure 2B, panel 3).
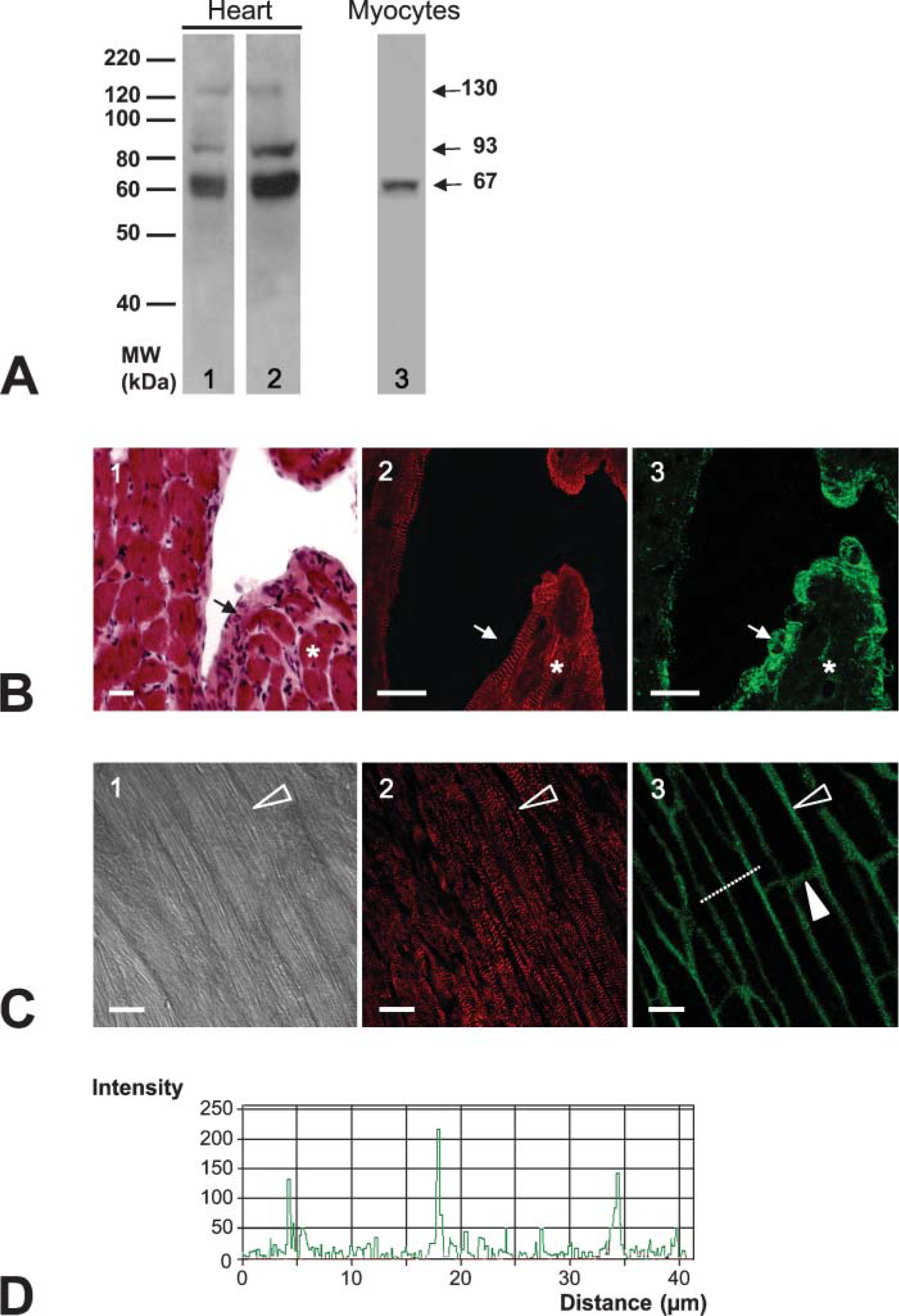
Expression and localization of CXCR4 in normal rat heart and isolated myocytes.
To demonstrate specific expression of the 12-kDa CXCL12 protein in cardiac myocytes, we compared CXCL12 expression in infarct scar area (ISA) and non-infarcted area (NIA) in rat heart 4 months after myocardial infarction. At this time the infarct scar was clearly deprived in cardiac myocytes in contrast with the NIA (Pfeffer and Braunwald 1990; Milliez et al. 2005). These two areas, NIA and ISA, are illustrated in Figure 3A (upper panel). Picrosirius red staining showed an extensive distribution of collagen in the ISA with only very few myocytes (poor yellow coloration) as compared with the NIA (Figure 3A, lower panels). We first localized CXCL12 in these two ventricular areas by confocal microscopy at high magnification after immunofluorescence staining. In the NIA, CXCL12 was present at the myocyte plasmalemma with a regular striated pattern of staining corresponding to the sarcomere (Figure 3B, left panels). In the ISA, all scar tissue and endothelial cells bordering the vessel lumen were labeled by the anti-CXCL12 antibody (Figure 3B, right panels). Western blot analysis showed that only the 8-kDa CXCL12 protein was detected in the ISA, consistent with the absence of myocytes in this region, whereas both the 8- and 12-kDa CXCL12 proteins were present as expected in the NIA (Figure 3C). The smallest CXCL12 isoform was also more strongly expressed in the ISA than in the NIA. Absence of the 12-kDa CXCL12 protein most probably corresponding to the γ isoform in the infarct scar supports our data obtained in normal isolated cardiac myocytes.
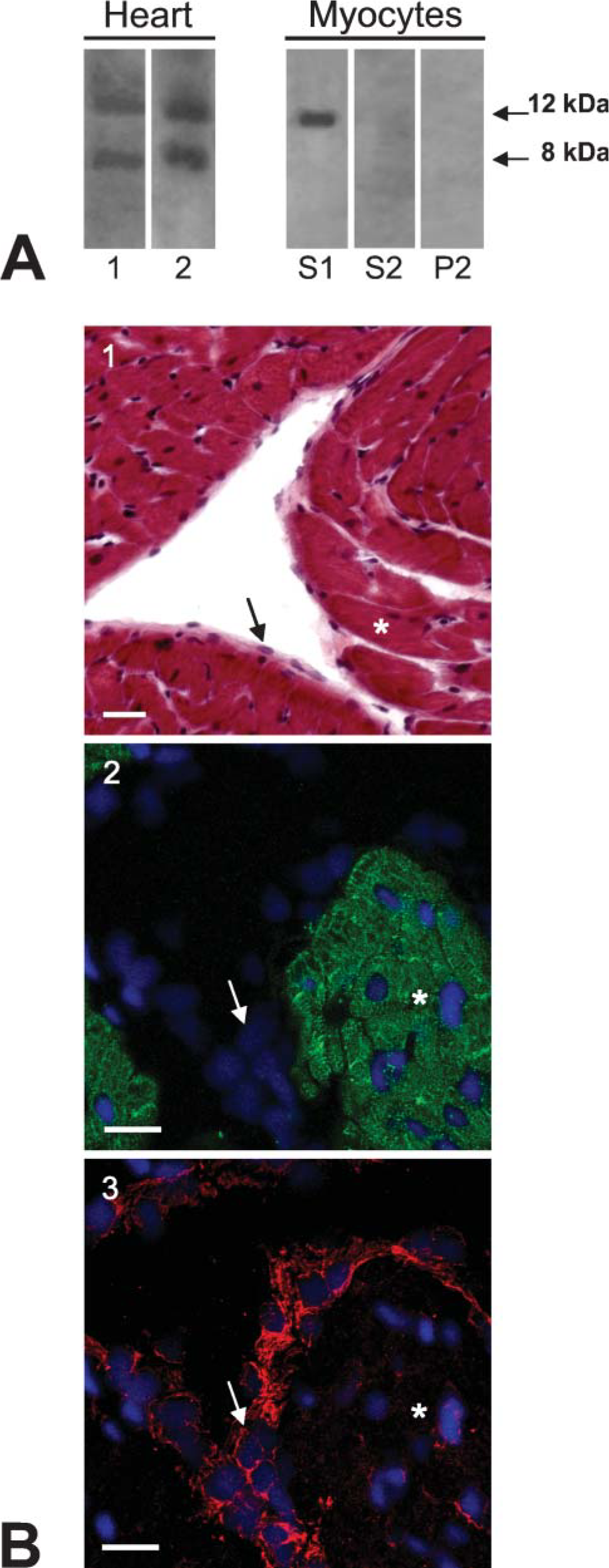
Expression and localization of CXCL12 protein in normal rat heart and isolated myocytes.
Together these results clearly demonstrate that cardiac myocytes specifically express the γCXCL12 protein.
CXCL12 Is Present at the Level of T-tubules in Rat Cardiac Myocytes
We analyzed the subcellular expression of CXCL12 in cardiac myocytes further, using both confocal laser microscopy (CLM) with high z-axis spatial resolution and transmission electronic microscopy (TEM). These techniques enable high-resolution immunocytochemical analysis of the two-dimensional topographic distribution of proteins. For CLM, ventricular cryosections were immunostained with the polyclonal anti-CXCL12 antibody, and F-actin was detected with phalloidin. For each sample, three successive longitudinal planes (superficial, intermediate, and internal) and a transverse plane were defined. In the superficial plane (Figure 4A), only CXCL12 was detected in myocytes, displaying well-aligned and regular striated labeling (every 2.27 ± 0.47 μm). In the intermediate plane (Figure 4B), located ∼2.5 μm below the membrane surface, myocytes displayed regular striated CXCL12 staining and the typical hatched pattern of F-actin staining corresponding to the Z-line. The two types of fluorescence showed the same periodicity across several striations: 1.98 ± 0.07 μm and 2.05 ± 0.44 μm for CXCL12 and F-actin, respectively. This overlapping of the fluorescence intensity curves suggests that CXCL12 is located close to actin in this plane. In the internal plane (Figure 4C), CXCL12 was detected only at the cell periphery, whereas F-actin was detected throughout the myocyte. In the transverse plane (Figure 4D), staining for CXCL12 and F-actin was continuous with no overlap, as confirmed by fluorescence intensity distribution along one Z-line. All these results suggest that CXCL12 is located at the plasmalemma level of cardiac myocytes, in continuity with the Z-line.
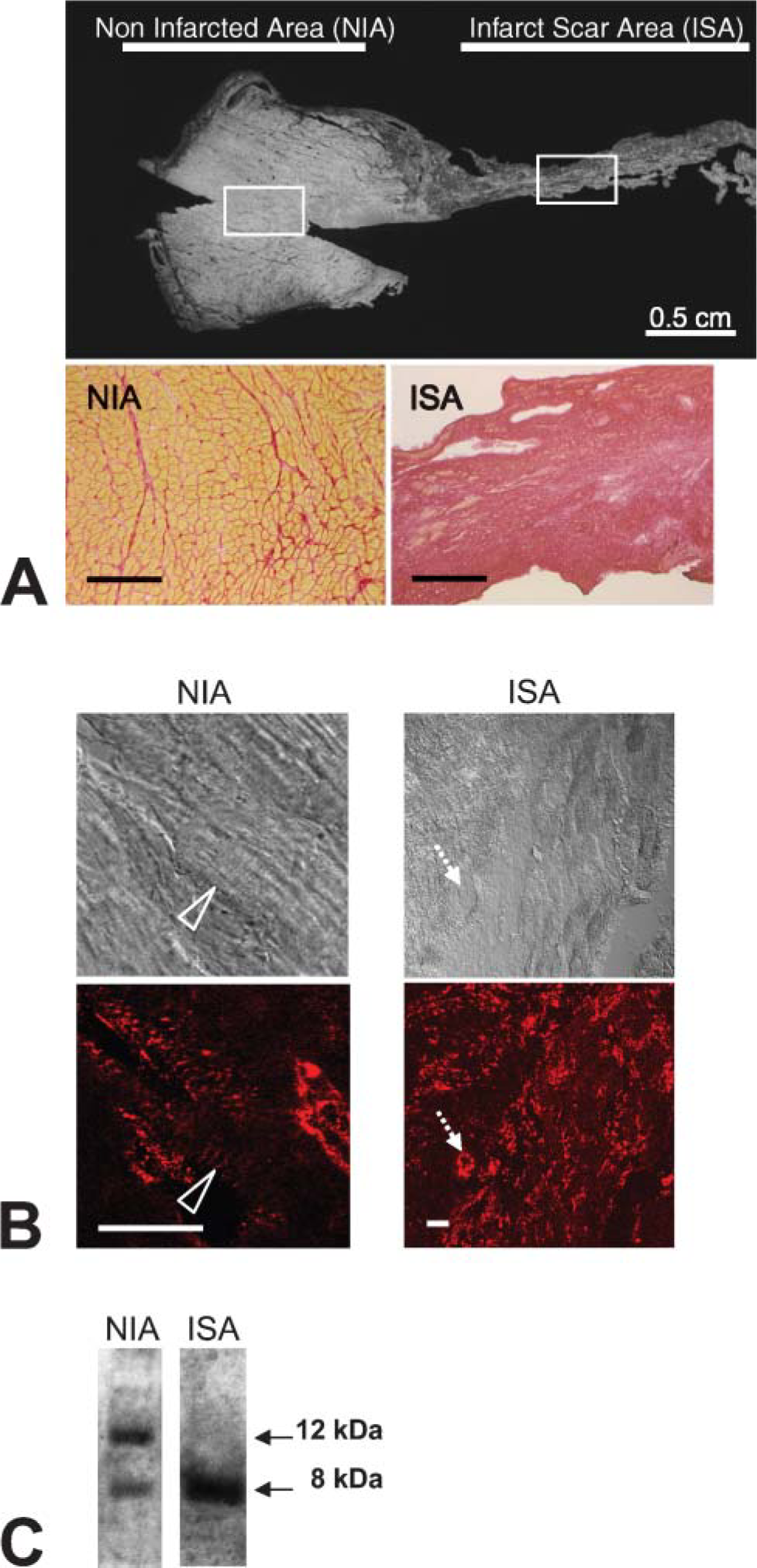
Expression and localization of CXCL12 protein in myocardial infarcted rat heart: comparison between non-infarcted area (NIA) and infarct scar area (ISA).
Several relevant structures near the Z-line could have been involved; we focused on the sarcoplasmic reticulum and carried out double immunolabeling for both RyRs and CXCL12 on isolated cardiac myocytes (Figure 5, A1). In these cells, RyRs displayed the longitudinal and transverse distribution corresponding to the T-tubule pattern (Figure 5, A2), as previously reported (Thomas et al. 2003). Punctuate staining for CXCL12 was observed along the plasmalemma (Figure 5, A3). An analysis of CXCL12 and RyR staining, based on the distribution of fluorescence intensities, demonstrated a mean overlap of 590 ± 27 nm (in yellow on the histogram), well above the limits of CLM resolution (Figure 5B). The scatterplot of individual pixels from paired images showed that 86.7% of red pixels corresponding to CXCL12 labeling overlapped with 6.7% of green pixels corresponding to RyRs labeling (Figure 5C). This strongly suggests that the cardiac myocyte CXCL12 pool is located close to RyRs.
To refine ultrastructural domain of CXCL12 expression, TEM with CXCL12 gold immunolabeling was carried out. Several gold particles were detected in T-tubules in front of the Z-line, near the plasmalemma mitochondrial pool (Figure 5D). No labeling was observed deeper in the myocyte (data not shown).
These confocal and electron microscopy data clearly indicate that CXCL12 is located in the proximal part of the T-tubules.
Discussion
We report for the first time the expression and subcellular localization of CXCR4 and CXCL12 proteins in normal rat cardiac myocytes. γCXCL12 protein was identified as the only isoform present in this cell type.
We detected CXCR4 as a major 67-kDa band in normal rat whole heart and isolated cardiac myocytes. This molecular mass is higher than predicted from the primary amino acid sequence (NBCI nucleotide accession number: AAL47855) and may correspond to a particular isoform of CXCR4. However, this result is not surprising given the heterogeneous pattern of expression previously reported for this receptor (Sloane et al. 2005). CXCR4 localization was previously reported in cardiac myocytes using immunochemical staining and transmitted light microscopy (Damas et al. 2000). In our study, we specified the expression of CXCR4 in cardiac myocytes using immunofluorescence labeling and confocal microscopy and showed that this receptor was located principally at the plasmalemma, like all other known receptors with seven transmembrane domains.
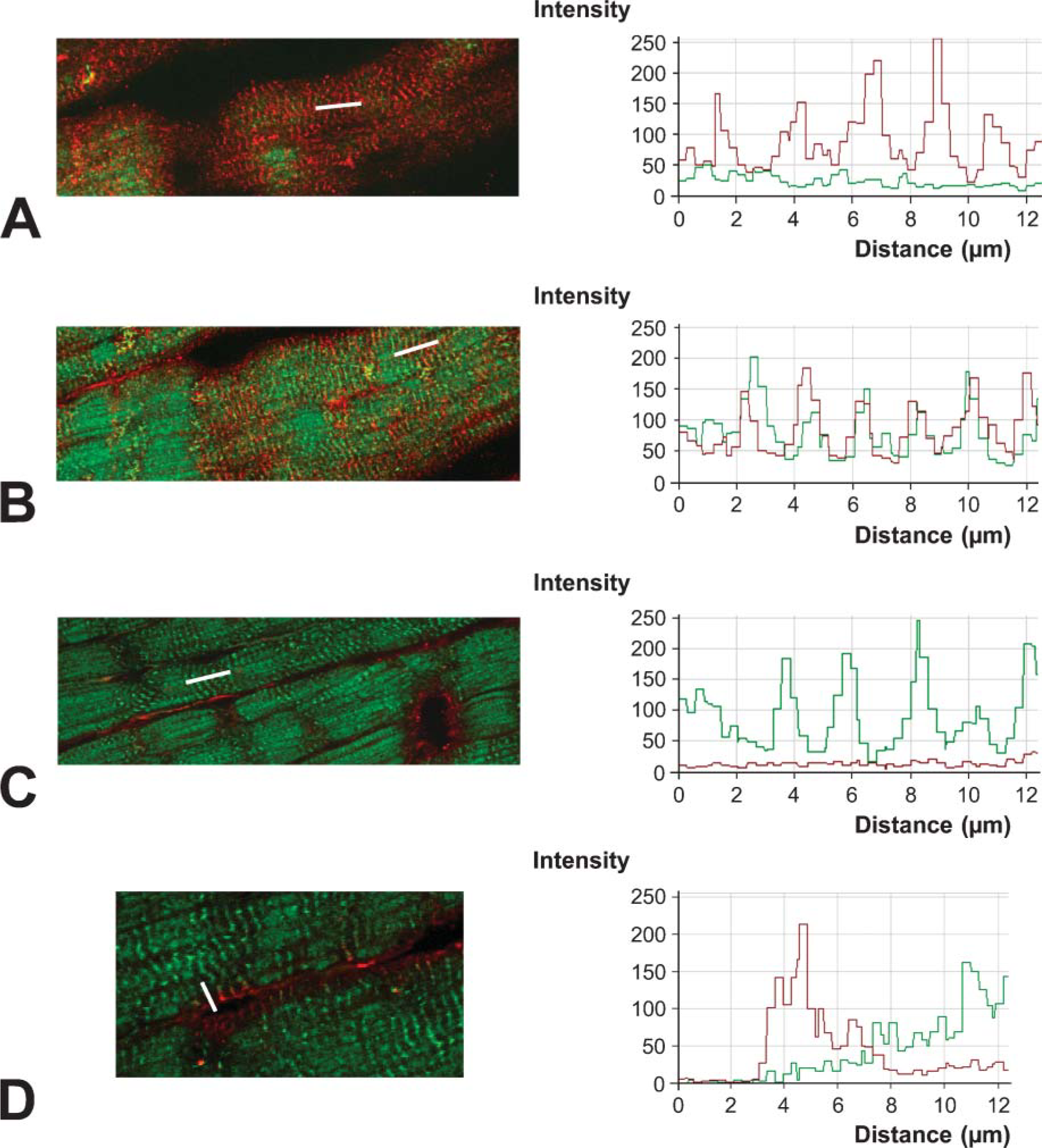
Localization by confocal laser microscopy of CXCL12 in normal ventricle sections.
We detected the α/β and γCXCL12 proteins in normal rat heart, consistent with previous mRNA studies (Gleichmann et al. 2000; Pillarisetti and Gupta 2001). Our immunofluorescence experiments identified two major cell types expressing CXCL12 in normal heart: cardiac myocytes and endothelial cells. In infarcted scar, CXCL12 was expressed only in endothelial and inflammatory cells. These observations are consistent with a potential differential expression of CXCL12 isoforms in the heart according to cell type (Belperio et al. 2000; Ceradini et al. 2004). Our immunoblotting results provide the first demonstration that cardiac myocytes express essentially the γCXCL12 protein. In addition, γCXCL12 transcripts were also detected by quantitative RT-PCR in isolated ventricular myocytes (data not shown), suggesting that γCXCL12 may be secreted by cardiac myocytes.
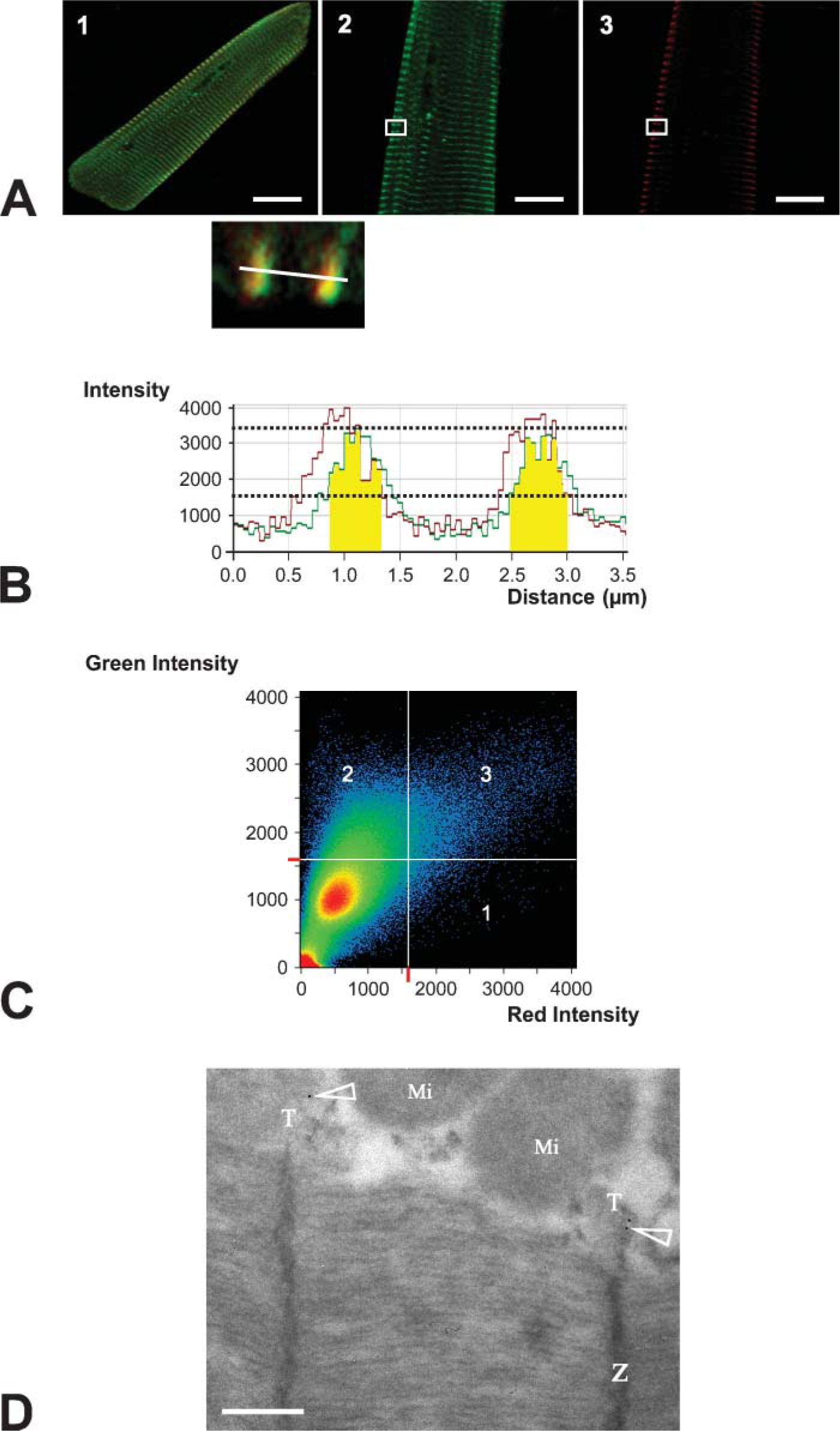
Structural and ultrastructural localization of CXCL12 in rat ventricular myocytes.
Our confocal and electron microscopy studies showed that the CXCL12 pool was present in the continuity of the Z-line, close to some of the RyRs and in the proximal part of the T-tubular system of the cardiac myocytes. The T-system consists of both invaginations of the cardiac sarcolemma, mostly located at the Z-line, and longitudinal extensions. This complex system of transverse and longitudinal branching tubules extends deep into the cytoplasm and is found in close apposition with the sarcoplasmic reticulum. Key proteins involved in regulating calcium homeostasis, particularly during excitation-contraction coupling, are known to be concentrated in these structures (Brette and Orchard 2003). For this reason it will be of particular interest to study a potential colocalization of CXCL12 or its receptor with other proteins involved in excitation-contraction coupling (dihydropyridine receptors, Serca ATPase, Na+/Ca2+ exchanger, …). Unfortunately, in our study, commercial polyclonal antibodies directed against these different candidate proteins did not give satisfactory results. Our results, nevertheless, raise questions about the potential role of the CXCL12/CXCR4 axis in cardiac myocyte calcium homeostasis. This issue was previously addressed by the group of Schecter (Pyo et al. 2005), who observed a decrease in calcium transients and fractional shortening in adult rat cardiac myocytes in the presence of exogenous CXCL12 and increasing extracellular [Ca2+]. Both of these findings and those presented here suggest that CXCL12 may modulate calcium homeostasis and contraction of cardiac myocytes via an autocrine and/or paracrine mechanism. We suggest that this new role of the CXCL12/CXCR4 axis should be added to its well-known role in chemoattraction in the heart, in both physiological and physiopathological contexts. During cardiac or hindlimb muscle ischemia, CXCL12 recruits circulating tissue-committed CXCR4-positive stem cells, thereby contributing to vascular and tissue remodeling (Askari et al. 2003; Yamaguchi et al. 2003; Wojakowski et al. 2004). Improvements in cardiac function have been reported following transplantation of cardiac fibroblasts overproducing CXCL12 into infarcted myocardium. This recovery of function has been attributed to beneficial reverse remodeling following CXCL12-induced stem-cell homing (Askari et al. 2003; Wojakowski et al. 2004). We suggest that the increase in CXCL12 levels may also contribute to the recovery of cardiac function via direct effects on cardiac myocytes. In the current state of knowledge, an attractive hypothesis would be to assign a specific role to a given CXCL12 protein isoform. However, to date, it seems too early: first, because the best-known CXCL12 cellular action, namely, chemoattraction, appears to be a common property of all CXCL12 isoforms (Yu et al. 2006) and second, only the αCXCL12 protein had been used to study the different actions of CXCL12 ligand on cellular functions. To answer this major question, specific tools such as recombinant peptides or antibodies directed against the different CXCL12 isoforms will be necessary.
This study describes the expression and localization of CXCL12 and CXCR4 proteins in cardiac myocytes and highlights the value of deciphering the role of the CXCL12/CXCR4 axis in both normal and pathological hearts in more detail. Such work would be useful in at least two therapeutic fields in which the CXCL12/CXCR4 axis is currently being considered as a possible target. First, in the promising field of cell therapy, our data highlight the need to investigate further the functional effects of increase or decrease in the amount of CXCL12 on cardiac myocytes. Second, in the field of HIV and cancer treatment, such studies would facilitate evaluation of the potential secondary effects of CXCR4 antagonists on cardiac function.
Footnotes
Acknowledgements
Audrey Segret and Jeanne Flavigny were supported by a grant from the Ministère de l'Education Nationale, de la Recherche et de la Technologie and the Association Française contre les Myopathies, respectively.
We thank Valérie Nicolas (Service Imagerie-Microscopie Confocale, IFR-141, Châtenay Malabry, France) for expert assistance with confocal microscopy and Christine Longin and Sophie Chat (INRA Unité; GPL-Plateau de microscopie électronique, Jouy-en-Josas, France) for TEM preparation and assistance. We thank Dr. Fernando Arenzana-Seisdedos for kindly providing CXCL12 antibodies and Dr. Isabelle Marty for RyRs antibodies. We thank Marc Derive for technical assistance.
