Abstract
Recombinant human (rh) TNF-related apoptosis-inducing ligand (TRAIL) harbors potential as an anticancer agent. RhTRAIL induces apoptosis via the TRAIL receptors TRAIL-R1 and TRAIL-R2 in tumors and is non-toxic to nonhuman primates. Because limited data are available about TRAIL receptor distribution, we performed an immunohistochemical (IHC) analysis of the expression of TRAIL-R1, TRAIL-R2, the anti-apoptotic TRAIL receptor TRAIL-R3, and TRAIL in normal human and chimpanzee tissues. In humans, hepatocytes stained positive for TRAIL and TRAIL receptors and bile duct epithelium for TRAIL, TRAIL-R1, and TRAIL-R3. In brains, neurons expressed TRAIL-R1, TRAIL-R2, TRAIL-R3 but no TRAIL. In kidneys, TRAIL-R3 was negative, tubuli contorti expressed TRAIL-R1, TRAIL-R2, and TRAIL, and cells in Henle's loop expressed only TRAIL-R2. Heart myocytes showed positivity for all proteins studied. In colon, TRAIL-R1, TRAIL-R2, and TRAIL were present. Germ and Leydig cells were positive for all proteins studied. Endothelium in liver, heart, kidney, and testis lacked TRAIL-R1 and TRAIL-R2. In alveolar septa and bronchial epithelium TRAIL-R2 was expressed, brain vascular endothelium expressed TRAIL-R2 and TRAIL-R3, and in heart vascular endothelium only TRAIL-R3 was present. Only a few differences were observed between human and chimpanzee liver, brain, and kidney. In contrast to human, chimpanzee bile duct epithelium lacked TRAIL, TRAIL-R1, and TRAIL-R3, lung and colon showed no TRAIL or its receptors, TRAIL-R3 was absent in germ and Leydig cells, and vascular endothelium showed only TRAIL-R2 expression in the brain. In conclusion, comparable expression of TRAIL and TRAIL receptors was observed in human and chimpanzee tissues. Lack of liver toxicity in chimpanzees after rhTRAIL administration despite TRAIL-R1 and TRAIL-R2 expression is reassuring for rhTRAIL application in humans. (
A
Repeated rhTRAIL administration to cynomolgus monkeys appeared remarkably safe and non-immunogenic (Ashkenazi et al. 1999; Kelley et al. 2001; Lawrence et al. 2001). It was reassuring that also chimpanzees, with the highest genetic homology with humans, showed no toxicity after rhTRAIL administration (Kelley et al. 2001; Lawrence et al. 2001). The observation that histidine- or Flag-tagged rhTRAIL resulted in apoptosis of cultured human hepatocytes (Jo et al. 2000) and cells in human brain slices (Nitsch et al. 2000) has caused concern about the clinical applicability of rhTRAIL. Lawrence et al. (2001), however, showed that human and cynomolgus monkey hepatocytes are sensitive in vitro to histidine- and crosslinked Flag-tagged versions of rhTRAIL but not to the native sequence, recombinant soluble ligand. Comparable results were obtained in normal human keratinocytes (Qin et al. 2001).
The broad TRAIL expression and the inability of rhTRAIL to induce apoptosis in normal cells suggest that these cells contain mechanisms that protect them from TRAIL-induced apoptosis (LeBlanc and Ashkenazi 2003). One possible mechanism could be restricted TRAIL receptor expression. Five human TRAIL receptors have now been identified. Two of them are death-inducing receptors, TRAIL-R1 (DR4) and TRAIL-R2 (DR5/TRICK2), containing a cyto-plasmic death domain to transmit the apoptotic signal (Chaudhary et al. 1997; MacFarlane et al. 1997; Pan et al. 1997a, b; Schneider et al. 1997; Sheridan et al. 1997; Walczak et al. 1997). Two additional decoy receptors, TRAIL-R3 (DcR1/TRID), and TRAIL-R4 (DcR2/TRUNDD), have either truncated or absent intracellular domains and are unable to transduce the death signal (Degli-Esposti et al. 1997a, b; MacFarlane et al. 1997; Marsters et al. 1997; Pan et al. 1997a,1998; Schneider et al. 1997; Sheridan et al. 1997). The fifth receptor is the soluble osteoprotegerin (OPG), a regulator of osteoclastogenesis (Emery et al. 1998).
TRAIL, TRAIL-R1, TRAIL-R2, TRAIL-R4, and OPG show 84–99% extracellular protein sequence identity between cynomolgus monkey and humans (Lawrence et al. 2001). Moreover, rhTRAIL binds to cynomolgus monkey receptors with an affinity comparable to that for human receptors, demonstrating good crossreactivity. The extracellular domains of all four chimpanzee TRAIL receptors even show 97–99% sequence identity with the human receptors (Lawrence et al. 2001). Because of this high homology and the absence of rhTRAIL toxicity in chimpanzees, knowledge about the balance between TRAIL death and decoy receptors in tissues in human and chimpanzee will further support clinical studies. In addition, more insight may be obtained in the physiological role of TRAIL and its receptors knowing their normal distribution. Until now, only limited data are available about the expression of TRAIL receptors in normal tissues. Several reports have demonstrated mRNA expression of the TRAIL receptors in various human tissues (Chaudhary et al. 1997; Degli-Esposti et al. 1997a, b; MacFarlane et al. 1997; Marsters et al. 1997; Pan et al. 1997a, b, 1998; Schneider et al. 1997; Sheridan et al. 1997; Walczak et al. 1997). The observed mRNA expression, however, does not reveal the cell type in which the receptor is expressed, which can be easily visualized by immunohistochemical (IHC) techniques. Therefore, the aim of the present study was to perform an IHC inventory of TRAIL receptors and TRAIL in relevant normal human tissues and to compare this with chimpanzee tissues.
Materials and Methods
Tissue Collection
Paraffin-embedded normal human tissue specimens were retrieved from the files of the Department of Pathology of the University Hospital Groningen. Tissues were obtained from biopsy and resected material of the liver, kidney, colon, testis, lung, brain, and heart, of which the last two were obtained from autopsy material. Frozen normal chimpanzee tissues were purchased from the Biomedical Primate Research Center (TNO; Rijswijk, The Netherlands). Frozen tissues from kidney, heart, liver, brain, colon, lung, and testis of two chimpanzees were fixed in 10% formalin and paraffin-embedded. Histological classifications were performed on hematoxylin and eosin (H&E)-stained slides and in most cases two or three samples without inflammation were examined for expression of TRAIL and its receptors.
Reverse Transcription Polymerase Chain Reaction (RT-PCR)
Total RNA was isolated by guanidine thiocyanate-phenol-chloroform extraction from frozen pulverized chimpanzee and human liver tissues. The quality of the samples was checked by agarose gel electrophoresis. Before cDNA synthesis, RNA was treated with DNase I (Roche Diagnostics; Almere, The Netherlands). cDNA was synthesized as described by the manufacturer's protocol (Life Technologies; Breda, The Netherlands) using oligo dT primers and MMLV transcriptase. The PCR programs and primers used to amplify TRAIL, TRAIL-R1, TRAIL-R2, and TRAIL-R3 were described elsewhere (Griffith et al. 1998; Rieger et al. 1999; Strater et al. 2002). Primer sequences for GAPDH were 5′-CACCACCATGGAGAAGGCTGG-3′ and 5′-CCAAAGTTGTCATGGATGACC-3′, which resulted in a 200-bp fragment after 24 cycles. To exclude amplification of any remaining genomic DNA, reverse transcriptase was omitted during the reverse transcriptase step. In addition, a negative control without template cDNA was run with every PCR to control for cross-contamination between tubes. No specific PCR signal was detected under these control conditions. The colon carcinoma cell line SW948 was used as a positive control (van Geelen et al. 2003). PCR products were electro-phoresed in a 2% agarose gel in 1 × Tris-borate EDTA buffer and checked for the right size with restriction enzymes.
Western Blotting Analysis
Frozen chimpanzee and human liver tissues were pulverized and dissolved in cold PBS (6.4 mM Na2HPO4, 1.5 mM KH2PO4, 0.14 mM NaCl, 2.7 mM KCl, pH 7.2). After centrifugation at 23,000 × g for 1 min to remove debris, the supernatant was collected, an equal volume of 2 × standard Western blotting sample buffer (0.5 M Tris-HCl, pH 6.8, 4% SDS, 20% glycerol, 0.002% bromophenol blue, 10% 2-β-mercaptoethanol) was added and the samples were boiled for 5 min. Protein concentration was determined according to Bradford (1976). All samples were size-fractionated on SDS-PAGE and transferred onto activated polyvinylidene difluoride (PVDF) membranes (Millipore; Bedford, MA). After blocking for 1 hr in Tris-buffered saline supplemented with 5% milk powder (Merck; Darmstadt, Germany) and 0.05% Tween-20 (Sigma-Aldrich Chemie; Zwijndrecht, The Netherlands), immunodetection of TRAIL, TRAIL-R1, TRAIL-R2, and TRAIL-R3 was performed using the following antibodies: a goat polyclonal IgG specific for TRAIL (1:100, clone K18; Santa Cruz Biotechnology, Santa Cruz, CA), a goat polyclonal IgG specific for TRAIL-R1 (1:500, clone C-20; Santa Cruz Biotechnology), a rabbit polyclonal IgG specific for TRAIL-R2 (1:500; Oncogene Research, Cambridge, MA), and a goat polyclonal IgG specific for TRAIL-R3 (1:250; Calbiochem, Breda, The Netherlands). Binding of these antibodies was determined using horseradish peroxidase (HRP)-conjugated secondary Abs (all from DAKO, Glostrup, Denmark) and visualized with the ECL chemiluminescence kit of Roche Diagnostics. The SW948 cell line served as a positive control for TRAIL-R1 and TRAIL-R2. Soluble rhTRAIL [made according to Ashkenazi et al. (1999)] and TRAIL-R3-Fc (Research and Diagnostic Systems; Uithoorn, The Netherlands) were used as controls for TRAIL and TRAIL-R3 staining, respectively.
Immunohistochemistry
For IHC staining, 3-μm-thick sections were cut from paraffin blocks. Before staining, slides were deparaffinized and rehydrated. For TRAIL-R2, antigen retrieval was performed by microwave treatment of the slides for 8 min at 700 W in 0.01 M citrate buffer, pH 6.0. For TRAIL, TRAIL-R1, and TRAIL-R3, no antigen retrieval was performed. Endogenous peroxidase was blocked with 0.3% H2O2 in PBS (6.4 mM Na2HPO4.H2O, 1.5 mM KH2PO4, 0.14 M NaCl, 2.7 mM KCl, pH 7.8) for 30 min, followed by incubation with avidin and biotin blocking solutions (Vector Laboratories; Burlingame, CA). For TRAIL-R1, TRAIL-R2, and TRAIL-R3, the slides were preincubated with 1% AB serum, 1% bovine serum albumin (BSA; Life Technologies, Breda, The Netherlands) in PBS for 15 min. For TRAIL, 5% normal rabbit serum was used instead of AB serum. The primary antibodies described earlier were applied for 1 hr at room temperature diluted 1:25, 1:100, 1:200, and 1:50 for TRAIL, TRAIL-R1, TRAIL-R2, and TRAIL-R3 respectively, in PBS/1% BSA. After washing with PBS, slides were incubated with a 1:300 dilution of a biotinylated rabbit anti-goat or a swine anti-rabbit antibody (DAKO), followed by addition of streptavidin-conjugated peroxidase (DAKO). Peroxidase activity was visualized by incubating the slides for 10 min in 3,3-diaminobenzidine tetrahydrochloride (DAB; Sigma–Aldrich Chemie) solution (0.05% DAB, 0.1% imadizol, 0.03% H2O2 in PBS). Counterstaining was performed with hematoxylin for 2 min. As negative controls, the primary antibodies were substituted with non-immune normal goat or normal rabbit IgGs. For TRAIL, TRAIL-R1, and TRAIL-R3, IHC was performed in the presence or absence of a 10-fold excess of a corresponding blocking peptide. Positive controls were (tumor) tissue samples, found on previous occasions to stain positive: normal liver, first trimester placenta, and colon or ovarian cancer. All sections were reviewed by two examiners (DCS and HH) and evaluated in a semiquantitative way for intensity of staining: no staining (0), weakly positive staining (1), moderate staining (2), or strong positive staining (3). The localization of staining (nuclear, membranous, or cytoplasmic) was also recorded. In contrast to a previous report (Koornstra et al. 2003), we used intensity of staining to compare the expression of TRAIL and the receptors in different cell types present in each organ.
Results
Detection of TRAIL-R1, TRAIL-R2, TRAIL-R3, and TRAIL mRNA and Protein in the Liver
To validate the specificity of the antibodies used in the IHC staining, additional analyses were performed on chimpanzee and human liver. First, the TRAIL-R1, TRAIL-R2, TRAIL-R3, and TRAIL mRNA expression was evaluated by RT-PCR. Figure 1A shows that both chimpanzee and human liver expressed mRNA of the three TRAIL receptors and TRAIL. Next, the protein expression was checked in the liver by immunoblotting analysis using the same antibodies for TRAIL-R1, TRAIL-R2, TRAIL-R3, or TRAIL detection as used for IHC stainings. The TRAIL antibody is directed against the extracellular carboxy terminus of the TRAIL protein and therefore recognizes both soluble and membrane-bound TRAIL. As shown in Figure 1B, the antibodies detected protein products corresponding to the predicted molecular mass. In SW948, the antibody used to detect TRAIL-R1 showed several bands at approximately 50 kD and higher. Because TRAIL-R1 has an N-linked glycosylation site (Pan et al. 1997b), these bands could represent differently glycosylated forms of the TRAIL-R1 protein, which was also observed in chimpanzee and human liver. TRAIL-R2 protein was detectable in chimpanzee and human liver at approximately 46–50 kD as described by Walczak et al. (1997). The TRAIL-R3 antibody detected TRAIL-R3-Fc at approximately 66 kD. TRAIL-R3 protein was present in chimpanzee and human liver at approximately 32–35 kD. The slightly higher band in the chimpanzee liver in comparison to human liver could be explained by difference in glycosylation of the TRAIL-R3 protein (Sheridan et al. 1997). The antibody used for TRAIL detection recognized the soluble rhTRAIL at approximately 20 kD (Ashkenazi et al. 1999; Lawrence et al. 2001). In chimpanzee and human liver, several bands were detected at approximately 20, 40, and 60 kD corresponding to monomer, dimer, and trimer forms of soluble TRAIL (Ashkenazi et al. 1999; Lawrence et al. 2001).
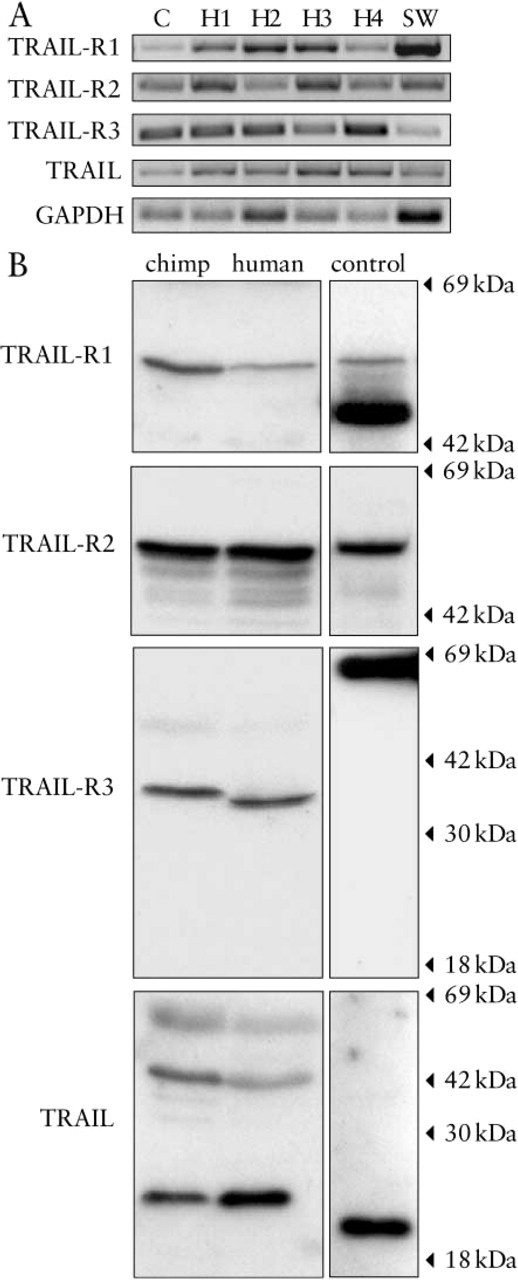
mRNA and protein expression of TRAIL and TRAIL receptors in liver tissue. TRAIL-R1, TRAIL-R2, TRAIL-R3, and TRAIL mRNA and protein expression in chimpanzee (C) and human (H1-H4) liver were determined by RT-PCR (
TRAIL-R1, TRAIL-R2, TRAIL-R3, and TRAIL Expression in Chimpanzee and Human Tissues
Detailed data on TRAIL-R1, TRAIL-R2, TRAIL-R3, and TRAIL expression in human and chimpanzee tissues are given in Table 1. Expression of these four proteins was cytoplasmic and no clear membrane staining was detected. Preincubation of the TRAIL-R1, TRAIL-R3, or TRAIL antibodies with the corresponding blocking peptide completely abolished the immunostaining, further confirming the specificity of the antibodies. Moreover, no staining was observed in all tissues when the primary antibodies were replaced by non-immune normal goat (TRAIL-R1, TRAIL-R3, TRAIL) or rabbit (TRAIL-R2) IgGs.
As shown in Figure 2, both chimpanzee and human hepatocytes stained positive for all three tested TRAIL receptors and TRAIL. The TRAIL-R3 expression was lower in chimpanzee hepatocytes than in human hepatocytes, where TRAIL-R3 was sometimes even strongly expressed. TRAIL-R1, TRAIL-R3, and TRAIL (arrows in Figures 2B, 2F, and 2H, respectively) stained positive in human but not in chimpanzee bile duct epithelium. The vasculature in chimpanzee and human livers lacked expression of the four examined proteins. Only TRAIL-R1 was weakly positive in sinusoidal endothelial cells. However, because these cells were difficult to distinguish, other methods, such as immunogold electron microscopy, should be used to analyze the expression of the receptors in sinusoidal endothelial cells.
In the human and chimpanzee brain, neurons were positive for TRAIL-R1, TRAIL-R2 (Figure 3A), and TRAIL-R3. Most human neurons lacked TRAIL, except for a few that revealed weak immunoreactivity for TRAIL. No TRAIL expression was observed in chimpanzee neurons. Endothelial cells in the arachnoid and capillaries of the human brain were positive for TRAIL-R2 and TRAIL-R3 (Figure 3C, inset), whereas vascular smooth muscle cells were negative. Only TRAIL-R3 was present in chimpanzee brain vascular endothelium of capillaries.
TRAIL-R1, TRAIL-R2, and TRAIL expression was relatively low in the kidney compared to some other tissues, such as the liver. No TRAIL-R3 expression was observed in chimpanzee or human kidney. TRAIL-R1 staining was detected in the tubuli contorti in chimpanzee and human. Human tubuli contorti also expressed TRAIL-R2. In both chimpanzee and human, tubuli contorti were positive for TRAIL. Cells in the Henle's loop were positive only for TRAIL-R2. No expression of the four examined proteins was detectable in the glomeruli or renal vasculature.
Expression pattern of TRAIL and its receptors in chimpanzee and human tissues determined by immunohistochemistry
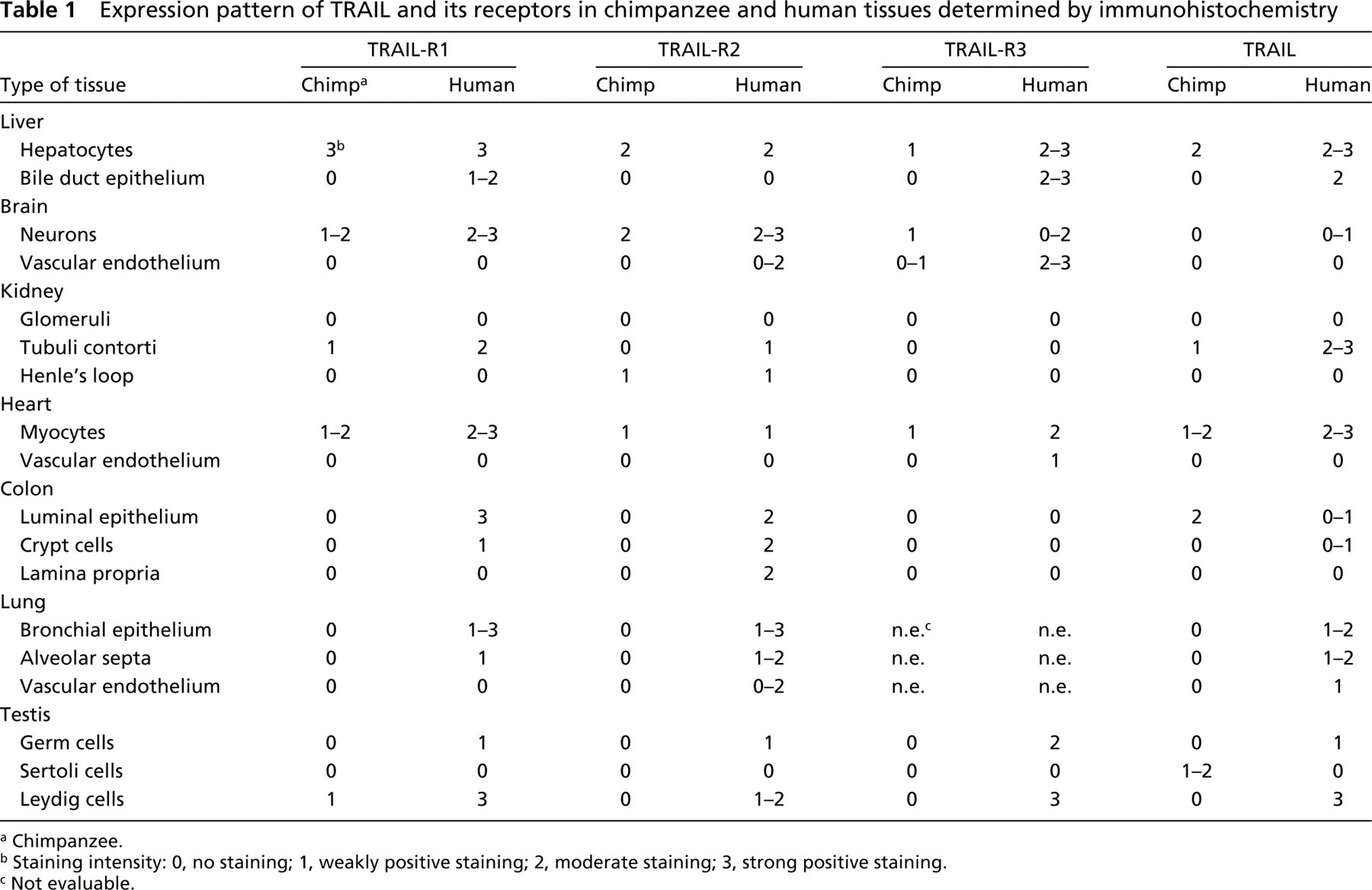
Chimpanzee.
Staining intensity: 0, no staining; 1, weakly positive staining; 2, moderate staining; 3, strong positive staining.
Not evaluable.
Chimpanzee and human heart myocytes stained positive for TRAIL-R1, TRAIL-R2, TRAIL-R3 (Figure 3D), and TRAIL. In contrast to chimpanzee, the human heart vascular endothelium in veins and arteries expressed TRAIL-R3 (Figure 3D, inset). No expression of the other two TRAIL receptors or TRAIL was found in the heart vasculature.
TRAIL-R1, TRAIL-R2, and TRAIL-R3 were not detectable in chimpanzee colon. However, the luminal epithelium in chimpanzee colon was positive for TRAIL. In human colon, TRAIL-R1 was highly expressed in the luminal epithelium and weak TRAIL-R1 expression was observed in crypt cells. TRAIL-R2 was homogeneously expressed in the luminal epithelium and crypt cells. Very faint but mostly no TRAIL-R3 staining was detectable in the human colon. Luminal epithelium and crypt cells were positive for TRAIL.
Human alveolar septa and bronchial epithelium expressed TRAIL-R1, TRAIL-R2 (Figure 3B), and TRAIL. Lung vascular endothelium in veins and arteries was positive for TRAIL-R2 (Figure 3B, inset) and TRAIL. Unfortunately, TRAIL-R3 staining was not evaluable because of high background staining in lung only. No staining of the four proteins was detectable in chimpanzee lung.
In the testis, human germ cells were weakly positive for TRAIL-R1, TRAIL-R2, and TRAIL, while TRAIL-R3 was strongly expressed. No staining was found in chimpanzee germ cells. In contrast to the human, chimpanzee Sertoli cells showed TRAIL expression. Chimpanzee and human Leydig cells expressed TRAIL-R1, but only the latter also expressed TRAIL-R2, TRAIL-R3, and TRAIL. Testis vasculature showed no expression of these proteins.
Discussion
In this study we examined by IHC the protein expression of the death ligand TRAIL and its receptors TRAIL-R1, TRAIL-R2, and TRAIL-R3 in normal chimpanzee and human tissues. No immunostaining for TRAIL-R4 was performed because no reliable antibody for paraffin-embedded specimens was available. Several reports studied the mRNA expression TRAIL and its receptors in a variety of human tissues (Wiley et al. 1995; Chaudhary et al. 1997; Degli-Esposti et al. 1997a, b; MacFarlane et al. 1997; Marsters et al. 1997; Pan et al. 1997a, b, 1998; Schneider et al. 1997; Sheridan et al. 1997; Tan et al. 1997; Walczak et al. 1997). Although our data are generally consistent with these mRNA data, IHC is a better technique to study in more detail which cells express the protein of interest. A few tissues revealed no detectable protein, although mRNA expression had been described earlier. In the kidney, for example, no expression of TRAIL-R3 protein was detectable in our study despite its presence at the mRNA level observed in some but not all studies (MacFarlane et al. 1997; Pan et al. 1997a; Schneider et al. 1997; Sheridan et al. 1997). An explanation might be the fact that mRNA levels are not necessarily predictive of protein expression levels due to protein-specific post-transcriptional and post-translational regulatory mechanisms. In the present study a few tissues revealed protein expression, but no mRNA was observed in other studies. For example, human liver showed immunoreactivity for TRAIL (Figure 2H), whereas others did not detect TRAIL mRNA expression by Northern blotting analysis (Wiley et al. 1995; Tan et al. 1997). However, we were also able to detect TRAIL mRNA by RT-PCR analysis (Figure 1A), implying that Northern blotting analysis is not sensitive enough when tissues express low levels of the mRNA of interest.
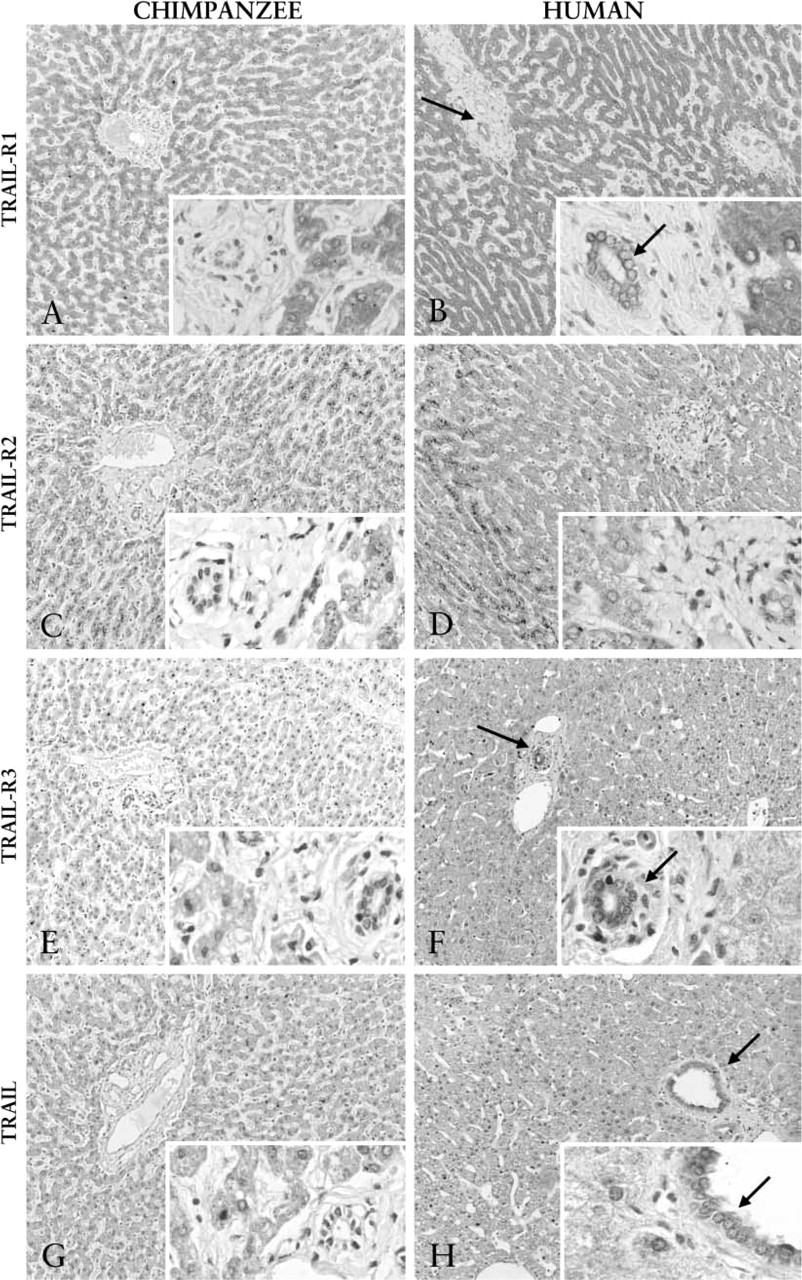
IHC analysis of TRAIL and TRAIL receptors in the liver. Chimpanzee (
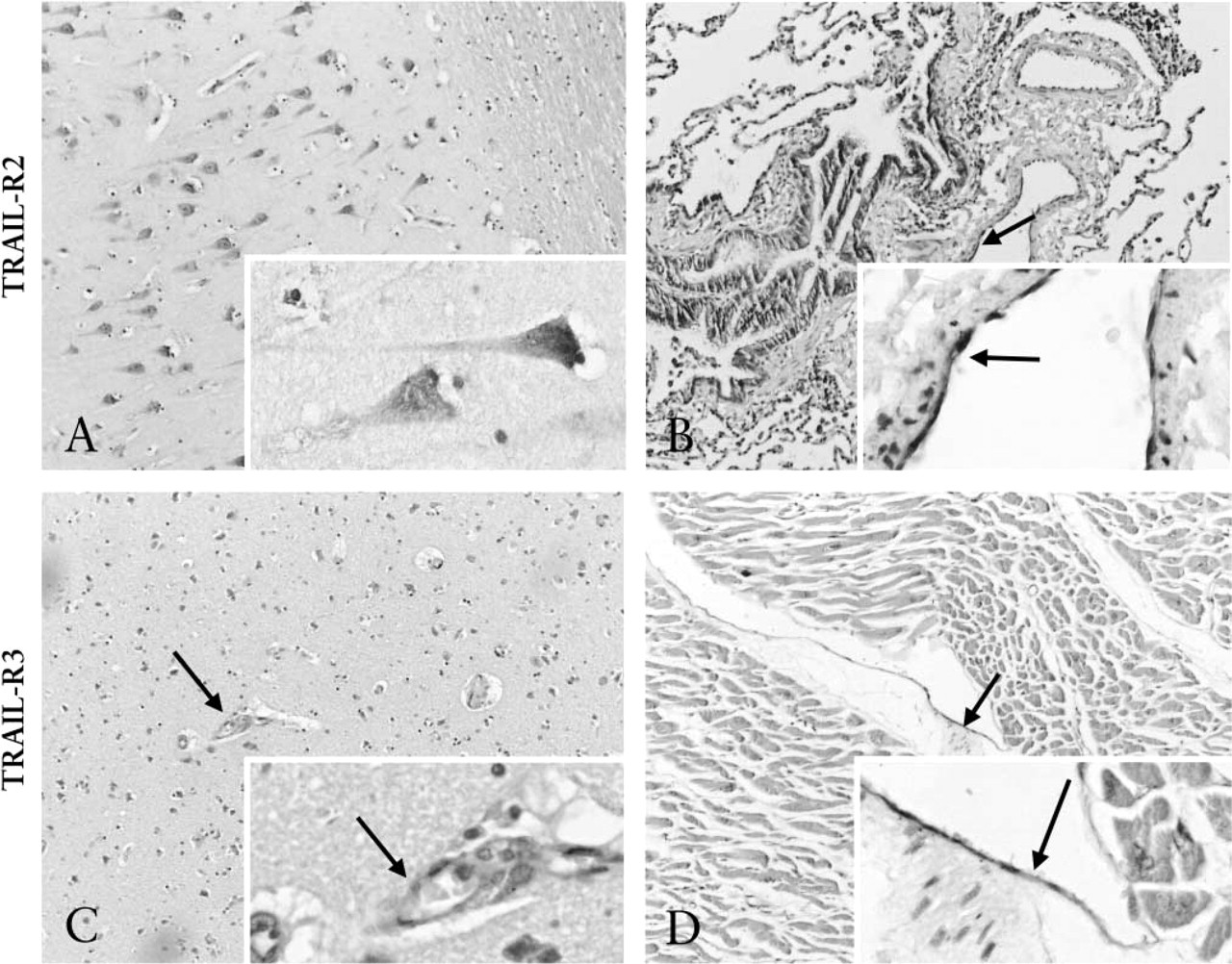
Expression of TRAIL-R2 and TRAIL-R3 in several human tissues. IHC staining was performed for TRAIL-R2 (
The absence of TRAIL protein and the presence of TRAIL-R1, TRAIL-R2, and TRAIL-R3 proteins in the human brain are in line with results obtained by others (Rieger et al. 1999; Dorr et al. 2002). Colon epithelium showed TRAIL, TRAIL-R1, and TRAIL-R2 expression while TRAIL-R3 was not detectable as defined by our method based on intensity of staining. In general, this is in agreement with Strater et al. (2002) and Koornstra et al. (2003). However, our results are not in accordance with a study performed by Ichikawa et al. (2001), which might be explained by the use of a different antibody to detect TRAIL-R2. Their anti-human TRAIL-R2 monoclonal antibody, TRA-8, could not detect TRAIL-R2 protein in liver, lung, kidney, spleen, testes, ovary, heart, or pancreas. In contrast, the TRAIL-R2 antibody used in the present study showed immunoreactivity in all of these tissues. For example, in human liver we could not only detect TRAIL-R2 protein by Western blotting analysis and IHC (Figures 1B and 2D, respectively) but we were also able to show TRAIL-R2 mRNA by RT-PCR (Figure 1A). These results strongly support the presence of TRAIL-R2 in human liver.
Interestingly, the death receptor Fas is not constitutively expressed in several organs in which our study revealed TRAIL-R1 and/or TRAIL-R2 expression (Leithauser et al. 1993). For example, all cells in the nervous system appeared to be Fas-negative, whereas we and others (Dorr et al. 2002) observed TRAIL-R1 and TRAIL-R2 expression. In addition to the nervous system, absence of Fas but presence of TRAIL death receptors was observed in cardiac muscle and testicular germ cells. Regarding FasL, the ligand for the Fas receptor, Lee et al. (1999) showed that it is widely expressed in both lymphoid and non-lymphoid tissues. Expression of FasL and TRAIL was comparable in the tissues examined.
Although the presence of multiple TRAIL receptors suggests that TRAIL may be involved in multiple processes, the precise physiological function of TRAIL and its role in human disease remains to be established. Recent reports indicate that TRAIL, like TNF and FasL, may play a role in the modulation of host defense mechanisms (Clarke et al. 2000; Song et al. 2000; Vidalain et al. 2000; Hilliard et al. 2001; Lum et al. 2001; Cretney et al. 2002; Sedger et al. 2002; Strater et al. 2002; Takeda et al. 2002). Studies using TRAIL-deficient mice demonstrated that TRAIL is important in controlling tumor growth (Cretney et al. 2002; Sedger et al. 2002; Takeda et al. 2002). Several effector cells, including lymphocytes (Dorothee et al. 2002), natural killer cells (Smyth et al. 2001; Takeda et al. 2002), monocytes (Griffith et al. 1999), and dendritic cells (Fanger et al. 1999; Liu et al. 2001), have been shown to play a critical role in TRAIL-mediated suppressing of tumor development or metastasis. In addition to tumor surveillance, TRAIL may play an important role in preventing autoimmunity. For example, chronic blockade of TRAIL in mice exacerbated autoimmune arthritis, whereas intra-articular TRAIL gene transfer prevented the disease (Song et al. 2000). In addition, TRAIL inhibits experimental autoimmune encephalomyelitis (EAE), because blockade of TRAIL exacerbated EAE induced by myelin oligodendocyte glycoprotein (Hilliard et al. 2001). In contrast to FasL, TRAIL does not induce activation-induced cell death in autoreactive lymphocytes but rather inhibits their activation and expansion via cell cycle arrest (Song et al. 2000; Lunemann et al. 2002). Furthermore, several studies have demonstrated that the TRAIL/TRAIL receptor system may play an important role in the elimination of virus-infected cells. Cells infected by human cytomegalovirus, adenovirus, reovirus, measles virus, or human immunodeficiency virus revealed increased TRAIL, TRAIL-R1, and TRAIL-R2 protein levels rendering them more sensitive to TRAIL-induced apoptosis by autocrine or T-cell-derived TRAIL (Clarke et al. 2000; Vidalain et al. 2000; Lum et al. 2001; Strater et al. 2002).
The lack of toxicity of rhTRAIL administration to chimpanzees (Kelley et al. 2001; Lawrence et al. 2001) and the high homology between the human and chimpanzee TRAIL and TRAIL receptors prompted us to examine the expression of these proteins in chimpanzee tissues. Interestingly, only a few differences in TRAIL, TRAIL-R1, TRAIL-R2, or TRAIL-R3 expression were observed between human and chimpanzee liver, brain, and kidney. However, TRAIL-R1 and TRAIL-R2 were not detectable in chimpanzee colon, lung, and testis, whereas these proteins were expressed in their human counterparts. In addition to these death receptors, the human testis also expressed the decoy receptor TRAIL-R3, which was absent in the chimpanzee testis. These differences might have some consequences for rhTRAIL administration in the clinic. However, taking into account that the most severe cell damage after administration of agonistic anti-Fas antibodies to mice was observed in their livers, the comparable expression of TRAIL-R1, TRAIL-R2, and TRAIL-R3 in both human and chimpanzee hepatocytes is reassuring. One has to question whether hepatocytes will be the primary target of rhTRAIL. Several studies have recently demonstrated that the severe hemorrhagic liver injury observed after administration of Fas agonists is most consistent with a microvascular injury beginning at the level of sinusoidal endothelial cells rather than a primary hepatocyte injury (Wanner et al. 1999; Janin et al. 2002; Jodo et al. 2003). In addition to the liver, disseminated vascular endothelial cell apoptosis was also observed after Fas agonist administration in organs such as kidney, heart, and brain (Janin et al. 2002). Therefore, the potential toxicity of rhTRAIL might be to human endothelial cells. However, several in vitro experiments using human umbilical vein endothelial cells, human microvascular endothelial cells, or human aortic endothelial cells have shown that these cells were relatively resistant to rhTRAIL (Sheridan et al. 1997; Gochuico et al. 2000; Zhang et al. 2000; Secchiero et al. 2003). Protection of these endothelial cells for rhTRAIL-induced apoptosis appeared to be dependent on TRAIL-R3 membrane expression, activation of Akt and ERK1/2 survival signaling pathways, and an unknown intra-cellular inhibitor(s) of apoptosis. In the present study, no TRAIL-R1 or TRAIL-R2 was detectable in liver, renal, testicular, or heart endothelium, of which the latter did express TRAIL-R3. Lung and brain endothelium expressed TRAIL-R2 and TRAIL-R3 but no TRAIL-R1. This might be of interest considering that TRAIL-R1 can induce apoptosis on binding of soluble TRAIL derivates, whereas TRAIL-R2 signals predominantly by membrane-bound TRAIL, but not or only modestly by soluble TRAIL derivates (Muhlenbeck et al. 2000; Wajant et al. 2001). Toxicity of rhTRAIL will, among others, depend on the presence of the death receptors on the cell surface, which has been demonstrated in human tumor cell lines (Zhang et al. 1999; Mitsiades et al. 2001a). In the present study, no membranous staining pattern was observed for TRAIL-R1 and TRAIL-R2. However, this does not preclude the presence of TRAIL-R1 or TRAIL-R2 on the cell surface because IHC is not the optimal technique to detect membranous staining, especially when there is abundant cytoplasmic expression. In addition, the decoy receptor TRAIL-R3 is able to prevent rhTRAIL-induced apoptosis only when present on the cell surface. In our study, TRAIL-R3 was detectable but no membranous staining pattern could be observed.
In summary, only a few differences were observed in expression of TRAIL and receptors between human and chimpanzee tissues. Lack of liver toxicity after administration of rhTRAIL to chimpanzees is not due to the absence of TRAIL death receptors, which is relevant for the toxicity profile in case of the clinical application of rhTRAIL in humans.
Footnotes
Acknowledgements
Supported by a grant from the Therapeutic Proteins for Chronic Diseases program of the University of Groningen, grant RUG 2000–2286 from the Dutch Cancer Society, and a grant from the Nijbakker–Morra Stichting.
