Abstract
Jawbones and long bones, despite their shared skeletal lineage, frequently exhibit distinct origins and developmental pathways. Identifying specific progenitor subsets for mandibular osteogenesis remains challenging. Type II collagen is conventionally associated with cartilaginous structures, yet our investigation has identified the presence of type II collagen positive (Col2+) cells within the jawbone development and regeneration. The role of Col2+ cells in jawbone morphogenesis and repair has remained enigmatic. In this study, we analyze single-cell RNA sequencing data from mice jawbone at embryonic day 10.5. Through fate-mapping experiments, we have elucidated that Col2+ cells and their progeny are instrumental in mandibular osteogenesis across both fetal and postnatal stages. Furthermore, lineage tracing with a tamoxifen-inducible CreER system has established the pivotal role of Col2+ cells, marked by Col2-CreER and originating from the primordial Meckel’s cartilage, in jawbone formation. Moreover, our research explored models simulating jawbone defects and tooth extraction, which underscored the osteogenic differentiation capabilities of postnatal Col2+ cells during repair. This finding not only highlights the regenerative potential of Col2+ cells but also suggests their versatility in contributing to skeletal healing and regeneration. In conclusion, our findings position Col2+ cells as essential in orchestrating osteogenesis throughout the continuum of mandibular development and repair.
Introduction
Contemporary theories suggest that the jawbone primarily arises through intramembranous ossification, a process characterized by the condensation of mesenchymal cells followed by their differentiation into osteoblasts. Conversely, the proximal region and the condylar process of the jawbone are subject to endochondral ossification, a distinct developmental pathway.1–3 Meckel’s cartilage (MC) plays a pivotal role in the embryonic development of the jawbone, serving as a critical support structure during the formative stages of jawbone development before its postnatal regression.4–7
Recent advances in skeletal biology have illuminated that certain cartilage markers can identify mesenchymal cells possessing multi-lineage differentiation potential. These versatile cells can differentiate into chondrocytes, osteoblasts, and early perichondrial precursors via distinct regulatory pathways. 8 Type II collagen α1 (Col2a1), a hallmark chondrocyte marker, has been at the forefront of these discoveries. Lineage-tracing studies have subsequently demonstrated that Col2+ cells are progenitors capable of differentiating into chondrocytes, osteoblasts, stromal cells, and adipocytes within the context of long bones.9–12 Within cranial bone, particularly in the canonical endochondral bone formation of the cranial base, Col2+ cells have been identified as key progenitors in the skeletal lineage.10,13 These lineage-tracing experiments have unequivocally established that Col2+ cells function not as intermediary cells but as direct precursors within the osteogenic lineage. Our investigations have also confirmed the presence of Col2+ cells within the jawbone, yet the specific roles and potential functions of these cells in jawbone development remain to be fully elucidated. The fate and function of Col2+ cells in the context of jawbone development present a compelling avenue for further research.
In this study, our objective was to delineate the fate and elucidate the potential functionalities of Col2+ cells within the context of jawbone development, employing sophisticated methodologies including single-cell RNA sequencing (scRNA-seq) analyses14,15 and lineage-tracing techniques. 16 Our investigations have led to the revelation that Col2+ cells, alongside their progeny, assume a critical role in the processes of jawbone development, remodeling, and repair following injury, primarily through their differentiation into osteoblasts.
Materials and Methods
Re-analysis of scRNA-seq Data
The scRNA-seq data were obtained from the GEO database (GSE141173, GSM3084671).15,17 Mandibles from mouse embryos were quickly dissected in ice-cold PBS and minced to a fine paste. Tissue of 12.5 mg was placed in a sterile 1.5-mL tube containing 0.5 mL protease solution containing 125 U/ml DNase and Bacillus licheniformis. The samples were incubated at 4C for total of 10 min, with trituration using a wide boar pipette tip every minute after the first two. Protease was inactivated using ice-cold PBS containing 0.02% BSA and filtered using 30-µM filter. The cells were pelleted by centrifugation at 200 ×g for 4 min and resuspended in 0.02% BSA in PBS. Cell number and viability were assessed using a hemocytometer and trypan blue staining. A total of 9600 cells were loaded onto a well on a 10× Chromium Single Cell instrument (10× Genomics) to target sequencing of 6000 cells. The analysis was performed using Seurat v4.0 (Satija, New York, USA) and Monocle v.2.0 (Cole-Trapnell Lab, Washington, USA) with official scripts. Clusters were visualized using Uniform Manifold Approximation and Projection.
Mice
All mice used in this study were of C57BL background, and Col2-Cre, Col2-CreER, and R26RtdTomato mice were bought from the Shanghai Model Organisms Center. We crossed Col2-CreER and Col2-Cre mice with R26RtdTomato reporter mice and analyzed the patterns of tdTomato labeling in the jawbone at different time points (E15.5, P3, 4W, 8W, and 1Y). Each group consisted of 3 mice. Animals were bred in a specific pathogen-free condition under a 12/12 day/dark cycle. Animal studies were carried out according to the guidelines of the National Institutional Animal Care, approval number: [2022]-DW-06.
Histological Preparation and Staining
Mice jawbones were dissected and fixed with 4% paraformaldehyde for 24–48 hr, then decalcified in 10% EDTA at 4C for 3–4 weeks. The specimens were embedded in optimal cutting temperature compound (Sakura, Taizhou, China) and cut into 10- µm-thick sections. For morphological evaluation, sections were stained with H&E (Sangon Biotech, Shanghai, China). For immunofluorescence staining, antigen retrieval was performed using hyaluronidase at 37C for 1 hr. Sections were blocked with goat serum in PBS-T (0.1% Triton X-100 in PBS) at 37C for 1 hr, followed by incubation with primary antibodies overnight at 4C. The primary antibodies used were as follows: Runx2 (1:200, Abcam, Massachusetts, USA), and anti-type II collagen (1:200, Boster Biological Technology, Wuhan, China). Subsequently, sections were incubated with Alexa Fluor 488 IgG (1:1000, Invitrogen, California, USA) and counterstained with DAPI, which specifically labels the cell nucleus (Sigma-Aldrich, St Louis, USA). All images were captured using Eclipse Ni-U microscope (Nikon, Japan), and processed with the ImageJ software (US National Institutes of Health, USA).
Tamoxifen Administration
The tamoxifen (Sigma T5648-1G) was dissolved in sunflower seed oil (Sigma, St Louis, USA) at a concentration of 20 mg/ml to create a working solution. The tamoxifen–oil mixture was incubated at 60C in oven until completely dissolved and then stored at 4C. Mice were intraperitoneally injected with tamoxifen at 1.5 mg/10 g body weight.
Bone Repair and Tooth Extraction Model
Under anesthesia, an incision to the angle of the mandible was made, and muscles were bluntly dissected to expose the mandibles of the 8-week-old Col2-CreER; R26RtdTomato mice. A 1.5-mm diameter drill hole defect was made before the ramus of the mandible. After surgery, 1.5 mg/10 g tamoxifen was intraperitoneally injected into the mice at the first day after surgery and harvested at 7 and 14 days. A tooth extraction model was created in 8-week-old Col2-Cre; R26RtdTomato mice, which were anesthetized with 0.3% pentobarbital sodium. The mouth was rinsed with sterile saline, and the mandibular first molars were extracted with forceps under a stereomicroscope. Bleeding was controlled by local pressure, and harvest was done at 7 and 14 days. Mice were recovered on a heating pad and then housed in independent cages with a soft food diet.
Statistical Analysis
Statistical analyses were performed using Prism 8 software (GraphPad Software). Data are presented as mean ± SD. Differences between two groups were determined by Student’s t-test. Differences between three groups were determined by one-way ANOVA. The significance level was set at p<0.05
Result
scRNA Sequencing Analysis Finds a Pseudo-Differentiation Trajectory From Jawbone-Derived Chondrocyte Progenitors to Osteoblasts
To elucidate the role of chondrocytes within MC, we delved into the cellular hierarchy of the mouse jawbone, leveraging insights from a recent single-cell atlas study of the mouse jawbone.15,17 Embryonic jawbone cells at E10.5 were categorized into six distinct clusters including epithelial cells, endothelial cells, erythrocyte, pericyte, neural crest (NC) cells 1 and NC cells 2 (Fig. 1A). Among the six clusters, cluster NC cells 2 present bone-forming potential (Fig. 1B), thus we further clustered the cells into three clusters including NC-mesenchyme stem cells, expressing Prrx1, Msx1; MC-chondrocyte, expressing Col2a1, Sox9; and pre-osteoblasts, expressing Col1a1 and Rgs5 (Fig. 1C to E). Through further pseudo-temporal analysis using Monocle 2, we found that NC-mesenchyme stem cells are situated on the starting point of differentiation, MC-chondrocytes are positioned in the middle, and the differentiation direction of MC-chondrocytes points towards pre-osteoblasts (Fig. 1F). Consequently, our scRNA-seq analysis delineated a pseudo-differentiation trajectory from jawbone-derived chondrocyte progenitors toward osteoblasts.
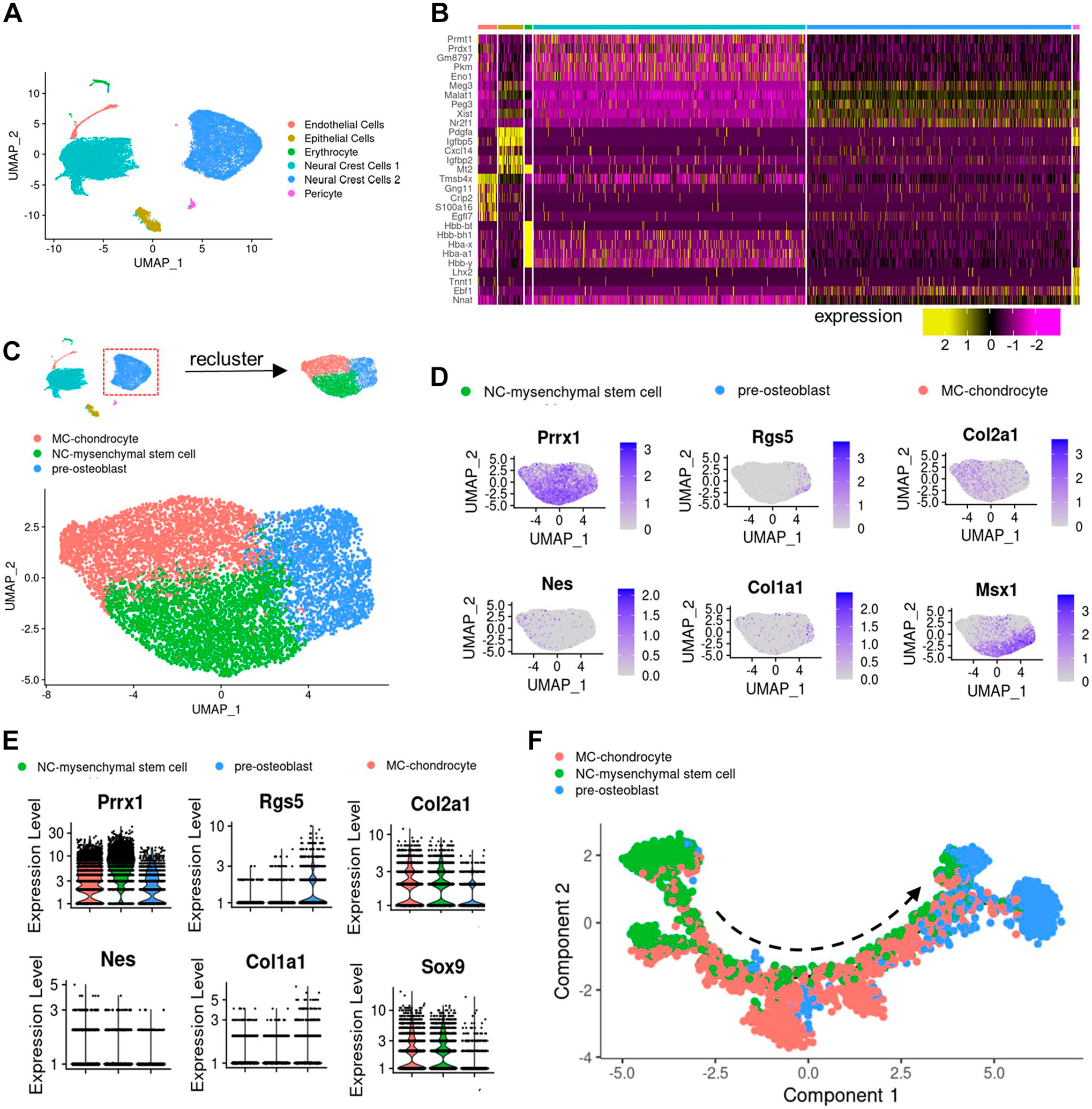
scRNA-RNA sequencing analysis of mice jawbone embryonic progenitors. (A) Dimensional reduction (Uniform Manifold Approximation and Projection [UMAP]) of scRNA-seq results of E10.5 jawbone cells, colored according to the clusters. (B) Heatmap showing the representative markers for each cluster across cells in the six clusters. (C) Diagram of further clustering. Dimensional reduction (UMAP) of scRNA-seq results of neural crest (NC) cells, colored according to the clusters. (D) Feature plots showing normalized expression levels of the selected stem cells. (E) Violin plots showing normalized expression levels of the selected stem cells. (F) Single-cell trajectory obtained by unsupervised ordering based on a pseudo-temporal order from left to right, colored by cell clusters. Abbreviations: MC, Meckel’s cartilage.
MC-Derived Col2+ Cells Facilitate Jawbone Development
Investigating the role of MC-derived Col2+ cells in jawbone development during embryonic and postnatal stages, we employed Col2-Cre; R26RtdTomato mice for fate mapping. These mice exhibit tomato fluorescence in Col2+ cells from the embryonic stage. In addition, Col2-CreER; R26RtdTomato mice were used, where Col2+ cells express tomato fluorescence following tamoxifen treatment at specified postnatal times. Analysis of Col2 lineage cell distribution in Col2-Cre; R26RtdTomato reporter mice revealed tdTomato+ cells in the jawbone’s trabecula and periosteum across embryonic and postnatal stages (Fig. 2A to E). The proportion of Col2+ cells in the periosteum increased significantly from 51.09 ± 6.64% at E15.5 to 89.53 ± 4.04% by 4 weeks, with nearly all periosteal cells being Col2+ in 8-week and 1-year-old mice (Fig. 2F). It indicates that Col2-Cre+ cells gathered in periosteum during the development of jawbone. Administering tamoxifen at E15.5, prior to MC degeneration, and analyzing at P0, we observed Col2-CreER+ cells migrating from MC throughout the embryonic jawbone (Fig. 2G, H). Colocalization of tdTomato and Runx2 immunostaining in the jawbone periosteum indicated these osteogenic cells originated from Col2-CreER+ cells, highlighting MC-derived Col2+ cells as primary constituents of periosteal and osteoblastic cells.
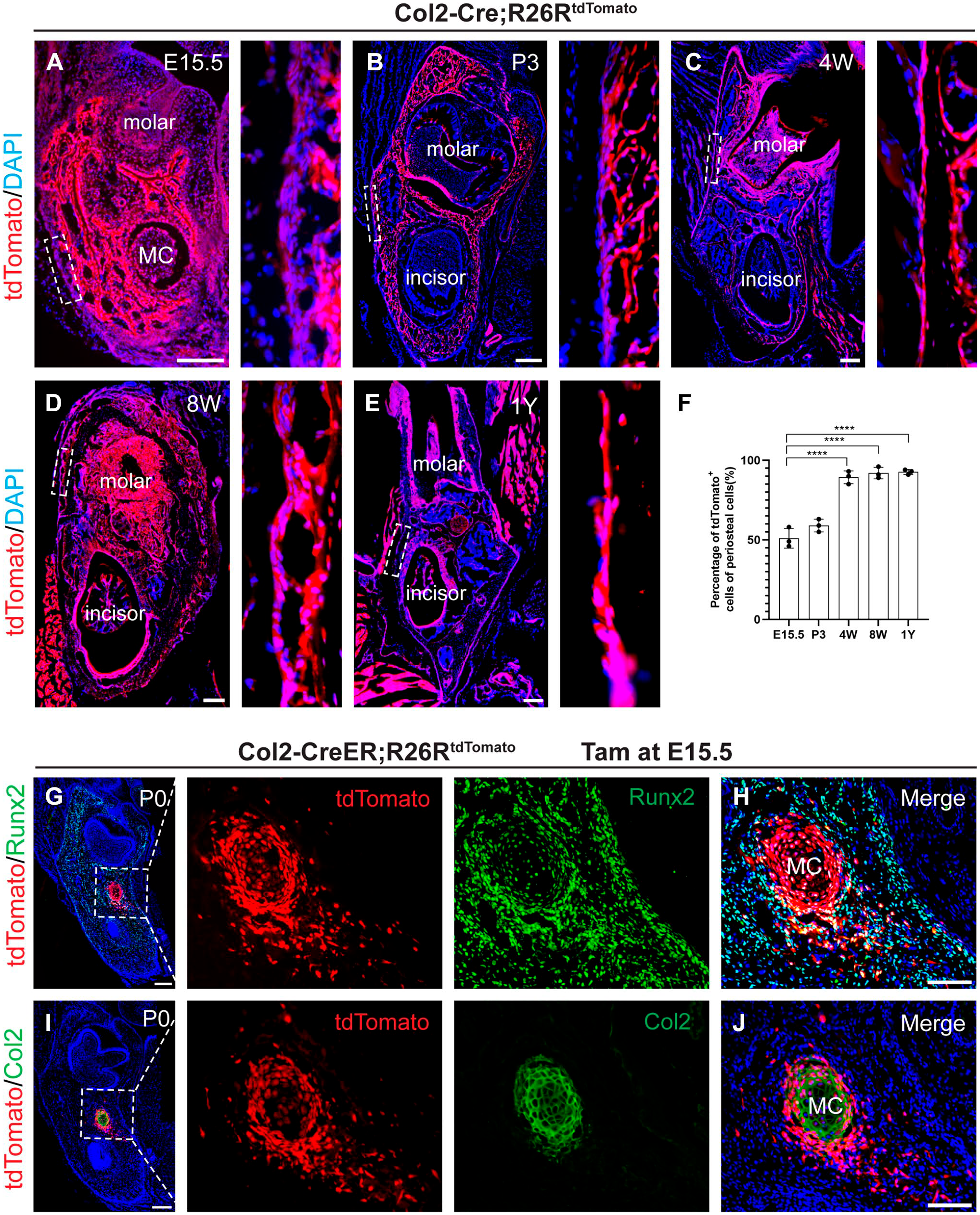
The distribution of Col2-Cre+ (Col2-Cre-labeled cells) cells in jawbone. (A to E) Representative immunofluorescence images from the lineage tracing of Col2-Cre+ cells in jawbone at different time points (E15.5, P3, 4W, 8W, and 1Y). (F) Percentage of tdTomato+ cells in the periosteum of jawbone (n=3). (G, H) Col2-CreER; R26RtdTomato mice were administered tamoxifen at E15.5 and harvested at P0. Representative immunofluorescence images of Col2+Runx2+ cells in the jawbone of P0 mice. (I, J) Immunofluorescence images of tdTomato+ cells surround the Meckel’s cartilage (MC). Green: chondrocyte, red: tdTomato, blue: nucleus. Data are presented as the mean ± SD. ****p<0.0001. Bars A-E = 100 μm; bars G-J = 50 μm.
Col2+ Cells: Osteoblast Precursors in Jawbone
Utilizing Col2-Cre; R26RtdTomato reporter mice, we observed numerous Col2-Cre+ cells in MC and newly forming bone at E15.5 (Fig. 3A). Quantitative data show that Col2+ cells participate as preosteoblasts cells and most of Runx2+ cells originate from Col2+ cells. Postnatally, as MC degenerated, no Col2-Cre+ chondrocytes remained in the jawbone, whereas Col2+ cells proliferated throughout (Fig. 3F to J). It suggests that Col2-Cre+ cells in the mandible of postnatal mice originate from chondrocytes in the embryonic mandible. At E15.5, 52.98 ± 11.94% of Col2+ cells in the periosteum expressed Runx2, a figure that rose to 75.53 ± 5.53% by postnatal week 4 (Fig. 3K). In adult and aged mice, nearly all Runx2+ osteoblasts were Col2+ (Fig. 3C to E), confirming the expansion and osteogenic differentiation potential of Col2-Cre+ cells in jawbone development. It indicates that Col2-Cre+ cells maintain relatively osteogenic in jawbone of aged mice.
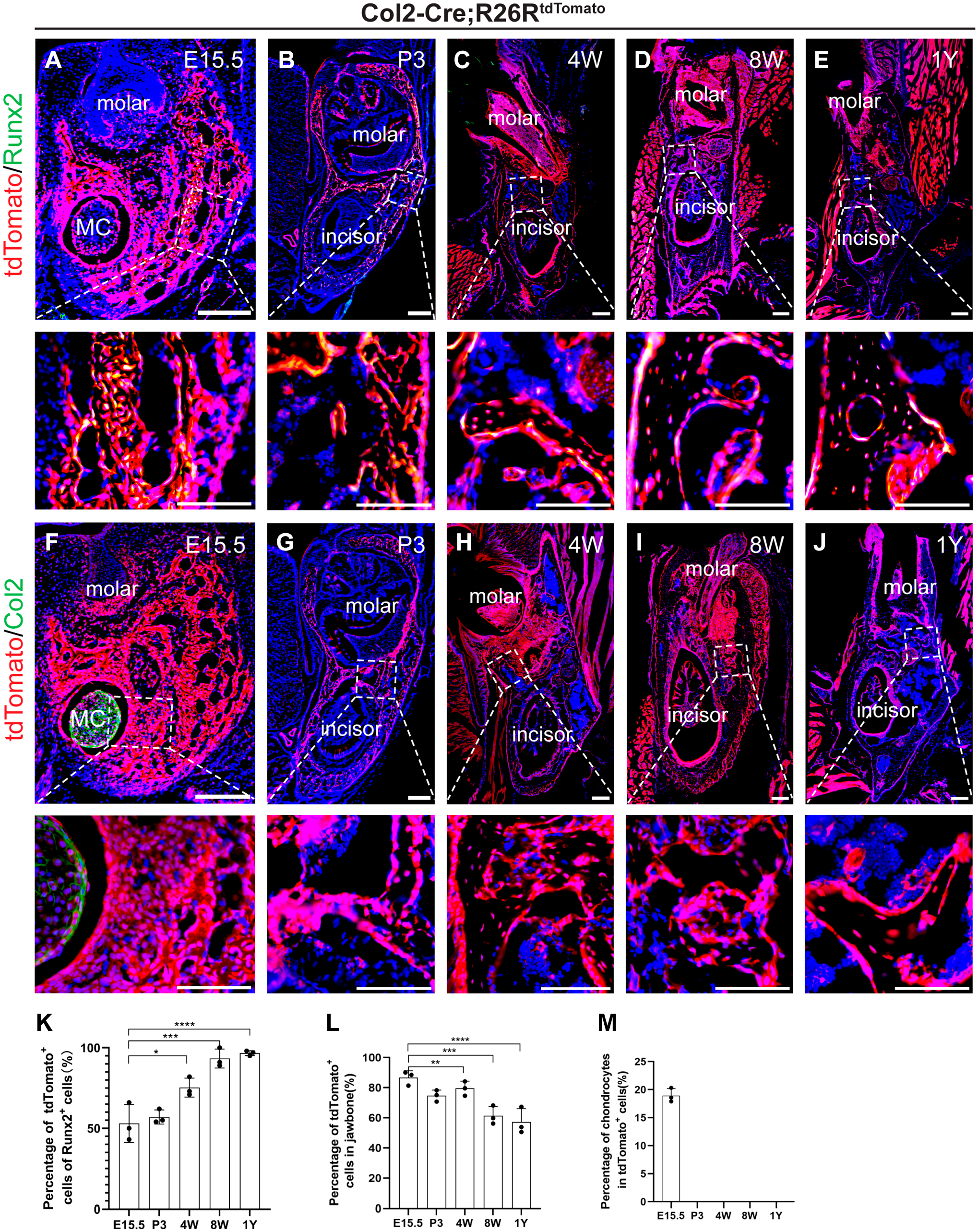
Col2-Cre+ cells (Col2-Cre-labeled cells) exist in jawbone from embryonic to aged mice. (A to E) Lineage tracing analysis of Col2-Cre+ cells was performed using Col2-Cre; R26RtdTomato. Representative immunofluorescence images of Col2+Runx2+ cells in the jawbone in E15.5, P3, 4W, 8W, and 1Y mice. Green: Runx2, red: tdTomato, blue: nucleus. (F to J) Representative immunofluorescence images of chondrocyte in the jawbone in E15.5, P3, 4W, 8W, and 1Y mice. Green: chondrocyte, red: tdTomato, blue: nucleus. (K) Percentage of tdTomato+Runx2+ cells in the jawbone (n=3). (L) Percentage of tdTomato+ cells in jawbone (n=3). (M) Percentage of chondrocytes in tdTomato+ cells (n=3). Data are presented as the mean ± SD. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001. Bars A-J = 100 μm.
Col2+ Cells in Bone Injury and Tooth Extraction Socket Healing
Exploring Col2+ cells participation in bone injury and tooth extraction socket healing, we created a bone injury model in 2-month-old Col2-CreER; R26RtdTomato mice and tracked Col2-CreER+ cells post-injury (Fig. 4A, B). One-week post-injury, Col2-CreER+ cells congregated in the callus, significantly contributing to early-stage bone repair as evidenced by their differentiation into Runx2+ osteogenic cells (Fig. 4E to H). By the second week, the number of Col2-CreER+ cells increased. The proportion of Col2+ cells in the callus increased significantly from 5.01 ± 0.01% to 16.67 ± 3.51% by 1 week (Fig. 4C). Similarly, following tooth extraction, Col2-Cre+ cells were notably present in the extraction sockets. Although the proportion of Col2+ cells in the sockets decreased significantly from 57 ± 1.73% to 33.67 ± 6.65% by 1 week, the Col2+ cells are still involved in the osteogenic process (Fig. 4K, L). These data indicated the osteogenic capability and role of the Col2+ cells in alveolar socket repair (Fig. 4M to P).
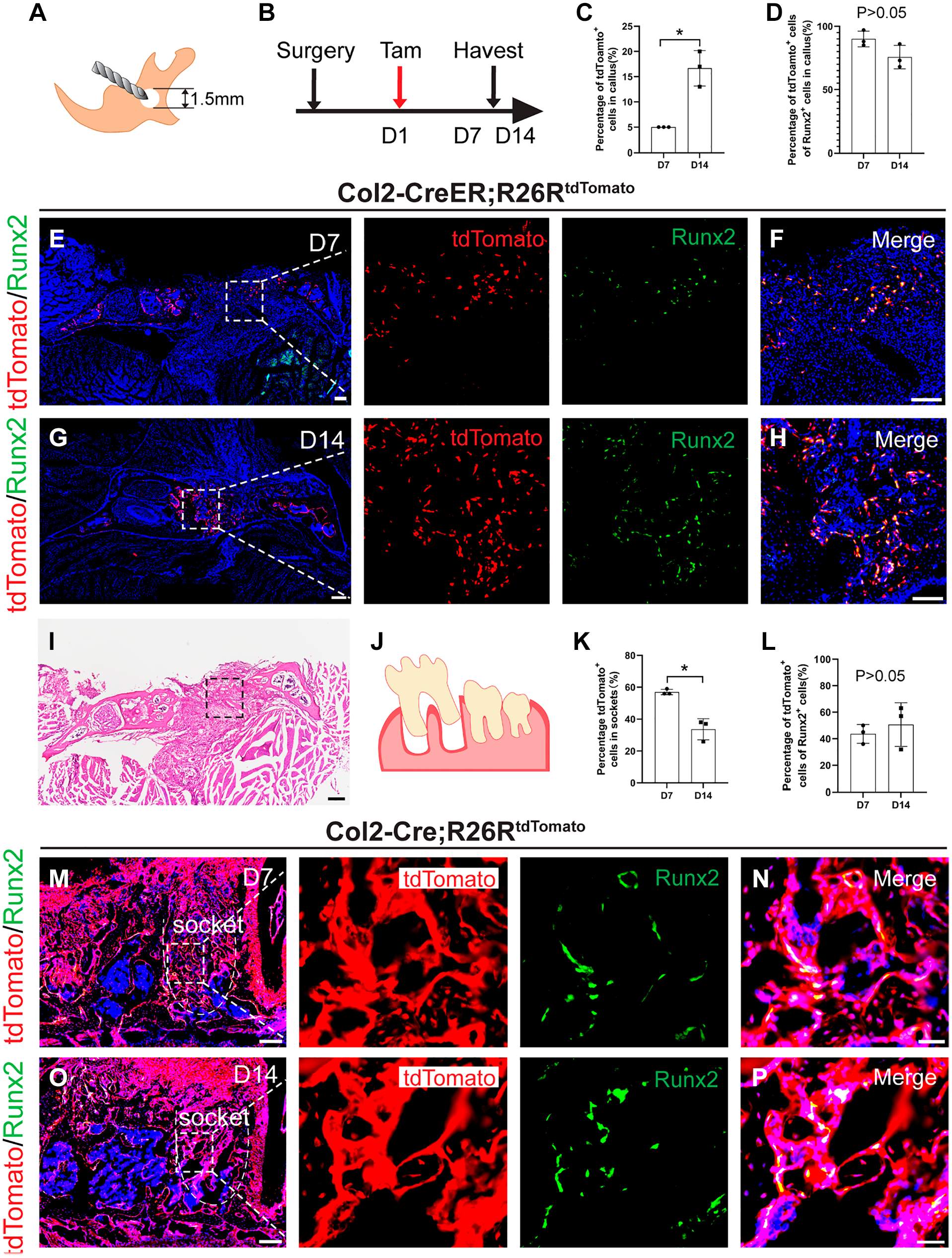
Col2+ cells are responsive toward jawbone injury and tooth extraction. (A) Schematic diagram of bone defect model in mouse jawbone. (B) Illustration of the experimental design. This analysis was performed by injecting tamoxifen into mice at the first day after surgery and harvesting the tissues after 7 and 14 days of healing. (C) Percentage of tdTomato+ cells in jawbone callus (n=3). (D) Percentage of tdTomato+Runx2+ cells in the jawbone callus (n=3). (E to H) Representative immunofluorescence images of tdTomato+Runx2+ cells in the jawbone callus at 7 and 14 days after surgery in Col2-CreER; R26RtdTomato mice. (I) Representative H&E image in the jawbone callus at 7 days after surgery. (J) Illustration of the extraction of the first molar in mouse jawbone. (K) Percentage of tdTomato+ cells in extraction sockets (n=3). (L) Percentage of tdTomato+Runx2+ cells in extraction sockets (n=3). (M to P) Representative immunofluorescence images of tdTomato+Runx2+ cells in the sockets at 7 and 14 days after extraction in Col2-Cre; R26RtdTomato mice. Data are presented as the mean ± SD. *p<0.05. Bars E, G, I, M, O = 200 μm; bars F, H = 50 μm; bars N, P = 20 μm.
Discussion
Recent investigations have elucidated the pivotal role of Col2+ cells as progenitors within the skeletal lineage, significantly contributing to bone development and homeostasis across various anatomical sites, including limbs and calvaria.8,10,13,18 MC, a transient embryonic structure critical for mandibular development, undergoes a well-documented degenerative process postnatally. 4 Our research has identified a notable migration and participation of Col2+ cells from MC in the formation of the periosteum and subsequent bone formation within the jawbone. Despite these observations, the extent to which Col2+ cells derived from MC contribute to jawbone development and its healing processes remains to be fully elucidated. 10
To address this gap, we employed Col2-Cre; R26RtdTomato mice to meticulously study the distribution and fate of Col2+ cells throughout jawbone development and repair phases. Our methodology encompassed scRNA-seq analysis to achieve single-cell resolution mapping of embryonic jawbone cells. This analysis unveiled a pseudo-temporal differentiation trajectory indicating a propensity of Col2+ cells toward osteogenic differentiation, aligning with prior findings in long bones. 8 Furthermore, genetic lineage-tracing studies furnished evidence of Col2+ cells’ involvement in jawbone development and osteogenesis as early progenitors, persisting beyond the degeneration of MC. Notably, Col2+ cells were found to mark similar early skeletal lineage cells in the jawbone, where intramembranous bone formation occurs. From the embryonic period through to old age, Col2+ cells and their progeny predominantly occupied the jawbone, demonstrating their capacity to differentiate into preosteoblasts. 19 During the embryonic development of the mandible, we conducted a focused tracing of Col2+ cells, mapping the fate of chondrocytes amid the degeneration of MC. Col2+ cells migrate from MC and acquire osteogenic capabilities, suggesting their substantial contribution to mandibular osteogenesis beyond merely serving as a structural template. Moreover, our investigation extended to the role of postnatal Col2+ cells in the production of both chondrocytes and osteoblasts during the healing processes of bone injuries and sockets.20,21 Notably, the jawbone callus was enriched with transient Col2+ cells. These cells undergo a transformation from chondrocytes to osteoblasts during mandibular fracture repair, significantly aiding in the healing process. In addition, Col2+ cells were observed to play a crucial role in alveolar socket repair, likely originating from periosteal stores and participating in the repair process.
In summary, this study illuminates the multifaceted contributions and heterogeneity of the Col2+ osteogenic lineage within the jawbone, underscoring the presence and functional significance of Col2+ cells in mandibular development and defect healing. These findings advance our understanding of mandibular development and regeneration plasticity.
Footnotes
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
H. Chen contributed to the acquisition, analysis, and interpretation of data, and drafted the manuscript. Y. Ding contributed to the design of the work, acquisition of data, and revised the manuscript for important intellectual content. Y. Sun contributed to conception and design and drafted and critically revised the manuscript. Y. Wang revised the manuscript. All authors approved the final version of the manuscript and agree to be accountable for all aspects of the work.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by National Key R&D Program of China (grant no. 2022YFA1103200), Shanghai Education Commission Scientific Research and Innovation Program (grant no. 2023ZKZD29), National Natural Science Foundation Projects of China (grant nos. 81822012, 82061130222, 81771043), Shanghai Academic Leader of Science and Technology Innovation Action Plan (grant no. 20XD1424000), the Shanghai Experimental Animal Research Project of Science and Technology Innovation Action Plan (grant nos. 8191101676, 201409006400), and Shanghai International Scientific Collaboration Fund (grant no. 21520710100).
