Abstract
Human intestinal spirochetosis (HIS) is a colorectal bacterial infection caused by the Brachyspira species. Griffonia simplicifolia-II (GS-II) is a lectin specific to terminal α/βGlcNAc residues. Here, we investigated terminal βGlcNAc residues in the context of HIS infection using GS-II-horseradish peroxidase staining and HIK1083 immunostaining specific to terminal αGlcNAc residues. Fourteen of 15 HIS cases were GS-II-positive on the bacterial body. No cases showed HIK1083 positivity. The percentage of bacterial bodies staining positively for GS-II based on comparison with anti-Treponema immunostaining was ≤30% in seven cases, 30–70% in two, and >70% in six. Of 15 HIS cases analyzed, none were comorbid with tubular adenomas, and three were comorbid with sessile serrated lesions (SSLs). To determine the species of spirochete infected, the B. aalborgi-specific or B. pilosicoli-specific NADPH oxidase genes were amplified by PCR. After direct sequencing of the PCR products, all nine cases in which PCR products were observed were found to be infected with B. aalborgi alone. These results indicate that the HIS bacterial body, especially of B. aalborgi, is characterized by terminal βGlcNAc and also indicate that terminal βGlcNAc on the HIS bacterial body is associated with HIS preference for SSLs:
Introduction
Human intestinal spirochetosis (HIS) is a colorectal bacterial infection caused by Brachyspira aalborgi (B. aalborgi) or B. pilosicoli. Although HIS is often discovered as an incidental finding,1,2 it can underlie colitis and cause diarrhea in some individuals. 3 Sessile serrated lesions (SSLs) are precursor lesions of serrated pathway-associated colorectal carcinomas. HIS infection rates are reportedly higher in cases with SSLs in the Japanese population.4,5 Griffonia simplicifolia-II (GS-II) is a lectin specific to terminal α/βGlcNAc residues. 6 Using GS-II-horseradish peroxidase (HRP) staining and HIK1083 immunostaining specific to terminal αGlcNAc residues, we found that increased terminal βGlcNAc is an SSL hallmark. 7 Altered glycosylation of mucin core proteins reportedly can alter the composition of microbiota associated with the mucosa in the gastrointestinal tract. 8 Therefore, the increase in terminal βGlcNAc in SSLs may be associated with HIS preference for SSLs. Here, we investigated terminal βGlcNAc residues in HIS infection to clarify the relationship between glycosylation and HIS colonization in the intestinal tract.
Materials and Methods
Case Selection
We used all 15 HIS cases identified in the pathology archive of Shinshu University Hospital from 2014 to 2021 and performed pathological analysis of coexisting lesions such as SSLs and tubular adenomas (TAs). Use of retrospective tissue samples was approved by the Ethics Committees of Shinshu University School of Medicine (no. 5352).
Immunohistochemistry (IHC) and GS-II-HRP Staining
Hematoxylin and eosin (H&E) staining was performed on 3-μm-thick sections of formalin-fixed, paraffin-embedded (FFPE) tissue blocks. Reagents and procedures used for IHC and GS-II-HRP staining are listed in Table 1. For anti-Treponema immunostaining, we used an antibody reported to cross-react with Brachyspira species.4,9,10 Staining was performed as previously described. 7 Briefly, GS-II-HRP staining was generally performed in the same way as IHC, except that slides were incubated for 30 min with a 0.1% amylase solution to block GS-II reactivity to glycogen after endogenous peroxidase activity blocking. Biotinylated GS-II (Vector Laboratories, Newark, CA) and HRP-conjugated streptavidin (DAKO, Santa Clara, CA) were used instead of primary and secondary antibodies, respectively. Negative controls were: for GS-II-HRP staining, lectin solutions containing 0.1 M GlcNAc, and for IHC, staining solutions lacking primary antibodies. No specific staining was seen in either negative control.
Reagents and Procedures Used in Immunohistochemistry and GS-II-HRP Staining.
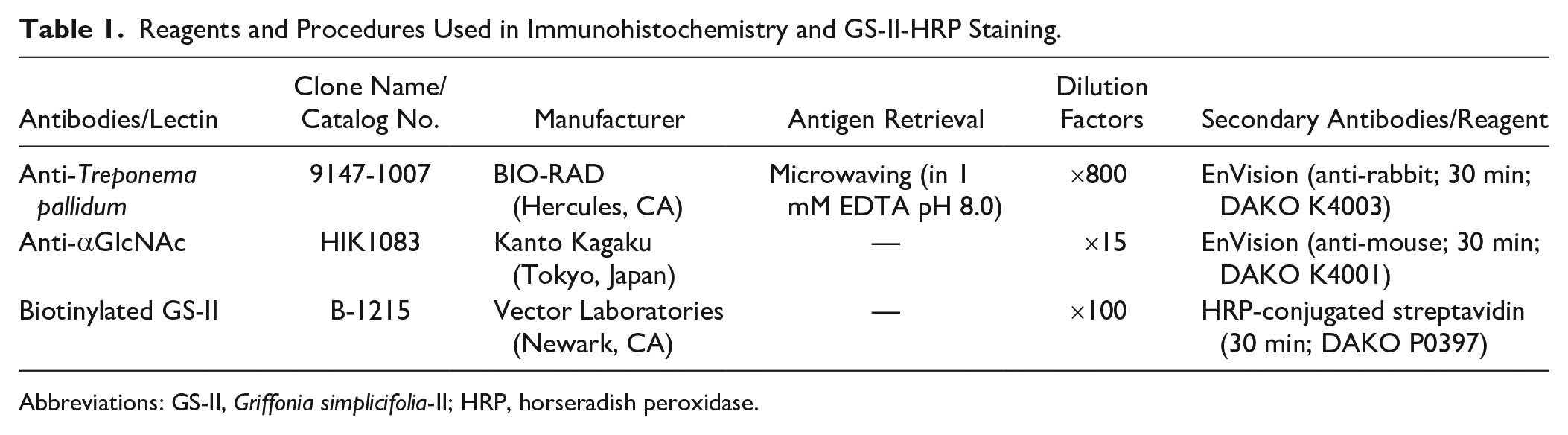

Abbreviations: GS-II, Griffonia simplicifolia-II; HRP, horseradish peroxidase.
Evaluation of IHC and GS-II-HRP Staining
We judged βGlcNAc to be present if cases were GS-II-positive and HIK1083-negative. The percentage of bacterial bodies staining positively for GS-II was evaluated by comparing GS-II-HRP staining and anti-Treponema immunostaining and was classified as three levels: ≤30%, 30–70%, or >70%. In cases with neoplastic lesions, such as SSLs and TAs, GS-II positivity of these lesions was also assessed. The coexistence of SSLs or TAs with HIS was evaluated as coexistent only when the organisms adhered to the mucosa of the lesion itself, not to the surrounding mucosa. Two authors (H.M., M.I.) reviewed immunohistochemical staining and GS-II-HRP staining and reached a consensus for each case.
DNA Extraction and PCR Amplification of NADPH Oxidase (nox) Genes
DNA was extracted from FFPE tissue sections of HIS cases using a QIAamp DNA FFPE Tissue kit (Qiagen, Hilden, Germany). Primes used for PCR amplification of nox genes of B. aalborgi and B. pilosicoli are shown in Table 2. For B. pilosicoli nox gene, the amplification protocol was as follows: initial denaturation at 98C for 15 sec; 35 cycles of denaturation at 98C for 10 sec, annealing at 54C for 30 sec, and extension at 72C for 20 sec, followed by a final extension at 72C for 4 min using ExTaq (Takara Bio, Kusatsu, Japan). Using the initial PCR product as a template, nested PCR was performed on the same protocol using primers inside it. For B. aalborgi nox gene, the PCR amplification protocol was generally similar, but annealing temperature was 51C and Takara Taq (rTaq) was used as it was difficult to amplify with ExTaq. PCR products were electrophoresed on an ethidium bromide-containing 1.0% agarose gel.
Sequences of the Primers Used in PCR Amplification and Direct Sequencing of nox Genes.
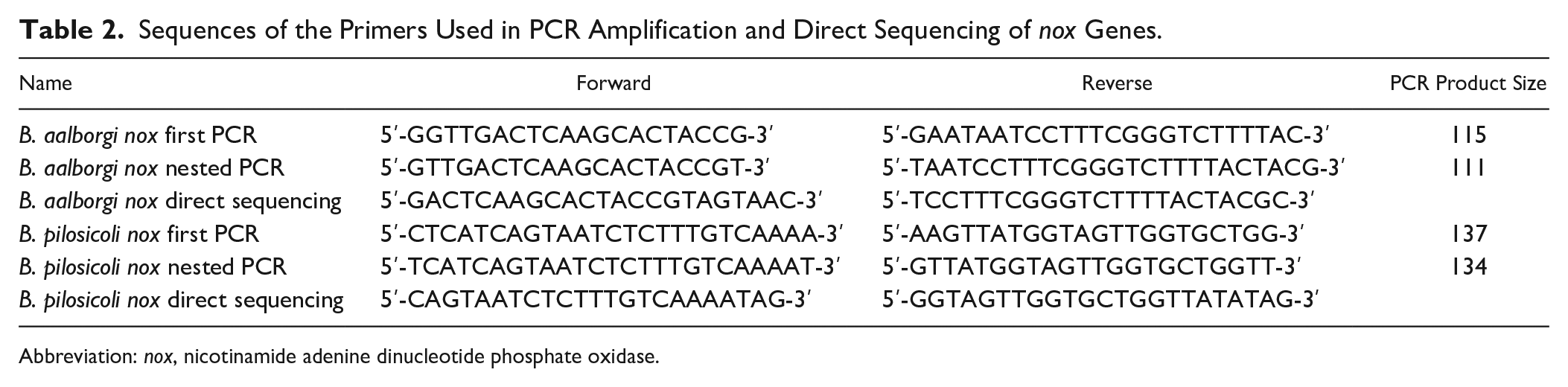
Abbreviation: nox, nicotinamide adenine dinucleotide phosphate oxidase.
Direct Sequencing of PCR Products of nox Genes
Primers used for direct sequencing of nox genes are shown in Table 2. PCR products of nox genes were electrophoresed on agarose gels and purified using a QIAquick Gel Extraction Kit (Qiagen). Subsequently, DNA sequencing reaction was performed using the forward or reverse primer (Table 2) with a BigDye Terminator v1.1 Cycle Sequencing Kit (Thermo Fisher Scientific, Waltham, MA). Sequences were determined using a capillary automatic sequencer ABI 3130XL PRISM Genetic Analyzer (Thermo Fisher Scientific).
Results
GS-II-HRP Staining in HIS Cases
All 15 HIS cases were positive in anti-Treponema immunostaining, consistent with the bacterial bodies identified by H&E staining (Fig. 1). No cases showed HIK1083 positivity (data not shown). Bacterial bodies were GS-II-positive in 14 cases, and the remaining case showed no positivity (Fig. 1, Table 3). We concluded that terminal βGlcNAc was present in these 14 cases since they were GS-II-positive and HIK1083-negative. The percentage of bacterial bodies staining positively for GS-II, which were calculated by comparison with anti-Treponema immunostaining, was ≤30% in seven cases, 30–70% in two, and >70% in six (Table 3). Of 15 HIS cases analyzed, none were comorbid with TAs, and three were comorbid with SSLs (Table 3, Fig. 2). Of the three cases where both HIS and SSLs were present, one showed decreased GS-II positivity on the bacterial body at the SSL site, despite overall high (90%) GS-II positivity in surrounding mucosa (case 15; Table 3, Fig. 2A to F). In this case, the amount of HIS bacterial bodies itself also reduced at the SSL site (Fig. 2A to F). The remaining two cases showed low (20% and 5%) GS-II positivity on the bacterial body at SSL sites (respectively, cases 6 and 12; Table 3, Fig. 2G to I). However, both cases contained almost no surrounding mucosa, and comparisons of GS-II staining on bacterial bodies between in surrounding mucosa and at the lesion site could not be made. In SSLs of these three cases, GS-II-HRP staining was positive in lesions, as we previously reported (Fig. 2C, I). 7
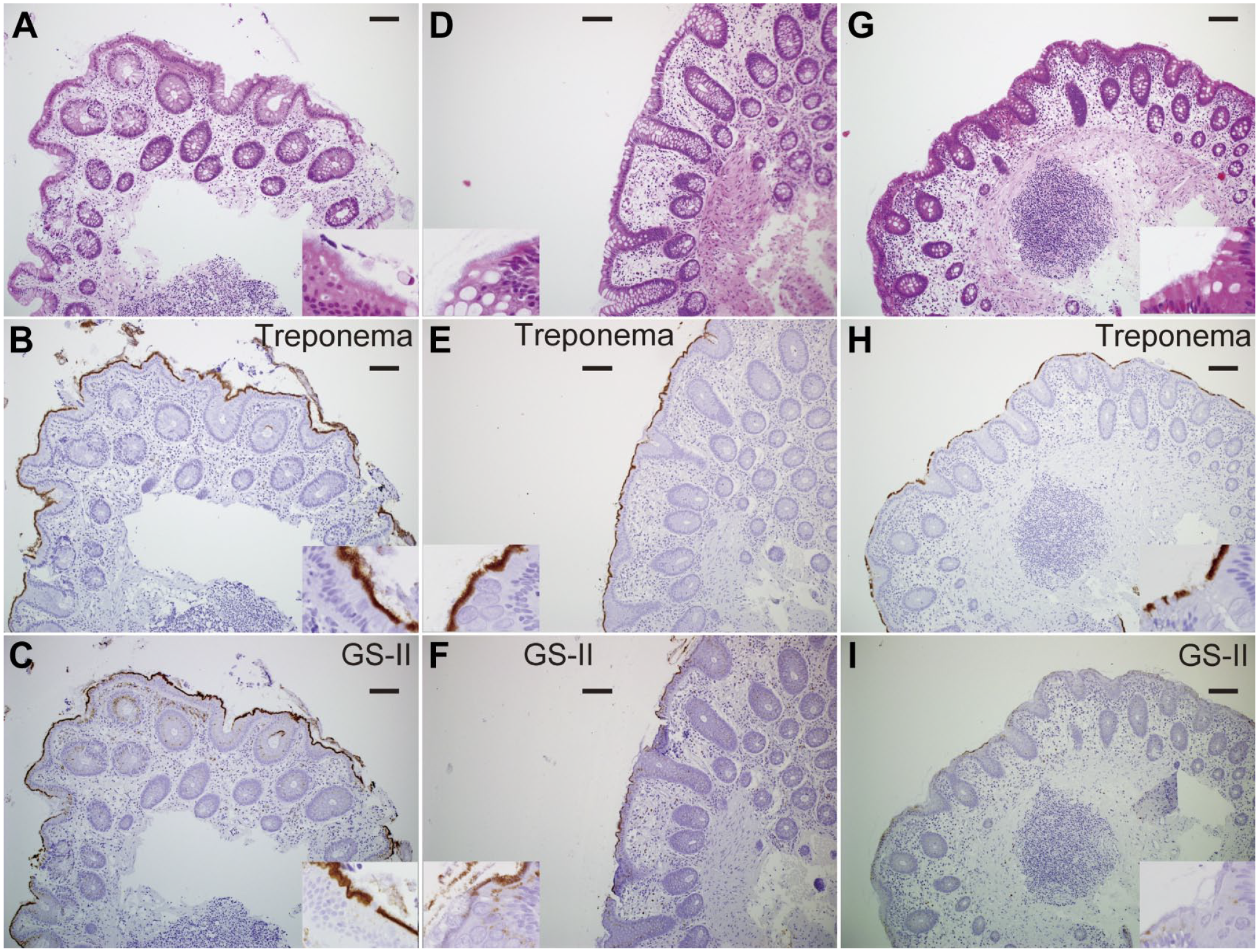
Griffonia simplicifolia-II-horseradish peroxidase (GS-II-HRP) staining and anti-Treponema immunostaining of human intestinal spirochetosis (HIS) cases with high, moderate, and low percentage of bacterial bodies staining positively for GS-II. Hematoxylin and eosin (H&E) staining (A), anti-Treponema immunostaining (B), and GS-II-HRP staining (C) of a HIS case with high level of GS-II staining (case 10). H&E staining (D), anti-Treponema immunostaining (E), and GS-II-HRP staining (F) of a HIS case with moderate level of GS-II staining (case 9). H&E staining (G), anti-Treponema immunostaining (H), and GS-II-HRP staining (I) of a HIS case with low level of GS-II staining (case 13) (Bars A-I = 100 µm).
The Percentage of Bacterial Bodies Staining Positively for GS-II and Coexisting Lesion of 15 HIS Cases.
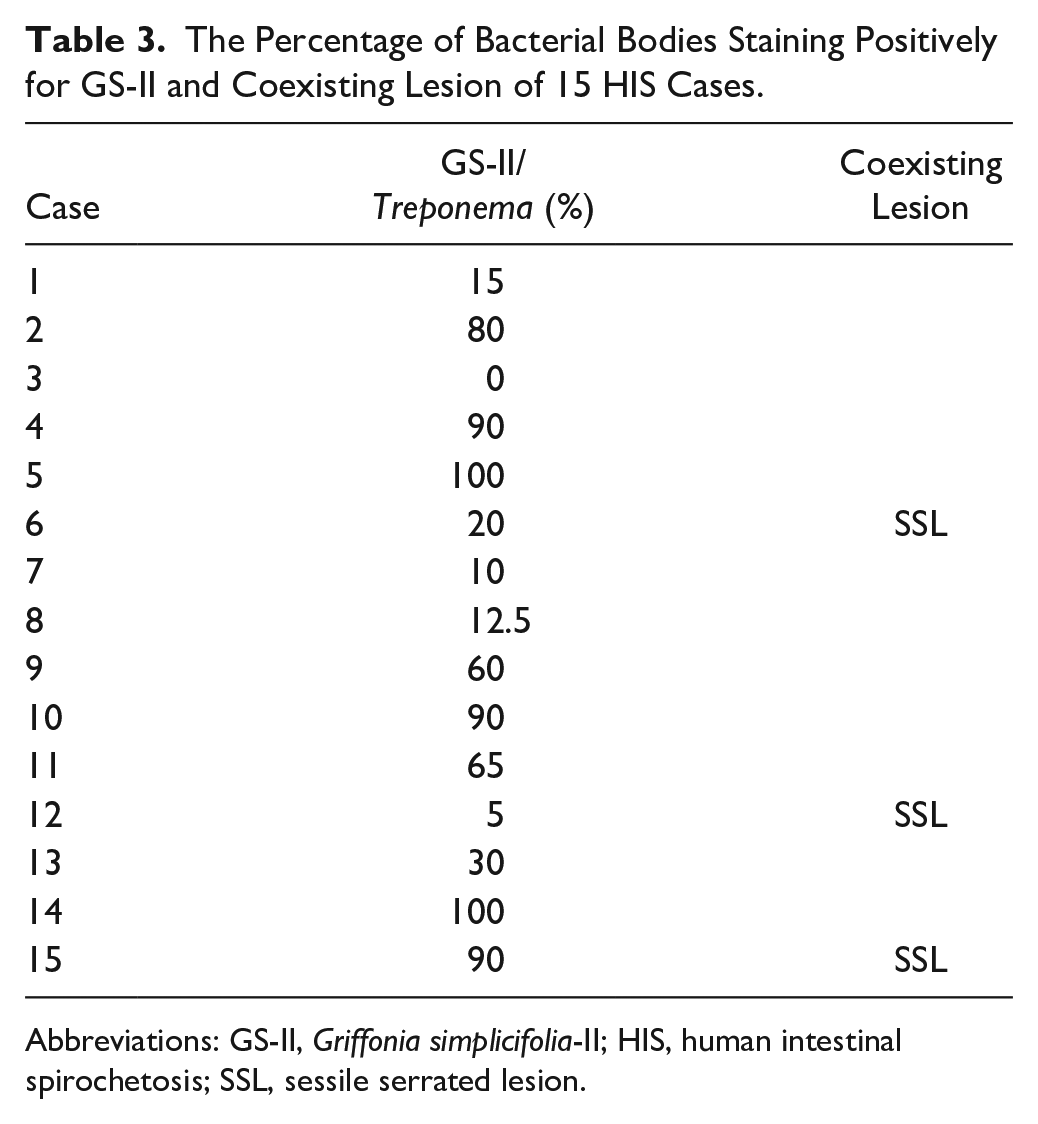
Abbreviations: GS-II, Griffonia simplicifolia-II; HIS, human intestinal spirochetosis; SSL, sessile serrated lesion.
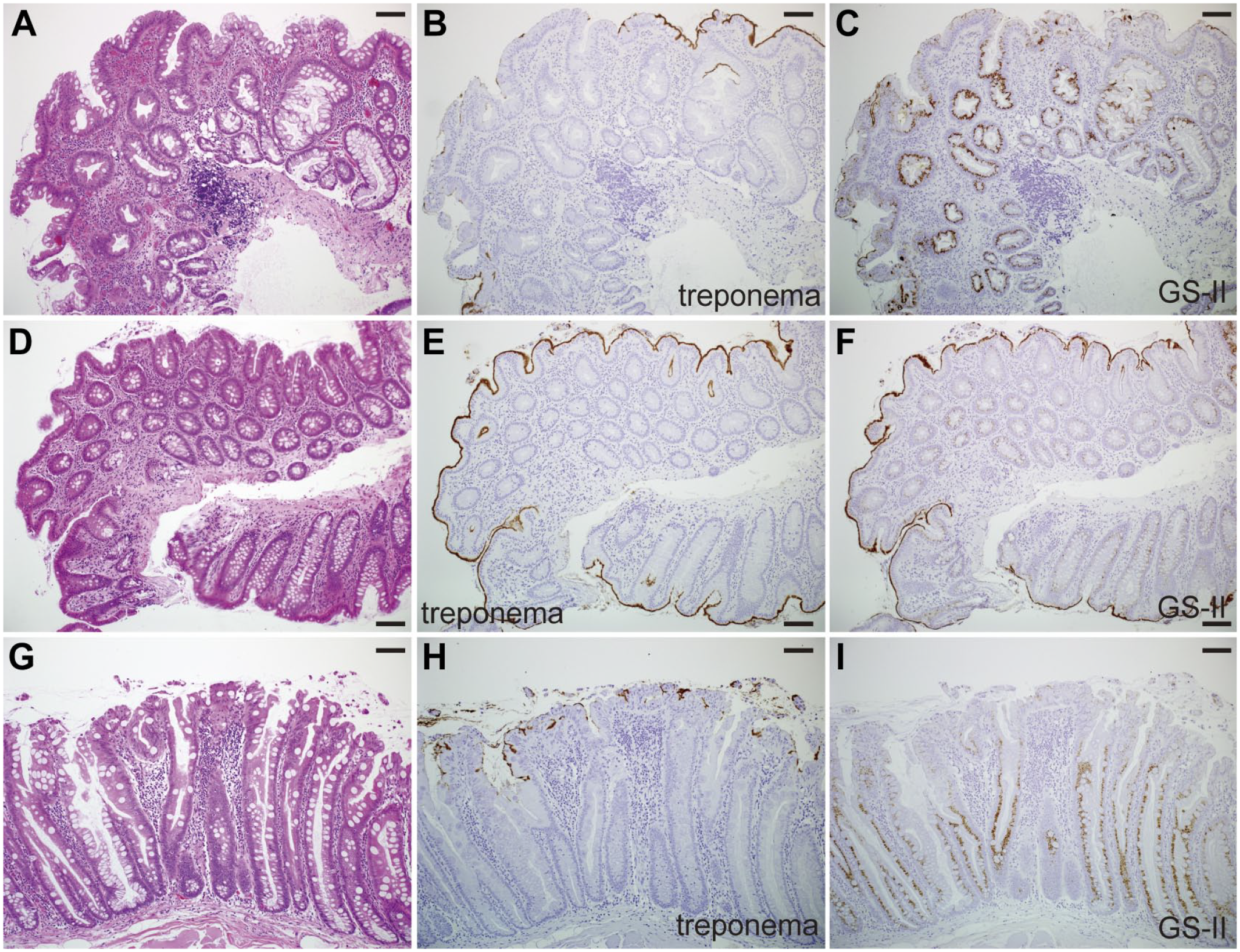
Griffonia simplicifolia-II-horseradish peroxidase (GS-II-HRP) staining and anti-Treponema immunostaining of human intestinal spirochetosis (HIS) cases coexisting with sessile serrated lesions (SSLs). Hematoxylin and eosin (H&E) staining (A and D), anti-Treponema immunostaining (B and E), and GS-II-HRP staining (C and F) at the SSL site (A-C) and in its surrounding mucosa (D-F) of a HIS case (case 15). H&E staining (G), anti-Treponema immunostaining (H), and GS-II-HRP staining (I) at the SSL site of a HIS case (case 12) (Bars A-I = 100 µm).
Detection of Two Strains of Spirochetes by Direct Sequencing After PCR Amplification of nox Genes
All 15 HIS cases were subjected to molecular detection of two strains of spirochetes (B. aalborgi and B. pilosicoli) by PCR amplification of species-specific portion of nox gene. Of the 15 HIS cases, PCR experiments showed distinct and reproducible electrophoretic bands in the recent six cases (cases 10–15). PCR products with B. pilosicoli nox primes were detected in these six cases by electrophoresis (Fig. 3A). PCR products with B. aalborgi nox primers were detected distinctively in four cases (cases 10, 12, 13, and 15) and electrophoretic bands were somewhat obscure in the remaining two cases (cases 11 and 14; Fig. 3A). Direct sequencing of PCR products with B. aalborgi nox primers showed that sequences of five cases (cases 10, 11, 12, 13, and 15) were 100% matched with previously reported B. aalborgi nox gene sequence (accession number AF060816.1, NCBI database of GeneBank) and it was difficult to determine DNA sequence in the remaining one case (case 14). However, direct sequencing of PCR products with B. pilosicoli nox primes revealed that PCR products of all six cases actually have the B. aalborgi nox gene sequence (accession number AF060816.1, NCBI database of GeneBank, as described above) except for primer sites at both ends (Fig. 3B). This sequence differed in seven bases from B. pilosicoli nox gene sequences (accession numbers QQ728895.1, OQ728852.1, OQ728904.1, and JF430756.1, NCBI database of GeneBank; Fig. 3B). Therefore, we judged that these six PCR products were generated by the binding of B. pilosicoli nox primers to B. aalborgi nox gene and that a single infection with B. aalborgi occurred in these six cases because of the lack of B. pilosicoli nox gene as a PCR template. In the remaining nine cases (cases 1–9), distinct and reproducible electrophoretic bands were not observed unlike above six cases. However, when repeated PCR was applied, electrophoretic bands from B. pilosicoli nox primes were detected in three cases (cases 6, 8, and 9; Fig. 3C), although the results were not always reproducible. Direct sequencing of PCR products revealed that PCR products of these three cases actually have the B. aalborgi nox gene sequence (accession number AF060816.1, NCBI database of GeneBank, as described above) except for primer sites at both ends (Fig. 3B) as well as above six cases. Therefore, we judged that a single infection with B. aalborgi also occurred in these three cases for the same reason, despite the absence of PCR products with B. aalborgi nox primers.
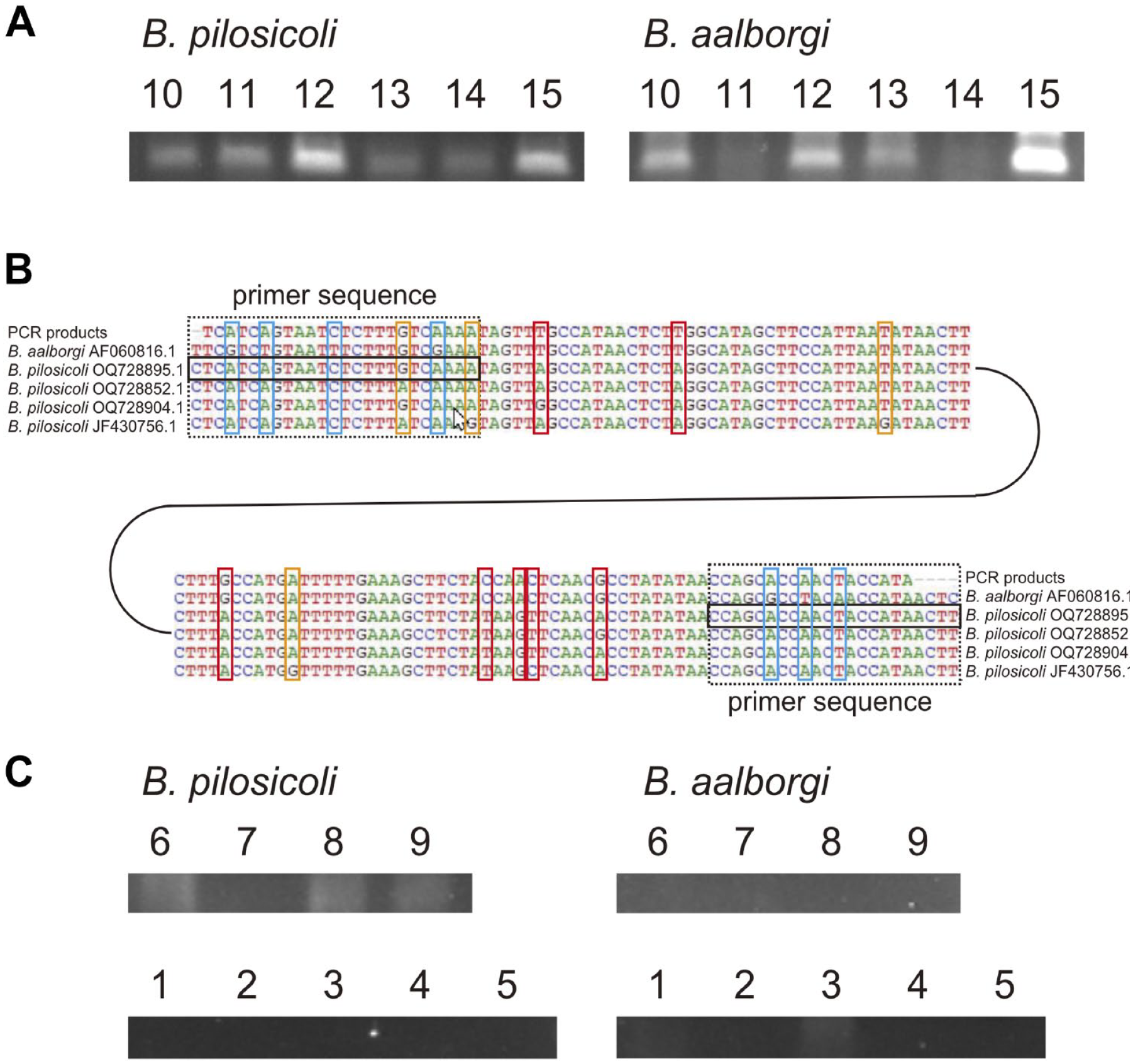
Direct sequencing after PCR amplification of nicotinamide adenine dinucleotide phosphate oxidase (nox) genes in B. aalborgi or B. pilosicoli. Electrophoretic bands of PCR products of nox genes of the two strains derived from six human intestinal spirochetosis (HIS) cases (cases 10–15). Direct sequencing of PCR products described in (A) with B. pilosicoli nox primers (B). PCR products of all six cases have the same sequence corresponding to the top one in (B). Black dotted rectangles at both ends indicate primer sites, and black solid rectangles indicate the actual primer sequences. The blue rectangles indicate bases where the PCR product sequence differs from the B. aaloborgi nox gene sequence at primer sites. The red rectangles indicate bases where the PCR product sequence differs from the B. pilosicoli nox gene sequence in the middle portion. The orange rectangles indicate bases where there are differences between the reported several B. pilosicoli nox gene sequences. Electrophoretic bands of PCR products of nox genes of the two strains derived from remaining nine HIS cases (cases 1–9; C). Results of these nine cases are not always reproducible.
Discussion
HIS is a colorectal bacterial infection with Brachyspira species and found as an incidental finding or with colitis and diarrhea. Here, we reveal that almost all HIS cases showed terminal βGlcNAc on bacterial bodies. These results indicate that terminal βGlcNAc can serve as a biomarker of this microorganism and have implications for the colonization to the intestinal tract or preference for SSLs of this microorganism.
There have been various reports regarding the attachment of HIS to neoplastic lesions such as TAs and SSLs. Some reports indicate that the degree of HIS attachment does not change in TA lesion sites,11–13 whereas some reports indicate a decrease in the amount of HIS attachment in TAs.14–16 Electron microscopic analysis suggests that HIS becomes entrapped between the microvilli of the colonic surface epithelium, and the shortened and obscured microvilli in TAs are thought to inhibit HIS attachment.14,15 Higher rates of HIS infection have been reported in Japanese patients with SSL.4,5 However, it has not been reported whether the degree of HIS adherence differs between the SSL site and the surrounding mucosa. Here, we demonstrated that the relative frequency of HIS in SSL was higher than in TA, and the amount of HIS in one lesion of SSL was apparently reduced compared with the surrounding mucosa. In addition, we also showed that the positive rate of GS-II on HIS bacterial body was low at SSL sites (see Table 3, Fig. 2). Altered glycosylation on mucin has been suggested to alter mucosal-associated microbiota in the gastrointestinal tract. 8 As we previously reported 7 and further confirmed here, SSL secrets mucins carrying terminal βGlcNAc residues. Considering these facts, one possible mechanism for the higher relative frequency of HIS in SSL, despite the reduced degree of HIS attachment in tumorigenic lesions such as TA and SSL, is that exogenous βGlcNAc secreted from SSL promotes HIS colonization and compensates for the reduced attachment in tumorigenic lesions. βGlcNAc on HIS bacterial bodies may be intrinsically required for HIS colonization and survival in the intestinal tract, but this compensation by exogenous βGlcNAc secreted from SSL may result in a decrease in terminal βGlcNAc on HIS bacterial bodies (see Fig. 4). However, further analysis is needed to test this hypothesis.
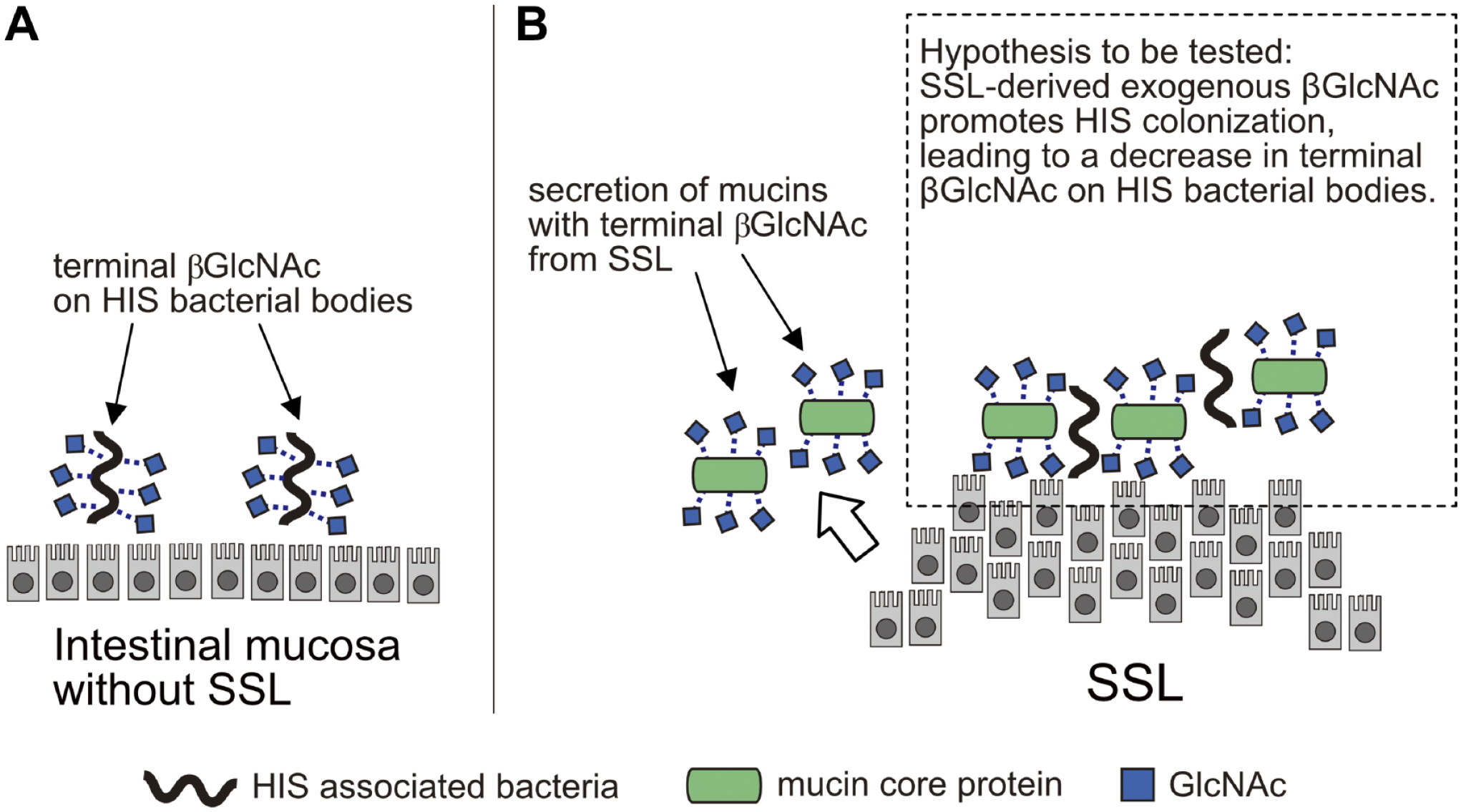
Possible mechanism of human intestinal spirochetosis (HIS) preference for sessile serrated lesions (SSLs) through terminal βGlcNAc. (A) HIS colonization to intestinal mucosa without SSL. (B) SSL secrets mucins carrying terminal βGlcNAc residues. It is speculated that exogenous βGlcNAc from SSL may have promoted HIS colonization, resulting in a decrease in terminal βGlcNAc on HIS bacterial bodies.
We also note that the HIS preference for SSLs reported in the Japanese population4,5 is not as apparent in a study of an Australian population. 10 That Australian study used electron microscopy to analyze the structure of HIS bacterial bodies and questioned the specificity of the antibody used in a series of studies,4,10 because the structure of the microorganisms was not seen in cases with focally positive immunostaining for Treponema. Because that same antibody has been used in these studies,4,10 including this one, its specificity is likely to be a concern. However, it seems unlikely that this antibody cross-reacts with βGlcNAc, as it recognized HIS bacterial bodies but not βGlcNAc in SSLs (see Fig. 2). Further investigation is needed to determine the cause of this difference in HIS preference for SSLs observed in Japanese and Australian populations.
Direct sequencing after PCR amplification of nox genes revealed that a single infection with B. aalborgi occurred in nine HIS cases (see Fig. 3). These results are similar to previous studies where B. aalborgi infection was more prevalent than B. pilosicoli infection in Japan and Western countries including Australia.9,17–20 Although B. pilosicoli induces spirochetosis in many kinds of animals, including humans, primates, various mammals, and birds, B. aalborgi is only known to infect humans and higher primates.21–23 Therefore, these results suggest that B. aalborgi have a stronger preference for humans than B. pilosicoli regardless of location. In addition, of the six cases in which PCR products were not obtained, especially cases 1–5 were from 2015 or earlier, suggesting that deterioration or degradation of DNA in the paraffin sections was the cause of the failure to obtain adequate results.
In conclusion, we have newly identified terminal βGlcNAc as a biological feature of HIS bacterial bodies, especially bacterial bodies of B. aalborgi. This feature can serve as a biomarker of HIS and may explain HIS preference for SSLs.
Footnotes
Acknowledgements
We would like to express our gratitude to Dr Elise Lamar for her critical review and editing of the manuscript.
Author Contributions
HM designed the research, performed immunostaining, analyzed histopathological data, constructed figures and tables, and drafted the manuscript. MI analyzed histopathological data and provided suggestions for experimental design and editing the manuscript. CF provided suggestions for molecular detection of spirochetes and editing the manuscript. JN provided suggestions for the experimental design and editing the manuscript. All authors approved the final version of the manuscript.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by Grants-in-Aids for Scientific Research 21K15400 from the Japan Society for the Promotion of Science (Hisanori Matoba).
