Abstract
Retinal astrocytes are vital for neuronal homeostasis in the retina. Together with Müller glia, they provide retinal cells with neurotrophic factors, antioxidative support, and defense mechanisms such as the formation of the blood-retinal barrier. Substantial heterogeneity of astrocyte morphology and function represents a challenge for identification of distinct subtypes which may be potential targets for therapeutic purposes. Hence, identification of novel markers of astrocyte subpopulations is highly relevant to better understand the molecular mechanisms involved in retinal development, homeostasis, and pathology. In this study, we observed that the cell cycle regulator, p16INK4a, is expressed in immature astrocytes in the mouse retina. Immunohistochemical analysis showed p16INK4a expression in the optic nerve of wild-type mice from 3 days to 3 months of age and in the nerve fiber layer of the adult mouse retina. Colocalization of p16INK4a expression and glial fibrillary acidic protein (immature/mature astrocyte marker) tends to decrease with age. However, colocalization of p16INK4a expression and vimentin (immature astrocyte marker) remains high in the optic nerve from the early postnatal period to adulthood. The observations from this study provide a valuable tool for further investigations of ocular astrocytes in the developing retina as well as in degenerative retinopathies.
Introduction
Retinal macroglia consist of two classes of cells: astrocytes and Müller cells. While Müller cells span the neural retina from the outer to the inner limiting membrane, retinal astrocytes are predominantly confined to the nerve fiber layer (NFL) and ganglion cell layer. 1 Diverse and complex functions have been identified for retinal glial cells. Astrocytes and Müller cells are both essential for overall retinal health and are particularly important for trophic and metabolic support of neurons including neuronal synaptic activity and neurotransmitter homeostasis.2 –4 These cells, as immune cells, can also proliferate and migrate in response to different types of damage to defend and protect the neurons. 5 However, when this activation is sustained over time, reactive gliosis can represent a key pathological feature. Reactive gliosis is thus considered an inherent element in many retinal pathologies including proliferative diabetic retinopathy, glaucoma, age-related macular degeneration, and retinitis pigmentosa.6,7 Consequently, ocular astrocytes play a central role in retinal health and disease and thus provide a potential target for novel therapeutic approaches.
Astrocytes are generally considered the most abundant glial cell in the central nervous system (CNS) but are morphologically as well as functionally somewhat heterogeneous,2,8 and the current understanding of the individual subtypes, morphology, and function is very limited. Furthermore, retinal astrocytes differ morphologically between species; for example, in mice and rats, astrocytes are classified as star-shaped in contrast to the human retina where two morphologically distinct types of astrocytes have been identified.6,9 Consequently, identifying astrocyte subtypes based on morphology is far from ideal, and there is a pressing need for selective and accurate markers enabling identification of astrocyte subtypes.
In rodents, during the late embryogenesis/early postnatal period, a mix of astrocyte precursor cells and immature astrocytes migrate into the retina from the optic nerve1,8,10,11 and spread peripherally across the NFL toward the margins of the neuroretina. During their migration, the astrocyte precursor cells undergo sequential differentiation through three defined stages. In the first differential stage, immature perinatal astrocytes express glial fibrillary acidic protein (GFAP) in addition to paired box gene 2 (PAX2) and vimentin. In mature perinatal astrocytes, the second differential stage, loss of vimentin expression is observed, and eventually in the final stage, mature astrocytes have lost PAX2 expression while still presenting a robust GFAP expression.8,12 Due to the vast heterogeneity of astrocytes and their complex maturation, new and additional selective markers are needed to improve our limited understanding of the functions and roles of astrocyte subtypes and astrocyte precursor cells in retinal development, health, and disease.
The Cdkn2a gene encodes for two different proteins that are translated from alternative first exons, p16INK4a and p14ARF, which act as cell cycle regulators and tumor suppressors. 13 p16INK4a is a key player in the retinoblastoma protein pathway controlling the G1 to S transition. 14 Through binding to cyclin-dependent kinases 4 and/or 6 (CDK4/6), p16INK4a inhibits cyclin D–CDK4/6 complex formation, ultimately leading to G1 cell cycle arrest, 14 which is associated with several physiological processes such as cancer, aging, and cellular senescence. p16INK4a was established as a tumor suppressor gene decades ago and beautifully illustrates the direct link between cell cycle control and cancer as it is often seen inactivated/downregulated in a variety of tumor types. 15 However, overexpression of p16INK4a has also been reported in some tumor types, demonstrating the complex role of p16INK4a in cancer pathology. 15 p16INK4a is a hallmark of aging and is widely used as a biomarker for detecting senescent cells.16 –18 Cells may enter into a protective state of irreversible cell cycle arrest, known as senescence, in response to cellular replicative exhaustion as well as a number of other stressors including genomic and epigenomic damage, irradiation, oxidative stress, and genotoxic drugs. 19 But several studies have shown that p16INK4a may be involved in other biological processes other than cancer and senescence. 20 p16INK4a has been found to be crucial for the differentiation of different cell types, such as myofibroblasts and keratinocytes in processes of tissue repair and wound healing.21,22 p16INK4a was also detected in postnatal mouse primary brain astrocytes nearly two decades ago 23 ; however, the functional role in these cells still remains to be established.
The findings of p16INK4a in mouse postnatal brain astrocytes prompted us to investigate the presence and localization of p16INK4a in the normal mouse retina from the early postnatal period through to the fully developed adult retina. In this study, we show that p16INK4a is extensively expressed in the cytoplasm of a subpopulation of ocular astrocytes in the mouse retina as early as postnatal day 3 (PND3), and that expression persists throughout retinal development although to a more limited extent. Immunohistochemical colocalization studies of p16INK4a and markers for immature differentiation stages of astrocytes further suggest that p16INK4a is expressed in immature astrocytes in the optic nerve as well as in a small population within the NFL of the adult mouse retina.
Traditional protein and RNA quantitative techniques provide valuable quantitative data on gene expression and protein levels but do not offer multifaceted details on cellular localization and/or migration. Due to the overall cellular morphological complexity and heterogeneous composition of the retina, immunohistochemical investigations are an essential tool for detailed studies of exactly that. The results presented here suggest that p16INK4a may be a marker for selective and specific identification of immature/incompletely differentiated ocular astrocytes in the retina and thus may prove a novel and important tool for further investigation of the roles of ocular astrocytes in retinal development and disease.
Methods
Animals
Animal experiments and breeding were approved by the Oxford Animal Ethics Committee in accordance with the UK Home Office Guidelines on the Animal (Scientific Procedures) Act, 1986 as well as the Association for Research in Vision and Ophthalmology statements on the care and use of animals in ophthalmic research.
B6.Cg-Tg(Nrl-EGFP)1Asw/J mice expressing an enhanced green fluorescent protein (EGFP) under the control of the neural retina leucine zipper (Nrl) promoter (kind gift from Anand Swaroop, National Eye Institute, Bethesda, MD), hereinafter called Nrl-EGFP mice, were used throughout the study.
Quantitative Real-time PCR Analysis
cDNA was synthesized by reverse transcription from 0.8 µg of total RNA extracted from the retina of Nrl-EGFP mice at ages PND3, 3 weeks, and 3 months (n=6 per cohort). Taqman chemistry-based quantitative polymerase chain reaction (qPCR) was used to quantify the relative expression of Cdkn2a (Taqman Gene Expression Assay Mm01257348_m1, specific for Cdkn2a transcript NM_009877.2 exon 1–2 boundary, Life Technologies, Inchinnan, Scotland, UK). Glyceraldehyde 3-phosphate dehydrogenase (Gapdh) was used as a housekeeping gene control (Taqman Gene Expression Assay Mm99999915_g1, Life Technologies, Inchinnan, Scotland, UK).
Immunohistochemistry
Retinal cells expressing p16INK4a, GFAP, vimentin, and PAX2 were identified by immunofluorescence imaging of ocular cryosections from Nrl-EGFP mice aged 3 days, 3 weeks, and 3 months (n=3–5 for all ages). Eyes were enucleated and fixed in 4% paraformaldehyde for 30 min at room temperature (RT). Eyes from animals aged 3 weeks and 3 months were subsequently dissected, and cornea, lens, and iris were removed. Due to size, no dissection was performed on enucleated eyes from animals aged 3 days. Eyes were cryoprotected sequentially in a 10%–30% sucrose gradient and embedded in an optimal cutting temperature compound (VWR Chemicals, Leicestershire, UK) and frozen on dry ice using ice cold isopentane. Sections measuring 18 μm in thickness were prepared using a cryostat (Leica Biosystems, Newcastle upon Tyne, UK) and mounted on poly-L-lysine slides. Sections were permeabilized using 0.2% Triton X-100 (X100-100ML, Sigma-Aldrich, Dorset, UK) and blocked with 10% protein block (ab64226; Abcam, Cambridge, UK) in 0.1% Triton X-100 for 10 min and subsequently incubated with primary antibody over night at 4°C before final incubation with fluorophore-coupled secondary antibody for 2 hr, followed by mounting with a ProLong Diamond antifade mount with DAPI (P36962; ThermoFisher Scientific, Inchinnan, Scotland, UK).
Images were acquired using the LSM-710 inverted confocal microscope system (Zeiss, Oberkochen, Germany) with a pinhole setting of 1 airy unit. All images were processed using ImageJ and Adobe Photoshop software.
Antibodies
Primary antibodies used were rabbit polyclonal anti-p16INK4a N-terminal (1:1000) (ab189034; Abcam, Cambridge, UK), chicken polyclonal anti-GFAP (1:2000) (ab4674; Abcam, Cambridge, UK), chicken polyclonal anti-vimentin (1:1000) (ab24525; Abcam, Cambridge, UK), rabbit polyclonal anti-PAX2 (1:200) (71–6000; Invitrogen, Inchinnan, Scotland, UK), and goat polyclonal anti-PAX2 (1:1000) (AF3364; R&D systems, Abingdon, UK).
Secondary antibodies used were (all Alexa Fluor conjugated antibodies from ThermoFisher Scientific, Inchinnan, Scotland, UK) goat antirabbit F(ab’)2 cross-absorbed secondary fluor plus 555 (1:1000) (A48283), goat anti-rabbit 568 (1:1000) (A11036), goat anti-chicken 647 (1:1000) (A21449), donkey anti-goat 647 (1:1000) (A21447), or donkey anti-goat 568 (1:1000) (A11057).
p16INK4a Antibody Validation
To validate the rabbit polyclonal anti-p16INK4a antibody, liver tissue from the p16-Cre/R26-mTmG mouse model kindly provided by Dr. Dmitry V. Bulavin and Dr. Laurent Grosse was used. This knockin mouse model, fully characterized by Grosse et al., 24 was generated for continuous labeling of p16. Immunofluorescence staining of paraffin sections of a liver from a 1+-year-old mouse was performed to detect p16-positive cells: intrinsic expression (EGFP) and by using the rabbit polyclonal anti-p16INK4a N-terminal (ab189034; Abcam, Cambridge, UK). Briefly, 8-µm sections were deparaffinized and rehydrated before proceeding with a heat-induced antigen retrieval step. Sections were then blocked for 1 hr at RT using the serum-free protein block (ab64226; Abcam, Cambridge, UK) and incubated with the primary antibody overnight at RT (dil 1:50). After several short washes, sections were incubated with the secondary antibody goat antirabbit AF647 (#A21245; ThermoFisher Scientific, Inchinnan, Scotland, UK) (dil 1:500) for 1 hr at RT and counterstained with Hoechst (#62249, Life Technologies, Inchinnan, Scotland, UK). Slides were mounted using SlowFade Gold Antifade Mountant (#S36936, Life Technologies, Inchinnan, Scotland, UK) and imaged using the LSM-710 inverted confocal microscope system (Zeiss, Oberkochen, Germany).
Colocalization Analyses
Analyses were performed on individual confocal sections of acquired z-stacks using a 40× oil immersion objective with a pinhole of 1 Airy Unit. Co-occurrence analyses were performed using the Celleste 4.1 software’s (Invitrogen) “colocalization overlap function” with automated thresholding of both fluorophores as this method does not take individual pixel intensity into account. The colocalization overlap coefficient is calculated for each individual confocal section in a given z-stack using the general equation
Three-dimensional rendering
Confocal data (lsm format) was opened in Fiji (ImageJ) and separated into individual channels before saving in tiff format. Subsequently, the tiff files were opened in Chimera (UCSF) and rendered in 3D. Partial transparency was set for the vimentin and GFAP signals to allow the colocalization/overlap with p16INK4a to be observed.
Statistical Testing
The GraphPad Prism software (version 8.2.0; GraphPad Inc., San Diego, California, USA) was used to analyze colocalization and RNA expression data. The nonparametric method, Kruskal–Wallis test, was used to compare data that are not normally distributed. The parametric equivalent test, one-way analysis of variance (ANOVA), was used to analyze differences between normally distributed data obtained from the three groups (PND3, 3 weeks, and 3 months).
RNA Sequencing Data Analyses
Transcript per million (TPM) normalized bulk RNA sequencing data, and sample annotations of human embryonic stem cells (hESCs) differentiated to astrocytes 25 were downloaded from https://github.com/3D-Neural-NBME/Bulk-RNA-seq-analysis/tree/master/Data. TPM values were log2-transformed with an offset of 1. For each gene, values were mean centered with a standard deviation of 1 (Z score). Heatmap visualization was performed using the R package ComplexHeatmap. 26 Data handling was performed in R version 3.6.1. 27
Results
p16INK4a Is Expressed in the Optic Disk and the NFL in the Mouse Retina
The antibody used in this study to detect p16INK4a protein has been thoroughly and repeatedly validated on mouse tissue and mouse cell lysates using immunohistochemistry and western blotting in over 30 peer-reviewed publications. 28 Nevertheless, we have validated its specificity on liver samples from the p16-Cre/R26-mTmG knockin mouse model with intrinsically EGFP-labeled p16. Fluorescent images showed colocalization between EGFP and p16 protein detected with the rabbit polyclonal anti-p16INK4a N-terminal (ab189034; Abcam). As negative control, a sample not incubated with the primary antibody only showed EGFP-positive cells (Appendix Fig. 1).
Immunohistochemical analyses of mouse ocular histological cross-sections showed a strong cytoplasmic p16INK4a signal in stellate cells resembling astrocytes in the optic nerve at ages PND3, 3 weeks, and 3 months (Fig. 1A, D, and G). p16INK4a was furthermore detected in sporadic stellate cells throughout the NFL in mice aged 3 weeks and 3 months but only detected in stellate cells in the NFL close to the optic disk area in PND3 retinas (Fig. 1B, C, E, F, H, and I).
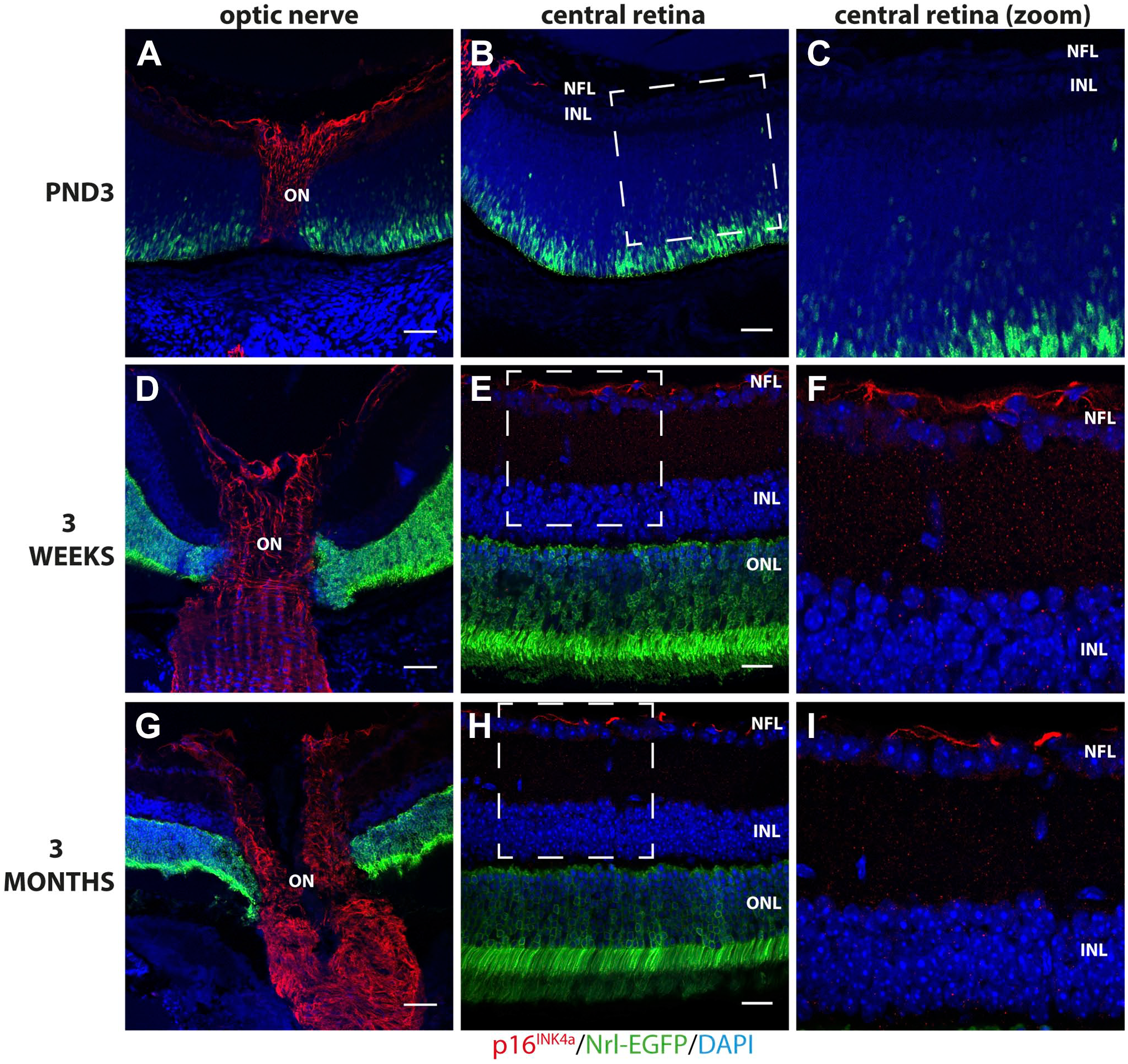
Immunohistochemical detection of p16INK4a in the optic nerve (ON) and NFL of Nrl-EGFP mouse retina sections aged 3 days (PND3), 3 weeks, and 3 months. A strong cytoplasmic p16INK4a signal (red) was detected in stellate cells in the ON at all ages investigated (A, D, G) but only detected in stellate cells in the NFL optic disk area in PND3 retinas (B and C). In contrast, p16INK4a was detected in sporadic stellate cells throughout the NFL in mice aged 3 weeks (E and F) and 3 months (H and I). Rabbit polyclonal anti-p16INK4a detected with Alexa donkey anti-rabbit-568 (red) in mouse retinal tissue expressing EGFP under the control of the Nrl-promoter (green). Tissue is counterstained with Hoechst nuclear stain (blue). Abbreviations: EGFP, enhanced green fluorescent protein; PND3, postnatal day 3; NFL, nerve fiber layer; Nrl, neural retina leucine zipper; INL, inner nuclear layer; ONL, outer nuclear layer. Scale bars: optic nerve panel (A, D, G) 100 µm, central retina panel (B, E, H) 50 µm.
To validate the p16INK4a signal observed in the immunohistochemical investigations of mouse ocular sections, p16INK4a retinal mRNA expression was analyzed using quantitative reverse transcription PCR (RT-qPCR). p16INK4a mRNA expression was detected at all ages investigated using a Taqman probe specific for the p16INK4a transcript. The level of expression was significantly lower in 3-week and 3-month-old retinas than in PND3 retinas (*p<0.0001, one-way ANOVA) (Fig. 2A). The amount of p16INK4a cDNA detected at all the ages investigated was moderate—the cycle threshold (Ct) values ranged between 28.92 and 38.36. The specific 55 base pair band corresponding to p16INK4a cDNA amplified during the qPCR was visualized in an agarose gel (Fig. 2B).
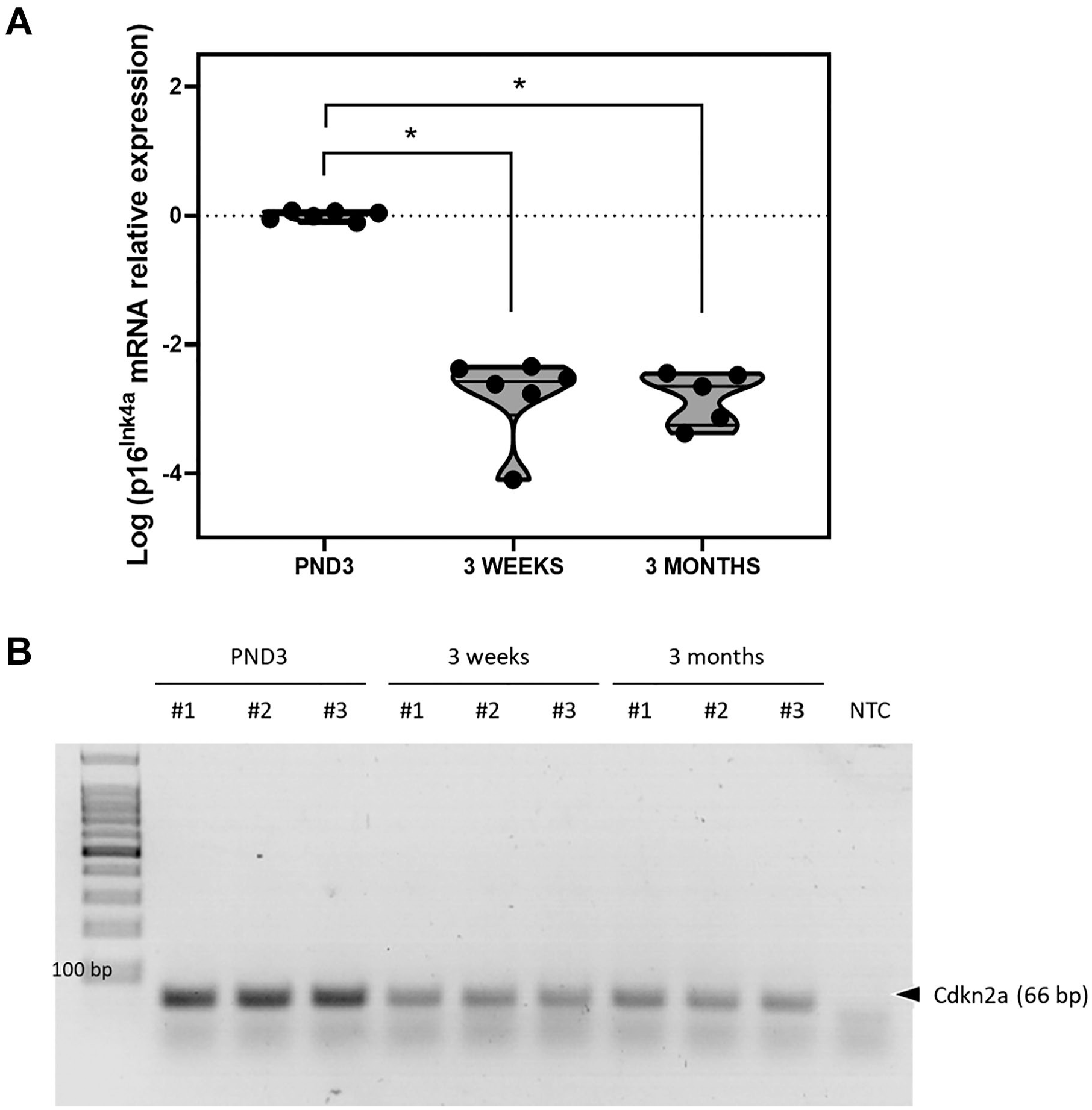
p16INK4a mRNA expression decreases from postnatal day 3 to 3 months in the mouse retina. (A) Scatter plot representing the relative expression of p16INK4a as log-transformed data in the three cohorts tested. p16INK4a expression is significantly lower in 3-week and 3-month-old retinas than in 3-day-old retinas (*p<0.0001, one-way ANOVA) (n=5–6). (B) Agarose gel electrophoresis showing the 66-base pair p16INK4a band amplified in three samples per cohort. Abbreviation: NTC, no template control; ANOVA, analysis of variance.
p16INK4a Is Expressed in Mouse Retinal Astrocytes
To determine if p16INK4a was expressed in the cytoplasm of astrocytes, ocular sections were coincubated with antibodies against p16INK4a and GFAP, an established astrocyte marker. Substantial colocalization between p16INK4aand GFAP was observed in most cells in the optic nerve in all ages investigated (Fig. 3A–E [3 weeks], Appendix Fig. 2A–E [PND3], and Appendix Fig. 3A–E [3 months]). Colocalization of p16INK4aand GFAP was also detected in a subset of stellate cells in the NFL of the area surrounding the optic nerve head at PND3 (consistent with astrocyte migration and population of the postnatal mouse retina) and along the full NFL in ages 3 weeks and 3 months (in a pattern not consistent with activated Müller cells) (Fig. 3F–J [3 weeks], Appendix Fig. 2F–J [PND3], and Appendix Fig. 3F–J [3 months]).
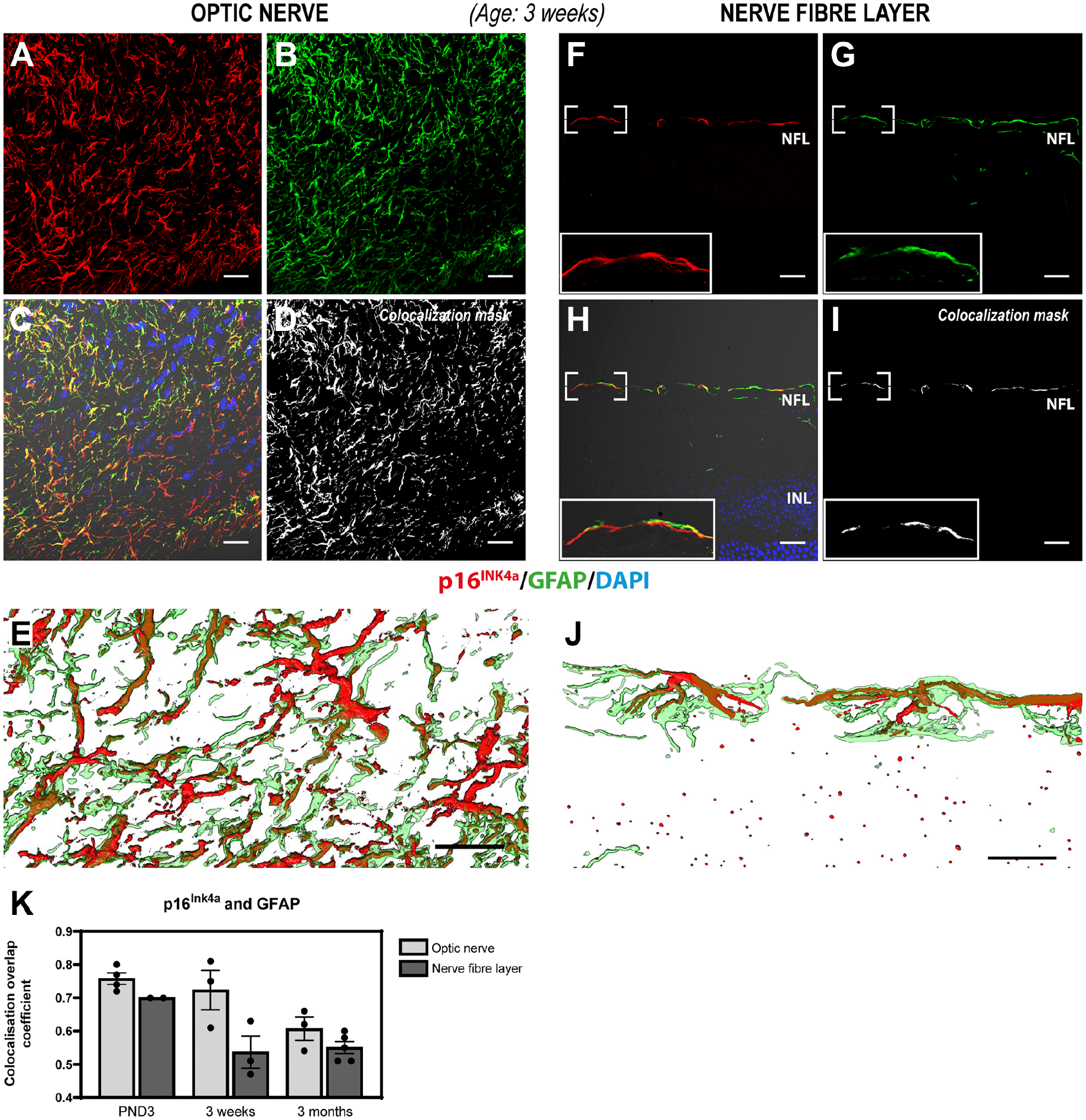
Immunohistochemical detection of p16INK4a (red) and GFAP (green) in the optic nerve (A–E) and NFL (F–J) of mouse retinal sections aged 3 weeks. Substantial overlap between p16INK4aand GFAP was observed in stellate cells in both the optic nerve and the NFL in a pattern not consistent with activated Müller cells, suggesting that p16INK4a is expressed in the optic nerve and a subpopulation of NFL astrocytes. Panels A–C depict the p16INK4a (A), GFAP (B) signals, and the merged image (C) including differential interference contrast (DIC) and Hoechst nuclear stain from a single confocal section of the optic nerve. (D) Colocalization mask for p16INK4a and GFAP in the optic nerve. (E) Zoom 3D-rendering of the substantial overlap of p16INK4a and GFAP signals in the optic nerve from a series of acquired confocal sections within the depicted area. Panels F–H depict the p16INK4a (F) and GFAP (G) signals, as well as the merged image (H) including DIC and Hoechst nuclear stain from a single confocal section of the retina NFL. (I) Colocalization mask for p16INK4a and GFAP in the NFL. (J) Zoom 3D-rendering of the substantial overlap of p16INK4a and GFAP signals in the NFL from a series of acquired confocal sections within the depicted area. (K) Scatter plot with a bar graph representing the colocalization overlap coefficient for each individual sample. No significant differences were found between the different areas analyzed (optic nerve vs NFL) or between the three ages investigated (p>0.5, Kruskal–Wallis test, n=2–5). Nevertheless, the overlap coefficient values range between 0.5 and 0.8, indicating that there is a moderate to strong colocalization between p16INK4a and GFAP, mainly at PND3. Rabbit polyclonal anti-p16INK4a detected with Alexa donkey anti-rabbit-568 (red) and chicken polyclonal anti-GFAP detected with goat anti-chicken-647 (green) in mouse retinal tissue expressing EGFP under the control of the Nrl-promoter (EGFP signal not shown). Tissue is counterstained with the Hoechst nuclear stain (blue). Abbreviations: EGFP, enhanced green fluorescent protein; GFAP, glial fibrillary acidic protein; PND3, postnatal day 3; NFL, nerve fiber layer; Nrl, neural retina leucine zipper; INL, inner nuclear layer. Scale bars: 20 µm.
Quantitative analysis of p16INK4a and GFAP colocalization shows a greater level of colocalization in the optic nerve than in the NFL at all ages investigated (average overlap coefficient ranging from 0.76 at PND3 to 0.61 at 3 months of age) (Fig. 3K). Although there is no statistically significant difference, there was a trend of decreased colocalization in PND3 retinas compared with that in 3-month-old retinas (p>0.05, Kruskal–Wallis test, n=3–4). In the NFL, the colocalization between p16INK4a and GFAP was more substantial at PND3 but showed a tendency to decrease from this point (p>0.05, Kruskal–Wallis test, n=2–5) (Fig. 3K). Interestingly, 3D-rendering of the z-stacks revealed that p16INK4a appears to be largely located to the radial cellular processes of the astrocytes (Fig. 3J and Supplemental Videos 1 and 2). These observations are in agreement with the timeline of astrocyte migration and population of the postnatal mouse retina. 8 These results suggest that p16INK4a is expressed in optic nerve astrocytes as well as in a subset of NFL astrocytes. In contrast to the widespread p16INK4a expression observed in the optic nerve (in all ages investigated), it appears that the adult mouse retina only retains a small population of p16INK4a-expressing astrocytes in the NFL (Fig. 3F–J [3 weeks] and Appendix Fig. 3F–J [3 months]).
p16INK4a Expression Is Confined to Immature Perinatal Astrocytes Present in the Optic Disk of Young and Adult Mice and in the NFL Only at Early Postnatal Age
To determine if p16INK4a is expressed in immature astrocytes, we coincubated them with antibodies against p16INK4a and vimentin, a marker for immature perinatal astrocytes. 8 Colabeling of p16INK4a and vimentin showed a substantial overlap of expression in the optic nerve at all ages investigated (Fig. 4A–E [3 weeks], Appendix Fig. 4A–E [PND3], and Appendix Fig. 5A–E [3 months]). This suggests that the mouse optic nerve retains immature perinatal astrocytes in the adult optic nerve and, furthermore, that p16INK4a is expressed in immature astrocytes.
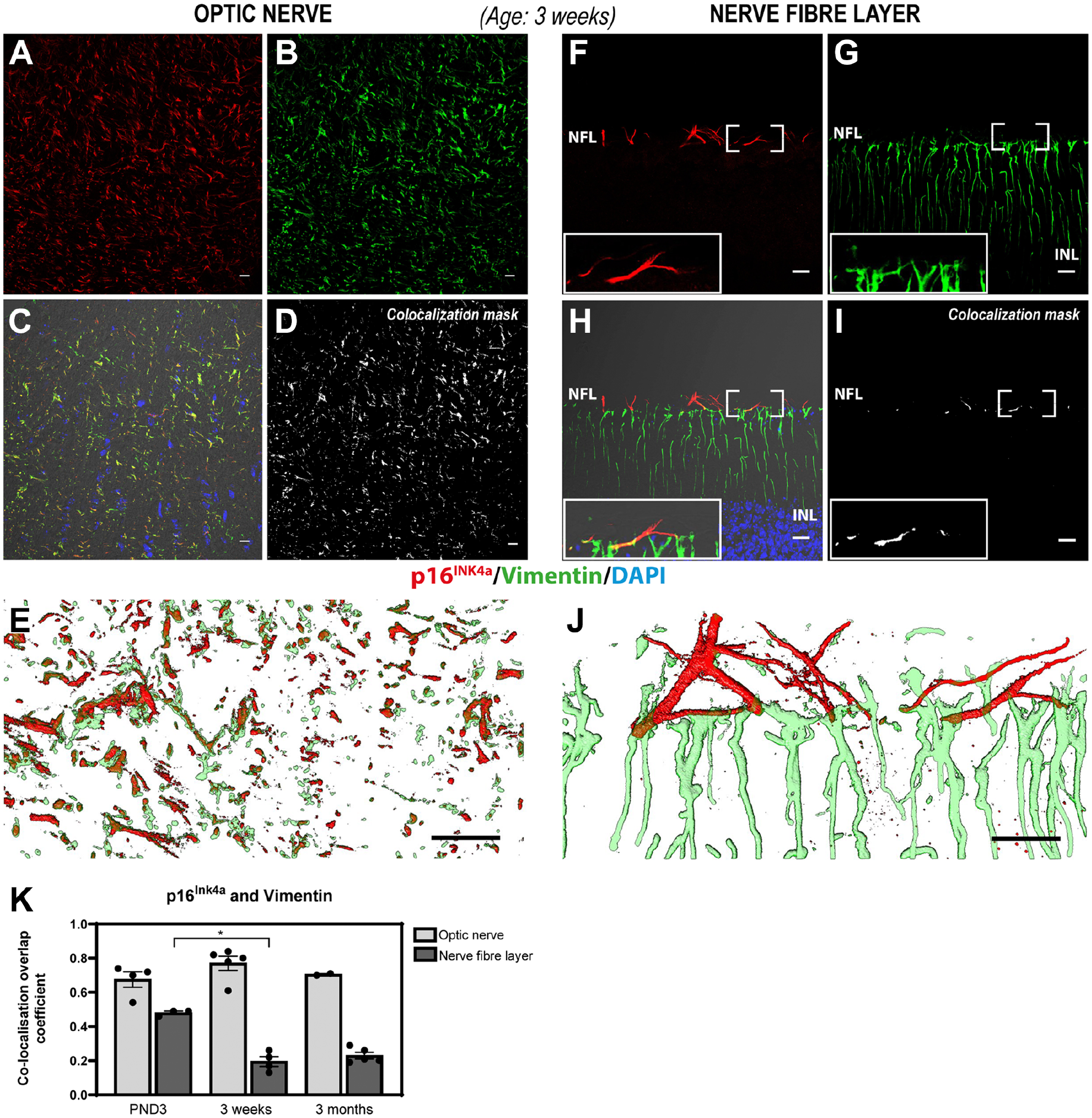
Immunohistochemical detection of p16INK4a (red) and vimentin (green) in the optic nerve (A–E) and the NFL (F–J) of mouse retinal sections aged 3 weeks. A substantial overlap between p16INK4a and vimentin was observed in stellate cells in the optic nerve but not in the NFL suggesting that p16INK4a is expressed in optic nerve astrocytes but not in NFL astrocytes. Panels A–C depict the p16INK4a (A) and vimentin (B) signals, as well as the merged image (C) including DIC and Hoechst nuclear stain from a single confocal section of the optic nerve. (D) Colocalization mask for p16INK4a and vimentin. (E) Zoom 3D-rendering of the substantial overlap of p16INK4a and vimentin signals in the optic nerve from a series of acquired confocal sections within the depicted area. Panels F–H depict the p16INK4a (F) and vimentin (G) signals, as well as the merged image (H) including DIC and Hoechst nuclear stain from a single confocal section of the retina NFL. (I) Colocalization mask for p16INK4a and vimentin. (J) Zoom 3D-rendering of both p16INK4a and vimentin signals in the NFL from a series of acquired confocal sections within the depicted area. (K) Scatter plot with a bar graph representing the colocalization overlap coefficient for each individual sample. No significant differences in the overlap coefficient value for p16INK4a and vimentin were found in the optic nerve between all ages investigated (p>0.5, Kruskal–Wallis test), n=2–5). There is a statistically significant decrease in the colocalization of p16INK4a and vimentin in the NFL from PND3 to week 3 (p<0.05, Kruskal–Wallis test, n=3–5). Rabbit polyclonal anti-p16INK4a detected with Alexa donkey anti-rabbit-568 (red) and chicken polyclonal anti-vimentin detected with goat anti-chicken-647 (green) in mouse retinal tissue expressing EGFP under the control of the Nrl-promoter (EGFP signal not shown). Tissue is counterstained with Hoechst nuclear stain (blue). Abbreviations: EGFP, enhanced green fluorescent protein; PND3, postnatal day 3; NFL, nerve fiber layer; Nrl, neural retina leucine zipper; INL, inner nuclear layer. Scale bars: 20 µm.
Colocalization analyses of vimentin and p16INK4a in the NFL showed very little overlap in 3-week-old and 3-month-old retinas (Fig. 4F–J [3 weeks] and Appendix Fig. 5F–J [3 months]). At these ages, vimentin was detected in cells spanning the entire retina and not in NFL stellate cells as expected since vimentin is a widely used marker for Müller cells. However, a substantial overlap between vimentin and p16INK4a was observed in the NFL in the optic disk area at PND3 but not peripherally in the NFL (Appendix Fig. 4F–J). Three-dimensional rendering of acquired z-stacks supports the colocalization overlap analysis and clearly illustrates the difference in expression patterns of vimentin in the optic nerve and NFL astrocytes (Fig. 4 [3 weeks] and Supplemental Videos 3 and 4 [3 weeks]).
Quantitative analysis of p16INK4a and vimentin colocalization in the optic nerve shows a strong colocalization between both proteins at all ages (average overlap coefficients ranging from 0.68 to 0.77) (Fig. 4K) but shows no statistically significant difference in the colocalization overlap coefficient between the groups of the ages investigated (p>0.05, Kruskal–Wallis test, n=2–5). In the NFL, a significant decrease in the colocalization of p16INK4a and vimentin from PND3 to 3 weeks of age (p<0.05, Kruskal–Wallis test, n=3–5) was observed. Colocalization in the NFL remains low at 3 months of age but is not significantly reduced compared with that at 3 weeks of age (Fig. 4K). Loss of vimentin expression in NFL astrocytes suggests that there are no immature perinatal astrocytes in the mouse retina past the age of 3 weeks.
To further dissect the differentiation stage of the p16INK4a-expressing astrocytes in the mouse eye, we incubated retinal sections with antibodies against PAX2, a marker for immature and mature perinatal astrocytes.8,12 A strong nuclear signal was observed for PAX2 in the optic nerve in all ages investigated (Fig. 5A, D, and G), further supporting our observations of immature astrocytes. A strong nuclear signal was also observed in numerous cells in the NFL from the optic nerve head to the periphery of the retina in PND3 mice (Fig. 5B and C). In contrast, only sporadic cells in the NFL showed nuclear expression of PAX2 in mouse retinas aged 3 weeks and 3 months (Fig. 5E, F, H, and I). Coincubation with antibodies against p16INK4a (cytoplasmic) and PAX2 (nuclear) shows that both proteins appear to be expressed in these cells in the adult mouse retina (Fig. 5J). Combined, these observations suggest that both the adult mouse optic nerve and NFL have immature astrocytes although these may constitute only a small population of cells. In addition, PAX2 nuclear signal was also observed in the inner nuclear layer in retinas aged 3 weeks and 3 months (Fig. 5E, F, H, and I). This could represent Pax2 expressed in the nuclei of Müller glia, since previous studies have shown that PAX2 is not only a marker of astrocytes in the NFL but also a marker of central Müller glia, whose soma aligns with the inner nuclear layer. 29
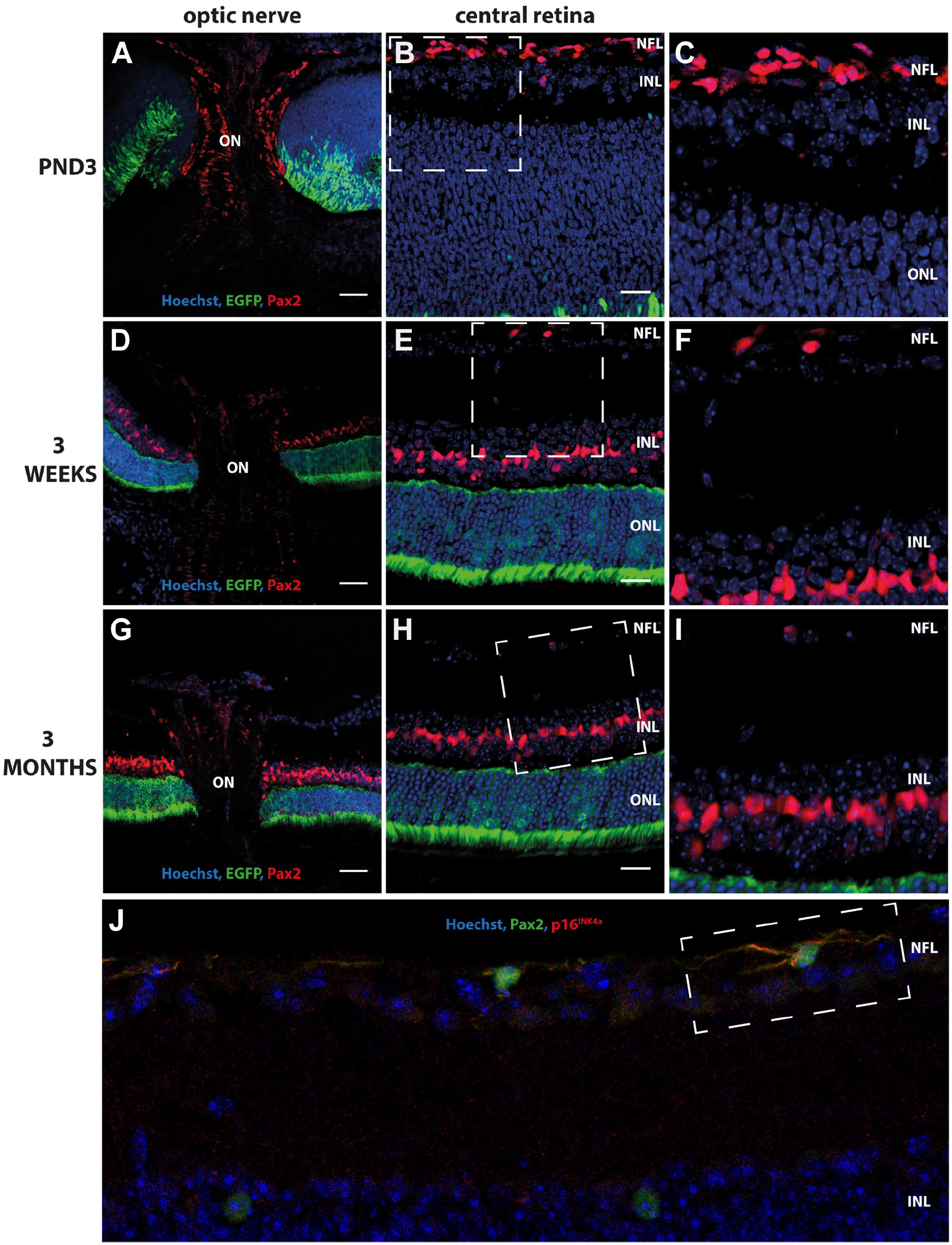
Immunohistochemical detection of PAX2 in the ON and NFL of Nrl-EGFP mouse retina sections aged 3 days, 3 weeks, and 3 months. A strong nuclear signal was observed for PAX2 (red), a marker for mature perinatal astrocytes, in the ON in all ages investigated (A, D, G). In addition, a strong nuclear signal was also observed in numerous cells in the NFL of the central retina in PND3 mice (B and C). In contrast, only sporadic cells in the NFL showed nuclear expression of PAX2 in mouse retinas aged 3 weeks (E and F) and 3 months (H and I). Rabbit polyclonal anti-PAX2 detected with Alexa donkey anti-rabbit-568 (red A–I). Overlapping expression of PAX2 (green) and p16INK4a(red) was observed in the NFL of adult mouse retina (J) (highlighted by the square with a white dotted line) using a goat anti-PAX2 antibody detected with Alexa donkey anti-goat 647 and rabbit polyclonal anti-p16INK4a detected with goat anti-rabbit F(ab’)2 cross-adsorbed secondary Alexa fluor plus 555. Tissue is counterstained with Hoechst nuclear stain (blue). Abbreviations: EGFP, enhanced green fluorescent protein; PAX2, paired box gene 2; PND3, postnatal day 3; ON, optic nerve; NFL, nerve fiber layer; Nrl, neural retina leucine zipper; INL, inner nuclear layer; ONL, outer nuclear layer. Scale bars: optic nerve panel (A, D, G) 100 µm, central retina panel (B, E, H) 50 µm.
Astrocyte RNA Sequencing Data Analyses
To verify our immunohistochemical findings using a different method, we accessed previously published single-cell RNA sequencing data of murine retina. 30 However, we were challenged by the low number of astrocytes detected in these samples, making up 56 out of 46,808 total cells. Further transcripts of the p16INK4a encoding gene Cdkn2a were detected in just eight cells in the data set, suggesting that Cdkn2a expression levels are below the limit of detection for this single-cell RNA sequencing platform.
Therefore, we turned to bulk RNA sequencing data of hESCs induced to differentiate to astrocytes in vitro. 25 CDKN2A was not detected in hESCs but was induced at day 15 of the astrocyte differentiation protocol, concurrent with other astrocytic marker genes, and expression was maintained for the remainder of the protocol (Appendix Fig. 6A). Importantly, CDKN2A transcripts were also detected in primary astrocytes isolated from the human brain (cerebral cortex) (Appendix Fig. 6A). Furthermore, in 3D co-cultures of induced neurons and astrocytes differentiated from hESCs, CDKN2A expression was specific to astrocytes (Appendix Fig. 6B). The observed CDKN2A expression in both astrocytes isolated from the human brain and induced from hESCs could suggest general expression of CDKN2A in the astrocytic lineage, which would also make plausible the expression of CDKN2A in retinal astrocytes. However, caution must be taken, as there could be transcriptional differences between astrocytes of different origin.
Discussion
In this study, we show p16INK4a cytoplasmic expression in astrocytes of the optic nerve as well as in a subpopulation of ocular astrocytes in the NFL of the mouse retina and demonstrate that p16INK4a colocalizes with established markers for immature differentiation stages of astrocytes.
The Cdkn2a gene encodes two distinct proteins, p16INK4a and p14ARF, using alternative reading frames. 13 A previous study showed that both gene products are similarly expressed in the human and rat retina and optic nerve, 31 complicating expression analyses of either. In the present study, we therefore used a Cdkn2a-specific Taqman probe to quantify the expression levels of p16INK4a mRNA in the mouse retina and optic nerve, detecting low expression levels comparable to the reported levels in human and rat eyes. One of the limitations of this assay is that the expression of p16INK4a mRNA was quantified in the whole retina by qPCR and not specifically in astrocytes. The detection of the p16 transcript by in situ hybridization on retinal cryosections from the Nrl-EGFP mouse was unsuccessfully attempted (data not shown), probably due to the low level of expression (p16 cDNA amplified at Ct 28.92–38.36 in the qPCR) or to the fact that standard histological techniques such as formaldehyde fixation or paraffin embedding damage the RNA.
Similarly, a commercially available and well-validated polyclonal antibody against the p16INK4a N-terminal region (specific for p16INK4a) was used to exclude unspecific detection of p14ARF in our study. The extent of astrocyte functional and morphological diversity has become increasingly evident over the last century, and novel roles and subtypes are continuously emerging. 32 Astrocytes are essential for normal functioning of the entire CNS and thus have also been appointed central roles in a vast number of CNS pathologies. 33 Localization and interaction partners appear to be important factors for the functional and structural properties of the individual astrocyte. Thus, special attention must be paid on elucidating the unique functional and structural properties of retinal astrocytes.
Based on morphology, astrocytes are traditionally divided into two major classes: protoplasmic and fibrous. Recently, single-cell RNA sequencing studies have suggested an unexpected diversity of prospective brain cellular subtypes, including additional subpopulations of astrocytes. 34 However, these findings must be further investigated and validated through anatomical integration, as well as assessment of morphological and physiological criteria to define distinct subpopulations. More specific classification of cellular types is commonly based on the expression of established cellular markers in addition to morphology although markers that specifically label astrocyte subtypes are rare. Expression of GFAP has become the most widely used marker for immunohistochemical detection of astrocytes in general, but unfortunately, it is not an exclusive marker for astrocytes. Furthermore, it does not distinguish individual astrocyte subtypes nor does it label the finer processes or the cell body. 2 Likewise, the use of vimentin and/or PAX2 expression, established markers for immature differentiation stages of astrocytes, for histological reliable identification of early-stage retinal astrocytes, is also problematic. Vimentin is an intermediate filament protein and a major part of the cytoskeleton in mesenchymal cells, but more importantly, it is also widely used as a histological marker for Müller cells, the other retinal glial cell type. 35 PAX2 on the other hand is a transcription factor and, accordingly, predominantly localizes to the nucleus, prohibiting any cellular morphological information to be recorded when applied. Thus, identification of novel, more discerning markers of astrocyte subpopulations is paramount to advance our understanding of the functional and structural roles of ocular astrocytes in health and disease.
Through coincubation of antibodies against GFAP and p16INK4a, we identified expression of the widely used senescent marker p16INK4a in the cytoplasm of optic nerve astrocytes and in a subpopulation of NFL astrocytes in normal mice from the postnatal period through to adulthood (aged 3 months). Colocalization investigations of p16INK4a and the established markers for immature stages of astrocyte differentiation, vimentin (immature perinatal astrocyte marker) and PAX2 (immature and mature perinatal astrocytes marker), furthermore showed that p16INK4a appears to be expressed in the immature astrocyte population of the optic nerve and that there is no significant overlap in p16INK4a and vimentin expression in the NFL in the 3-week-old and 3-month-old mice. In contrast, nuclear PAX2 expression was identified in the NFL in all ages investigated although only sporadically in the later stages. Coincubation with antibodies against p16INK4a showed that both proteins appear to be expressed in these cells. In support of these observations, a recent comparative study of PAX2 expression in retinal glial cells and optic nerve of birds and mammals showed PAX2 expression in GFAP-expressing astrocytes in normal mouse optic nerve and NFL. 36 Combined, this suggests that p16INK4a is expressed in the immature perinatal astrocytes found in the optic nerve but also that the expression is sustained past the immature perinatal astrocyte differentiation stage as p16INK4a is expressed in PAX2-expressing NFL astrocytes that no longer express vimentin. Interestingly, p16INK4a does not appear to be expressed in the mature astrocyte either, or at least only in a subset of them, as only a small population of NFL astrocytes show colabeling for p16INK4a and GFAP. Thus, this suggests that the adult mouse retina appears to retain a population of undifferentiated astrocytes in the NFL and that these cells could potentially represent a pool of astrocyte progenitor cells for maintaining the astrocyte population throughout life. As PAX2 is also only identified in a small subset of cells in the NFL, it could suggest that the p16INK4a-expressing astrocytes in the NFL could be mature perinatal astrocytes. To our knowledge, the detection of cytoplasmic expression of p16INK4a in neonatal/young ocular astrocytes has not been reported previously and is an interesting observation. However, to establish fully the exact stage of astrocyte differentiation of the p16INK4a-expressing cells would require further investigations that are out of the scope of the present study. For now, our targeted analysis of published transcriptomic studies of primary and hESC-derived astrocytes complements and supports our immunohistochemical data and provides a solid foundation for further investigations.
The involvement of p16INK4a in cell cycle control is generally believed to be carried out by nuclear p16INK4a, but accumulating evidence suggests that p16INK4a may be expressed and functional in both the nucleus and the cytoplasm.37,38 This disputes the long-held assumption that the cytoplasmic staining observed in immunohistochemical studies is merely an unspecific background. Interestingly, 2D gel electrophoresis analysis of nuclear and cytoplasmic fractions of selected cell lines suggests that a key difference between the cytoplasmic and nuclear p16INK4a populations may be phosphorylation. 37 In addition to cell cycle control, p16INK4a has been suggested to be involved in a number of other processes including angiogenesis, apoptosis, and cell invasion/migration. 15
Many aspects of p16INK4a function and regulation are still not fully understood. In particular, the functional roles of cytoplasmic p16INK4a remain elusive. p16INK4a was previously reported to interact with several cytoskeleton proteins including α-β-γ actin, α-β tubulin, and CDK4/6. 39 However, the cytoplasmic interaction of p16INK4a with CDK4/6 was later suggested to represent a regulatory mechanism for the inhibitory effects of p16INK4a on the cell cycle. 37 Consequently, one might speculate that the cytoplasmic ocular astrocyte p16INK4a expression identified in our study may function in a similar way and, thus, could suggest that these cells retain proliferative characteristics due to the lack of the inhibitory effects of p16INK4a in the nuclei. This would be in stark contrast to the quiescent status of in situ astrocytes under normal physiological conditions. Interestingly, both astrocyte precursor cells and immature astrocytes have been reported to show proliferative and migratory potential.8,12,40 It is unlikely that the p16INK4a expression we observed in some ocular astrocytes reflects a population of senescent astrocytes. First, we observed cytoplasmic expression of p16INK4a, and it is usually the increased nuclear expression of p16INK4a that is a recognized indicator of cellular senescence. 19 Second, we observed a similar cytoplasmic expression of p16INK4a in optic nerve astrocytes in wild-type adult mice and mice only three days old. In support of this, no apparent senescent-associated β-gal staining is observed in inner retinal flat mounts of 4.5-month-old rats. 41 Collectively, this indicates that the cytoplasmic expression of p16INK4a may be a novel marker for immature astrocytes. Our data thus suggest that the retina retains a population of immature/precursor astrocyte cells that may support astrocyte turnover into adulthood.
In conclusion, our immunohistochemical investigations of p16INK4a in the wildtype young mouse retina have identified p16INK4a as a potential novel marker of immature astrocytes in ocular health and pathology. However, the authors acknowledge that future investigations should further develop and confirm these initial findings by demonstrating that these p16INK4a-expressing astrocytes indeed proliferate during the early postnatal stage and do not enter senescence. It would also be of potential interest to investigate the relationship between the expression pattern of p16INK4a and the astrocytes morphology and migration in different ocular disorders including reactive gliosis, a central feature of many retinal disorders.
Footnotes
Appendix
Acknowledgements
The authors thank Ross Campbell and Federica Staurenghi, Nuffield Laboratory of Ophthalmology, University of Oxford, for technical assistance and Professor Clare Futter, Institute of Ophthalmology, University College London, for useful discussions and help improving the manuscript.
Competing Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: AJR and SGD are founders and minor shareholders of OxStem Ltd. REM receives grant funding from OxStem Ltd. All other authors have no competing financial interests.
Author Contributions
All authors have contributed to this article as follows: quantitative real-time PCR (CMFC), statistical analyses (CMFC), experimental work for revised version (CMFC), drafting the revised manuscript (CMFC), design of the study (TS, AJR, SGD, ARB, REM), analyses of data (CMFC, TS), immunohistochemistry and microscopic analyses (TS), drafting the manuscript (TS), antibody validation (AS), 3D-rendering analysis (TB), manuscript preparation (CMFC, TS, TB, MQR, REM), RNA sequencing data analysis (MQR), preparation of ocular tissue for IHC (HO), data analysis (ARB, REM), and all authors have read and approved the final manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a grant from OxStem Ltd.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
