Abstract
Here, we report the first molecular cytogenetic characterization of the BALB/cAnN mouse derived B-cell non-Hodgkin lymphoma (B-cell NHL) cell lines A-20. Even though previously used as a model for testing of, for example, dexametason, up to present, no data in the genetic properties of A-20 were available. The present study closed this gap and provides evidence that A-20 is a model for B-cell NHL subgroup sporadic Burkitt’s lymphoma. C-myc oncogene is involved in a translocation and copy number alterations as gain of murine 14q material could be observed. Interestingly, the cell line showed the karyotype 39,X,-X or -Y,t(2;15)(qE5;qD2),del(6)(qB3qC3),del(9)(qA3qA4),dup(14)(qE1qE4) in ~95% of the cells, being exceptionally stable for cell lines being established 38 years ago. Still, ~5% of the cells showed polyploidization followed by chromothripsis. It remains to be determined if this can be observed also in other cell lines, just has not been reported yet, and/or if it is a unique feature of A-20. Overall, finally here, the necessary genetic data to identify A-20 as a model for human sporadic Burkitt’s lymphoma are provided.
Introduction
B-cell non-Hodgkin lymphomas (B-cell NHL) comprise a heterogeneous group of acquired malignancies. According to Cuneo, 1 they can be classified as lymphoplasmacytic lymphoma, follicle center cell lymphoma, diffuse large-cell lymphoma, Burkitt’s lymphoma (nowadays subdivided by World Health Organization into endemic, sporadic, and HIV-associated Burkitt’s lymphoma), Burkitt-like lymphoma, mantle cell lymphoma, marginal zone B-cell lymphoma, and splenic lymphoma with villous lymphocytes. B-cell NHL can be characterized best by IHC, but also by fluorescence in situ hybridization (FISH).2,3
As a simple PubMed search shows, murine cell lines are widely used for lymphoma research: more than 9050 hits are found. 4 The here-studied murine B-cell NHL A-20 is among this list. It has been applied in only about 5 published studies; yet, this limited number is most likely due to the fact that up to the present, there was no (cyto)genetic study done for it. Thus, genetic content was unknown, and it was widely unclear for which lymphoma type A-20 is a model for.
A-20 is a BALB/c B-cell lymphoma line, derived from a spontaneous reticulum cell neoplasm found in an “old” BALB/cAnN mouse. According to Kim et al., 5 the modal chromosome number of A-20 cell line was between 33 and 38, and no gender of the mouse was given from which the cells were derived from. Besides some immunophenotypic data,6–9 no further basic information could be found for A-20 in the literature. Nonetheless, even a test of medication meant for lymphoma treatment was done on this cell line, for example, for dexametason.10,11
Here, we present the first detailed molecular cytogenetic analyses of the cell line A-20. Multicolor-FISH using whole chromosome painting (wcp) probes and the FISH banding-approach “murine multicolor banding (mcb)” as well as array-comparative genomic hybridization (aCGH) were used, as previously reported. 12 The obtained data on imbalances and chromosomal breakpoints in A-20 were translated into human genome to find out the human lymphoma subtype corresponding to A-20.
Materials and Methods
Murine Lymphoma Cell Line A-20
The cell line A-20 was obtained from American Type Culture Collection (ATCC TIB-208; Middlesex, UK) as a not further characterized B-lymphocyte line, and grown in RPMI-1640 medium with 2mM L-glutamine, 25 mM HEPES (4-[2-hydroxyethyl]-1-piperazineethanesulfonic acid), and 5% fetal calf serum in the presence of antibiotics. Cells were split in two portions and worked up cytogenetically (flask 1; see below); from flask 2, whole genomic DNA was extracted using Blood & Cell Culture DNA Midi Kit (Qiagen; Hilden, Germany) according to manufacturer’s instructions. For cytogenetic workup of cells from flask 1, colcemid at the concentration of 0.1µg/ml was added for 2–3 hr. Subsequently, the cells were hypotonicly treated in 0.075 M KCl for 20 min and fixed/cytogenetically worked up, and metaphase spreads were prepared according to standard procedures. 13
Molecular Cytogenetics
FISH was performed as described previously.12,14 For multicolor-FISH with wcp probes, the SkyPaint DNA Kit M-10 for Mouse Chromosomes (Applied Spectral Imaging; Glasgow, UK) was used according to manufacturer’s protocol. For FISH-banding, previously established murine chromosome-specific mcb probe mixes were applied. 15 Image acquisition was done using a Zeiss Axioplan microscope equipped with ISIS software (MetaSystems; Altlussheim, Germany). At least 29 metaphases were analyzed for each probe set.
ACGH was performed and evaluated according to standard procedures using SurePrint G3 Mouse CGH Microarray, 4 × 180 K (Agilent Technologies; Ratingen, Germany). 12
Data Analysis
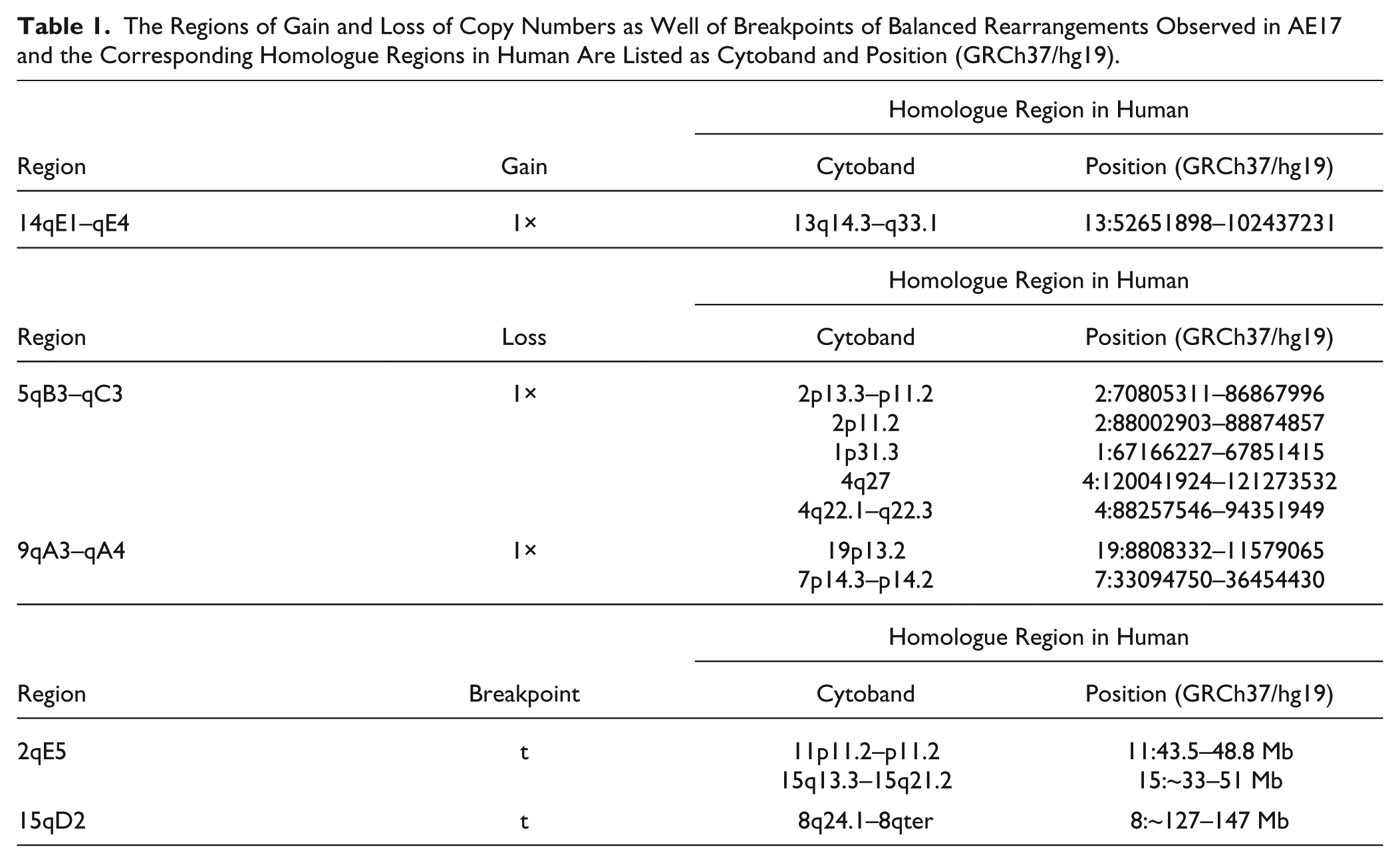
The regions of imbalances present in A-20, as defined after analyses of aCGH and mcb data, were aligned with their human homologous regions using Ensembl and the University of California, Santa Cruz (UCSC) Genome Browser as previously described. 14 Also, potential cancer-related genes for observed breakpoint regions were identified via the “Atlas of Genetics and Cytogenetics in Oncology and Haematology.” 16 The alignments are summarized in Table 1.
The Regions of Gain and Loss of Copy Numbers as Well of Breakpoints of Balanced Rearrangements Observed in AE17 and the Corresponding Homologue Regions in Human Are Listed as Cytoband and Position (GRCh37/hg19).
To align the obtained data with the human B-cell NHLs, data were collected from Cuneo, 1 Ota et al., 17 Ferreira et al., 18 Tirado et al., 19 Zhao et al., 20 Thelander and Rosenquist, 21 García et al., 22 and Neat et al. 23
Results
The murine cell line A-20 revealed the following final karyotype after using all 21 wcp probes in one experiment (Fig. 1A), all chromosome-specific murine mcb probe sets (Fig. 1B), and aCGH (data not shown):
39,X,-X or -Y,t(2;15)(qE5;qD2),del(6)(qB3qC3),del (9)(qA3qA4),dup(14)(qE1qE4).
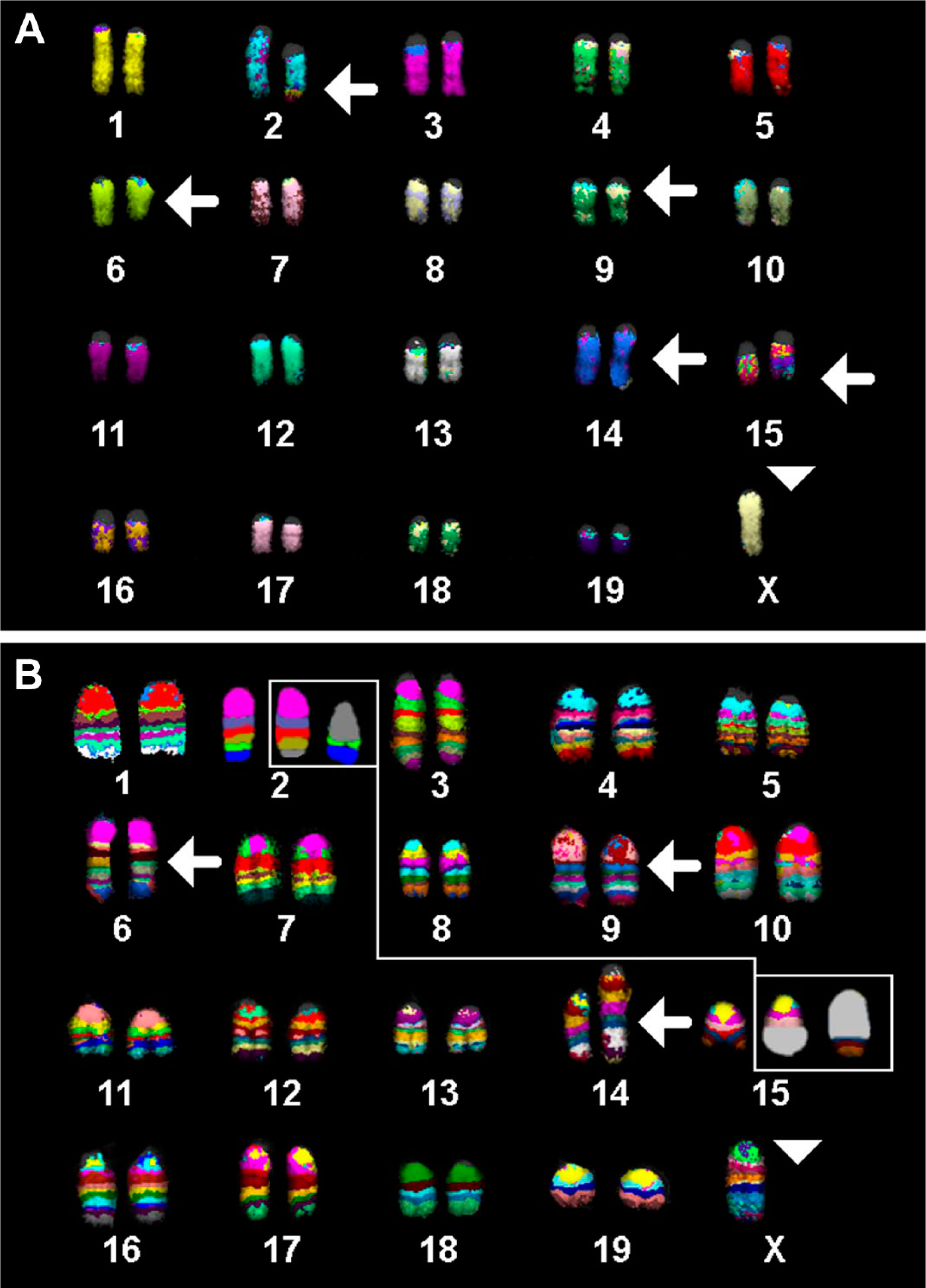
(A) Multicolor-FISH using all 21 murine whole chromosome paints as probes was applied on chromosomes of the cell lines A-20. Results for the main clone are depicted. (B) Murine multicolor banding results for all chromosomes confirmed and refined the results of multicolor-FISH from Fig. 1A. Abbreviation: FISH, fluorescence in situ hybridization.
This karyotype was found in more than 95% of the cells; of those cells with only 1.5% of the cells showed single cell aberrations, such as loss or gain of single chromosomes or structural rearrangements. Also ~0.2% of the A-20 cells were tetraploid without any additional alterations. In total, ~5% of the overall 639 analyzed metaphases of cell line A-20 showed highly complex rearranged karyotypes, which could not be analyzed in detail as shown in Fig. 2.
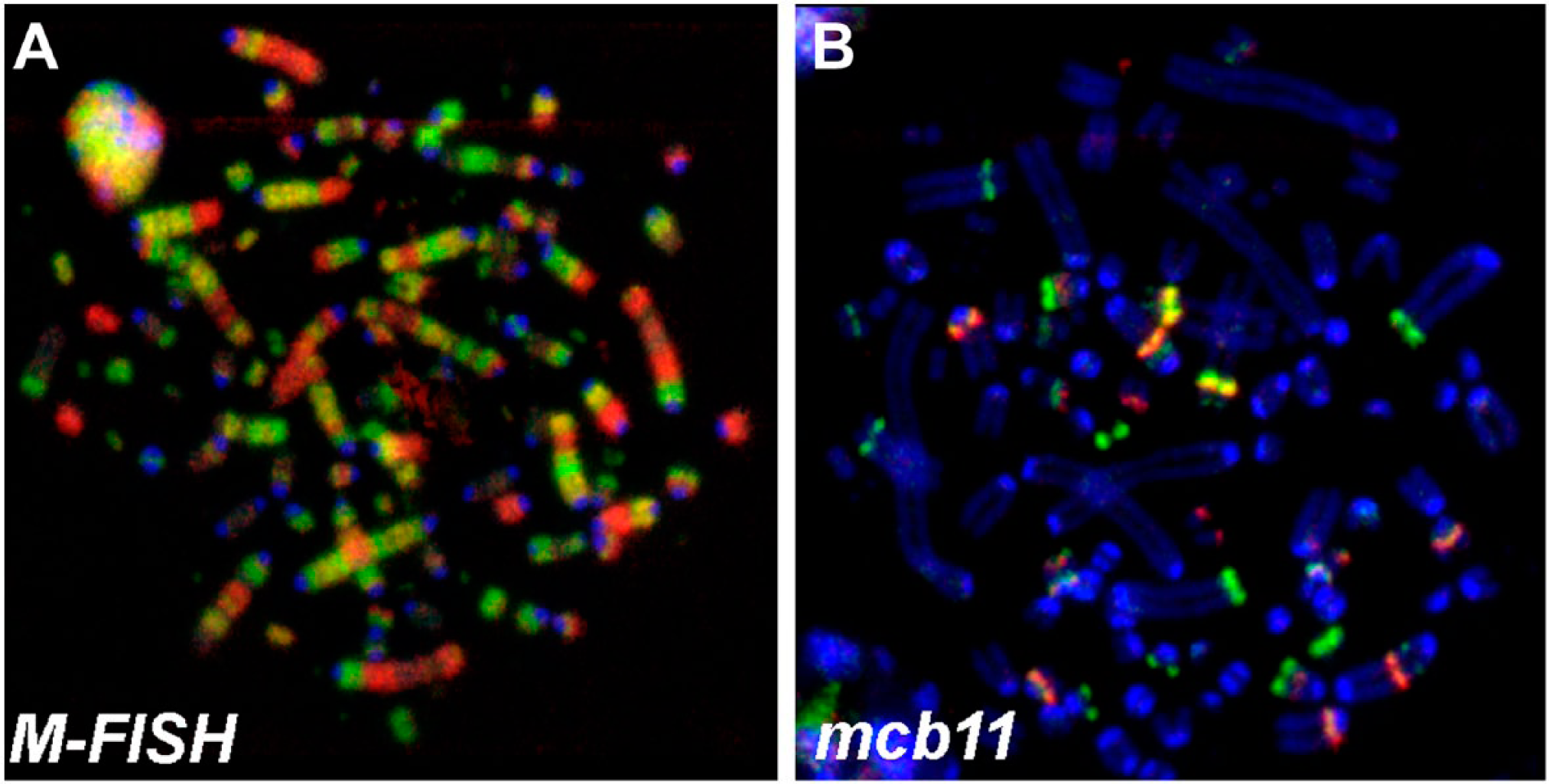
2/31 detected cells with complex aberrant karyotypes—in (A) a metaphase after multicolor-FISH (M-FISH) and (B) mcb11 are shown. Abbreviation: FISH, fluorescence in situ hybridization; mcb, murine multicolor banding.
Overall, the resulting chromosomal imbalances observed in aCGH were in agreement with the FISH results of the main clone (summarized in Fig. 3). According to aCGH, six breakpoints could be characterized in more detail as follows (according to the National Center for Biotechnology Information Build 37 [UCSC mm9, July 2007]):
6qB3: 58.56 Mb
6qC3: 84.63 Mb
9qA3: 20.79 Mb
9qA4: 26.13 Mb
14qE1: 88.15 Mb
14qE4: 116.82 Mb
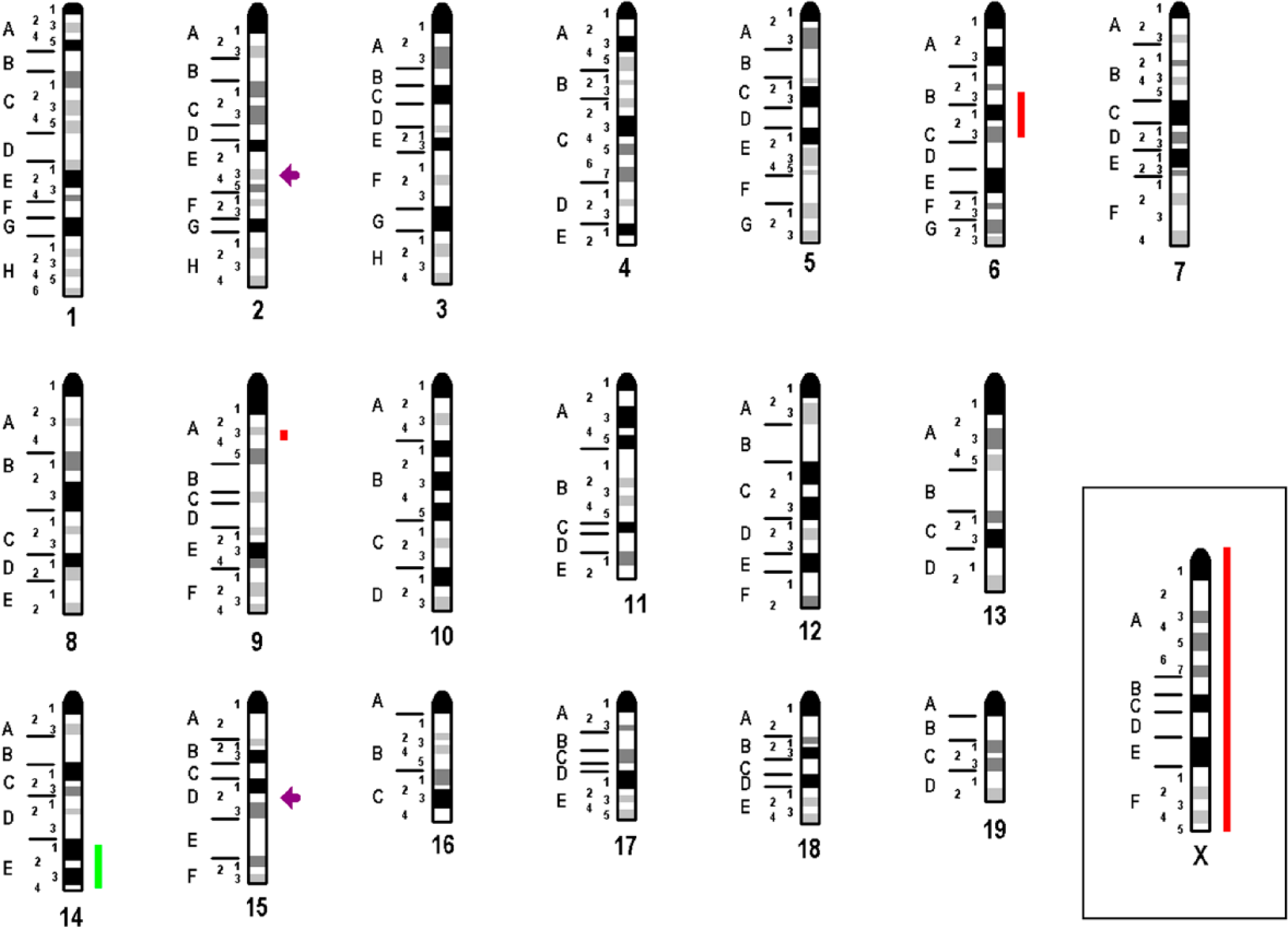

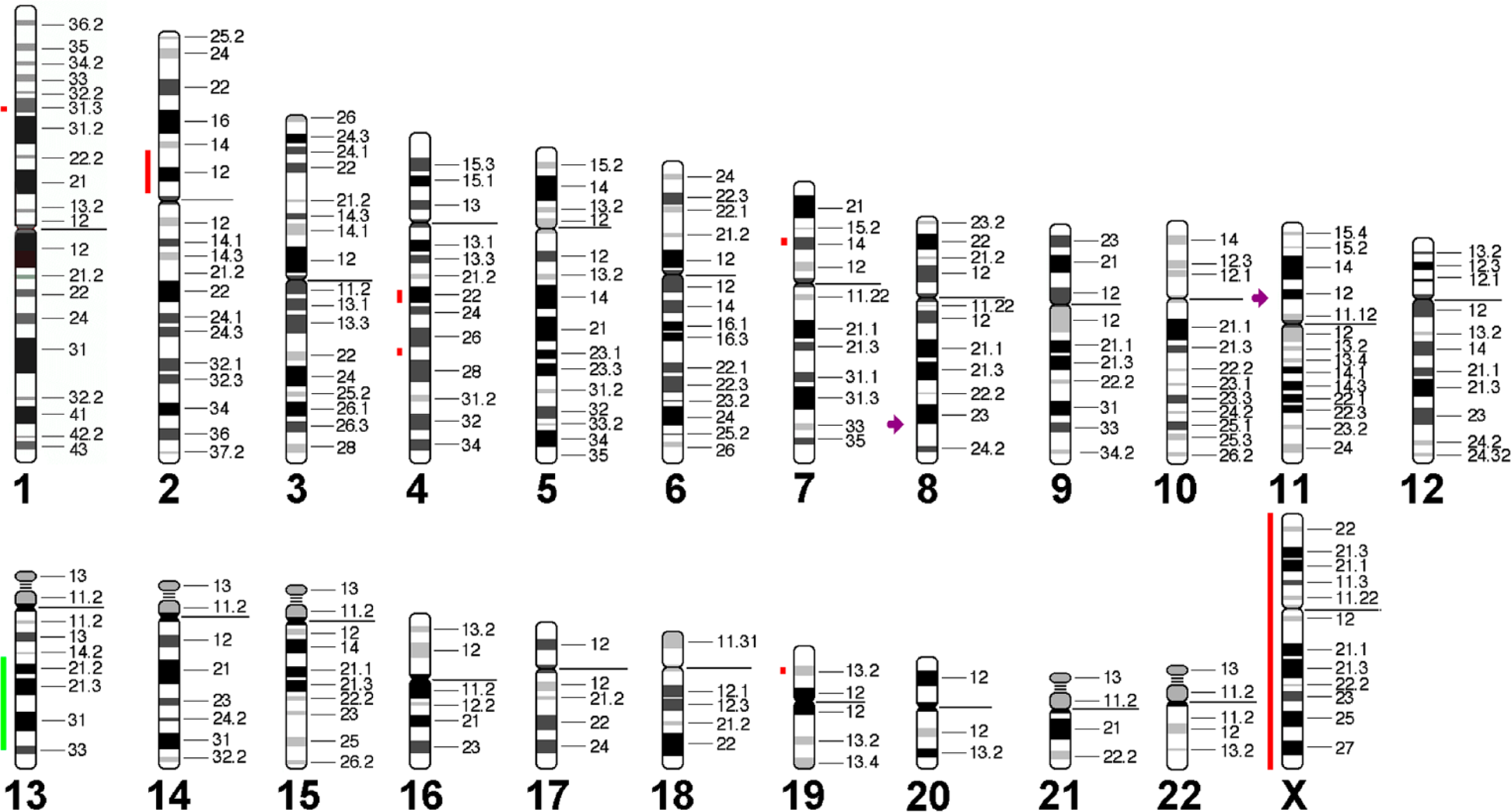
Imbalances and breakpoints observed in A-20 are summarized here. Gains are depicted as light green bars, losses as red bars, breakpoints as violet arrow.
Imbalances smaller than 3.5 Mb detected in aCGH were not considered (see discussion).
The imbalances of A-20 were translated to human genome (Fig. 4) and aligned with their homologous regions in human (Table 1) and compared with human lymphomas. The highest concordance was found with sporadic Burkitt’s lymphomas (Table 2).
Comparison of “Translated” Imbalances of A-20 and Human B-Cell NHLs.
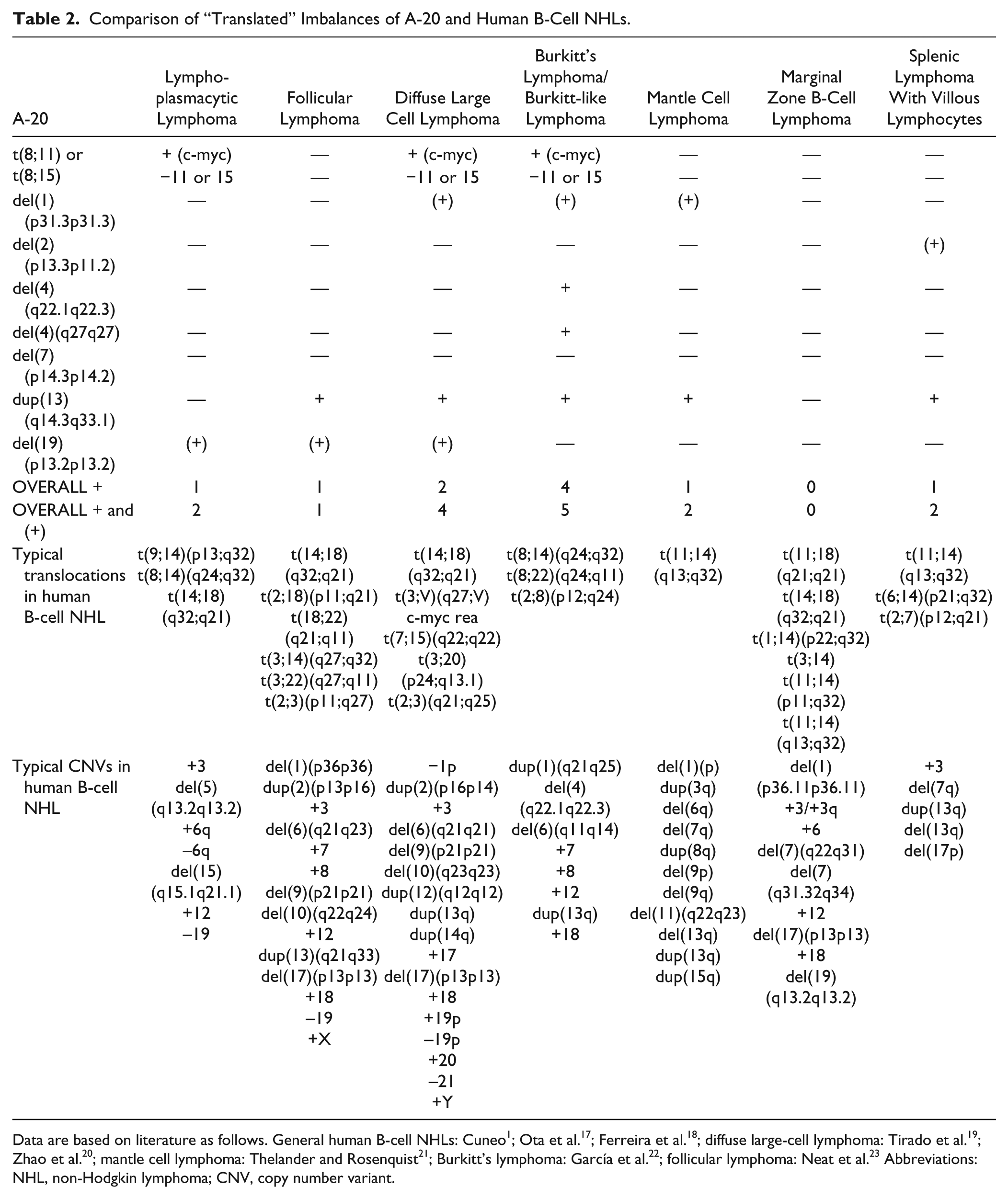
Data are based on literature as follows. General human B-cell NHLs: Cuneo 1 ; Ota et al. 17 ; Ferreira et al. 18 ; diffuse large-cell lymphoma: Tirado et al. 19 ; Zhao et al. 20 ; mantle cell lymphoma: Thelander and Rosenquist 21 ; Burkitt’s lymphoma: García et al. 22 ; follicular lymphoma: Neat et al. 23 Abbreviations: NHL, non-Hodgkin lymphoma; CNV, copy number variant.
Discussion
Genetically, murine tumor cell lines are not well characterized; this also holds true for cell line A-20. Here, the karyotype of A-20 was studied and defined in detail based on aCGH and mcb.
We found that the cell line is still diploid, even though most human and many murine cell lines become tetraploid over time,14,24 including human Burkitt’s lymphoma cell lines.25,26 As outlined by Davoli and de Lange, 24 tetraploidization could be connected with telomere disruption; however, for A-20, there are no studies available yet on telomere length and/or telomerease activity that could support or contradict this theory. Overall, when comparing A-20 with other decade-old cell lines such as NIH3T3, 14 it is amazing that initially chromosome numbers between 33 and 38 5 were reported, and now A-20 still presents a stable chromosome number of 39 in ~95% of the cells. Still, in the remainder ~5% of the cells, chromothripsis was observed (Fig. 2). This could reflect a chromothripsis-related mechanism to avoid tetraploidization of the malignant cells. 27 Thus, other long-established cell lines with and without tetraploidization should be studied for small percentages of chromothriptic cells, as it could well be that such small cell populations were simply ignored previously, as suggested by others.28,29 By application of aCGH and mcb, it was possible to resolve 8 chromosomal breakpoints involved in (un)balanced chromosomal rearrangements in A-20. However, resolution of mcb is limited to 5–10 Mb. 14 Here, imbalances smaller than 3.5 Mb detected in aCGH were not considered in the evaluation. Only aberrations seen by FISH-banding and aCGH were included here, that is, as such were proven by two independent approaches (see also Kubicova et al. 12 ).
When transforming the imbalances and breaks observed in A-20 to human genome using databases and bioinformatics tools (see Table 1) such as were done before for NIH3T3 and AE17,12,14 it turned out that A-20 is a model for sporadic Burkitt’s lymphoma. The underlying idea is that identical imbalances and rearrangements found in the murine tumor cell line pinpoint for which human tumor the cell line can serve as a model.12,14 The translocations found involve c-myc and a yet unknown partner-gene on murine chromosome 15. So, in summary: c-myc is the driving oncogene and, in addition, 4 imbalances found in A-20 are typically observed as well in sporadic Burkitt’s lymphoma. Thus, even though also similar imbalances as in A-20 can be found in diffuse large-cell lymphoma, overall, only 1 compared with 3 matching copy number variations supports the view that A-20 was a sporadic Burkitt’s lymphoma in the original BALB/cAnN mouse.
As there is only one X-chromosome in A-20, it cannot be clarified if originally, the individual mouse was male or female. Even though the finding of overall two metaphases with two X-chromosomes among 60 correspondingly targeted cells (data not included in section “Results”), this is not necessarily indicative for an originally female individual, as the two corresponding cells just could have doubled the remaining X-chromosome during division. It is well known that chromosome numbers in tumor cells are unstable in general.30,31
Overall, finally, 38 years after establishment of the cell line, the present work provides the necessary genetic data to characterize A-20 as a model for human sporadic Burkitt’s lymphoma. In agreement with this, the successful previous use of dexamethasone,10,32 concanavalin A,9,33 and ruthenium complexes10,34 for growth inhibition in this cell line and human sporadic Burkitt’s lymphoma was reported.
Footnotes
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
NK created the murine multicolor banding probes, cultivated the cell line, and prepared the chromosomes. KG and SSHA performed and evaluated the fluorescence in situ hybridization experiments, MR did the array-comparative genomic hybridization studies, and TL developed the study and wrote up the final results and the article draft. The article was approved by all authors.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Supported by grant # 2013.032.1 of the Wilhelm Sander-Stiftung.
