Abstract
Banding cytogenetics is still the gold standard in many fields of leukemia diagnostics. However, in chronic lymphocytic leukemia (CLL), GTG-banding results are hampered by a low mitotic rate of the corresponding malignant lymphatic cells. Thus, interphase fluorescence in situ hybridization (iFISH) for the detection of specific cytogenetic aberrations is done nowadays as a supplement to or even instead of banding cytogenetics in many diagnostic laboratories. These iFISH studies can be performed on native blood or bone marrow smears or in nuclei after cultivation and stimulation by a suitable mitogen. As there are only few comparative studies with partially conflicting results for the detection rates of aberrations in cultivated and native cells, this question was studied in 38 CLL cases with known aberrations in 11q22.2, 11q22.3, 12, 13q14.3, 14q32.33, 17p13.1, or 18q21.32. The obtained results implicate that iFISH directly applied on smears is in general less efficient for the detection of CLL-specific genetic abnormalities than for cultivated cells. This also shows that applied cell culture conditions are well suited for malignant CLL cells. Thus, to detect malignant aberrant cells in CLL, cell cultivation and cytogenetic workup should be performed and the obtained material should be subjected to banding cytogenetics and iFISH.
Keywords
Introduction
Chronic lymphocytic leukemia (CLL) is the most frequent leukemia in adults in Western countries. It is a heterogeneous disease which is characterized by specific chromosomal and genetic aberrations associated with different clinical outcomes and overall survival. 1 The most frequent recurrent cytogenetic abnormalities in CLL are deletions in 13q14, 11q22-q23, 17p13, trisomy 12, and chromosomal rearrangements in 14q32.33. 2
CLL patients could be classified into different groups according to the detected chromosomal aberrations; each group shows distinct prognosis and response to treatment. The group harboring 17p13 deletion has the worst prognosis with a median survival of only 32 months, whereas patients with 13q14 deletion show the longest life expectancy with a median survival of 133 months.3,4
Detection of such abnormalities by banding cytogenetic analysis is limited due to several reasons, such as low mitotic activity of CLL cells and submicroscopic nature of some of the typical chromosomal changes. Thus, the detection rate of karyotyping reaches not more than 50%, even after long-term cultivation and stimulation by different types of mitogens such as 12-O-tetradecanoylphorbol-13-acetate (TPA) or phytohemagglutinin (PHA). However, detection rate could be increased significantly by the addition of CpG oligodeoxynucleotides and interleukin 2 (IL-2) to the cells culture.5,6
Malignant CLL cells express many growth activation markers; however, they are arrested in G0/early G1 phase of the cell cycle. They grow and accumulate due to apoptosis resistance mediated by mechanisms such as overexpression of B-cell lymphoma 2 (gene) caused by depletion of inhibitory miR-15 and miR-16. Thus, cell cycle transition and differentiation of CLL cells are induced after cultivation and stimulation by TPA.7–10
Interphase fluorescence in situ hybridization (iFISH) has been emerged as an effective technique with a higher resolution than G-banding techniques; besides, iFISH can be done without the need of cell culture.11,12 iFISH procedure can directly be performed in blood or bone marrow smears, being time saving and closer to in vivo situation.
Moreover, applying iFISH on uncultured bone marrow aspirate gives more reliable information for the clinician during the course of a leukemic process or when monitoring reverse sex bone marrow transplantation.13,14 Nonetheless, it should be kept in mind that iFISH is restricted to specific targeted chromosomal regions according to the selected probes, and thus, genomic complexity of a case may be underestimated. 12 To the best of our knowledge, there were only few previous studies comparing the reliability of iFISH data performed in CLL smears and cultivated CLL suspension with in parts contradictory data; also, only a limited number of FISH probes was tested.13,15,16
In the present study, we compared the detection rates of iFISH in CLL cases with known aberrations when studying cell suspension after cultivation and stimulation by TPA to blood or bone marrow smears, and data for six loci were obtained.
Materials and Methods
Patients
The present study included 38 CLL patients diagnosed according to standard criteria. 17 The samples were obtained under informed consent of the corresponding patients and according to institutional ethical committee guidelines (ethical committee of the Friedrich Schiller University Jena, code 1105-04/13). Clinical data and GTG-banding results (only for 25/38 cases) of the 38 patients can be found in Tables 1 and 2.
Gender, Age, and Cytogenetic Results of the Studied Cases/Samples.
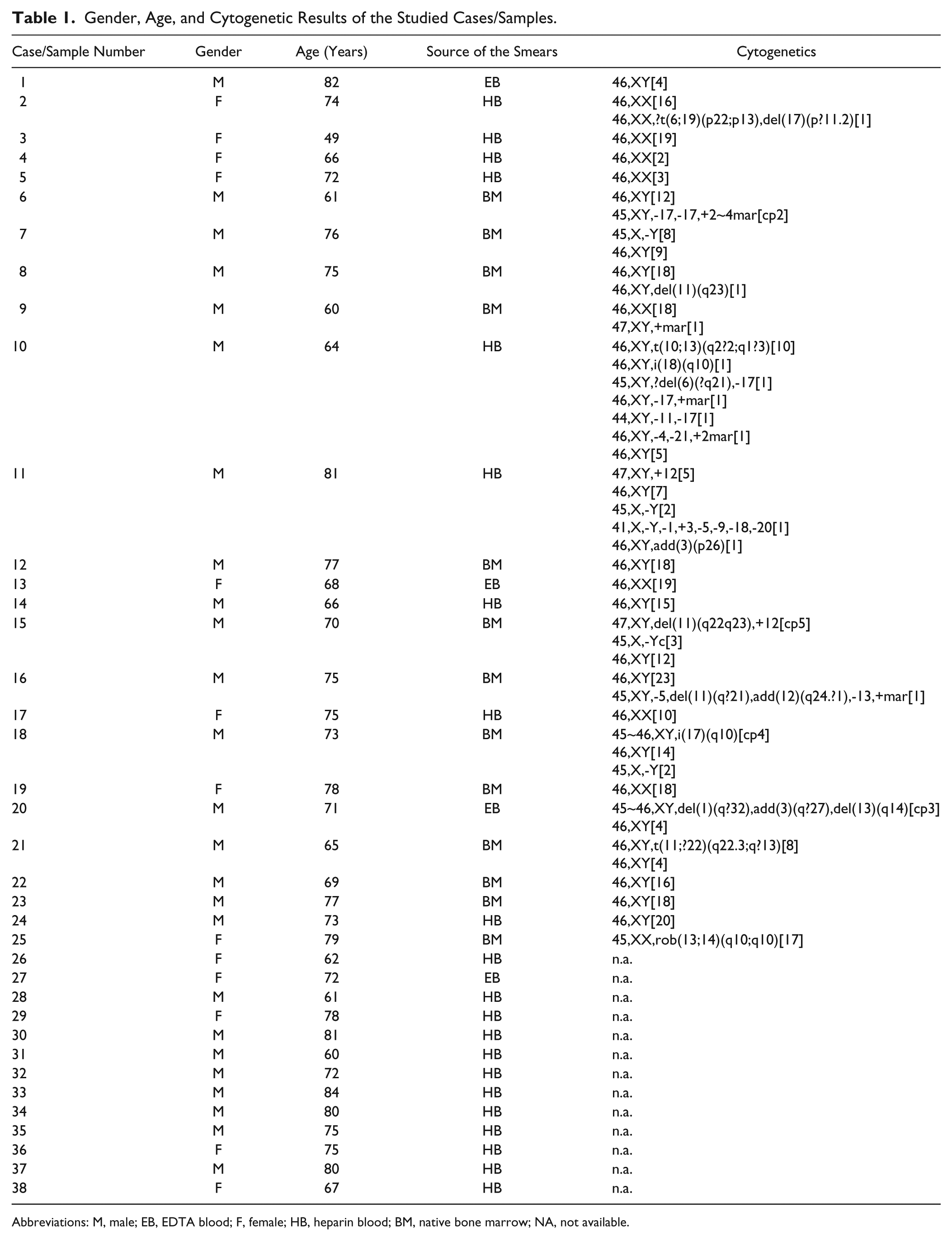
Abbreviations: M, male; EB, EDTA blood; F, female; HB, heparin blood; BM, native bone marrow; NA, not available.
The Chromosomal Aberrations Detected by iFISH in Both Cell Suspension and Blood or Bone Marrow Smears.

Abbreviation: iFISH, interphase fluorescence in situ hybridization.
Sample Preparation
Smears have been prepared according to standard procedures from different sources: 22 samples from heparinized blood, four samples from blood treated with EDTA, and 14 samples from heparinized bone marrow (see Table 1). The obtained slides were stored at room temperature (RT) for approximately 12 hr and then either used pretreated as described in the following or saved at −20C. The red blood cells (erythrocytes) were removed from the smears by incubating the slides in a fixative (methanol/acetic acid 3:1) for 15 to 60 min at RT and later in 100% methanol for 15 min. Then, the air-dried slides were transferred into 2× SCC for 15 min and dehydrated in an ethanol series (70%, 90%, 100%, 3 min each). After that, 200 µl of 1-M sodium thiocyanate solution was added per slide, covered with a 24 × 60-mm coverslip, and incubated at 37C for 1 to 2 hr in a humid chamber. Finally, the slides were rinsed in 1× PBS at RT for 5 min, dehydrated in an ethanol series (70%, 90%, 100%, 3 min each), and air dried.
The slides from cell suspensions were established by the standard air-drying method.
iFISH
iFISH analyses were done as previously reported in Alhourani et al. 11
For iFISH, the following probes were used: Abbott/Vysis (Wiesbaden, Germany): LSI p53/LSI ATM (in 17p13.1 and 11q22.3), LSI D13S319/LSI 13q34/CEP 12 (in 13q14.3, 13q34, and 12p11.1-q11.1), LSI IGH Dual Color, Break Apart Probe (in 14q32.33); Zytovision GmbH (Bremerhaven, Germany): ZytoLight SPEC BIRC3/MALT1 Dual Color Dual Fusion Probe (in 11q22.2 and 18q21.32).
For each iFISH analysis, 100,200 interphase nuclei were examined per sample and probe studied.
Statistical Analysis
The STATGRAPHICS Centurion Version 16.2 was used for statistical analysis. Paired Student’s t-test was used besides to evaluate the probability of significant differences between the groups. The chosen level of significance was 0.05.
Results
Overall, 61 chromosomal aberrations were detected in cell suspension and only 59 by iFISH in smears; in cases 3 and 5, the biallelic deletion in 13q13.4 could only be seen in cultivated material. The detected chromosomal aberrations in cell suspension and smears are listed in Table 2. Statistical analysis (multifactor ANOVA) revealed significantly (p<0.05) elevated rate of aberrations in suspension compared with the smears (Fig. 1). Significant difference was not found between individual groups (p>0.05) using t-test and ANOVA.
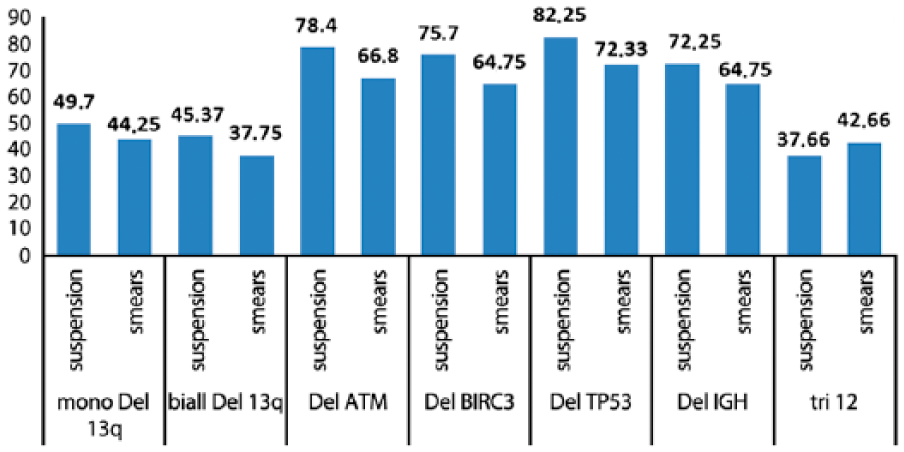
The mean number of the aberrant cells is given here as percentage for the chromosomal abnormalities which were detected in the studied cells derived from suspension and blood or bone marrow smears. The following groups are listed (see also Table 2): mono Del 13q, monoallelic deletion in 13q14.3; biall Del 13q, biallelic deletion in 13q14.3; Del ATM, deletion in 11q22.3; Del BIRC3, deletion in 11q22.2; Del TP53, deletion in 17q13.3; Del IGH, deletion in IGH locus; tri 12, trisomy 12.
However, two of the three cases with trisomy 12 (cases 11 and 12) showed a higher percentage of aberrant cells in the smears. Interestingly, in CLL cases with aberrations in less than 30% of the cells, the detection rate was alike in suspension and smears; student’s t-test gave a p value of 0.9714.
Discussion
iFISH is routinely applied in CLL diagnostics, and in many cases, follow-up studies of individual patients are done once on blood or bone marrow smears and the other time on previously cultivated material. In the literature, there are reports indicating for differences in detection rates, depending on the used material.13,15,16 In case of “iFISH-aberrations” in 11q22.3, centromere 12, 13q14.3, and 17p13, Sánchez and Aventín 15 found higher proportions of detectable aberrant clones in TPA-cultivated material than in smears. The same was found previously for 17p13 by Cano et al., 13 while the same authors could not confirm this effect for trisomy 12. Delgado et al. 16 did comparative analyses also with the same probes as Sánchez and Aventín 15 ; however, they did not specify the results and just reported higher detection rates for all used probes in TPA-cultivated material.
In this study, the same probes as previously studied13,15,16 and two additional loci (11q22.2 and 14q32.33) were tested on the two mentioned CLL-derived interphase materials. Overall, in this study, the number of aberrant cells detected was higher in cell suspension after cultivation and stimulation by TPA than in smears (Fig. 1). While in other leukemias, for example, chronic myeloid leukemia,18,19 the suggestion that smears are closer to in vivo situation than cultivated material is true, this is different for CLL. As TPA stimulation obviously promotes growth of aberrant malignant cells, 16 use of suspension instead of smears brings a real benefit for detection of CLL-associated aberrations.
However, when dividing the studied aberrations into such being present at percentages below (group 1) and in more than 30% (group 2), the differences were not statistically significant for group 1; in other words, both smears and cell suspension could be used to identify the chromosomal aberrations in the cases studied presenting with low percentage of aberrant cells. Also, Cano et al. 13 showed the ability to detect TP53 deletions and trisomy 12 in uncultivated smears at a low percentage of aberrant cells. However, as seen in cases 3 and 5, low-level presence of biallelic deletion of RB1 may be missed if only applying smears.
Surprisingly, the percentage of trisomy 12 aberrant cells was higher in the smears in two studied cases; also, Cano et al. 13 reported such an observation, as they found that the mean number of cells with trisomy 12 in the CLL samples after cultivation (40.9%) was less than in the same fresh uncultured samples (49.7%); just one out of eight cases with trisomy 12 showed more aberrant cells by 3% after 3 to 5 days cultivation and stimulation with TPA as a mitogen. Also, Sánchez and Aventín 15 and Delgado et al. 16 found that CLL cases with trisomy 12 have higher proportions of aberrant cells in smears rather than in TPA-cultivated suspension. These unexpected findings in the trisomic cases could be explained by TPA stimulating capacity or by technical problems which may affect the proliferation of neoplastic trisomic CLL cells during cultivation. 13 According to the present study including not only four but six loci, this effect may be restricted to trisomy 12 positive CLL cells.
Overall, this study showed that TPA-stimulated cells enable more reliable detection of aberrant clones in CLL, which could be explained by the influence of cultivation and simulation by TPA as a mitogen on cell cycle transition and differentiation of CLL cells. 8 Considering this is especially important in case of follow-up studies of patients and when comparing results from different laboratories using different material (TPA-stimulation derived interphase cells or smears). Wrong conclusions on clone sizes and/or development of the latter are possible, and the impact of the proportion of aberrant cells in CLL on the overall survival was reported in previous studies.20–22 Overall, according to our and previous results,13,15,16 using TPA stimulation and cell cultivation is to be preferred to iFISH in smears in CLL; also, the possibility to get banding cytogenetic result and by that to pick up unusual CLL-associated aberrations should not be neglected.
Footnotes
Competing Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
EA did the interphase fluorescence in situ hybridization (iFISH) work and drafted the manuscript; RA and TH did statistical evaluation, AG, CS, and BP performed the banding cytogenetic evaluation of the cases and provided the clinical data; and TL designed the study and finalized the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by a fellowship from the KAAD (Catholic Academic Exchange Service; to E.A.). There was no additional funding for this study.
