Abstract
Summary
Antibiotic sensitivity patterns of 21 bacterial isolates from some clinically ill New Zealand rabbits were evaluated against 12 commonly used antibiotics by the Kirby-Bauer disk-diffusion sensitivity testing method. The 21 isolates consisted of six Bordetella bronchiseptica, eight Pasteurella multocida, four Staphylococcus aureus and three Pseudomonas alcaligenes that were associated with snuffles, pneumonia, otitis media, genital infections and conjunctivitis in these groups of caged rabbits. The four bacteria species were all sensitive to kanamycin, gentamycin and enrofloxacin, with variable sensitivity to the other antibiotics tested. The results of this antibiogram could serve as a field guide in the treatment of very acute bacterial diseases of rabbit prior to the availability of the results of local sensitivity tests. Such sensitivity tests should be reviewed yearly to update this antibiogram.
The use of rabbits in toxicity studies has led to an abundance of information on the actions of drugs on rabbit tissues both in vivo and in vitro, but there is a dearth of data on their efficacy in the treatment of clinical diseases (Morris 2000, Harcourt-Brown 2002). In most countries of the world, there are only a few products licensed for use in rabbits. Toxic doses and teratogenic effects may be known, but not actual adverse effects. Young healthy rabbits are used for toxicity studies rather than aged, ill animals that have not been screened for other diseases. Therefore, the reputation of many antibiotic preparations in the clinical situation is based on limited publication (Morris 1995) and anecdotal evidence from practitioners, thus resulting in a confused picture with conflicting reports of safety and efficacy of products, especially antibiotics.
Yet, there is a temptation to prescribe antibiotics in any situation where there is a sick animal and no specific diagnosis has been made. However, because of the risk of antibiotic-associated diarrhoea (Carman 1993), it is preferable to reserve antibiotic therapy for situations where there is a definite indication for its use based on sensitivity tests. This is particularly necessary, given the peculiar digestive physiology of the rabbit, for example with ampicillin, where the beneficial caecotrophy could be hindered (Escoula et al. 1981), the aerobic to anaerobic bacteria ratio in the normal gut affected (Straw 1988) and antibiotic-associated enterotoxaemia observed (Carman 1993).
Clinically, a standardized test is needed to compile antibiotic sensitivity patterns of the more commonly isolated pathogens of laboratory rabbits in order to determine the prevalence of resistant strains and specifically to ascertain which antimicrobial agent could be recommended to treat a particular bacterial disease.
This report gives the antibiograms of some pathogenic bacteria isolated from some clinically ill rabbits in the Experimental Animal Unit (EAU) of the University of Ibadan, Ibadan, Nigeria.
Materials and Methods
Clinical history
Several reports of sudden death and general poor growth rate were received from the rabbit unit of the University's EAU in May 2006. Clinical examinations revealed rhinitis, conjunctivitis, otitis media, abscesses in some animals and dyspnoea, grunting and mucopurulent nasal discharges suggestive of snuffles in some other animals. In one section of the animal unit (no. 3), two older males had orchitis, three young males had severe conjunctivitis, while one doe had a suppurative genital infection, in addition to severe dyspnoea.
Incidental pneumonic lesions consisting of consolidation of the anteroventral lung area were observed at the postmortem examination of two apparently healthy rabbits that died suddenly on the day of sampling. One of the two rabbits also had a large thoracic abscess.
Bacterial sampling and identification
Swab samples were taken antemortem and postmortem from the sick and dead rabbits, respectively. Deep nasal swabs were taken from animals with pneumonia and rhinitis, using the commercial swabs (Stuart's transport medium, Oxoid, Basingstoke, UK). Identification of bacterial isolates was done by standard methods used routinely in the microbiology laboratory, including serological typing and biochemical tests as detailed by Carter and Cole (1990). In addition, control strains of Staphylococcus aureus, ATCC25953, were used as quality controls to check disk potency and other factors that might affect the inhibition zone size.
Susceptibility testing
Antibiograms were performed using the high-concentration single disk method of Bauer et al. (1966). The Mueller-Hinton agar (Schering Corporation, Bloomfield, NJ, USA) was used. The agar was enriched by the addition of 5% sheep blood for strains of bacteria that failed to grow using the regular Mueller-Hinton formula. The medium was prepared and poured into 15 × 150 mm petri dishes at a depth of 5-6 mm. This was refrigerated and used within one month as recommended (Dewees et al. 1970). The selection of antibiotics was based largely on whether isolates were Gram-positive or negative and varying combinations of 12 antibiotic disks were used with no more than 10 disks per plate (Baltimore Biological Laboratory, Cockeysville, MD, USA). Disk potency in micrograms or units is tabulated in Table 1.
Antibiograms of pathogenic bacteria isolated from sick rabbits
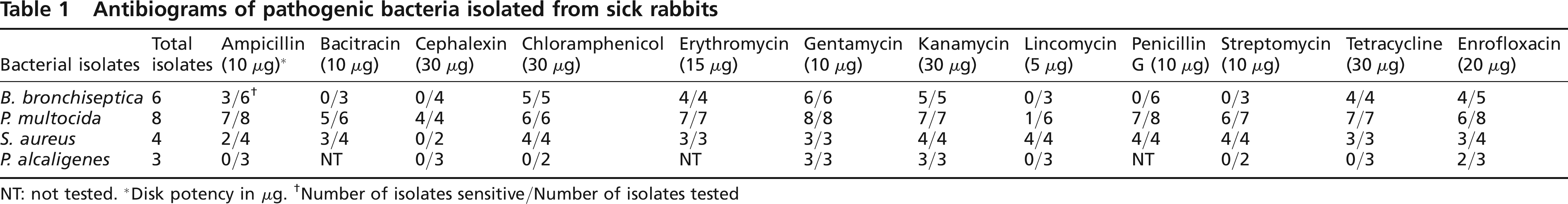
NT: not tested.
Disk potency in μg.
Number of isolates sensitive/Number of isolates tested
Measurement of zone size
The measuring template that came with the antimicrobial disks was used to evaluate the inhibition zone of each agent. The sensitivity of isolates to each agent was evaluated by comparing the zone sizes to a standard interpretative chart as recommended (Anoon 1972). Zone size for each antimicrobial agent was measured independently before comparison. The specific identification of each bacterial isolate was of utmost importance in the validation of each measurement (Shahiki & Ellner 1969).
Results
The sensitivity patterns of the pathogenic bacteria from the rabbit unit of the EAU are summarized in Table 1. The table presents the ratio of the number of isolates sensitive to the number of isolates tested against each antibiotic tested for each isolate. Bacterial isolates in general have uniform antimicrobial sensitivity patterns regardless of the specific source or disease syndrome (Ericsson & Sherris 1971); therefore they are not tabulated according to the organ or disease syndrome.
All isolates were sensitive to gentamycin, kanamycin and enrofloxacin, including eight isolates of Pasteurella multocida, from six rabbits with snuffles and pneumonia and two with rhinitis and otitis media. Bordetella bronchiseptica was tested in six rabbits, four with snuffles and two with pneumonia. Staphylococcus aureus was tested in four rabbits, two with conjunctivitis and two with genital infections, while Pseudomonas spp. were tested in three rabbits, two rhinitis and one septicaemia. Pseudomonas spp. were the least sensitive, being resistant to ampicillin, cephalexin, chloramphenicol, lincomycin, streptomycin and tetracycline, while P. multocida and S. aureus were the most sensitive, being sensitive to 11 of 12 antibiotics. B. bronchiseptica was intermediate in sensitivity, being resistant to five of the 12 antibiotics tested (Table 1).
Discussion
The results of disk-diffusion antibiotic sensitivity tests usually differ from one laboratory to another and, in the absence of standardization, it becomes difficult to compare laboratory results (Matseni et al. 1969). However, the Kirby-Bauer method adopted in this study, though old, is a well-controlled procedure that deserves wider application in animal disease diagnostic laboratories.
The choice of antimicrobial agents appropriate for treatment of a given condition involves the consideration of a number of factors, including the predicted susceptibility of the infecting organisms to antibiotics, the site and nature of the infections, the routes of administration in a species, species or individual idiosyncrasies and the pharmacodynamics of the agent (Blazevic et al. 1972).
The veterinarian in charge of the laboratory animal facility usually assesses these considerations using professional judgement, and a good understanding of the pathological processes characteristic of the disease being treated is also essential for the full appreciation and field application of any antibiogram. In the application of the results of this or other sensitivity studies, veterinarians must remember that discrepancies usually exist between in vitro and in vivo sensitivities. This may be because of poor absorption of antibiotic from the site of administration (e.g. neomycin and streptomycin from the gut), failure of some antibiotics to reach effective concentration in certain tissues, mixed infections, incorrect diagnosis of bacterial diseases, antibiogram information from different organisms or dead tissue and pus or fibrotic lesions preventing antibiotics from reaching sensitive organisms (Ericsson & Sherris 1971). In these situations prolonged administration or increased dosage of the selected antibiotics is advisable. If, however, the organisms acquire resistance through mutation or selection in the course of prolonged use, a change to other effective antibiotics based on the same antibiogram is recommended.
Ericsson and Sherris (1971) have reported that many organisms, especially staphylococcus and enteric pathogens, readily develop resistance to drugs and their sensitivity patterns change from time to time. Also, bacterial isolates from different locales may vary slightly in their sensitivity from these results. Ideally, an antibiogram should be performed on each bacterial pathogen prior to initiation of treatment, however, this is often impractical. In some instances of naturally occurring infectious diseases of laboratory animals, it may be helpful to begin treatment before the results of an antibiogram are available. When isolate-specific antibiograms are not available, the sensitivity patterns of the laboratory rabbits presented here may serve as a good field guide. This does however require periodic review for possible emerging microbial resistant strains.
Footnotes
Acknowledgement
The technical assistance of the staff of the Department of Veterinary Microbiology and Parasitology is gratefully acknowledged.
