Abstract
Background
WEQAS, one of the largest EQA (External Quality Assessment) providers in the UK, offers a bile acid EQA scheme, with linear serum pools containing cholic acid, deoxycholic acid and chenodeoxycholic acid, reflecting levels observed in obstructive cholestasis. Total bile acids are currently measured routinely by non-specific enzymatic methods. Traceability of results to the SI unit utilizing reference target values is the preferred method of comparison of returned results where available, ensuring the transfer of accuracy from definitive methods to routine methods.
Methods
Target values have been assigned to EQA material utilizing isotope dilution gas chromatography mass spectrometry (ID-GCMS). The methodology was based on published routine methods and adapted for use as a ID-GCMS target method. The total bile acid target value was reported as the sum of the three major bile acids measured: cholic acid, deoxycholic acid and chenodeoxycholic acid.
Results
The produced target values have been used to assess the performance of total bile acid methods. A degree of variability was observed between the third-generation enzyme-formazan methods and the fifth-generation thio-NADH methods. Additionally, the Sentinel results showed a positive bias in comparison to their peer formazan method group.
Conclusion
The use of ID-GCMS target results provides a common comparison for resumed results in EQA schemes, highlighting any method differences. Thus can then aid in the harmonisation of results observed for each of the different methods.
Introduction
Bile acids are one of the major components of bile, a secretion stored in the gall bladder. The primary bile acids (cholic acid and chenodeoxycholic acid) are synthesized from cholesterol by the liver, secreted into the gall bladder and then into the small intestine, where they aid in the absorption of fat and fat-soluble vitamins and also cholesterol homeostasis. Within the intestine, bacterial deconjugation and dehydroxylation of the primary bile acids may take place to form secondary bile acids such as deoxycholic acid, lithocholic acid and ursodeoxycholic acid. Bile acids are then reabsorbed and transported back to the liver to enter the enterohepatic circulation. More recently, a role as signalling molecules and an endocrine function has been identified for bile acids. 1–3 Obstetric cholestasis is one of the most common liver conditions during pregnancy, with analysis of bile acid levels the most sensitive test. 4–5 Other conditions requiring analysis of bile acids include acute fatty liver of pregnancy, hepatitis and liver cirrhosis.
Routine methods used to measure total bile acids in serum are non-specific and measure all of the major bile acids present in serum. The most common methods used are enzymatic, using 3-α hydroxysteroid dehydrogenase to convert bile acids to 3-ketosteroids and NADH. Most routine methods either monitor the rate of formation of Thio-NADH at 405 nm (e.g. Biostat, Stockport, UK; Diazyme, La Jolla, CA, USA; Randox 5th generation, Crumlin, UK) or measure the formation of formazan at 540 nm after addition of nitrotetrazolium blue and Diaphorase (e.g. Randox 3rd generation, Sentinel Diagnostics) as indicated in Figure 1.
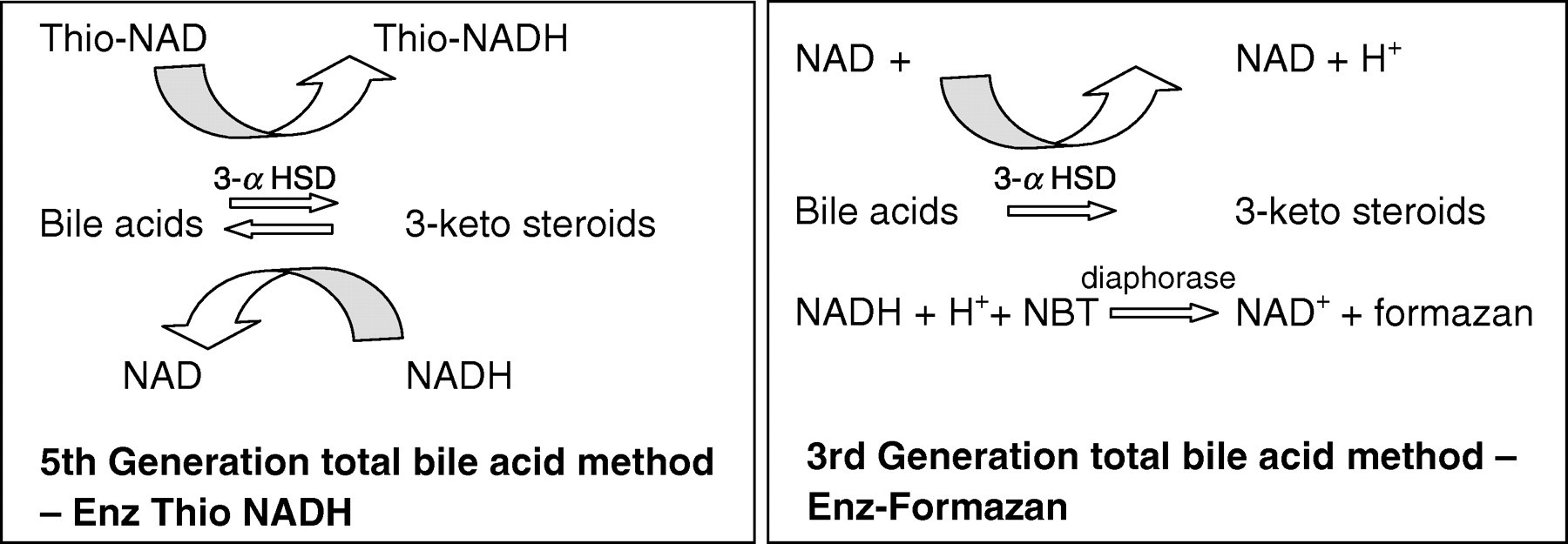
Current total bile acid enzymatic methods
Wide interlaboratory variation has been observed for the different methods within the WEQAS bile acid scheme when observing returns relative to an overall mean. The overall mean was thought to be inappropriate as a target value for this analyte as it is greatly influenced by the predominant method (Thio-NADH) and provides no information on the specificity and accuracy of the methods. Currently, specific methods for bile acids are available by both isotope dilution gas chromatography-mass spectrometry (ID-GCMS) 6–10 and tandem mass spectrometry. 11–14 Since the gold standard for reference target methodology for steroidal compounds is still recognized to be ID-GCMS, 15 a method for measurement of bile acids using this technology was therefore developed by WEQAS to assist in the harmonization of results for these assays. This ID-GCMS method was developed by the WEQAS Reference Laboratory, adapted from the method of Setchell et al. 6
Materials and methods
ID-GCMS method
An ID-GCMS method has been adapted from the method of Setchell et al. 6 The analysis was undertaken by the Reference Laboratory, a Division of the WEQAS Quality Laboratory based in Cardiff. Following extraction and delipidation, samples were chemically derivatized to form the methyltrimethylsilyl ether (Me TMS) derivatives of the bile acids prior to analysis by GCMS (Figure 2).
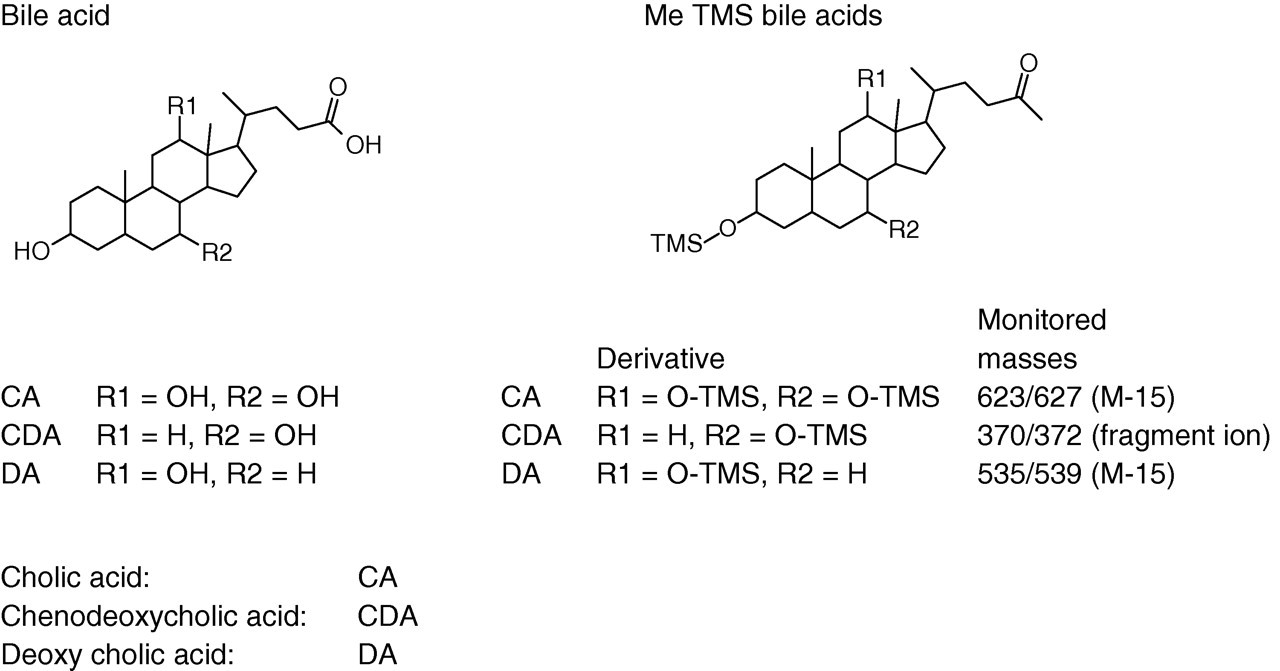
Structure of major bile acids and their methyltrimethylsilyl derivatives
Standards
Material of the purest form was used for standardization (Sigma-Aldrich, Dorset, UK; cholic acid and deoxycholic acid 99% pure, chenodeoxycholic acid 98% pure). All stocks were prepared gravimetrically in methanol (HPLC grade, Thermo-Fisher Scientific UK Ltd, Loughborough, UK). Top stocks of bile acids were prepared to provide three separate stocks designated A, B and C at a level of 1 mg/mL for each of the individual bile acids. These stocks were then diluted to provide mixed A, B and C working stocks containing each of cholic acid, deoxycholic acid and chenodeoxycholic acid at a concentration of 50 ng/μL. Standards were aliquoted into 2 mL deactivated reaction vials to produce a bracketed standard curve from each of the A, B and C working stocks containing 2.0 μg (S1), 2.5 μg (S2) and 3.0 μg (S3) of the mixed bile acids. A mixed deuterated internal standard (d 4-deoxycholic acid (2,2,4,4-d 4 (3α, 5β, 12α) – 3, 12-dihydroxy-5-cholan-24-oic acid), d 4-chenodeoxycholic acid (2,2,4,4-d 4–3α, 7α-dihydroxy-5β-cholanic acid) and d 4-Cholic acid ((2,2,4,4-d 4 (3α, 5β, 7α, 12α) – 3, 7, 12-trihydroxycholan-24-oic acid), QMX Laboratories Ltd, Thaxted, UK) was added to each standard (2.5 μg) and evaporated to dryness under a stream of nitrogen prior to chemical derivatization.
Quality control material
Since no traceable quality control material is currently available for bile acid analysis, in-house preparations were made. Quality control material was prepared using double charcoal stripped serum from pooled single donor serum, tested negative for HIV and hepatitis B and C at donor level. Cholic acid, chenodeoxycholic acid and deoxycholic acid was gravimetrically added from methanolic stocks to produce mixed bile acid concentrations of 48 (high), 25 (medium) and 10 μmol/L (low), respectively, of each of the major bile acids. This material was aliquoted into 1.5 mL cryovials and stored at −20°C.
Sample processing
The sample volume for each sample and quality control material was adjusted and aliquoted into stoppered B14 glass tubes such that the amount of bile acid present was 2.5 μg. The total volume was then adjusted to 3 mL with 1% sodium chloride. Prior to further sample processing 2.5 μg mixed deuterated internal standard (nominal concentration) was added to all samples, standards and quality control material.
All samples and quality controls were then hydrolysed by addition of 2 mL of 0.1 mol/L sodium hydroxide solution and heated at 64°C for 30 min. Bile acids were subsequently extracted by solid-phase extraction, 200 mg Isolute C18 columns (Kinesis, Cambs, UK). These columns were first conditioned with 4 mL of methanol, followed by 4 mL of purified water. Once samples were applied, they were then washed with 4 mL purified water and eluted with 2 mL of methanol. The eluants were dried under a stream of nitrogen and subsequent delipidation was undertaken by the use of Sephadex LH20 gel chromatography (GE Healthcare, Bucks, UK) using chloroform/methanol (60/40) as a running solvent. All samples, quality controls and standards were dried under a stream of nitrogen. The bile acids were first methylated by addition of 100 μL methanolic HCl (8.25 mL conc. HCl added to 100 mL of methanol) and allowed to incubate at room temperature for 30 min. Each was then evaporated to complete dryness under a stream of nitrogen at 60°C. Silylation was achieved by addition of 25 μL of trimethylsilylimidazole (TMSI, Sigma-Aldrich) at 80°C for 120 min to form the derivatives indicated in Figure 2.
All samples were injected into a Hewlett Packard 5973 Mass Selective Detector coupled to a Hewlett Packard HP6890 Gas Chromatograph with a Gerstel ptv injector. The column used was a 0.2 mm diameter, 0.33 μm film, 25 m J&W DB5-MS. Both injector and oven were set at an initial temperature of 75°C, with the ptv injector ramped to 270°C at a rate of 12°C/min and held for two minutes. The oven was ramped to 295°C at a rate of 25°C/ min with a gas flow of 0.5 mL/min. Gravimetric analysis was used throughout, allowing uncertainty measurements to be estimated according to GUM, 16 with traceability by use of gravimetrically prepared QC material (Table 1, certified material not available). All sample volumes were adjusted such that the ratio of the concentration of analyte to internal standard was approximately 1 (50 ng). Measured results were calculated using a bracketed standard curve with standards giving response ratios of 0.8 (40 ng bile acid), 1 (50 ng bile acid) and 1.2 (60 ng bile acid), respectively.
Reproducibility (internal QC material)
EQA sample preparation
The EQA samples were prepared to reflect the bile acids found in obstetric cholestasis, with cholic acid used as the predominant bile acid. Additional bile acids in the form of chenodeoxycholic acid and deoxycholic acid were also included. The overall ratio is usually 3.4:1 for cholic acid and the other bile acids, respectively.
EQA material
The base material was human serum, tested negative for HIV and hepatitis B and C at donor level. The samples were spiked with the major bile acids, since endogenous levels in normal subjects tend to be low. The pools were filtered aseptically and gentamicin added to maintain sterility prior to spiking. Other preservatives such as sodium azide are not appropriate as some of the enzymatic reactions used during routine analysis of bile acids may be inhibited. All samples were stored at −20°C prior to distribution to laboratories participating in the bile acid EQA scheme. All prepared samples had target values assigned using the ID-GCMS method.
For the recovery exercise, three additional samples were prepared (samples 4, 5 and 6), and spiked with approximately 100 μmol/L of one of each of the major bile acids. Sample 4 was spiked with 100 μmol/L of cholic acid, sample 5 with 100 μmol/L chenodeoxycholic acid and sample 6 with 100 μmol/L deoxycholic acid. These samples were targeted using the ID-GCMS method and distributed to all participating laboratories within the bile acid EQA scheme along with their normal sample distribution.
Results
ID-GCMS method
The total ion count of processed samples showed good separation of each of the major bile acids analysed. Further specificity for each is achieved by single ion monitoring of the respective ions as detailed in Figure 2. Each bile acid forms a fragment ion of m/z 370 (Figure 3, traces C1 and C2); therefore, the long GC oven programme was necessary to increase the retention time and give base peak separation between deoxycholic acid and chenodeoxycholic acid (Figure 3). Studies have shown no interference from other major bile acids (ursodeoxycholic acid and hyodeoxycholic acid, data not shown).
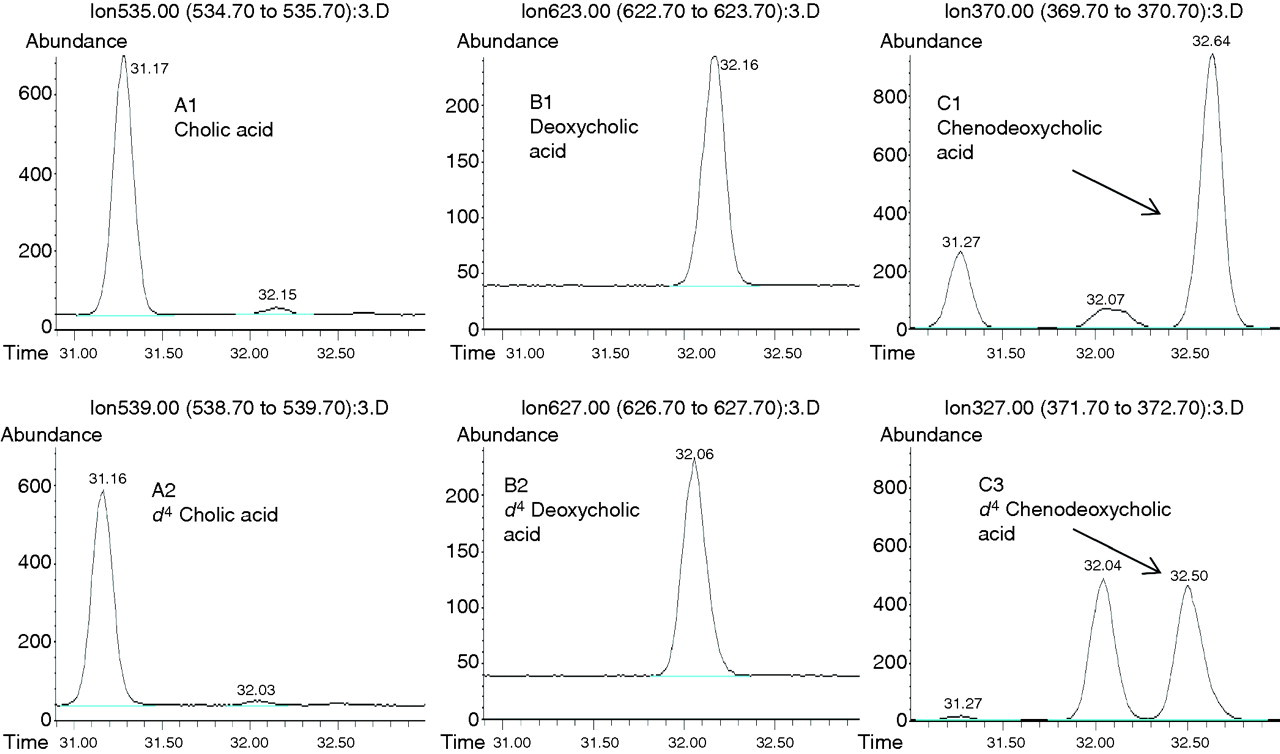
Bile acid chromatograms obtained from ID-GCMS analysis of cholic acid, chenodeoxycholic acid and deoxycholic acid. ID-GCMS, isotope dilution gas chromatography mass spectrometry
Reproducibility was assessed using quality control material consisting of charcoal stripped serum spiked with each bile acid (Table 1). These samples were analysed six times in duplicate in three separate analytical runs. The percentage coefficient of variation for each was less than 3.5% for each level tested.
For EQA purposes, target values were provided as the sum of each of the major bile acids measured by ID-GCMS.
EQA scheme
Observation of returned data for the WEQAS bile acid scheme indicated a marked difference in performance between the various analytical groups. The majority of participants are currently using the enzymatic Thio-NADH method and as a result this method would dominate the overall mean value if used to compare scheme participant data. Clearly, there was a need for the implementation of a common reference target point. The ID-GCMS method was therefore developed to address this issue. The major groups returning results as part of the scheme were enzymatic and used either Thio-NADH (enz-thio-NADH: 5th generation) or formazan (enz-formazan: 3rd generation) as the endpoint detection. Figure 4 shows the difference between these groups across the analytical range of the EQA scheme. The users of the Sentinel Diagnostics formazan method were assigned a separate group within the WEQAS EQA scheme due to the differences in both analytical bias and imprecision for this method when comparing them with their enz-formazan peer group. The observed bias was seen to increase at lower concentrations of sample (Figure 5). While all methods state that there are few interferences, at the lower concentrations of bile acid it is likely that any interference would result in a greater effect on the measured concentration. Other bile acids present (ursodeoxycholic acid and lithocholic acid) circulate at a much lower level than the three major bile acids chosen for use as the target (1–2% of the total bile acids present). The levels of these lesser bile acids in the EQA samples are therefore unlikely to exert an effect on the final analytical result of the routine methods. The Sentinel method shows a higher bias compared with that of the other formazan methods, which may indicate a calibration issue. As a result of these observed differences between the different methods, the recovery experiment was undertaken to observe the specificity of the methods to each of the major bile acids present.
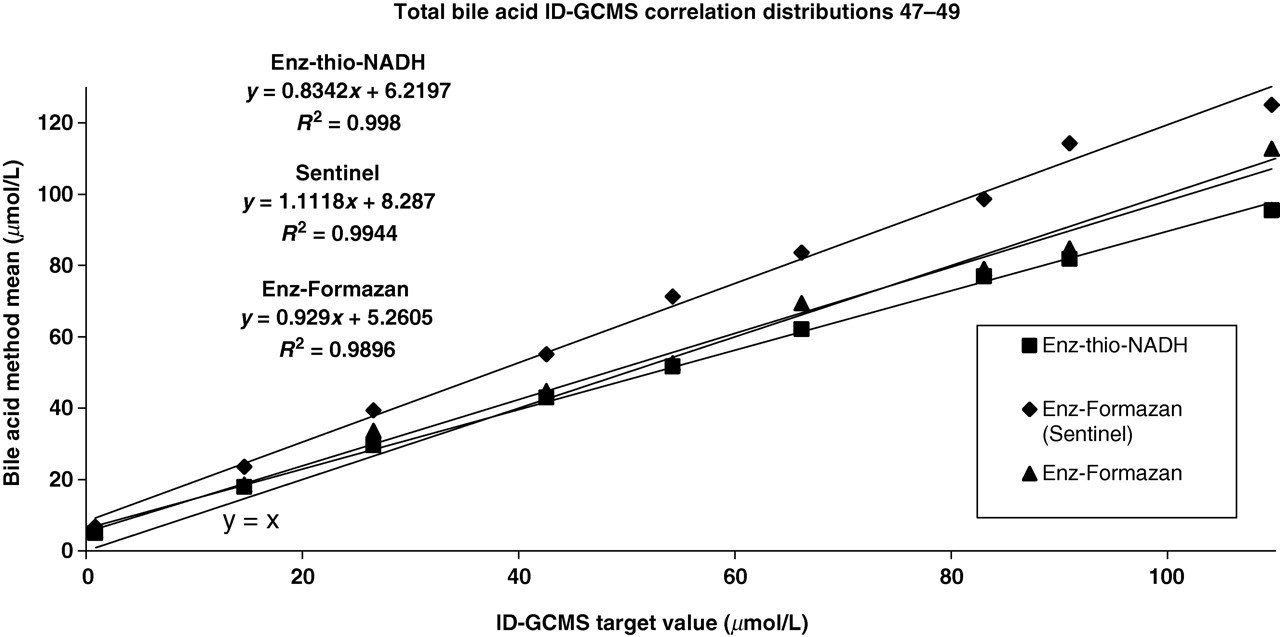
Comparison of returned results for total bile acids with an ID-GCMS target method (combined cholic, deoxycholic andf chenodeoxycholic acids). ID-GCMS, isotope dilution gas chromatography mass spectrometry
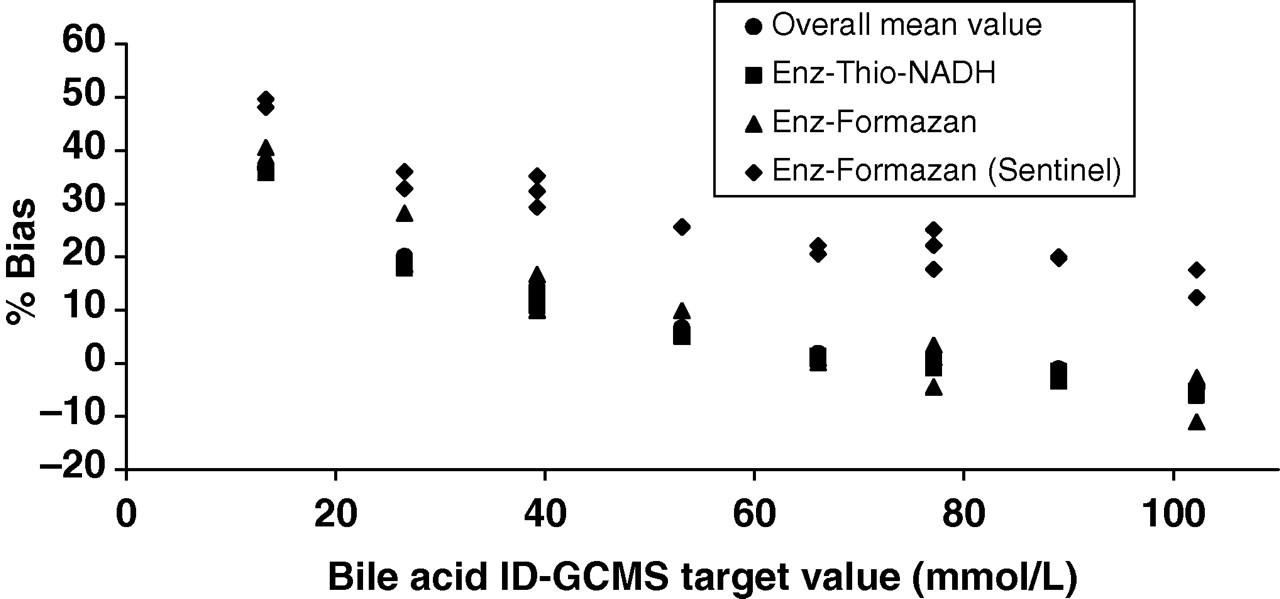
Bias plot for returned results for distributions 62–67
By spiking samples with individual bile acids, it was possible to assess the recovery of the total bile acid methods for each of cholic acid, deoxycholic acid and chenodeoxycholic acid. Comparing the recovery of the methods for each of bile acids with the ID-GCMS target method (Table 2), a wide variation in the performance of the commercially available methods for the analysis of total bile acids was observed. For cholic acid, the enz-thio-NADH and enz-formazan groups had a negative bias of −14% and −20%, respectively, while the Sentinel group had a positive bias of +21%. For chenodeoxycholic acid, these methods had a negative bias of −36% and −28%, respectively, with a positive bias for the Sentinel kit of +12%. For deoxycholic acid, good agreement with the ID-GCMS method was observed for the enz-thio-NADH method, with a +26% positive bias for the Sentinel method. The variability in the enz-formazan method for deoxycholic acid was too wide to determine bias where a standard deviation of 22 μmol/L was observed.
Bile acid recovery experiment: comparison with ID-GCMS targets
ID-GCMS, isotope dilution gas chromatography mass spectrometry
Poor recoveries were observed for chenodeoxycholic and cholic acid for all methods. However, as similar specificities were observed for all the bile acid compounds for the Sentinel method this could potentially be addressed through re-standardization. The variability in specificity of the other methods, however, suggests that harmonization of these methods may not be achieved easily.
Conclusion
A specific method for bile acids has been developed using ID-GCMS, suitable for the comparison of returned results from participating laboratories within the WEQAS bile acid EQA scheme. Such a comparison is deemed superior when compared with the use of trimmed laboratory means for comparison of EQA data. Since current routine bile acid methods are very non-specific, the major bile acids normally present were selected for use in the EQA scheme and each of cholic acid, chenodeoxycholic acid and deoxycholic acid were targeted by the ID-GCMS method. The ID-GCMS method has been used to compare all returned results for the bile acid scheme. Such a comparison may aid in the harmonization of results for bile acids produced by the different methods. Evidence on the variability in specificities of the methods for the different bile acids has been identified when comparing scheme participant data with a common target point.
DECLARATIONS
