Abstract
Background
Golgi protein-73 (GP73) is a newly identified candidate serum marker for liver diseases. The utility of this biomarker remains limited, largely due to the lack of quantitative information. The aims of this study were to quantify serum GP73 (sGP73) in healthy individuals and in patients with liver diseases, and to validate sGP73 as a biomarker for early diagnosis of liver disease.
Methods
Recombinant GP73 was used to generate monoclonal (mAb) and polyclonal antibodies (pAb). Using these antibodies in a quantitative enzyme-linked immunosorbent assay, GP73 was measured in serum from 263 patients with various forms of liver and other diseases.
Results
The median sGP73 in patients with liver disease was significantly higher (P < 0.001) than in healthy individuals and in patients with other diseases. When sGP73 was used to detect liver disease, it had a sensitivity of 82% and a specificity of 80% at the optimal cut-off value of 85.5 μg/L. The area under the receiver-operating characteristic curve was 0.9.
Conclusion
sGP73 concentration in patients with liver disease was three-fold higher than in healthy individuals. However, sGP73 concentrations did not differ significantly between patients from each liver disease group. Furthermore, sGP73 was not significantly elevated in patients with diseases other than liver disease compared with healthy individuals. These results suggest that sGP73 may be used as a serum marker for the diagnosis of liver disease.
Introduction
The spectrum of liver disease includes acute hepatitis, chronic hepatitis, liver fibrosis, cirrhosis and hepatocellular carcinoma (HCC), with liver cirrhosis as the most important factor in the development of HCC. 1 The clinical manifestations of liver disease are diverse, ranging from isolated and clinically silent laboratory abnormalities to dramatic and rapidly progressive liver failure. It is estimated that approximately 40% of patients with cirrhosis are asymptomatic. Early diagnosis of liver disease could provide more therapeutic options, and in patients with cirrhosis facilitate the prevention of HCC. 2
Golgi protein-73 (GP73) is a newly identified serum marker for HCC. 3–6 In normal human liver, it is expressed in biliary epithelial cells, but is barely detectable in hepatocytes. Its expression is, however, upregulated in hepatocytes from patients with viral and non-viral liver disease. 3,7–9 Marrero and co-workers 5,6 reported that GP73 is present in human serum, and sGP73 is also upregulated in HCC. After semi-quantification of sGP73 with densitometric analysis of immunoblots from 352 serum samples, they concluded that sGP73 is a better marker than α-fetoprotein (AFP) for the diagnosis of HCC. Subsequently, Block et al. 10 found that GP73 expression was elevated in the serum of the woodchuck model of HCC. However, the mechanism of GP73 regulation and the biological significance of elevated sGP73 expression remains unclear. Iftikhar et al. 7 proposed that there are two mechanisms to regulate GP73; the first of which is triggered during acute hepatocellular injury, and the second during the progression of chronic liver disease. More recently, Bachert et al. 11 reported that upregulated intracellular GP73 expression enhances its intracellular trafficking, most likely through the endosomal pathway, which provides the opportunity for endoproteolytic cleavage of GP73, resulting in the secretion of the truncated GP73. These studies provided a plausible model to explain the higher sGP73 concentration observed in patients with liver disease.
When detected by immunoblotting and measured semi-quantitatively by densitometry, sGP73 shows better sensitivity and specificity for HCC compared with AFP, 5 the only serum marker currently recommended for HCC. However, the use of GP73 as a clinical serum marker for the routine investigation of liver disease (HCC in particular) remains limited, 10 mainly due to the lack of a convenient quantitative assay. 12
In this study, we have generated antibodies against GP73 and have used them to develop a sandwich enzyme-linked immunosorbent assay (ELISA) for the quantification of GP73 in serum. The method was then used to determine the concentration of GP73 in serum samples from patients with liver and other diseases and also healthy individuals. We believe this to be the first report of the quantitative measurement of GP73 in serum.
Materials and methods
Cloning and expression of recombinant GP73
Recombinant GP73 proteins were prepared as described by Kladney. 3 Briefly, a 1083 bp fragment of the GP73 cDNA, corresponding to the intra-Golgi portion of the protein (amino acids 41–400), was amplified by polymerase chain reaction (PCR) using a primer pair (5′-CATAGGGATCCCTCCAGACACGGATCATGGAGCTGGAAGGC-3′) and (5′-GAGAGAAGCTTTCAGAGTGTATGATTCCGCTTTTCACGCTG-3′). The PCR fragment was cloned to the bacterial expression vector pTrcHis A with an N-terminal 6× histidine tag (Invitrogen, CA, USA), using the BamHI and HindIII restriction sites within the multicloning site. The GP73–His fusion protein was expressed in TOP10 bacteria (Invitrogen), and purified by Ni-NTA affinity chromatography, according to the manufacturer's protocols (Ni-NTA agarose, Qiagen, Germany).
Generation of GP73 antibodies
Polyclonal antibodies (pAb) were prepared in rabbits according to standard procedures. 13 Briefly, New Zealand white rabbits were immunized with purified recombinant human GP73 protein in complete Freund's adjuvant (Sigma Chemicals Co.) to prepare hyper immune sera. The sera were used for antibody purification, ELISA, and western blotting.
Monoclonal antibodies (mAb) were prepared in Balb/c mice according to methods published elsewhere. 14 Ascites were collected, pooled and isotyped with hybridoma subisotyping kit (CalbioChem, Germany).
Serum samples
The study protocol was approved by the Guangzhou Institute of Biomedicine and Health's Institutional Review Board with informed consent. Serum samples were randomly obtained from patients attending Guangdong Provincial People's Hospital during hospital examination. The diagnosis was made based on guidelines from the Chinese Society of Hepatology, the Chinese Society of Infectious Diseases and the Chinese Medical Association. 15–17 In total, 155 patients had liver disease (58 females, 97 males, mean age [range] 54 [15–86] years). Of the total, 57 had hepatitis, 69 had cirrhosis and 29 had HCC. Serum was also collected from 36 individuals with Chlamydia pneumoniae infection-related respiratory diseases (14 females, 22 males, mean age [range] 45 [20–65] years) and 72 healthy individuals (20 women, 52 men, mean age [range] 41 [18–52] years). All serum samples were stored at −80°C until analysis.
GP73 sandwich ELISA
ELISA plates (Costar, USA) were precoated with anti-GP73 mAb and blocked with phosphate-buffered saline (PBS) containing 2% (w/v) bovine serum albumin (BSA). Purified recombinant GP73 protein was used as a standard. Serum samples were added to the wells. Anti-GP73 pAb was then added, followed by horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG (Pierce, USA). The plates were washed five times with PBS (pH 7.4) containing 0.05% Tween-20. Absorbance was measured at 450 nm by a microplate reader (Bio-Tek, USA). 18–20
Statistical analysis
Descriptive statistics for sGP73 in different groups were compared by examination of histograms. Group differences were tested using the Kruskall-Wallis test and the independent t-test. A P value of <0.05 was used to determine statistical significance. To determine the optimal cut-off value for sGP73 in the diagnosis of HCC and liver diseases, receiver-operating characteristic (ROC) curves were constructed using all possible cut-offs for each assay and the area under the curves (AUC) determined. 21 All analysis was performed with SPSS software (SPSS Inc., Chicago, IL, USA).
Results
Establishment of a quantitative sandwich GP73 ELISA
GP73 is a type II Golgi resident membrane protein. The N-terminal 12 amino acid residues are predicted in the cytoplasm (Figure 1A). The intra-Golgi domain (ectodomain) of GP73 starts from amino acid 41. The secreted GP73 is proteolytically cleaved by proprotein convertase (Pc) at R52, 11 resulting in the secretion of GP73. We cloned and expressed the ectodomain of GP73 (GP73t41) (Figure 1B). Mouse mAb and rabbit pAb were generated against GP73t41. Both mAb (Figure 1C) and pAb (data not shown) recognized GP73 from Huh7 cell lysate (Lane 1, Figure 1C) and from the culture supernatant specifically (Lane 2, Figure 1C).
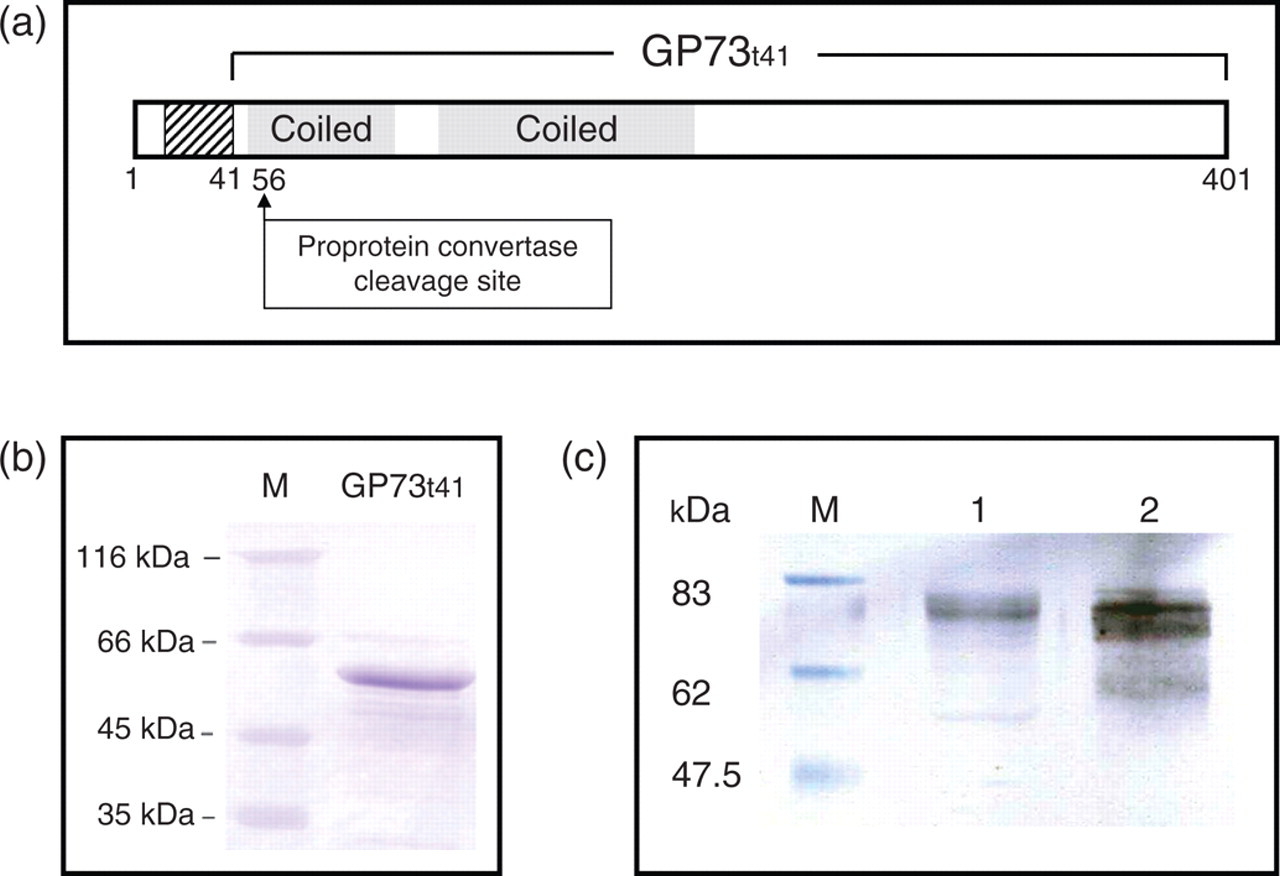
Truncated GP73t41 cloning, expression and antibody. (a) Schematic diagram of GP73. Striped box denotes trans-membrane domain; shaded boxes denote predicted coiled-coil domain; numbers indicate positions of amino acids. The predicted proprotein convertase cleavage site is indicated. (b) Purified recombinant GP73t41 expressed from Eschericia coli. M, protein size marker. (c) Specific recognition of GP73 with the anti-GP73 monoclonal antibody (mAb). GP73 in Huh7 cell lysate (Lane 1) and Hutpool cell culture supernatant (Lane 2) was detected with the anti-GP73 mAb after immunoblotting. M, Protein size marker
For quantitative determination of GP73, a sandwich ELISA was developed. This assay was performed using a combination of the anti-GP73 mAb and pAb. As illustrated in Figure 2A, polystyrene multiwell plates were precoated with mAb, testing samples were diluted and attached to the precoated plates. Captured GP73 was detected by pAb followed by the HRP-conjugated second Ab. A typical calibration curve for quantification of GP73 is shown in Figure 2B. The detection range of the ELISA permits a reliable assay of GP73 from 3 μg/L up to 250 μg/L.
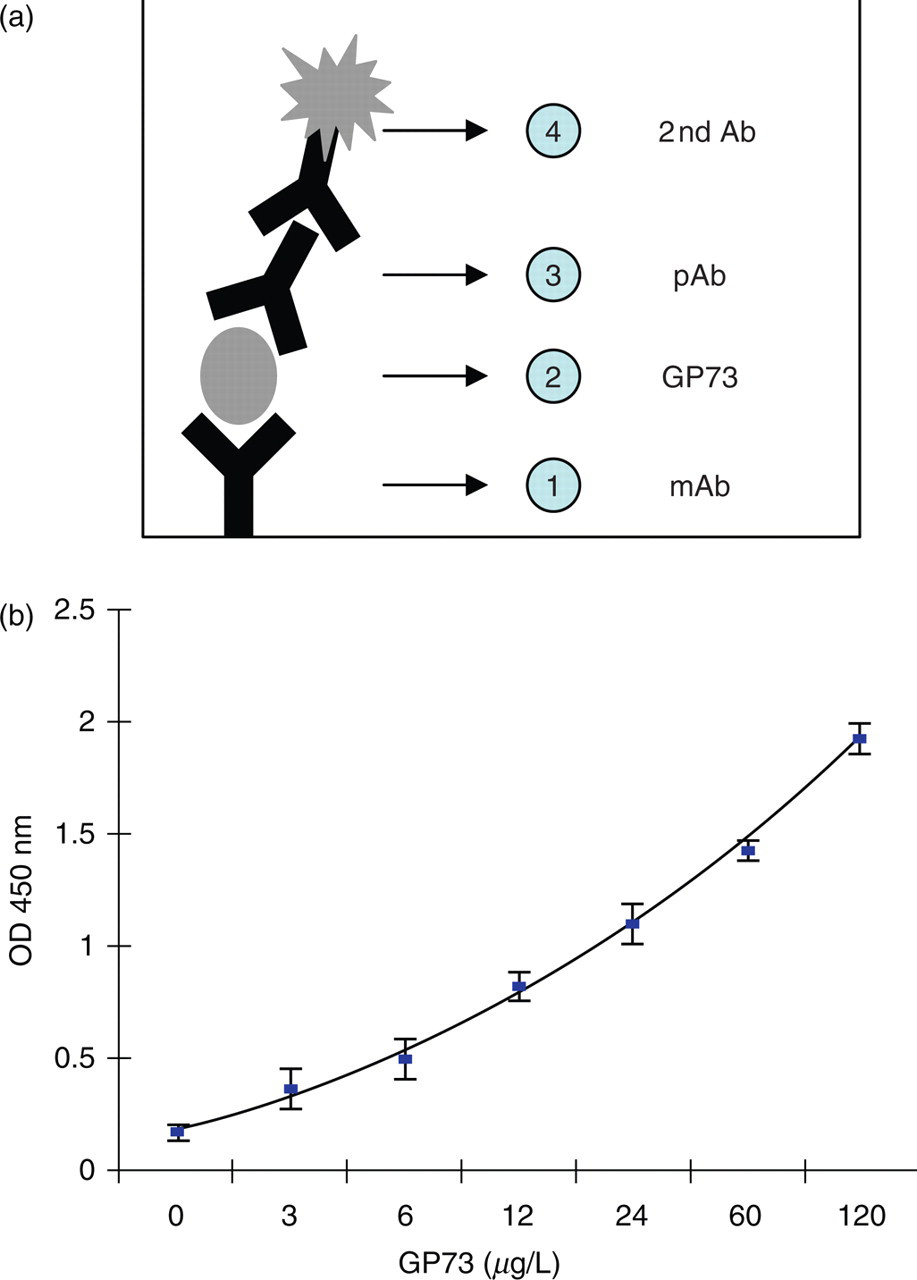
Glycoprotein73 (GP73) enzyme-linked immunosorbent assay (ELISA). (a) Schematic diagram of the assay. Monoclonal antibody was coated on the plate followed by adding the sample. The attached GP73 was detected by the polyclonal Ab. The colorimetric reaction was performed with horseradish peroxidase-conjugated anti-rabbit IgG. (b) A typical calibration curve for GP73 ELISA
Quantification of sGP73
With the GP73 ELISA available, we first determined the concentration range of sGP73 in patients with liver diseases and in healthy individuals. As summarized in Table 1, the median concentration of sGP73 in the 72 normal individuals was 54.3 μg/L – lower than that in patients with liver diseases (172 μg/L).
Serum GP73 value (μg/L) distribution in different categories
*Number of sample
†HBV DNA was measured by quantitative PCR, HBV DNA > 1000 copies/mL was considered as possible
‡Liver disease = hepatitis + cirrhosis + hepatocellular carcinoma
To determine whether elevated sGP73 was also observed in other non-liver diseases, a group of 36 serum samples from patients infected with Chlamydia pneumoniae were analysed. The median concentration of sGP73 in these patients was 63 μg/L, similar to that in healthy individuals and significantly lower than that in the patients with liver disease (P < 0.001).
Comparison of sGP73 in liver diseases
sGP73 concentration in all three diseased groups (hepatitis, cirrhosis and HCC) were significantly higher than in healthy subjects (P < 0.001), while sGP73 concentration in the non-liver diseased patients was not significantly different from that of the healthy subjects (P = 0.134) (Figure 3). Furthermore, the median concentration of sGP73 was not significantly (P = 0.379) higher in the HCC than in the hepatitis group.
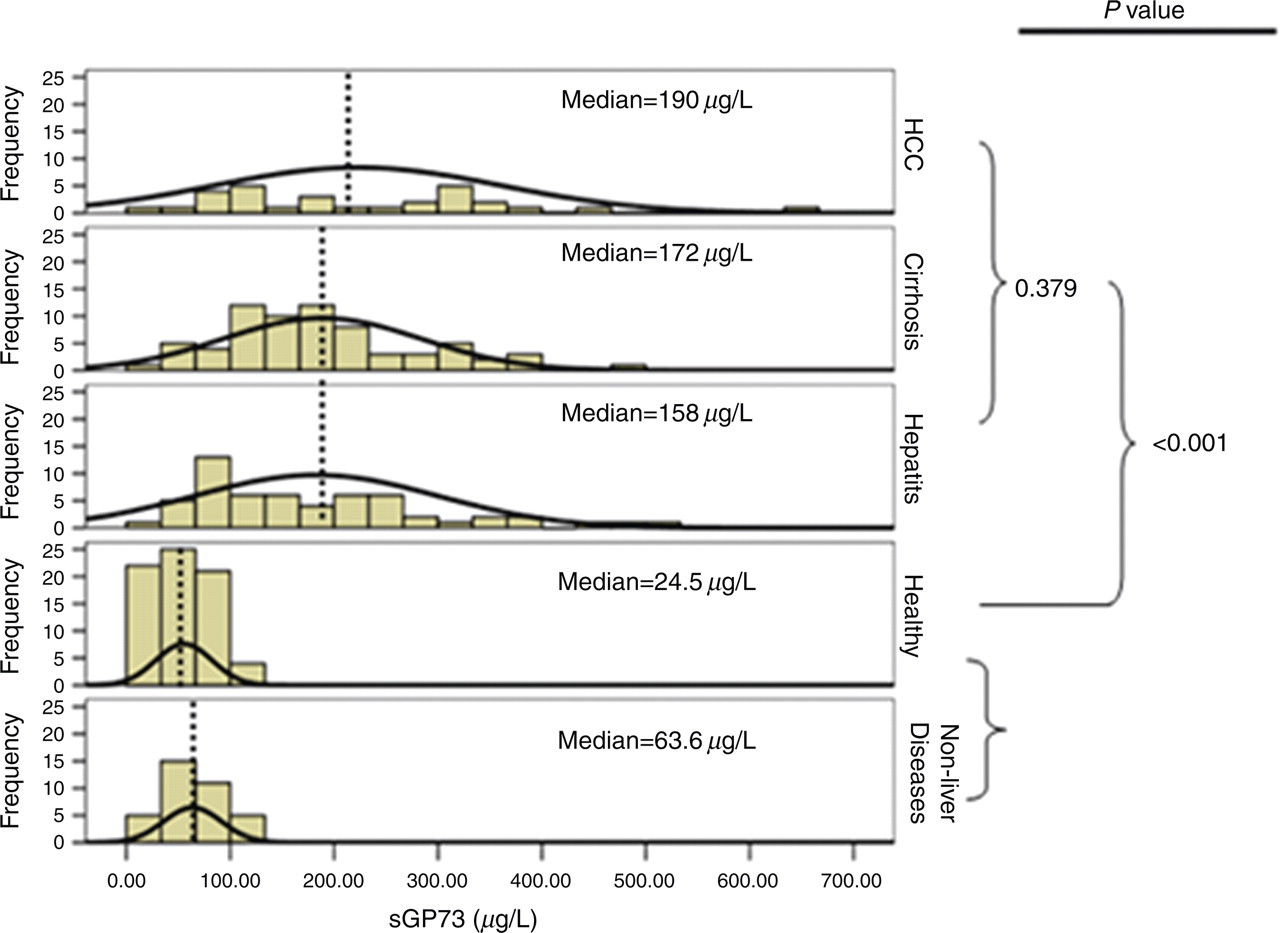
Histogram of the serum glycoprotein73 (sGP73) in different categories. P values between different groups are listed. Dashed lines indicate the mean value of sGP73 within each group
Since nearly 40% of the patients with liver disease were hepatitis B virus (HBV) DNA-positive (Table 1), and earlier studies have shown that GP73 expression is upregulated in HBV-replicating hepatocyte cells, 8 we questioned whether sGP73 in HBV-positive patients were different from those of HBV-negative ones. The concentrations of sGP73 in HBV-positive and -negative patients in each liver disease category were compared, and no significant sGP73 differences were observed.
sGP73 is a serum marker for liver diseases
The value of sGP73 as determined by the ELISA assay, in the diagnosis of liver disease and HCC, was determined through analysis of generated ROC curves. As shown in Figure 4, when sGP73 was used as a liver disease marker, the AUC was 0.91 (95% CI 0.88–0.95), predicting a good to excellent diagnostic value. Based on the ROC curve, sGP73 ELISA for liver disease had a sensitivity of 82% and a specificity of 80% at the optimal cut-off point of 85.5 μg/L. However, when sGP73 ELISA was used for HCC diagnosis, the AUC was 0.73 (95% CI 0.64–0.82), predicting a poor to fair diagnostic value. In this case, it had a sensitivity of 72.4% and a specificity of 61.5% at the optimal cut-off point of 121 μg/L.
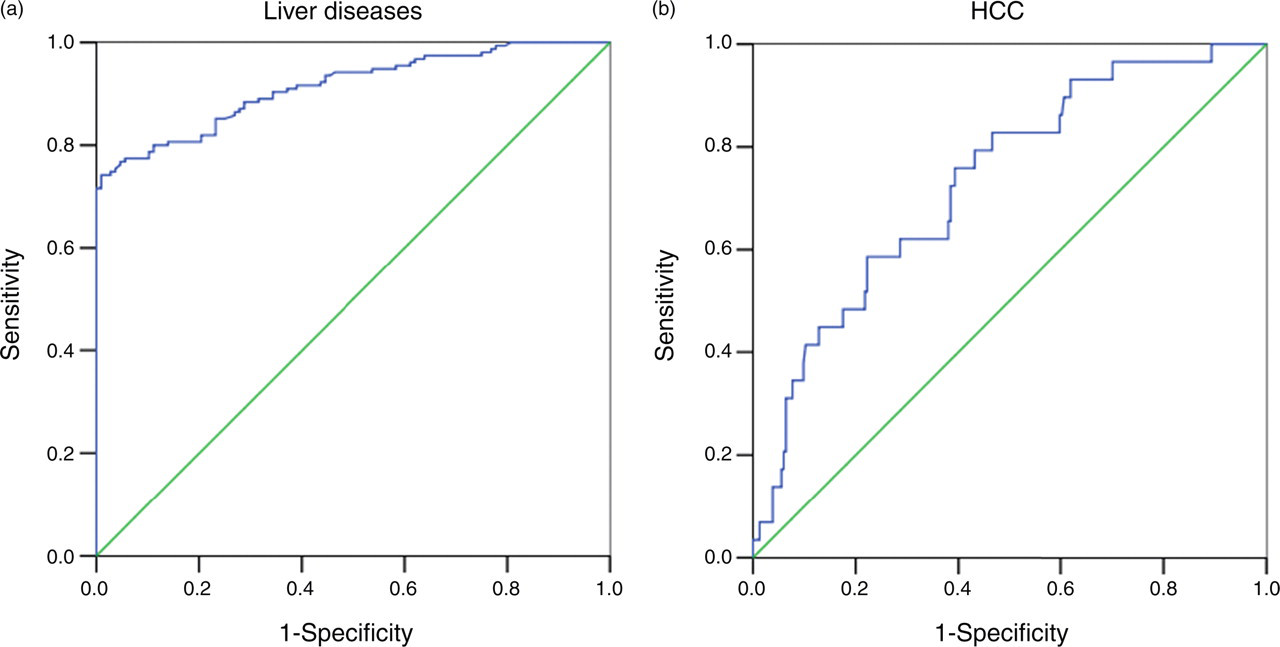
Receiver-operating characteristic curves. (a) Serum glycoprotein73 (sGP73) for all categories of liver diseases. (b) sGP73 for hepatocellular carcinoma patients
Serum GP73 was compared with other serum markers for liver diseases: alanine aminotransferase (ALT) and γ-glutamyltransferase (GGT) (Table 2). For hepatitis patients, ALT was most commonly increased with 81.4% of the samples above the cut-off value of 40 U/L. For both liver cirrhosis and HCC, however, sGP73 had the highest positive rates of 88.4% and 82.8%, respectively. When considering all the three liver diseases together, sGP73 had the highest positive rate (82%) among these three markers. These results suggest that sGP73 is a better serum marker for the diagnosis of liver disease.
Positive rates for alanine aminotransferase (ALT) and γ-glutamyltransferase (GGT) and serum glycoprotein73 (sGP73) tests
Discussion
GP73 is believed to be a new serum marker for liver disease. Studies have shown that its expression is upregulated in hepatocytes and in serum samples from patients with liver disease, with expression being lower in acute hepatitis and highest in late-stage HCC. 7 However, the methods used in these earlier studies were semi-quantitative immunohistochemistry, immunoflorescence or immunoblotting techniques, which were difficult to adapt for routine analysis. Consequently, the serum reference interval for sGP73 in healthy individuals remains unclear. In this study, we have developed a quantitative ELISA method for the determination of sGP73 and used it to measure sGP73 in a cohort of patients with and without liver disease, and in healthy controls.
GP73 is a resident Golgi membrane glycoprotein and it has been unclear as to why it should be present in serum. This question was addressed by Bachert et al. 11,22 In their report, they showed that the secretion of GP73 was directly associated with its upregulated expression, and the secreted GP73 is the proteolytic product of Pc, which recognizes R52VRR55 and cleaves at the C-terminal of R55. In our study, the secreted GP73 from Huh7 culture supernatant was purified and the N-terminal was sequenced. The N-terminal residues of the sGP73 were AAAERGAVEL (result not shown), matches the A56AAERGAVEL65 of GP73. This result confirmed that GP73 is cleaved after R55. These studies provided direct evidence that sGP73 was within the ectodomain of GP73.
To date, there have been several reports on the upregulated expression and disease association related to GP73. Results from these reports suggest that the upregulation of GP73 was a general feature of liver injury, and could be induced by viral or non-viral, acute or chronic liver diseases. In this report, sGP73 was quantitatively determined and values compared with healthy subjects and patients with various forms of liver disease. Our data indicated that the median concentration of sGP73 in healthy subjects was 54.3 μg/L, a value easily detectable with the GP73 ELISA. The median concentration in patients with liver disease was 172 μg/L, significantly higher than the median observed in the healthy group (P < 0.001). Interestingly, the median concentration of sGP73 for the group of non-liver disease patients observed in this study was not elevated, suggesting that elevation of sGP73 is specific for liver disease. However, more detailed work is required to investigate the molecular mechanisms behind the regulation of sGP73 expression and if this expression is disease-specific.
In this study, nearly 40% of the patients were HBV-positive (Table 1). After comparing sGP73 in patients with from those without HBV infection, no significant difference was found. This observation is consistent with the earlier report 8 in which HBV induced higher GP73 expression in liver tissue at a similar level as alcohol-induced liver disease. Again, these results suggest that upregulated sGP73 is a common consequence of liver damage.
ELISA is a more sensitive and superior method of quantification than immunoblotting. Use of this ELISA method should therefore result in a better comparison of GP73 values between different disease groups. For example, the immunoblot analysis might underestimate sGP73 concentration in the normal group and overestimated it in the HCC group. These differences in methodology could contribute to the different observations in our current report and the earlier report of Marrero et al. 5 In this study, sGP73 in HCC was three-fold higher than the normal group, while it was 9.4-fold higher in Marrero's report.
In summary, we have developed an ELISA for the measurement of GP73 in serum. Using this assay, we have shown that sGP73 is a sensitive and specific serum marker for liver disease. Our data has also indicated that sGP73 requires further evaluation as a marker for HCC. Use of this assay in the future will enable further studies to take place that will be able to determine any associations between sGP73 and stages of liver disease and also shed information on the regulation of GP73 expression. Results of such studies should lead to a greater understanding of the clinical diagnostic value of sGP73, together with its cellular function and role, if any, in pathogenesis.
Footnotes
Acknowledgements
We thank Changyou Chen, Yi Wang for helpful discussions. We also thank other members of the Viral Immunology Laboratory at GIBH for their support.
This work was supported by grants from Chinese Academy of Sciences (KSCX1-YW-10 to T.P.), and from Guangdong Provincial Government (no. 2007B030501012 to T.P.).
Yanli Gu and Wenli Chen made equal contributions to this work.
