Abstract
Multi-modality medical image classification aims to combine information from different modalities or devices to generate comprehensive and accurate diagnostic results. Existing research methods have ignored two characteristics of medical images across different phases: the highly redundant background and the low differentiation between different phases. Based on the idea of disentangled representation learning, we introduce a dual-branch network to disentangle images into shared features and modality-specific features. And based on the properties of different features, we propose a prototypical loss and a similar prototypical loss to constrain the two types of features, respectively. Our approach achieves strong performance in classification on the LLD-MMRI dataset and fusion on the ANNLIB dataset. Extensive ablation studies validate the contribution of each component of our framework. Code will be available at: https://github.com/wangzx-tech/explore_mmia
Introduction
Automatic classification of medical images can reduce the mental workload of physicians, improving the efficiency of diagnosis (Bhatnagar et al., 2013; Loening & Gambhir, 2003). However, previous research has been limited by the ability to collect datasets. Bring to assisted diagnosis on a single image (Li et al., 2014; Xiao et al., 2021), making it difficult to improve accuracy and real-world application. With the advancement of device capabilities, an increasing number of researchers have expanded their focus from single modality to multi-modality (Lou, 2023; Menze et al., 2015). Aiming to combine information from different medical image modalities to generate more comprehensive and accurate diagnostic results.
In the research of
However, they overlooked the unique characteristics of medical image processing tasks when incorporating this architecture (Zhao et al., 2023). Specifically, medical images tend to have highly redundant background information. For example, in multi-phase magnetic resonance imaging (MRI), the background features are similar across modalities despite phase differences. Under high inter-modality feature similarity, the capability of transformers to model relationships is impaired, which in turn inhibits the final fitting performance of the model.
Disentangled representation learning (Bengio et al., 2013) can decompose feature representations into multiple independent latent factors, allowing widespread applications in image editing and generation (Lee et al., 2018; Singh et al., 2019). Most existing image fusion methods have achieved improved performance by disentangling images into local and global features (Fu & Wu, 2021; Zhao et al., 2023). Therefore, we propose to introduce disentangled representation learning into the MMIC task by decomposing images into shared and modality-specific information. Attempt to address the problem of redundant background information and release the performance for the transformer architecture. More importantly, we designed dedicated loss functions for each branch. Through constraints such as fusion and contrast, these losses effectively guide the feature extraction tendencies of the dual branches, thereby significantly enhancing the model’s capability to extract information from different modalities.
However, existing disentanglement methods are based primarily on variational autoencoders, generative adversarial networks, and other latent variable models (Wang, Chen, et al., 2024), which are unsuitable for classification tasks. Disentangling methods used in image fusion adopt dual-branch architectures that can be adapted for classification, but most works (Fu & Wu, 2021; Vs et al., 2022) lack effective constraints on disentangled representations to ensure controllability and validity.
To address these issues, we propose corresponding constraints on the disentangled representations from the dual-branch architecture. We first validated our method for semantic extraction and fusion capabilities in the LLD-MMRI dataset, which contains eight-phase MRI images (Lou, 2023), and achieved promising results. Compared to excellent existing published methods (Gao et al., 2021), our approach can improve accuracy by 8.65%. Subsequently, we conducted experiments on medical image fusion tasks on AANLIB datasets and achieved near state-of-the-art (SOTA) performance (Zhan et al., 2022), demonstrating the versatility of our method. Figures 1(a) and 1(b) are the performance in classification and fusion, respectively, validating the capability of our method in fusing multi-modal information. Our main contributions can be summarized as: We propose a dual-branch feature extraction network. This architecture separately learns shared features and modality-specific features, effectively enhancing the model’s performance. A corresponding loss function is designed to work in synergy with our network architecture, specifically for the decoupling and fusion of different modal information in multi-modal tasks. It addresses the challenges where high-semantic information is difficult to extract and low-semantic information suffers from high redundancy. Considering the redundant nature of shared features, we propose a prototype loss to constrain them to aggregate common characteristics between modalities, thus enhancing the effectiveness of the features. For modality-specific features, in order to avoid over-constraint on similar modalities, which may lead to loss of modality information, we propose a prototypical loss based on a similarity prior, which ensures the effectiveness of distinctive features for similar modalities. We perform experiments on MMIC and medical image fusion for validation. The results achieve outstanding performance, demonstrating the feasibility of fusing multi-modalities via our method.
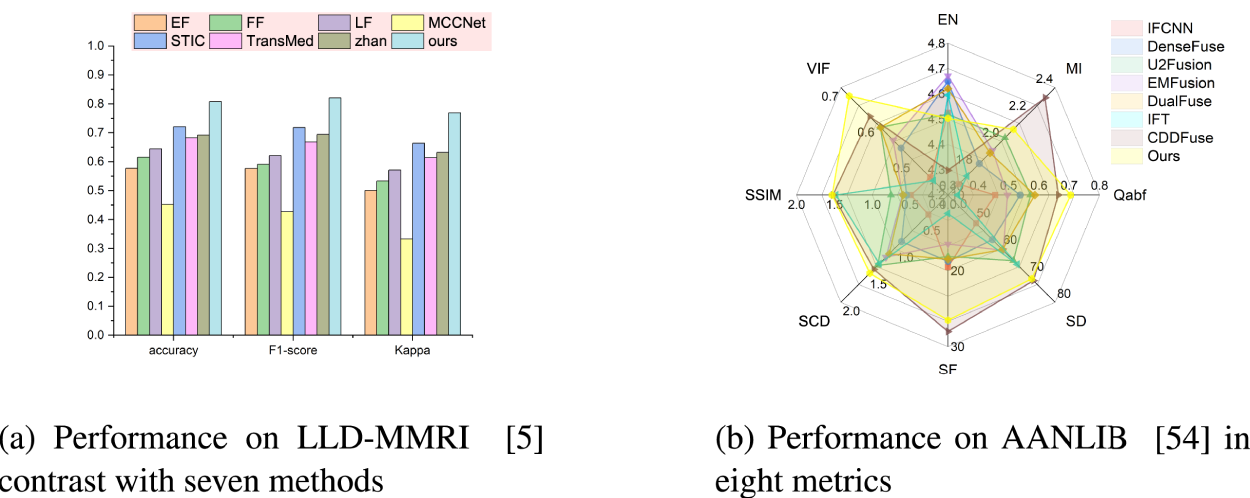
Performance comparison of our proposed method with existing classification and fusion approaches. (a) Performance on LLD-MMRI (Lou, 2023) contrast with seven methods and (b) performance on AANLIB (Zhang, 2022) in eight metrics.
This section will first introduce relevant background knowledge and previous work on medical image classification, followed by an overview of disentangled representation learning and application to image fusion. Finally, we briefly introduce the concept of prototype networks for few-shot learning.
Medical Image Classification
Common medical image analysis tasks include lesion classification and segmentation (Azam et al., 2022). Since segmentation focuses more on morphological changes while classification targets high-level semantics, which are higher requirements for multi-modal information utilization, this section elaborates on classification.
Motivated by recent advances in deep learning in natural image analysis, there has been a growing interest in leveraging deep learning for medical image diagnosis (Chan et al., 2020; Jiang et al., 2020). Li et al. (2014) employed a shadow neural network to classify interstitial lung disease in chest images. Xiao et al. (2021) proposed an ultra-lightweight end-to-end classification neural network for electrocardiogram classification. However, their networks were designed for single images, whereas in medical image analysis, advanced utilization of multi-phase data can promote better feature learning and representation, and further improve diagnostic performance and reduce uncertainty (Gao et al., 2021; Hamm et al., 2019; Panayides et al., 2020).
Although researchers have made preliminary attempts at multi-modal information fusion, such as Guo et al. (2019) utilizing multi-modal images including computed tomography (CT), MRI (T1 and T2), and positron emission tomography (PET) to identify soft tissue sarcomas, and Weibin et al. (2019) proposing a deep learning-based radiomic approach that uses multiphase CT images to predict early recurrence of hepatocellular carcinoma, these studies have limitations. The aforementioned method applies simple concatenation or averaging for fusion, which cannot fully explore complementary information across modalities.
Recently, researchers have favored transformers for MMIC (Shamshad et al., 2023). Xu et al. (2023) used attention mechanisms for feature extraction and fusion, along with utilizing prior knowledge of different lesions in the same patient, to guide accurate prediction in multiphase CT. Dai et al. (2021) and Zhan et al. (2022) both attempted to combine transformer and convolutional neural network (CNN) for MMIC, adopting the same pipeline of using CNN for feature extraction followed by transformer for feature fusion, achieving excellent classification performance.
However, their research overlooks constraints between modalities. Incorporating the dual-branch decoupling concept from image fusion, specific constraints can be designed for different features to minimize redundant information that repeatedly disturbs the model.
Disentangled Representation Learning and Image Fusion Applications
Disentangled representation learning aims to learn interpretable and controllable representations that capture the underlying explanatory factors behind the data (Bengio et al., 2013). In a disentangled representation, each dimension corresponds to a single generative factor while being invariant to changes in other factors.
Medical image fusion commonly involves modalities such as MRI, PET, CT, and single-photon emission computed tomography (SPECT), and is often jointly validated with natural image fusion (James & Dasarathy, 2014; Li et al., 2021). Recent studies on combining disentangled representation learning have achieved performance improvements. Fu and Wu (2021) proposed a dual-branch network, first decoupling features into detail and semantic ones, then fusing the decoupled features independently, before finally merging outputs of the two branches. Xu et al. (2022) proposed a unified unsupervised image fusion method that adaptively determines importance relations between images through multi-level features extracted by CNN. Subsequent studies have been largely based on two-branch architectures (Ding et al., 2020; Dong et al., 2022; Liu et al., 2021).
With the rise of transformers and their strong performance in cross-modal tasks, an increasing number of researchers have applied this architecture to image fusion (Dosovitskiy et al., 2020; Radford et al., 2021; Xu et al., 2023). For example, Vs et al. (2022) employ a two-branch design for fusion, where CNN captures local features and transformer models long-range dependencies. Zhao et al. (2023) adopt the transformer for basic feature extraction and fusion of dual-branch outputs, while the invertible neural network extracts detailed features.
However, existing loss functions for dual-branch tasks struggle to handle scenarios with more than two modalities (Azam et al., 2022; James & Dasarathy, 2014) and lack generalizability across different decoupling tasks (Chen et al., 2022). Therefore, it is necessary to design a tailored loss function specifically for the complementary information within our dual-branch framework.
Prototype Network
Prototype networks are a type of neural network that can perform effective representation learning from limited training data. The key idea is to learn a prototypical representation for each class, rather than explicitly learning a decision boundary as in traditional neural networks (Snell et al., 2017).
Prototype features help alleviate the impact of inter-class differences on the model. We transfer them to multi-modal scenarios and use them in a self-supervised manner to constrain the information deviation between different modalities, thereby reducing the influence of redundant information.
Method
This section provides a detailed explanation of the proposed method. The first subsection introduces the overall architecture of the model. The next subsection delineates the core design methodology. The last two subsections elaborate on the details of the feature fusion module and the overall training loss function, respectively.
Overview
As illustrated in Figure 2, our model architecture consists mainly of three parts. One is a dual-branch feature disentanglement module, which is used to decouple the original information into shared and specific information simultaneously. The second is a feature fusion module that fuses the shared and specific information. The last is a task decoding module.
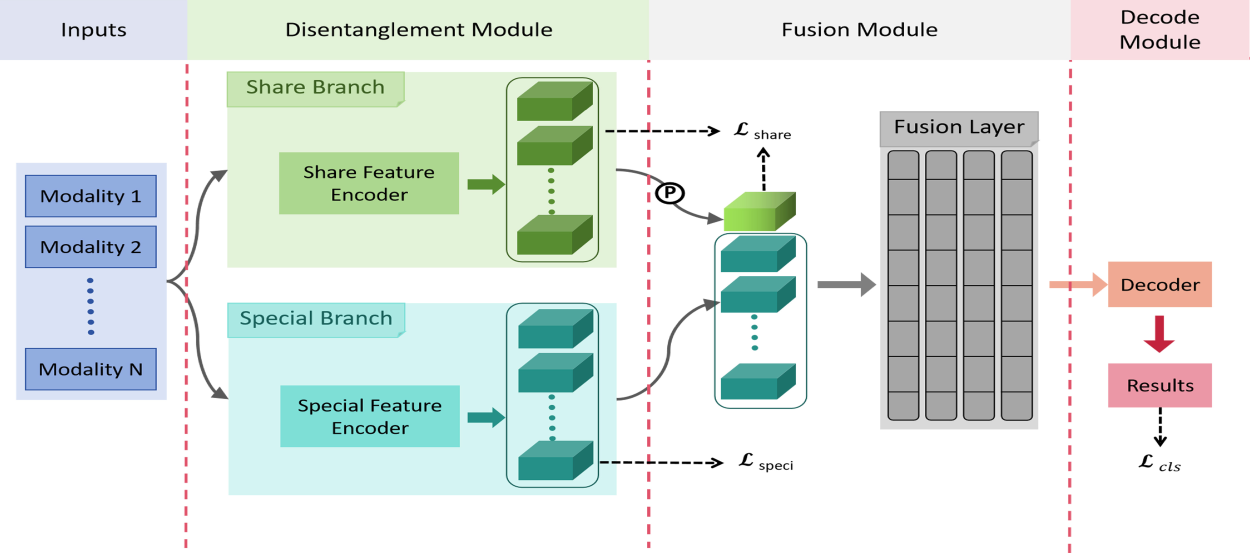
Overview of our proposed framework. The blue box represents input images of different modalities, and the overall architecture contains three parts: Disentanglement, fusion, and decoding. Where
Our core contribution lies in the feature disentanglement module, which mainly consists of a dual-branch network structure and constraint methods designed for each branch to extract disentangled representations, detailed in Section 3.2. For the feature fusion module, we adopt existing techniques with slight modifications to accommodate our disentanglement approach, elucidated in Section 3.3. As for the decoding module, a simple fully connected layer is used to obtain classification results. The fused features are flattened and then fed directly into a fully connected layer to output the final predictions.
Li et al. (2023) believe that in remote sensing image detection, most objects are small under a bird’s eye view and require environmental features such as object locations to aid identification. Similarly, in medical images, contextual information around lesion regions is also valuable for disease diagnosis. Therefore, we believe that global information can promote accuracy in MMIC. Hence, we utilize a dual-branch structure in the extraction stage to extract global and local features separately from the images. The global features primarily contain background information for the different modalities, which is weakly relevant to the lesions; we refer to these as shared features. Local features contain primarily modality-specific representations that are strongly relevant to local lesion regions. We refer to these as specific features.
Due to the fact that ResNet (He et al., 2016) shows great potential in deep learning and its simple structure, making it suitable as a benchmark network to verify the validity of the framework. For image classification tasks, simpler ResNet networks are adopted in both branches with weight sharing for all phases. The particular network configurations are visualized in Figure 3.
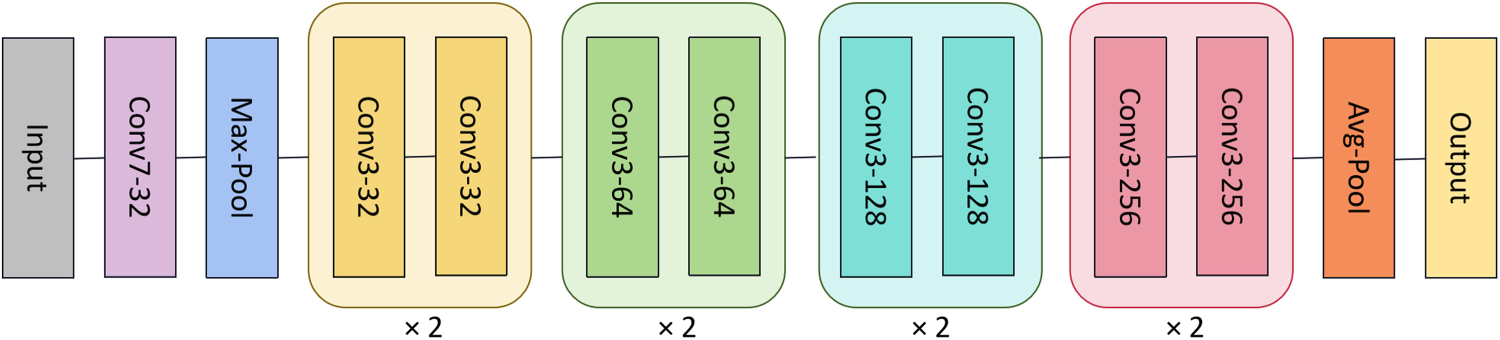
Architecture of the feature extractors in medical image classification.
Note that the adopted ResNet networks convert all the original 2D operations into 3D, in order to adapt to the 4D input and encode it into a feature vector for each modality.
However, most existing dual-branch fusion methods do not impose effective constraints on the disentangled representations, resulting in uncertain convergence of the features and making it difficult to interpret whether the model has learned the corresponding representations. Zhao et al. (2023) constrain the base and detail features between the two modalities for different branches by maximizing and minimizing the correlation coefficients, respectively, as shown in the following:

We extend the approach of equation (1) to constraint methods for an arbitrary number of modalities by adding the constraints equation (1) between every two modalities to accommodate the broader range of multi-modal model constraint methods being investigated:
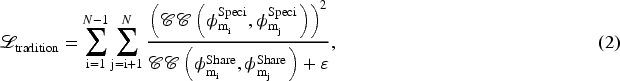
We formulate the final loss function as the sum of pairwise modality difference losses. In the next two subsections, we first analyze this simple extension approach, then propose our prototype loss to achieve effective disentanglement of multi-modal features.
First, as illustrated in Figure 4, we assume the shared feature of each modality is represented by a point. The optimization objective with Figure 4(a) is to gradually pull the feature points of all modalities closer together, which may eventually converge to a result similar to Figure 4(b). We can observe that the traditional optimization target is rather complex, making it difficult to ensure that the solutions will not bias toward any single modality, thus leading to unpredictable issues.
To address the limitations of the aforementioned constraint and considering the high redundancy among the shared features from different modalities, we can set a background feature to encompass the redundant parts across all modalities. This forces the level of importance to converge evenly across the modalities. Inspired by the prototype network in few-shot learning (Snell et al., 2017), which identifies prototype centers for each class in semantic space. We hypothesize that the shared features across modalities should have a prototype representation. Thus, we define a prototype for shared features:

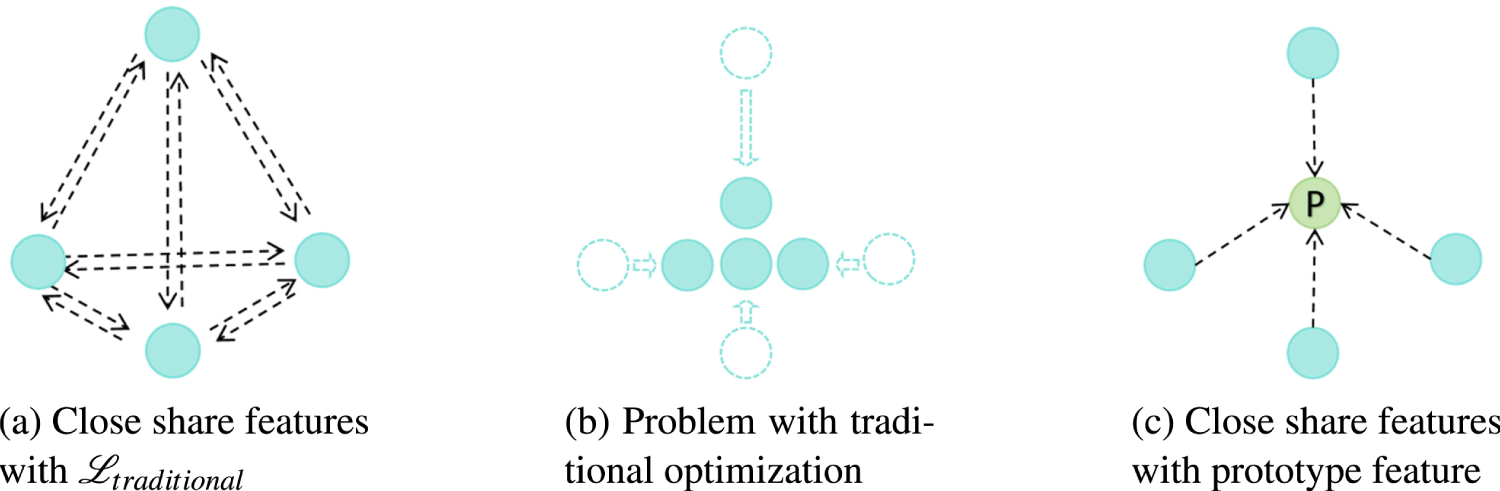
Illustrative comparison of the traditional method (equation (2)) and proposed prototype loss (equation (4)) for share features. Including multi-modality feature alignment and problems with the traditional method. Green nodes represent the computed prototype feature, blue nodes represent the original features, and arrows indicate the direction of optimization of the original features. (a) Close share features with
As illustrated in Figure 4(c), we utilize the green dot as the prototype feature. Based on this concept, we adopt the prototype feature as pseudo-labels to constrain the convergence direction of the features from each modality. Meanwhile, the constraint on each modality is independent, given the determined prototype, which encourages equal treatment of all modalities. It also mitigates the effects of noise or anomalies during modal acquisition by limiting the variance of shared features across modalities, enhancing stability while reducing the complexity of fusing shared features. The loss function is defined as:

The proposed approach visualizes the specific feature representations of each modality through differently colored data points, using matching colored arrows to denote inter-modal constraint effects (Figure 5). Infrared and visible spectrum images exhibit substantially dissimilar intrinsic feature spaces. To address this, Zhao et al. (2023) apart these modal representations by expanding the loss function to include additional regularization terms that explicitly enforce distribution apart between modal feature spaces, as shown in Figure 5(a). However, directly extending this methodology to more than two modalities significantly increases optimization difficulty due to the higher complexity of the problem.
Initially in Figure 5(b), prototype representations are leveraged to generate pseudo-label assignments for self-supervision. By imposing weighted losses that magnify the distances between encoded features and specific prototypical features, attentional mechanisms learn to focus on modality-specific patterns.
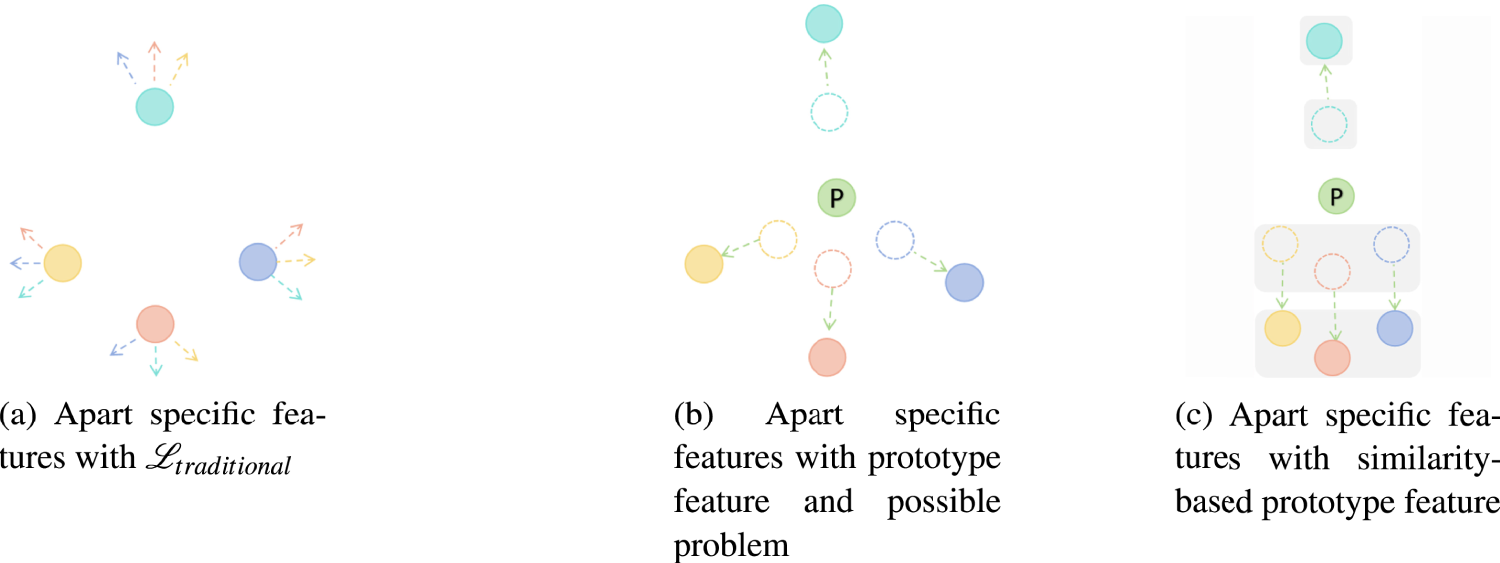
Illustrative comparison of the traditional method (equatio (2)) and proposed prototype loss (equations (7) and (9)) for specific features. Including multi-modality feature apart from the problem with the simply-prototype method. Green nodes represent the computed prototype feature, other color nodes represent the original features with different modalities, and arrows indicate the direction of optimization of the same color original feature. (a) Apart specific features with
However, both methods may overlook potential inter-modal commonalities during distancing—for example, in multi-phase MRI classification, liver cyst traits are not substantially enhanced in contrast phase, so their representations need not be far distanced, which can degrade performance or cause mistakes. Thus, we implement a similarity-preserving constraint to retain cross-modal generalizability showed in Figure 5(c). First, the cosine similarity between modal feature embeddings is quantified, then leveraged to cluster modalities based on a predefined threshold 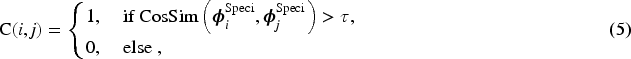
Then, we calculate the prototype features for each cluster, and the calculation function for single modality 
Subsequently, we provide cluster loss terms that enforce tight feature distributions between modalities assigned to common clusters and their respective prototypical representations. This preserves cross-modal generalizability for modalities deemed sufficiently similar by the initial thresholding process. Constraining modal feature deviations from highly representative prototypes compels generalized, transferable representations while sustaining discriminative power.

Concurrently, the cluster-specific prototypes are leveraged to extract more generalized representations for each modality. Specifically, prototypes derived from the joint cross-modal cluster distributions are utilized to initialize the modality-specific feature mappings. This framework enhances model stability while still enabling the dedicated pathways to capture intrinsic characteristics. The resultant independent prototypical representations for all modalities are obtained through an aggregation of the cluster-level prototypical vectors.

Finally, restricting the distance between prototype features and cluster-level prototypes is maximized, as shown in equation (9). It can lead to a large difference in the similarity of specific features of different modalities, thus ensuring that different modality-specific features can extract modality-specific information.
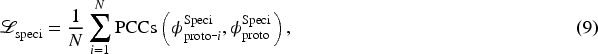
This completes an integrated regularization framework that balances (i) cross-modal generalizability through similarity-preserving constraints, while (ii) maintaining representational independence by separation from specialized prototypical conceptions. This joint optimization approach enhances model versatility across heterogeneous datasets by extracting both shared and unique information.
For image classification, the extracted shared and specific representations are fused separately. Unlike previous network fusion approaches (Fu & Wu, 2021; Vs et al., 2022; Zhao et al., 2023), we simply concatenated the shared prototype feature with specific features, input into the transformer (Vaswani et al., 2017) encoder for fusion, and the architecture is illustrated in Figure 6.
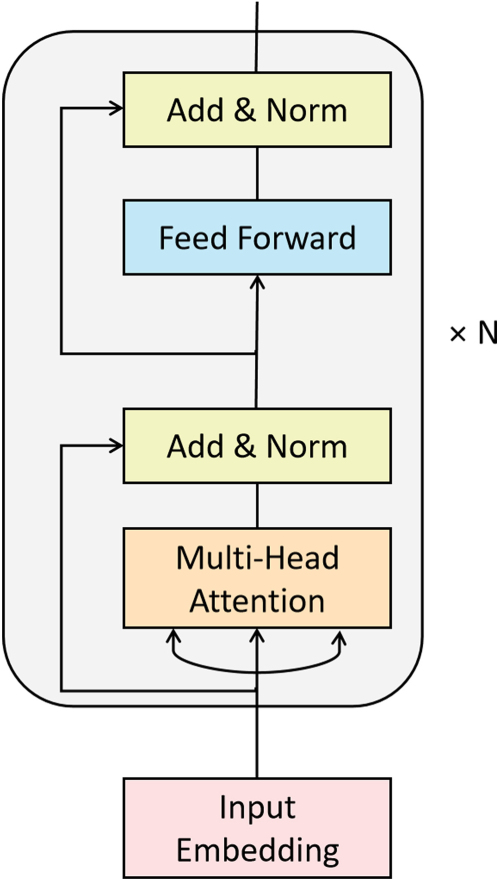
Basic architecture of the feature fusion module with
In order to validate the effectiveness of our approach, we employ the most basic transformer encoder architecture. And only retained the self-attention mechanism, which is important for multi-modal interaction, with no additional token settings. Since the information from different modalities is independent of each other, unlike the positional relationship between words or image patches, we discard the positional encoding scheme.
For classification tasks, the loss function includes the final classification loss additionally.

The final total loss function is:
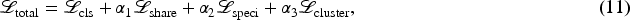
Datasets
Implementation Details
For the image fusion task, the baseline settings follow Zhao et al. (2023).
For image fusion evaluation, we adopt the same eight quantitative metrics as in CDDFuse. Specifically, entropy (EN; Roberts et al., 2008) measures the information contained in the fused image, mutual information (MI; Qu et al., 2002) evaluates the amount of information transferred from source images, and normalized weighted edge information (
All metrics used showed better performance with higher values.
Compared With Other Methods
We compare the proposed algorithm with existing state-of-the-art approaches.
First, we selected three baseline models from Guo et al. (2019), abbreviated as EF, FF, and LF, respectively, for classification. We also included four advanced approaches STIC (Gao et al., 2021), MCCNet (Xu et al., 2023), TransMed (Dai et al., 2021), and Zhan (Zhan et al., 2022) as comparison methods. Among them, except for TransMed and MCCNet, the other five approaches are based on 2D CNN, which we simply converted to 3D CNN with a conversion operation. Furthermore, since STIC uses diagnostic data as additional information and MCCNet utilizes different lesion images from the same patient to build complementary knowledge, we removed these components to adjust to the dataset.
As shown in Table 1, MCCNet performed poorly on the LLD-MMRI dataset, due to the lack of knowledge and greater inconsistency without coordination. The baseline model achieved around 60% accuracy, while existing SOTA models reach approximately 70% accuracy. Our method achieved 80.77% accuracy, improved by almost 8 percentage points over the best-performing TransMed model in this classification dataset. The F1 score increased from 0.7184 to 0.8202, a substantial gain of 0.1018. The Kappa score also improved by more than 0.1. These results from different metrics demonstrate the efficacy and powerful semantic fusion ability of the proposed method.
Quantitative Results of the Classification Task on LLD-MMRI.
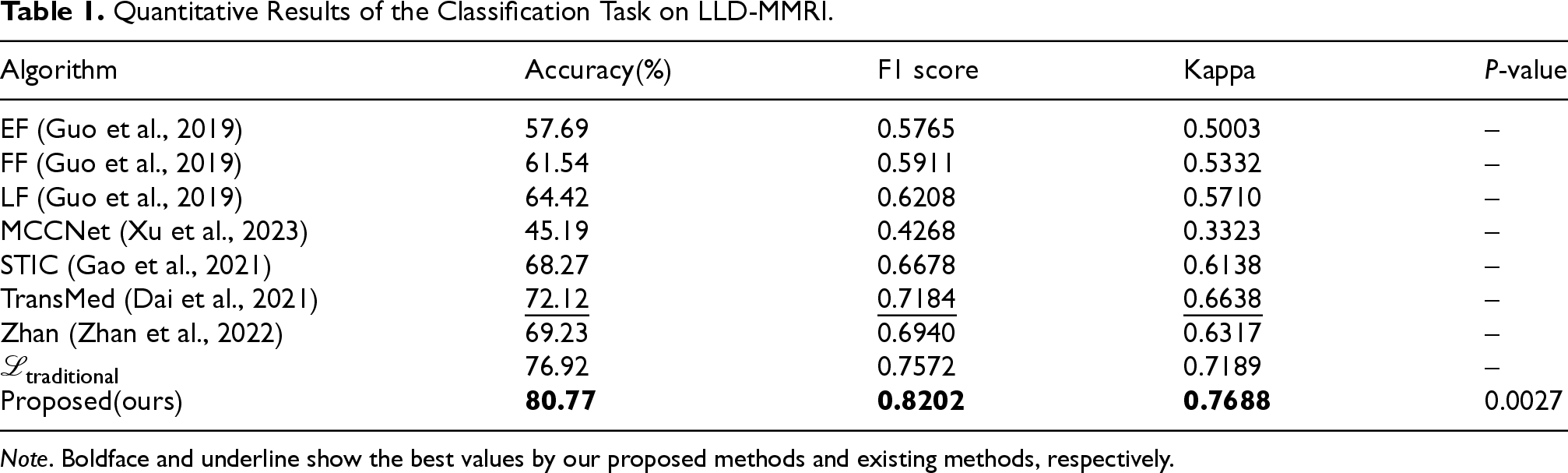
Note . Boldface and underline show the best values by our proposed methods and existing methods, respectively.
Quantitative Results of the Classification Task on LLD-MMRI.

Experiments were also conducted using the traditional loss function
To enhance the reliability of the experimental results, we selected methods employing traditional loss functions as the baseline and performed a paired sample
The visualization heatmaps are shown in Figure 7. It illustrates an example of the test set, where the first row displays the eight original MRI modalities. The three middle rows present heatmaps obtained by existing methods after training on LLD-MMRI and testing on the example images. The last row shows the results from our proposed method. It can be observed that STIC has larger attention regions for each modality, with only T2WI, out-phase, and venous phase focusing on the tumor area, while other modalities attend to peripheral background regions. In comparison, Zhan and our work leverage a transformer for feature fusion, which effectively avoids the problem of uneven attention across modalities in multi-modal situations. TransMed cuts images into patches before feeding them into the transformer, leading to unnecessary attention requirement for each patch by the network and, therefore, wasting computational resources. Although achieving good performance, Zhan presents overconcentration on the tumor area with attention regions covering the tumor for all modalities, which also results in inefficient usage of the network capacity. Our method demonstrates proper attention effects on both the tumor region and high-importance areas in different modalities, while significantly reducing redundant attention regions, hence enhancing the utilization efficiency of the network.
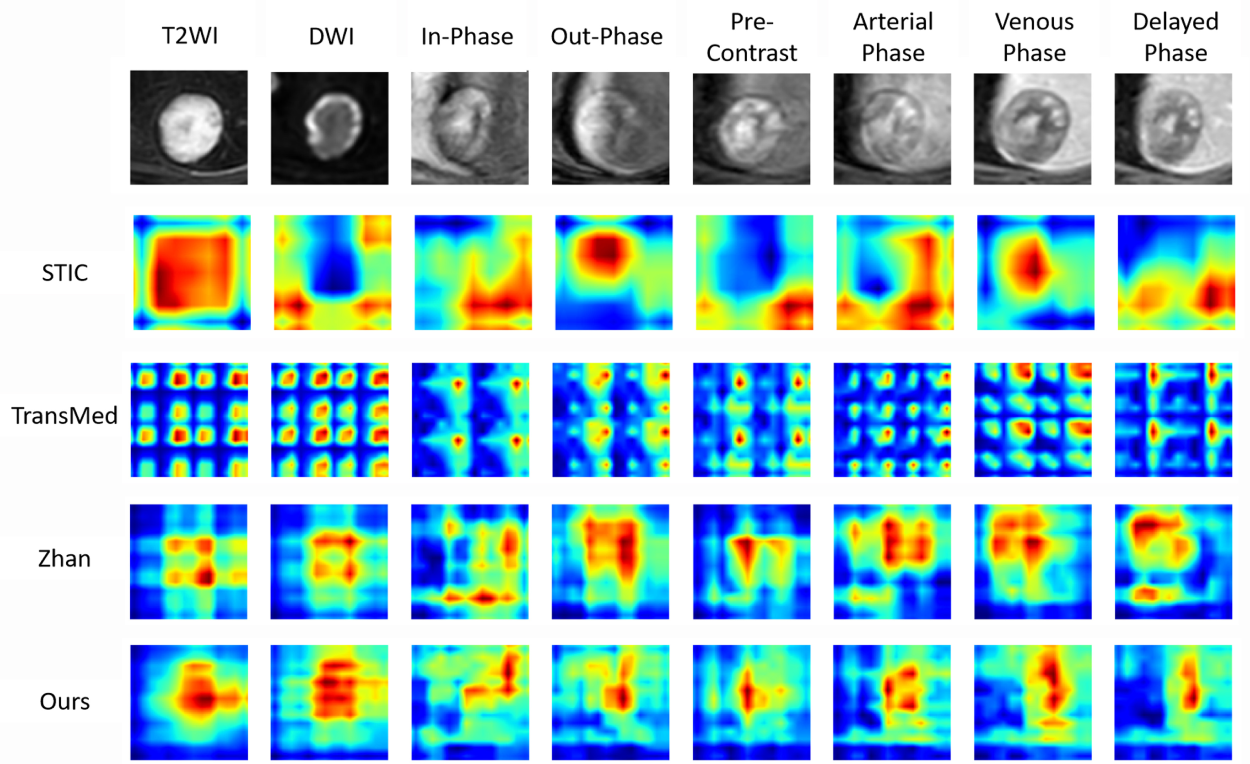
Comparison of heatmap visualizations on the LLD-MMRI dataset. Here, eight columns represent eight different phases. The first row shows the original image, the middle row shows the results of other methods, and the last row shows the results of our method.
To explore the impact of each module of our proposed method further, we conducted ablation studies on image classification.
Ablation Study Results of the Classification With Different Architectures on LLD-MMRI Dataset.
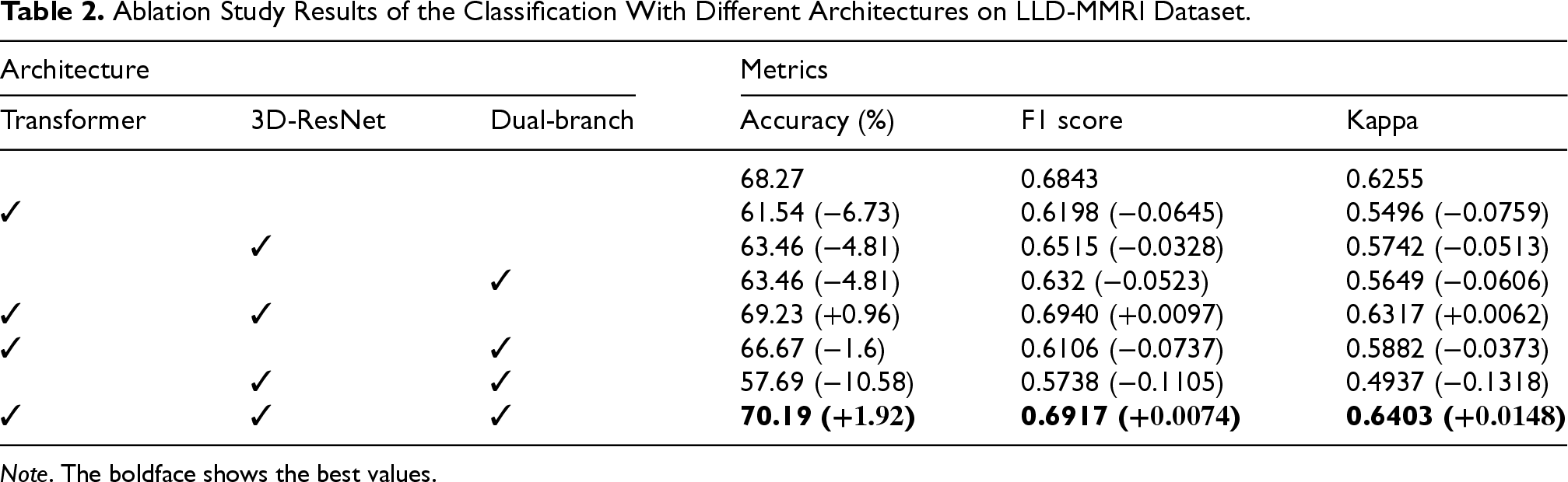
Note . The boldface shows the best values.
Ablation Study Results of the Classification With Different Architectures on LLD-MMRI Dataset.

Firstly, results represent we can have the best performance when combining all three architectures, and we can observe that using each architecture alone for feature fusion leads to a noticeable performance drop, especially on the transformer. However, when we utilize transformer architecture in collocation with others, performance could be improved. This indicates the complementary effects between different modules, and the transformer cannot fully exploit when models are inappropriate. Another phenomenon worth noting is that when we applied 3D-ResNet and dual-branch, the performance is the worst compared to other studies. This may mean that the transformer can effectively combine different modules to improve the overall performance of the model.
Ablation Study Results of Classification with Different Loss Functions on LLD-MMRI Dataset.
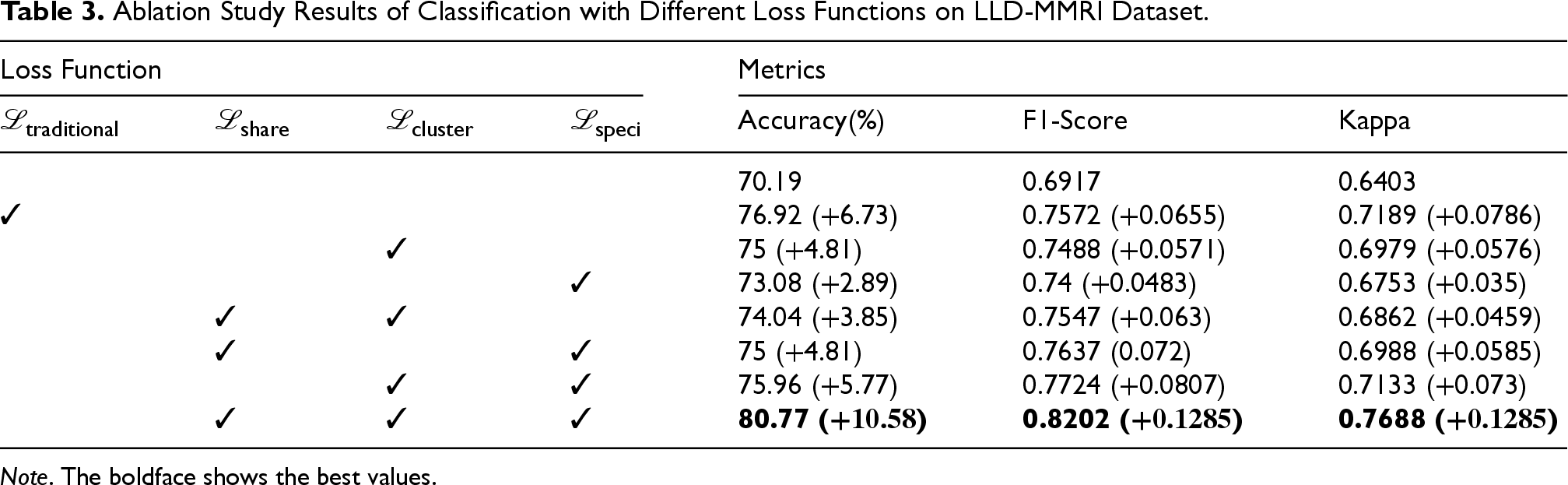
Analysis indicates performance deficits when excluding each of the proposed loss sub-modules relative to baseline metrics achieved using traditional loss functions, validating the synergistic interplay conferred by the introduced objective terms. The isolated use of
The first aspect, we conducted supplementary experiments on increased-modality scenarios to validate the rationality of the proposed approach from two dimensions: inference time per image expended and final classification results. Specifically, we expanded the LLD-MMRI into eight subsets with one to eight modalities, respectively, and performed training and testing under the corresponding modalities. The results are presented in Table 5.
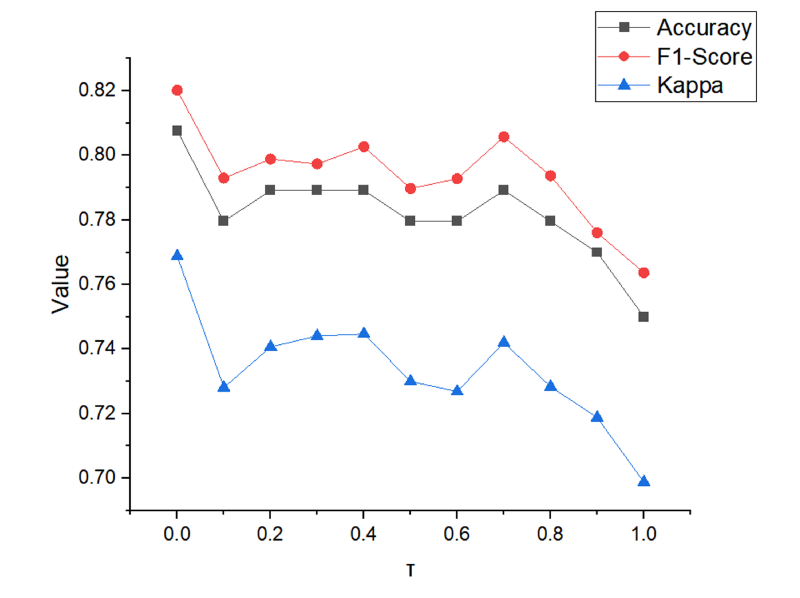
Results for different value of
Ablation Study Results of Classification With Different Features on LLD-MMRI Dataset.
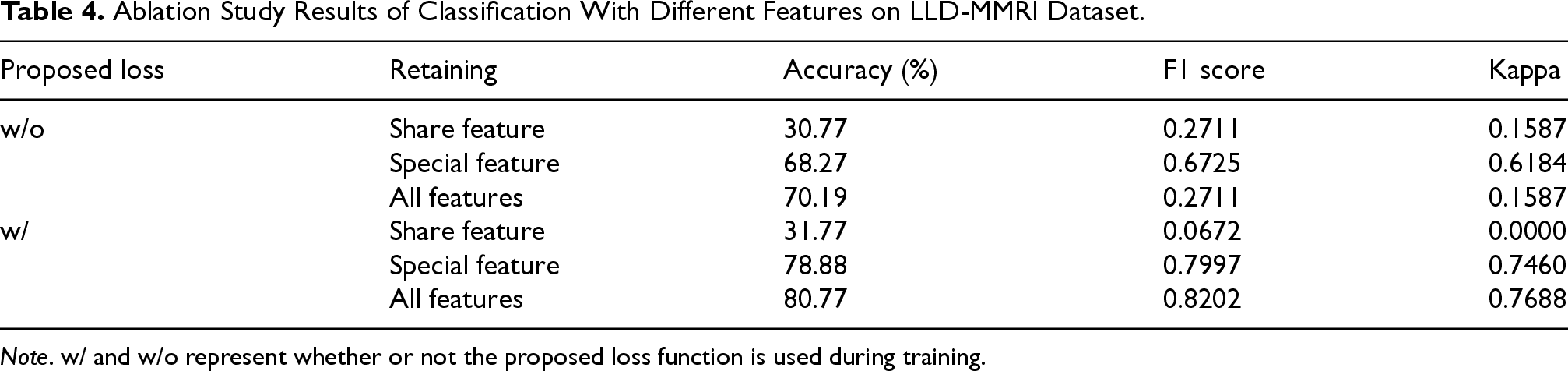
Quantitative Results of the Classification Task With Different Number of Modalities.
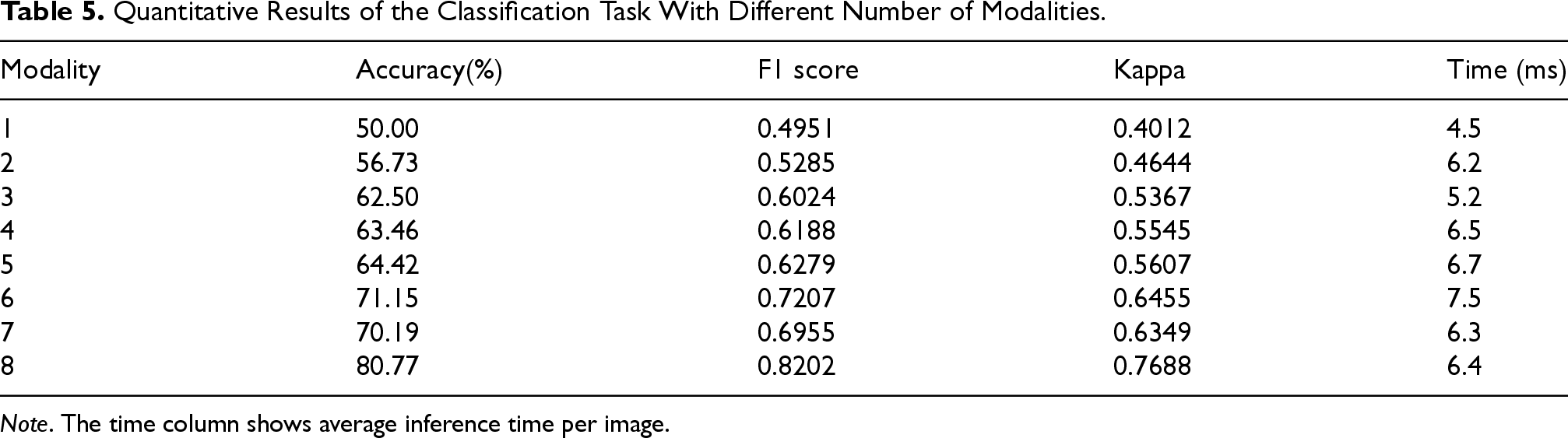
Comparing performance from one to eight modalities shows accuracy and other metrics improving steadily as the number of modalities increases, with accuracy boosted by 30.77% and other metrics improved by over 0.3 when reaching eight modalities. This validates the importance and necessity of multi-modal image fusion and demonstrates our model’s efficacy in fusing high-level semantic information across modalities. Also, the testing time shows a large increase only from one to two modalities, remaining stable as modalities further increase. This proves our model’s strong scalability with the number of modalities.
On the other hand, we test the robust performance of the model by adding Gaussian noise or a mask to the test image to simulate a low-quality image. The results are shown in Figure 9. It can be found that for low noise intensity, there is good robustness, but when the Gaussian noise intensity is >40, the performance decreases sharply. For adding masks, the robustness is poor, 5% mask also leads to a severe degradation in performance, probably because the model utilizes the detailed information to a high degree.
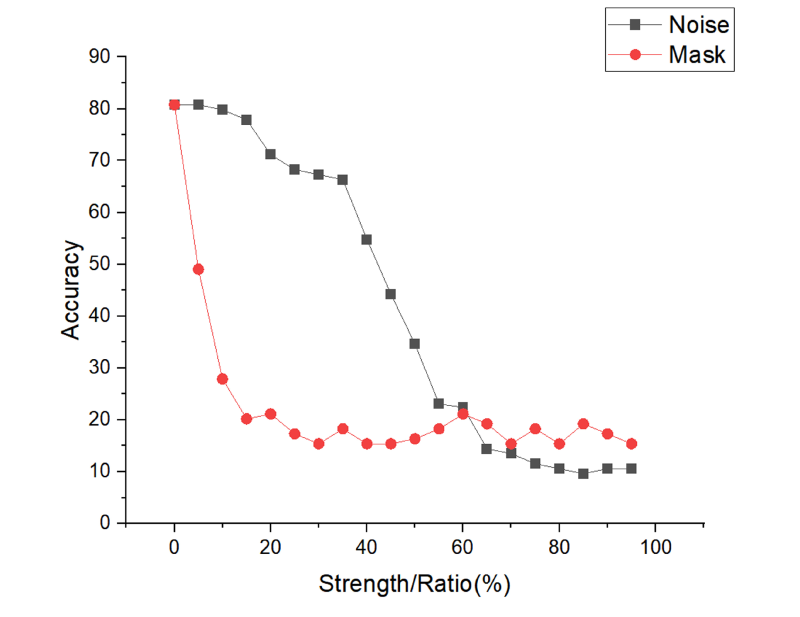
Results for different strengths of noise and different percentages of mask ratio.
For image fusion, we selected IFCNN (Zhang et al., 2020), DenseFuse (Li & Wu, 2019), EMFusion (Xu & Ma, 2021), U2Fusion (Xu et al., 2022), DualFuse (Fu & Wu, 2021), IFT (Vs et al., 2022), and CDDFuse (Zhao et al., 2023) as comparison methods. The first four are based on single-branch architectures, among which U2Fusion fuses features from different depths of the single branch and is categorized as a single-branch method. The last three share the dual-branch structure with our model, allowing comprehensive evaluation of our approach’s efficacy. Experiments were conducted by training and testing each model on the full AANLIB dataset, where similarity loss in EMFusion is only applicable to fusing red–green–blue images, we remove it in training.
As shown in Table 6, displaying fusion results between every modality pair would be lengthy, so we included them in the supplemental materials and only present the average performance on the test set after full testing in the main text. Ablation studies will also follow this format. It can be observed that the EN metric has little difference, since it was used as the loss function by all methods. We see that U2Fusion achieved excellent performance in the single-branch setting, but still lags behind the dual-branch networks. Among the dual-branch methods, the transformer structure outperformed CNN by a large SSIM margin. Without the share feature fusion module and constraints for specific features, our method remained comparable to the SOTA method CDDFuse. This validates that our proposed approach maintains strong multi-modal semantic fusion capacity without sacrificing low-level dual-modality fusion performance.
Quantitative Results of the Fusion Task on AANLIB.
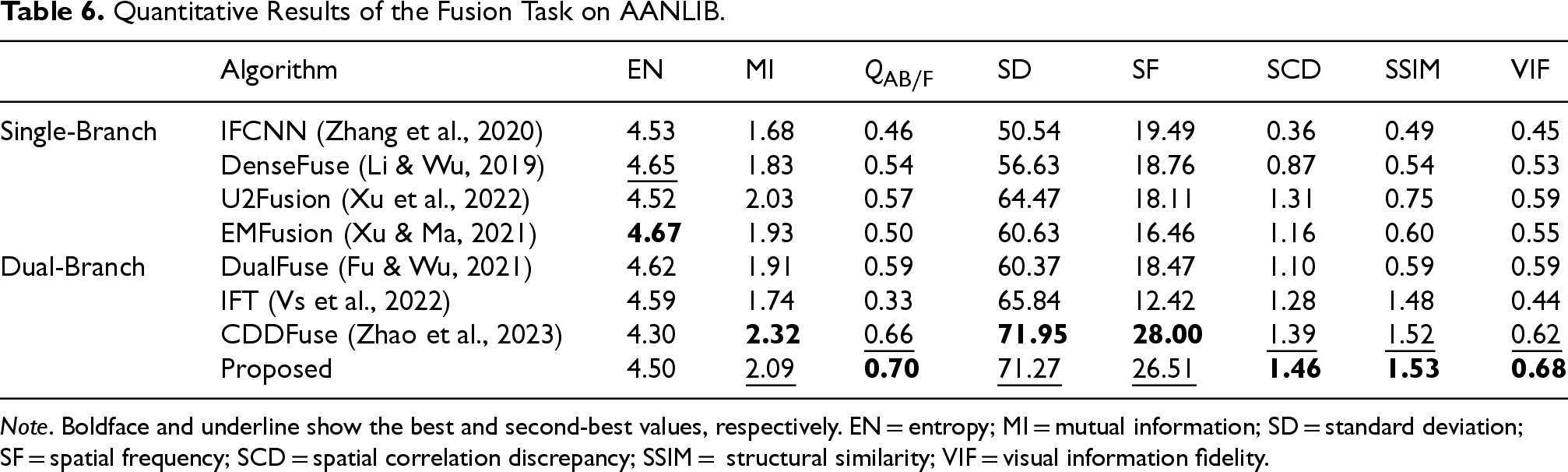
Note . Boldface and underline show the best and second-best values, respectively. EN = entropy; MI = mutual information; SD = standard deviation; SF = spatial frequency; SCD = spatial correlation discrepancy; SSIM = structural similarity; VIF = visual information fidelity.
Quantitative Results of the Fusion Task on AANLIB.

The visualization results are shown in Figure 10. Three subfigures depict the visualization of the fusion between MRI and CT, PET, and SPECT, respectively. The images demonstrate that our proposed method and CDDFuse achieve clearer and more detailed fusion results compared to other existing methods across different modality fusion tasks. Moreover, we can observe that CDDFuse suffers from loss of CT characteristics after MRI–CT fusion, and also leads to inversion of SPECT features from white to black after MRI–SPECT fusion, while our method still obtains better results on such samples, indicating higher preservation of semantic features after fusion. Furthermore, in MRI–PET fusion, the small colored region in the middle fused by our method exhibits more significant contrast compared to the surroundings than that of CDDFuse, suggesting that our method enables finer-grained fusion of color characteristics.
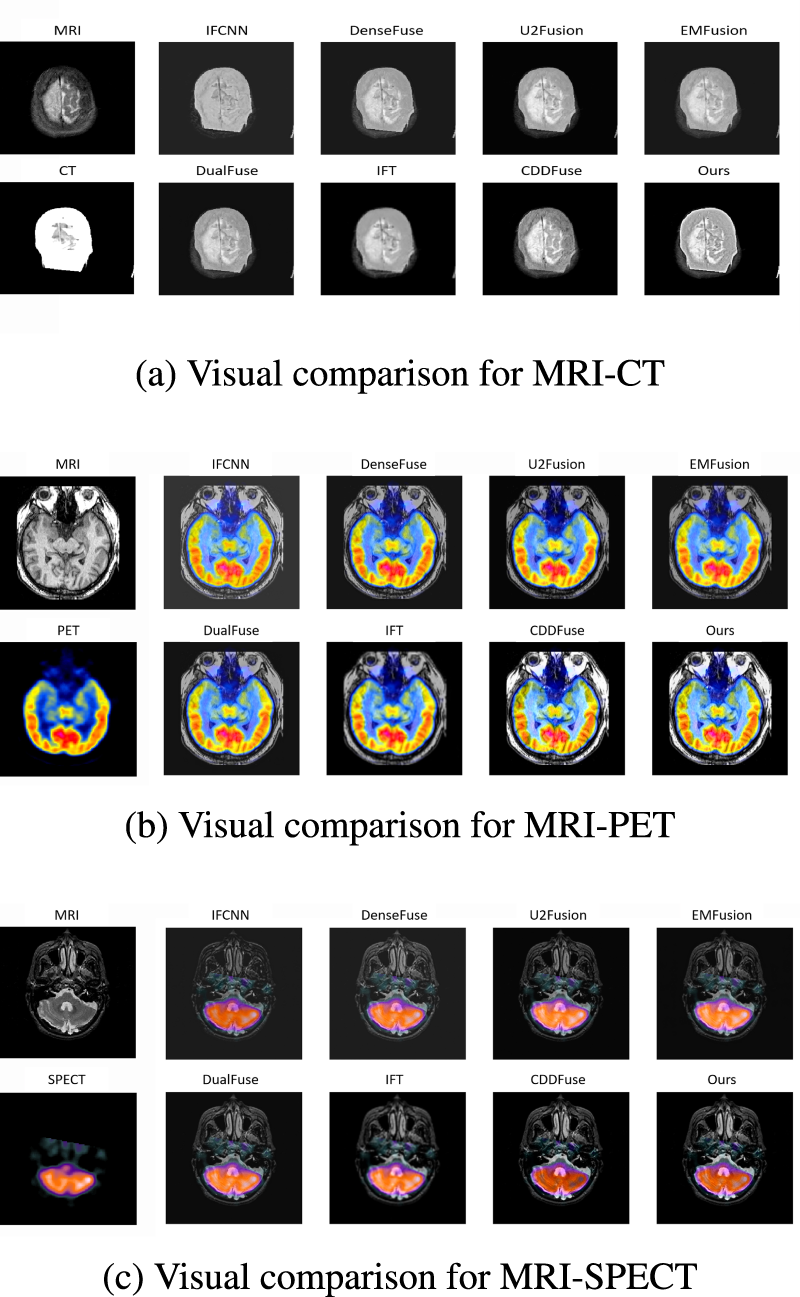
Visual comparison in medical image on the ANNLIB dataset with three combinations of different modalities. (a) Visual comparison for MRI–CT, (b) visual comparison for MRI–PET, and (c) visual comparison for MRI–SPECT. MRI = magnetic resonance imaging; CT = computed tomography; PET = positron emission tomography; SPECT = single-photon emission computed tomography.
In this paper, we propose a dual-branch architecture that can be used to efficiently diagnose multi-modal medical images. Using a basic network architecture and following the pipeline, we propose a prototype loss for different feature maps to efficiently consolidate multi-modal information when increasing the number of modalities. Experiments on both image classification and fusion tasks validate the fusion efficacy of the method. This provides an effective general solution for deep learning to utilize different modality information as the number of medical imaging modalities increases. However, due to limited time and resources, we did not conduct more experiments to identify the most effective architectures for medical images, nor validated on more multi-modal medical imaging tasks and datasets.
Footnotes
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
